Antimicrobial Susceptibility and Characterization of Resistance Mechanisms of Corynebacterium urealyticum Clinical Isolates
Abstract
1. Introduction
2. Results
2.1. Susceptibility of C. urealyticum to Antimicrobial Agents
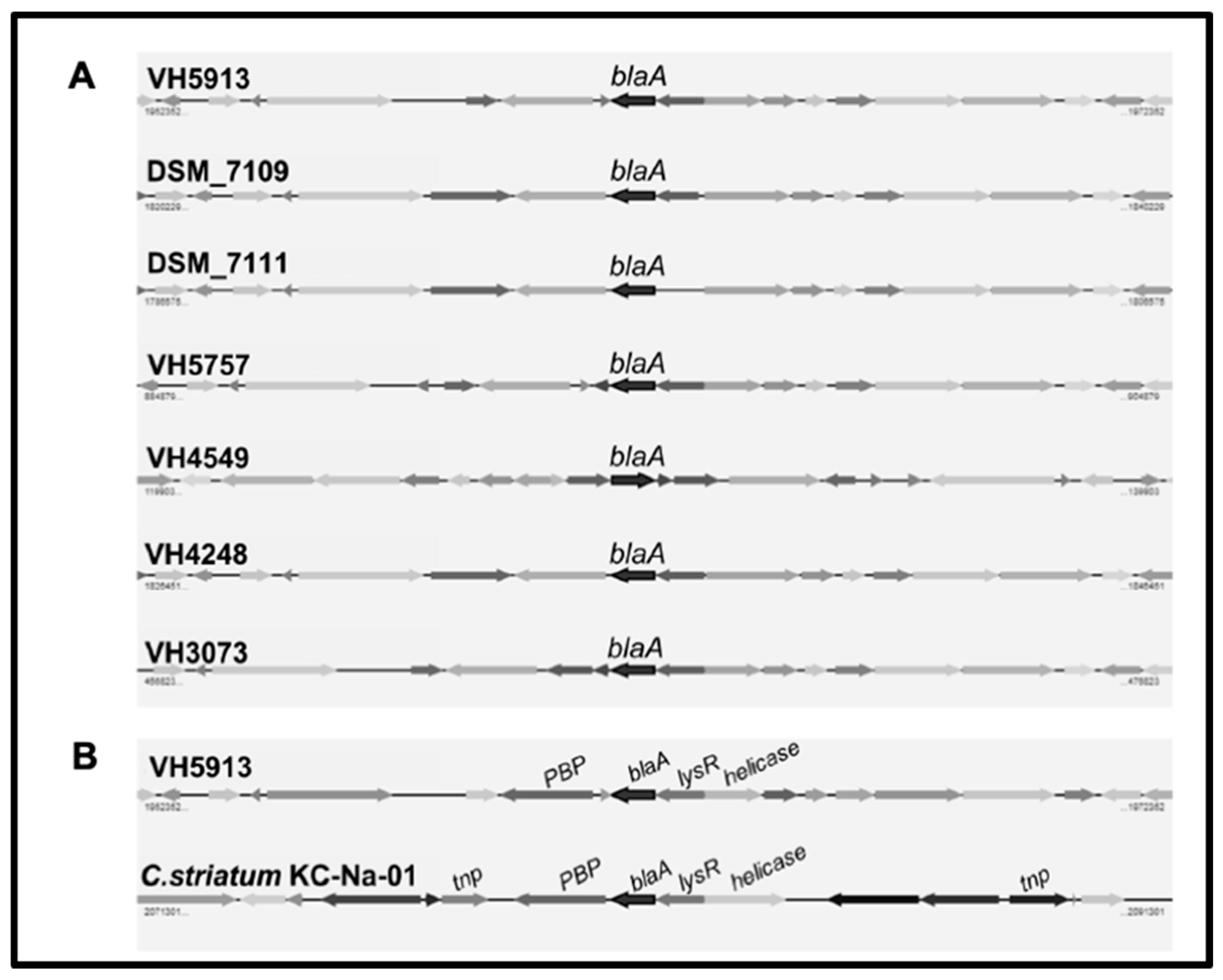
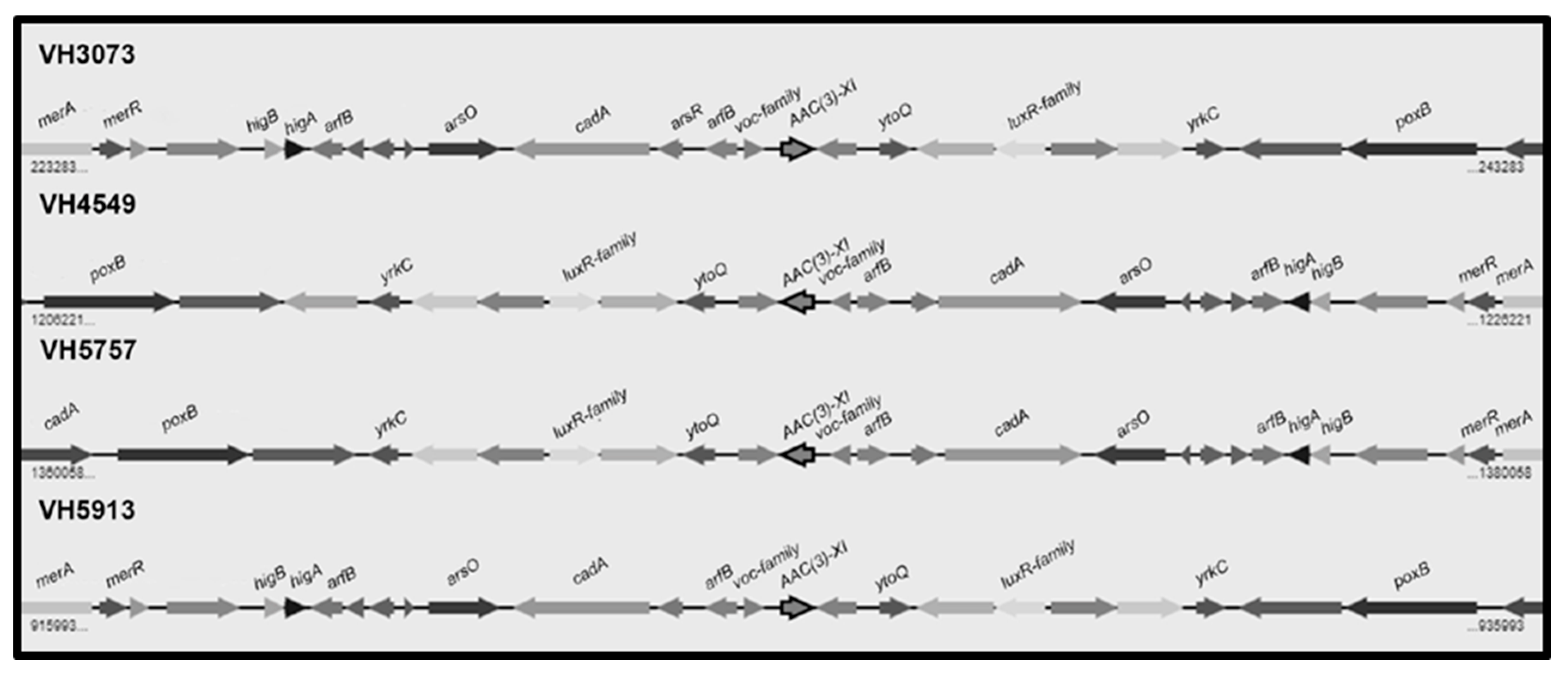
2.2. Detection of Resistance Genes by PCR and Genome Sequencing
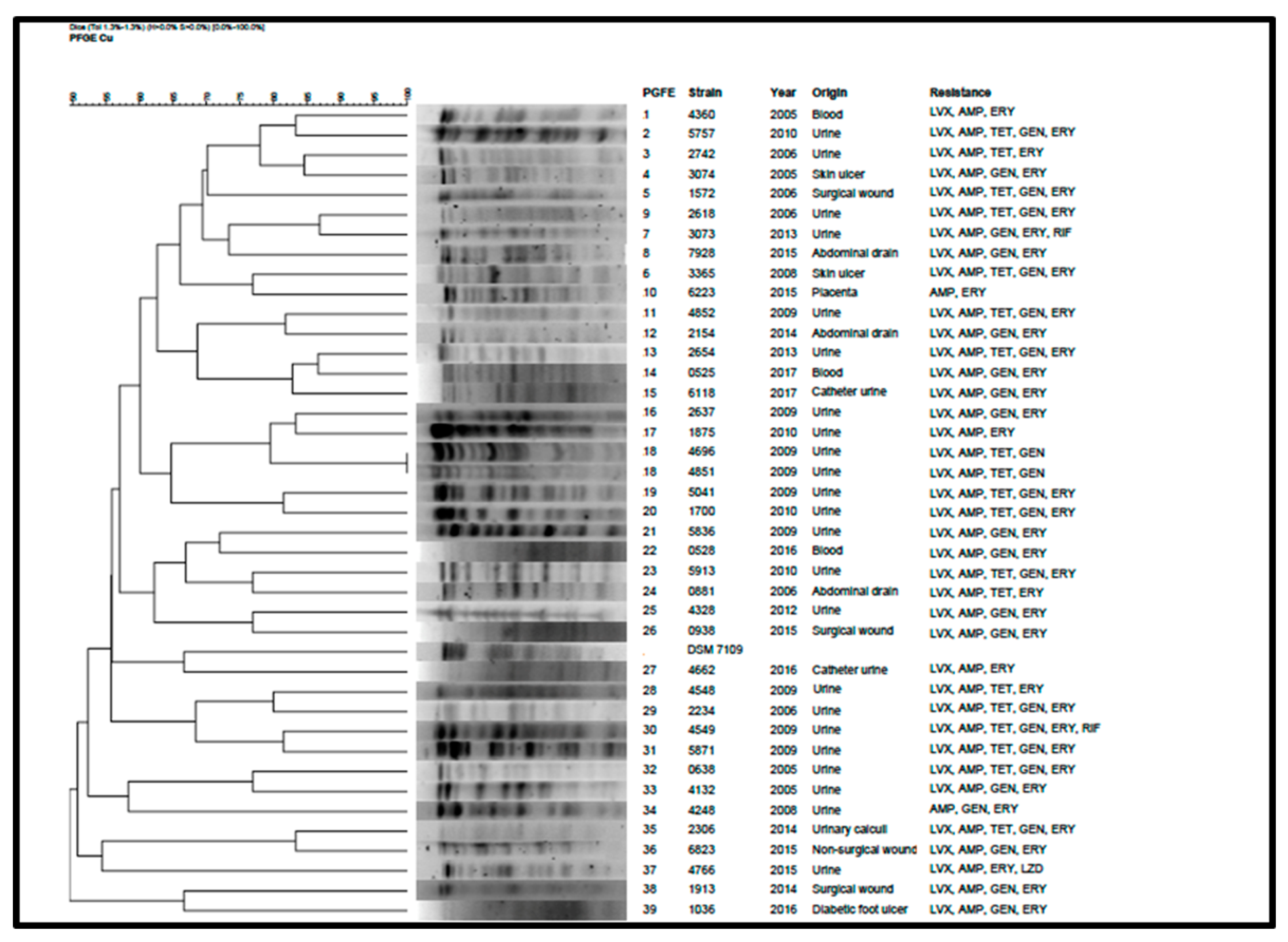
2.3. Molecular Epidemiology of the C. urealyticum Isolates
3. Discussion
4. Materials and Methods
4.1. Bacterial Strains and Growth Conditions
4.2. Antimicrobial Susceptibility Assays
4.3. Search of Resistance Genes by PCR
4.4. PCR Products Sequencing
4.5. Genome Sequencing and Analysis
4.6. Pulsed-Field Gel Electrophoresis (PFGE)
4.7. Data Availability
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Bernard, K. The genus Corynebacterium and other medically relevant coryneform-like bacteria. J. Clin. Microbiol. 2012, 50, 3152–3158. [Google Scholar] [CrossRef]
- Ramsey, M.M.; Freire, M.O.; Gabrilska, R.A.; Rumbaugh, K.P.; Lemon, K.P. Staphylococcus aureus Shifts toward Commensalism in Response to Corynebacterium Species. Front. Microbiol. 2016, 7, 1230. [Google Scholar] [CrossRef] [PubMed]
- Soriano, F.; Ponte, C.; Santamaria, M.; Castilla, C.; Fernández Roblas, R. In vitro and in vivo study of stone formation by Corynebacterium group D2 (Corynebacterium urealyticum). J. Clin. Microbiol. 1986, 23, 691–694. [Google Scholar] [CrossRef]
