Antimicrobial Susceptibility Testing for Salmonella Serovars Isolated from Food Samples: Five-Year Monitoring (2015–2019)
Abstract
1. Introduction
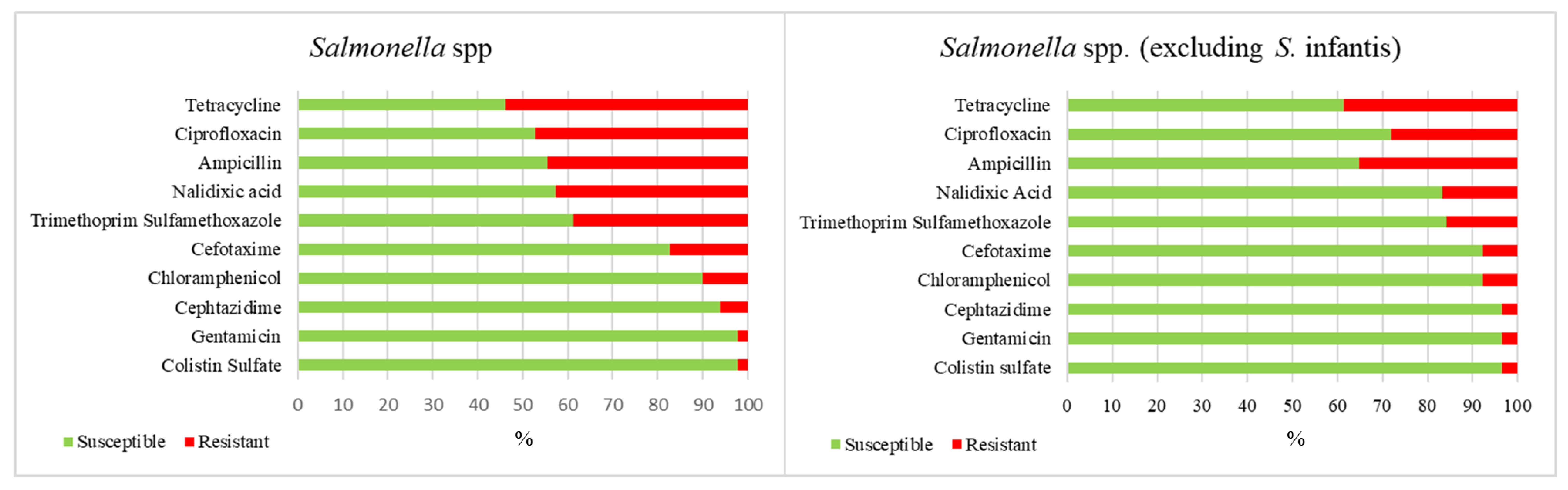
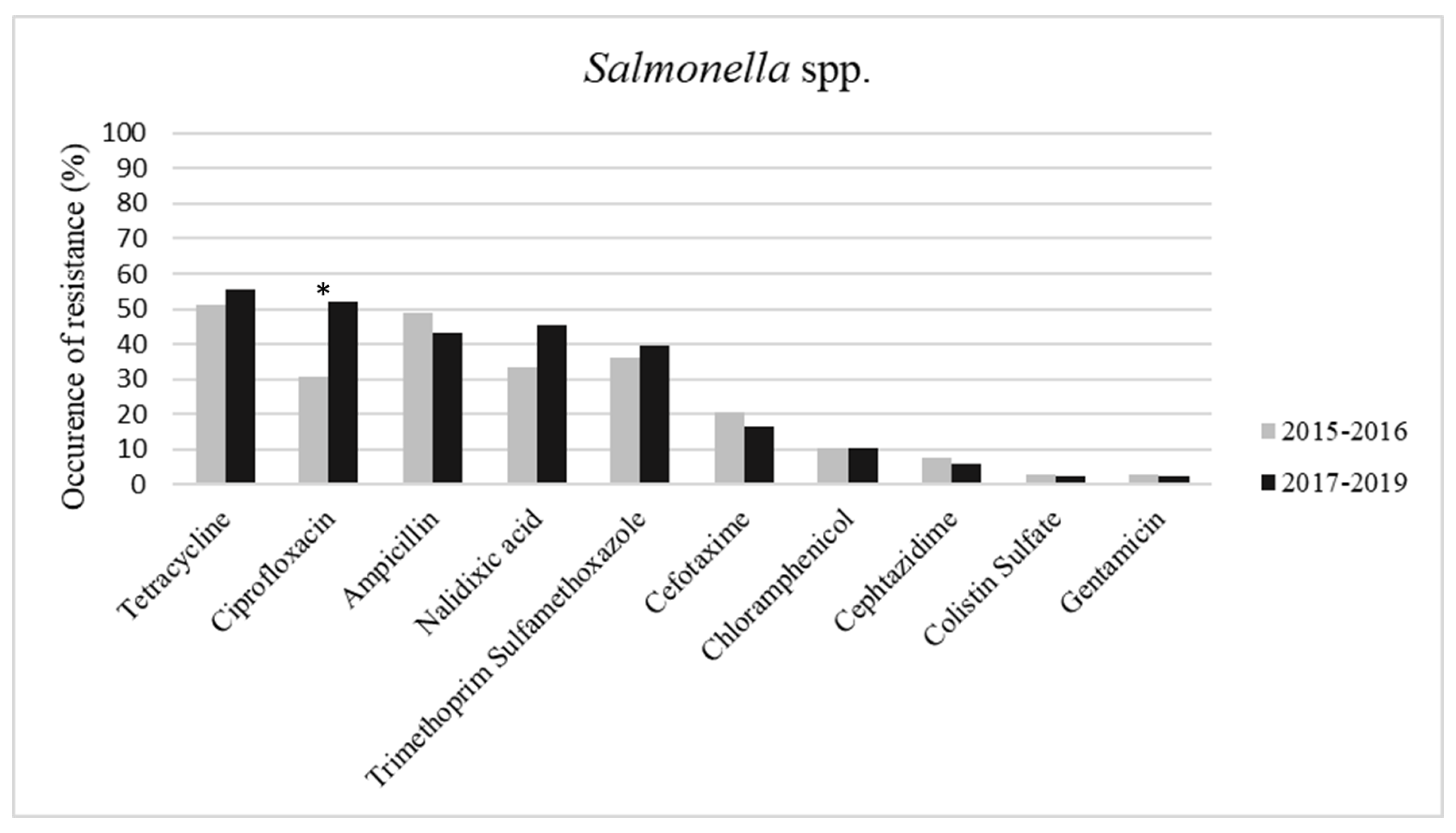
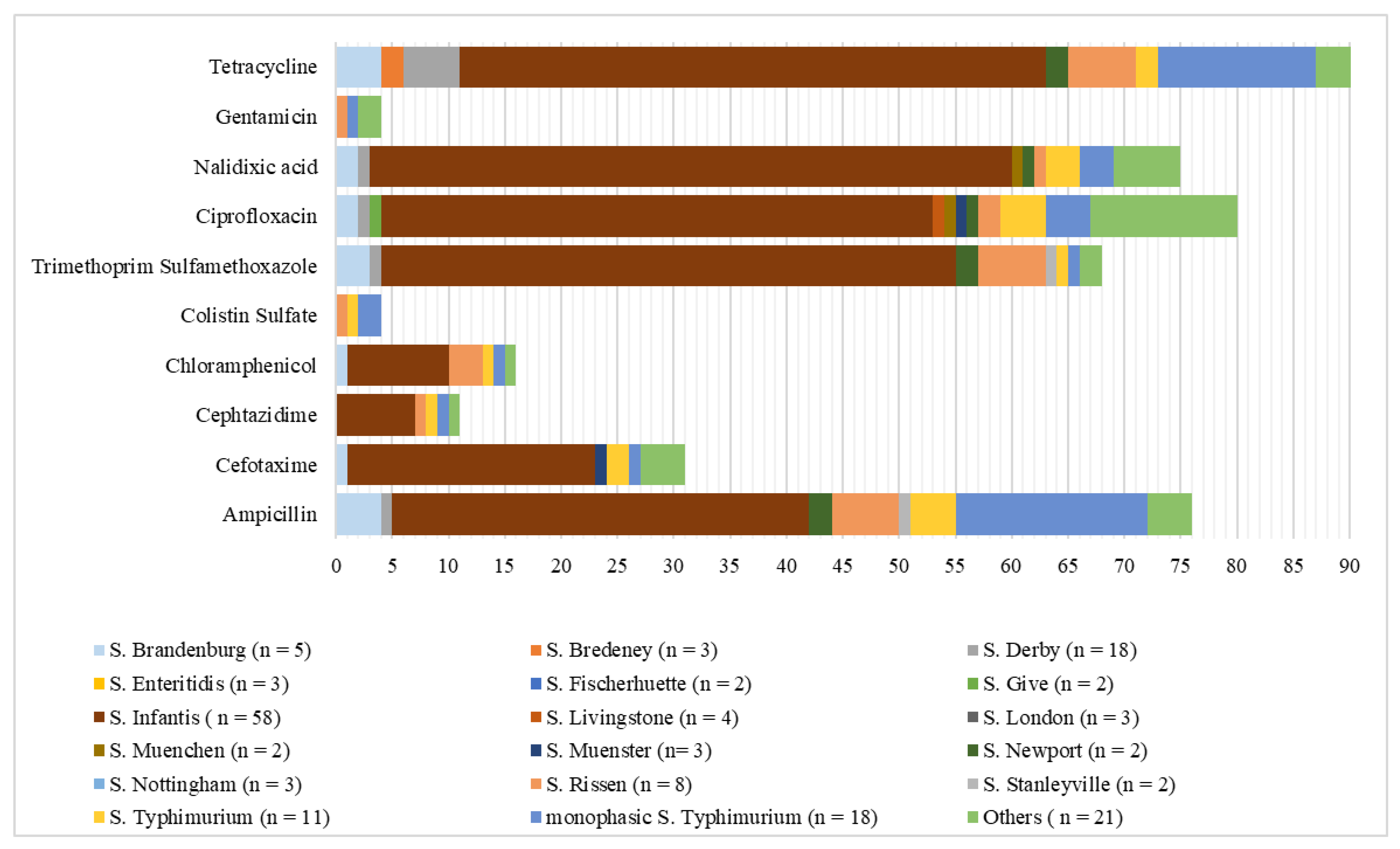
2. Results
3. Discussion
4. Materials and Methods
4.1. Strains
4.2. Antibiotic Susceptibility Testing
4.3. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- European Food Safety Authority; European Centre for Disease Prevention and Control. The European Union One Health 2018 Zoonoses Report. EFSA J. 2019, 17. [Google Scholar] [CrossRef]
- Mąka, Ł.; Popowska, M. Antimicrobial resistance of Salmonella spp. isolated from food. Rocz. Panstw. Zakl. Hig. 2016, 67, 343–358. [Google Scholar] [PubMed]
- Wang, X.; Biswas, S.; Paudyal, N.; Pan, H.; Li, X.; Fang, W.; Yue, M. Antibiotic resistance in salmonella typhimurium isolates recovered from the food chain through national antimicrobial resistance monitoring system between 1996 and 2016. Front. Microbiol. 2019, 10, 1–12. [Google Scholar] [CrossRef] [PubMed]