- Tauch, A.; Trost, E.; Tilker, A.; Ludewig, U.; Schneiker, S.; Goesmann, A.; Arnold, W.; Bekel, T.; Brinkrolf, K.; Brune, I.; et al. The lifestyle of Corynebacterium urealyticum derived from its complete genome sequence established by pyrosequencing. J. Biotechnol. 2008, 136, 11–21. [Google Scholar] [CrossRef] [PubMed]
- Pagnoux, C.; Bérezné, A.; Damade, R.; Paillot, J.; Aouizerate, J.; Le Guern, V.; Salmon, D.; Guillevin, D. Encrusting Cystitis Due to Corynebacterium urealyticum in a Patient with ANCA-Associated Vasculitis: Case Report and Review of the Literature. Semin. Arthritis. Rheum. 2011, 41, 297–300. [Google Scholar] [CrossRef] [PubMed]
- Funke, G.; Von Graevenitz, A.; Clarridge, J.E.; Bernard, K.A.; Funke, G. Clinical Microbiology of Coryneform Bacteria. Clin. Microbiol. Rev. 1997, 10, 125–159. [Google Scholar] [CrossRef]
- Fish, D.N.; Piscitelli, S.C.; Danzinger, L.H. Development of Resistance During Antimicrobial Therapy. Pharmacotherapy 1995, 15, 279–291. [Google Scholar] [CrossRef]
- Gómez-Garcés, J.L.; Alos, J.I.; Tamayo, J. In vitro activity of linezolid and 12 other antimicrobials against coryneform bacteria. Int. J. Antimicrob. Agents 2007, 29, 688–692. [Google Scholar] [CrossRef]
- Fernández-Roblas, R.; Adames, H.; Martín-de-Hijas, N.Z.; García Almeida, D.; Gadea, I.; Esteban, J. In vitro activity of tigecycline and 10 other antimicrobials against clinical isolates of the genus Corynebacterium. Int. J. Antimicrob. Agents 2009, 33, 453–455. [Google Scholar] [CrossRef]
- Neemuchwala, A.; Soares, D.; Ravirajan, V.; Marchand-Austin, A.; Kus, J.V.; Patel, S.N. In Vitro Antibiotic Susceptibility Pattern of Non-diphtheriae Corynebacterium Isolates in Ontario, Canada, from 2011 to 2016. Antimicrob. Agents Chemother. 2018, 62, e01776-17. [Google Scholar] [CrossRef]
- Magiorakos, A.P.; Srinivasan, A.; Carey, R.B.; Carmeli, Y.; Falagas, M.E.; Giske, C.G.; Harbarth, S.; Hindler, J.F.; Kahlmeter, G.; Olsson-Liljequist, B.; et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: An international expert proposal for interim standard definitions for acquired resistance. Clin. Microbiol. Infect. 2012, 18, 268–281. [Google Scholar] [CrossRef] [PubMed]
- Guimaraes, L.C.; Soares, S.C.; Albersmeier, A.; Blom, J.; Jaenicke, S.; Azevedo, V.; Soriano, F.; Tauch, A.; Trost, E. Complete Genome Sequence of Corynebacterium urealyticum Strain DSM 7111, Isolated from a 9-Year-Old Patient with Alkaline-Encrusted Cystitis. Genome Announc. 2013, 1, e00264-13. [Google Scholar] [CrossRef] [PubMed]
- Schröder, J.; Maus, I.; Meyer, K.; Wördemann, S.; Blom, J.; Jaenicke, S.; Schneider, J.; Trost, E.; Tauch, A. Complete genome sequence, lifestyle, and multi-drug resistance of the human pathogen Corynebacterium resistens DSM 45100 isolated from blood samples of a leukemia patient. BMC Genom. 2012, 13, 141. [Google Scholar] [CrossRef] [PubMed]