- European Food Safety Authority; European Centre for Disease Prevention and Control. The European Union Summary Report on Antimicrobial Resistance in zoonotic and indicator bacteria from humans, animals and food in 2017/2018. EFSA J. 2020, 18. [Google Scholar] [CrossRef]
- Caniça, M.; Manageiro, V.; Abriouel, H.; Moran-Gilad, J.; Franz, C.M.A.P. Antibiotic resistance in foodborne bacteria. Trends Food Sci. Technol. 2019, 84, 41–44. [Google Scholar] [CrossRef]
- Johnson, A.P.; Johnson, A.P. Surveillance of antibiotic resistance. Philos. Trans. R. Soc. B Biol. Sci. 2015, 370, 20140080. [Google Scholar] [CrossRef]
- Founou, L.L.; Founou, R.C.; Essack, S.Y. Antibiotic resistance in the food chain: A developing country-perspective. Front. Microbiol. 2016, 7, 1–19. [Google Scholar] [CrossRef]
- European Centre for Disease Prevention and Control (ECDC); European Food Safety Authority (EFSA); European Medicines Agency (EMA). ECDC/EFSA/EMA second joint report on the integrated analysis of the consumption of antimicrobial agents and occurrence of antimicrobial resistance in bacteria from humans and food-producing animals Joint Interagency Antimicrobial Consumption and Resis. EFSA J. 2017, 15. [Google Scholar] [CrossRef]