- Galimand, M.; Fishovitz, J.; Lambert, T.; Barbe, V.; Zajicek, J.; Mobashery, S.; Courvalin, P. AAC(3)-XI, a new aminoglycoside 3-N-acetyltransferase from Corynebacterium striatum. Antimicrob. Agents Chemother. 2015, 59, 5647–5653. [Google Scholar] [CrossRef] [PubMed]
- Soriano, F.; Tauch, A. Microbiological and clinical features of Corynebacterium urealyticum: Urinary tract stones and genomics as the Rosetta Stone. Clin. Microbiol. Infect. 2008, 14, 632–643. [Google Scholar] [CrossRef] [PubMed]
- Salas, C.; Calvo, J.; Martinez-Martinez, L. Activity of Tigecycline against Coryneform Bacteria of Clinical Interest and Listeria monocytogenes. Antimicrob. Agents Chemother. 2008, 52, 1503–1505. [Google Scholar] [CrossRef][Green Version]
- Navas, J.; Salas, C.; Calvo, J.; Martinez-Martinez, L. Activity of daptomycin and three comparator agents against non-diphtheriae Corynebacterium isolates of clinical interest. J. Antimicrob. Chemother. 2012, 67, 776–778. [Google Scholar] [CrossRef]
- Lopez-Medrano, F.; Garcia-Bravo, M.; Morales, J.M.; Andrés, A.; San Juan, R.; Lizasoain, M.; Aguado, J.M. Urinary tract infection due to Corynebacterium urealyticum in kidney transplant recipients: An underdiagnosed etiology for obstructive uropathy and graft dysfunction-results of a prospective cohort study. Clin. Infect. Dis. 2008, 46, 825–830. [Google Scholar] [CrossRef]
- Ortiz-Pérez, A.; Martín-De-Hijas, N.Z.; Esteban, J.; Fernández-Natal, M.I.; García-Cía, J.I.; Fernández-Roblas, R. High Frequency of Macrolide Resistance Mechanisms in Clinical Isolates of Corynebacterium Species. Microb. Drug Resist. 2010, 16, 273–277. [Google Scholar] [CrossRef]
- Ramos, J.N.; Valadao, T.B.; Baio, P.V.P.; Mattos-Guaraldi, A.L.; Vieira, V.V. Novel mutations in the QRDR region gyrA gene in multidrug-resistance Corynebacterium spp. isolates from intravenous sites. Antonie Van Leeuwenhoek 2020, 113, 589–592. [Google Scholar] [CrossRef]
- Noussair, L.; Salomon, E.; El Sayed, F.; Duran, C.; Bouchand, F.; Roux, A.L.; Gaillard, J.L.; Bauer, T.; Rottman, M.; Dinh, A. Monomicrobial bone and joint infection due to Corynebacterium striatum: Literature review and amoxicillin-rifampin combination as treatment perspective. Eur. J. Clin. Microbiol. Infect. Dis. 2019, 38, 1269–1278. [Google Scholar] [CrossRef]
- Shah, M.; Murillo, J.L. Successful treatment of Corynebacterium striatum endocarditis with daptomycin plus rifampin. Ann. Pharmacother. 2005, 39, 1741–1744. [Google Scholar] [CrossRef] [PubMed]
- Yang, B.; Koga, H.; Ohno, H.; Ogawa, K.; Fukuda, M.; Hirakata, Y.; Maesaki, S.; Tomono, K.; Tashiro, T.; Kohno, S. Relationship between antimycobacterial activities of rifampicin, rifabutin and KRM-1648 and rpoB mutations of Mycobacterium tuberculosis. J. Antimicrob. Chemother. 1998, 42, 621–628. [Google Scholar] [CrossRef] [PubMed]