- Adesiji, Y.O.; Shivakumaraswamy, S.K.; Deekshit, V.K.; Kallappa, G.S.; Karunasagar, I. Molecular characterization of antimicrobial multi-drug resistance in non-typhoidal Salmonellae from chicken and clam in Mangalore, India. J. Biomed. Res. 2018, 32, 237–244. [Google Scholar] [CrossRef]
- Mancin, M.; Barco, L.; Losasso, C.; Belluco, S.; Cibin, V.; Mazzucato, M.; Bilei, S.; Carullo, M.R.; Decastelli, L.; Di Giannatale, E.; et al. Salmonella serovar distribution from non-human sources in Italy; Results from the IT-Enter-Vet network. Vet. Rec. 2018, 183, 69. [Google Scholar] [CrossRef]
- Zhang, L.; Fu, Y.; Xiong, Z.; Ma, Y.; Wei, Y.; Qu, X.; Zhang, H.; Zhang, J.; Liao, M. Highly prevalent multidrug-resistant Salmonella from chicken and pork meat at retail markets in Guangdong, China. Front. Microbiol. 2018, 9, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Jajere, S.M. A review of Salmonella enterica with particular focus on the pathogenicity and virulence factors, host specificity and antimicrobial resistance including multidrug resistance. Vet. World 2019, 12, 504–521. [Google Scholar] [CrossRef] [PubMed]