- Horng, Y.T.; Jeng, W.Y.; Chen, Y.Y.; Liu, C.H.; Dou, H.Y.; Lee, J.J.; Chang, K.C.; Chien, C.C.; Soo, P.C. Molecular analysis of codon 548 in the rpoB gene involved in Mycobacterium tuberculosis resistance to rifampin. Antimicrob. Agents Chemother. 2015, 59, 1542–1548. [Google Scholar] [CrossRef] [PubMed]
- Salem, N.; Salem, L.; Saber, S.; Ismail, G.; Bluth, M. Corynebacterium urealyticum: A comprehensive review of an understated organism. Infect. Drug Resist. 2015, 8, 129–145. [Google Scholar] [CrossRef]
- Navas, J.; Fernández-Martínez, M.; Salas, C.; Cano, M.E.; Martinez-Martinez, L. Susceptibility to Aminoglycosides and Distribution of aph and aac(3)-XI Genes among Corynebacterium striatum Clinical Isolates. PLoS ONE 2016, 11, e0167856. [Google Scholar] [CrossRef]
- Tauch, A.; Krieft, S.; Pühler, A.; Kalinowski, J. The tetAB genes of the Corynebacterium striatum R-plasmid pTP10 encode an ABC transporter and confer tetracycline, oxytetracycline and oxacillin resistance in Corynebacterium glutamicum. FEMS Microbiol. Lett. 1999, 173, 203–209. [Google Scholar] [CrossRef][Green Version]
- Goering, R.V. Pulsed field gel electrophoresis: A review of application and interpretation in the molecular epidemiology of infectious disease. Infect. Genet. Evol. 2010, 10, 866–875. [Google Scholar] [CrossRef]
- European Committee on Antimicrobial Susceptibility Testing. Breakpoint Tables for Interpretation of MICs and Zone Diameters; Version 10.0; EUCAST: Växjö, Sweden, 2020; Available online: http://www.eucast.org (accessed on 25 January 2020).
- Sutcliffe, J.; Grebe, T.; Tait-Kamradt, A.; Wondrack, L. Detection of erythromycin-resistant determinants by PCR. Antimicrob. Agents Chemother. 1996, 40, 2562–2566. [Google Scholar] [CrossRef]
- Alibi, S.; Ferjani, A.; Boukadida, J.; Cano, M.E.; Fernandez-Martinez, M.; Martinez-Martinez, L.; Navas, J. Occurrence of Corynebacterium striatum as an emerging antibiotic-resistant nosocomial pathogen in a Tunisian hospital. Sci. Rep. 2017, 7, 9704. [Google Scholar] [CrossRef]
- Sierra, J.M.; Martínez-Martínez, L.; Vázquez, F.; Giralt, E.; Vila, J. Relationship between mutations in the gyrA gene and quinolone resistance in clinical isolates of Corynebacterium striatum and Corynebacterium amycolatum. Antimicrob. Agents Chemother. 2005, 49, 1714–1719. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; Mcgettigan, P.A.; McWilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, P.; et al. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef] [PubMed]
- Wattam, A.R.; Davis, J.J.; Assaf, R.; Boisvert, S.; Brettin, T.; Bun, C.; Conrad, A.; Dietrich, E.M.; Disz, T.; Gabbard, J.L.; et al. Improvements to PATRIC, the all-bacterial Bioinformatics Database and Analysis Resource Center. Nucleic. Acids. Res. 2017, 45, D535–D542. [Google Scholar] [CrossRef]
- Overbeek, R.; Olson, R.; Pusch, G.D.; Olsen, G.J.; Davis, J.J.; Disz, T.; Edwards, R.A.; Gerdes, S.; Parrello, B.; Shukla, M.; et al. The SEED and the Rapid Annotation of microbial genomes using Subsystems Technology (RAST). Nucleic. Acids. Res. 2014, 42, 206–214. [Google Scholar] [CrossRef]
- Blom, J.; Kreis, J.; Spanig, S.; Juhre, T.; Bertelli, C.; Ernst, C.; Goesmann, A. EDGAR 2.0: An enhanced software platform for comparative gene content analyses. Nucleic. Acids. Res. 2016, 44, W22–W28. [Google Scholar] [CrossRef]
- Moretti, S.; Armougom, F.; Wallace, I.M.; Higgins, D.G.; Jongeneel, C.V.; Notredame, C. The M-Coffee web server: A meta-method for computing multiple sequence alignments by combining alternative alignment methods. Nucleic. Acids. Res. 2007, 35, W645–W648. [Google Scholar] [CrossRef]


| Antimicrobial Agent | Range (mg/L) | MIC50 | MIC90 | S ≤ Breakpoint > R | Resistant (n) | Total (%) R | |
|---|---|---|---|---|---|---|---|
| Ampicillin | 0.016–256 | 1 | >256 | 0.05 | 2 | 40 | 100.0 |
| Erythromycin * | 0.016–256 | >256 | >256 | 1 | 2 | 38 | 95.0 |
| Gentamicin | 0.016–256 | 4 | 9 | 1 | 1 | 33 | 82.5 |
| Levofloxacin | 0.002–32 | >32 | >32 | 1 | 1 | 38 | 95.0 |
| Linezolid | 0.015–256 | 0.75 | 1 | 2 | 2 | 1 | 2.5 |
| Rifampicin | 0.002–32 | 0.016 | 0.04 | 0.06 | 0.5 | 2 | 5.0 |
| Tetracycline | 0.016–256 | 2 | 4 | 2 | 2 | 20 | 50.0 |
| Vancomycin | 0.016–256 | 0.5 | 0.75 | 2 | 2 | 0 | 0.0 |
| Antimicrobial Agent | VH3073 | VH4248 | VH4549 | VH5757 | VH5913 |
|---|---|---|---|---|---|
| Ampicillin | >256 | >256 | >256 | >256 | >256 |
| Erythromycin | >256 | >256 | >256 | >256 | >256 |
| Gentamicin | 6 | 6 | 6 | 4 | 4 |
| Levofloxacin | >32 | 0.19 | >32 | >32 | >32 |
| Linezolid | 1 | 0.38 | 0.75 | 1 | 1 |
| Rifampicin | >32 | 0.04 | >32 | 0.016 | 0.016 |
| Tetracycline | 1.5 | 0.25 | 3 | 3 | 6 |
| Vancomycin | 0.75 | 0.38 | 0.5 | 0.75 | 0.75 |
| Strain | Tetracycline MIC | Tetracycline + PAβN MIC |
|---|---|---|
| VH638 | >256 | 0.016 |
| VH1572 | 4 | <0.016 |
| VH1700 | 4 | 0.094 |
| VH2234 | 32 | 1 |
| VH4548 | 4 | <0.016 |
| VH5757 | 3 | 0.125 |
| VH5913 | 6 | <0.016 |
| DSM 7109 | 32 | 32 |
| Gene | Resistance | DNA Sequence (5′-3′) | Tª | Size | Reference |
|---|---|---|---|---|---|
| ermX | Erythromycin | AACCATGATTGTGTTTCTGAACG ACCAGGAAGCGGTGCCCT | 57 °C | 560 | [30] |
| ermB | Erythromycin | GAAAAGGTACTCAACCAAATA AGTAACGGTACTTAAATTGTTTAC | 52 °C | 639 | [30] |
| mef(A-E) | Erythromycin | GCAAATGGTGTAGGTAAGACAACT TAAAACAAATGTAGTGTACTA | 52 °C | 399 | [30] |
| blaA | Penicillin | CAGTCTAGCCACTTCGCCAAT TGACTGCACGGATGGAGATGG | 55 °C | 808 | [31] |
| gyrA | Levofloxacin | GCGGCTACGTAAAGTCC CCGCCGGAGCCGTTCAT | 60 °C | 337 | [32] |
| tetA | Tetracycline | TTAGCGTTCGGCGACCTGG GGTGGTCTTGTCTGCCCTCA | 60 °C | 552 | This study |
| tetB | Tetracycline | ACGGTGTTCAACGCCCTGTT AACTGGGTGCCTTCAGGGTC | 59 °C | 506 | This study |
| rpoB | Rifampicin | CTGATCCAGAACCAGGTCCG GACGTACTCCACCACACCAG | 55 °C | 811 | This study |
| aac(3)-XI | Gentamicin | GCGGCTACGTAAAGTCC CCGCCGGAGCCGTTCAT | 60 °C | 337 | [14] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chapartegui-González, I.; Fernández-Martínez, M.; Rodríguez-Fernández, A.; Rocha, D.J.P.; Aguiar, E.R.G.R.; Pacheco, L.G.C.; Ramos-Vivas, J.; Calvo, J.; Martínez-Martínez, L.; Navas, J. Antimicrobial Susceptibility and Characterization of Resistance Mechanisms of Corynebacterium urealyticum Clinical Isolates. Antibiotics 2020, 9, 404. https://doi.org/10.3390/antibiotics9070404
Chapartegui-González I, Fernández-Martínez M, Rodríguez-Fernández A, Rocha DJP, Aguiar ERGR, Pacheco LGC, Ramos-Vivas J, Calvo J, Martínez-Martínez L, Navas J. Antimicrobial Susceptibility and Characterization of Resistance Mechanisms of Corynebacterium urealyticum Clinical Isolates. Antibiotics. 2020; 9(7):404. https://doi.org/10.3390/antibiotics9070404
Chicago/Turabian StyleChapartegui-González, Itziar, Marta Fernández-Martínez, Ana Rodríguez-Fernández, Danilo J. P. Rocha, Eric R. G. R. Aguiar, Luis G. C. Pacheco, José Ramos-Vivas, Jorge Calvo, Luis Martínez-Martínez, and Jesús Navas. 2020. "Antimicrobial Susceptibility and Characterization of Resistance Mechanisms of Corynebacterium urealyticum Clinical Isolates" Antibiotics 9, no. 7: 404. https://doi.org/10.3390/antibiotics9070404
APA StyleChapartegui-González, I., Fernández-Martínez, M., Rodríguez-Fernández, A., Rocha, D. J. P., Aguiar, E. R. G. R., Pacheco, L. G. C., Ramos-Vivas, J., Calvo, J., Martínez-Martínez, L., & Navas, J. (2020). Antimicrobial Susceptibility and Characterization of Resistance Mechanisms of Corynebacterium urealyticum Clinical Isolates. Antibiotics, 9(7), 404. https://doi.org/10.3390/antibiotics9070404