- European Medicines Agency. Answer to the Request from the European Commission for Updating the Scientific Advice on the Impact on Public Health and Animal Health of the Use of Antibiotics in Animals-Categorisation of Antimicrobials Answer to the Request from the European Commissi; European Medicines Agency: London, UK, 2019; Volume 44, pp. 1–67. [Google Scholar]
- Kim, Y.J.; Chon, J.W.; Lim, J.S.; Song, B.R.; Seo, K.H.; Heo, E.J.; Park, H.J.; Wee, S.H.; Oh, D.H.; Moon, J.S. Traceback Investigation for Salmonella Contamination at Egg Processing Plants in South Korea: Prevalence, Antibiotic Resistance, and Epidemiological Tracing by Rep-PCR Fingerprinting. J. Food Sci. 2015, 80, M759–M764. [Google Scholar] [CrossRef] [PubMed]
- Bonardi, S.; Alpigiani, I.; Bruini, I.; Barilli, E.; Brindani, F.; Morganti, M.; Cavallini, P.; Bolzoni, L.; Pongolini, S. Detection of Salmonella enterica in pigs at slaughter and comparison with human isolates in Italy. Int. J. Food Microbiol. 2016, 218, 44–50. [Google Scholar] [CrossRef] [PubMed]
- Trevisani, M.; Diegoli, G.; Fedrizzi, G.C. Chemical Hazards and their Control. In Meat Inspection and Control in the Slaughterhouse, 1st ed.; Ninios, T., Lundén, J., Korkeala, H., Fredriksson-Ahomaa, M., Eds.; John Wiley & Sons, Ltd.: Oxford, UK, 2014; pp. 354–382. [Google Scholar]
- Proroga, Y.T.R.; Capuano, F.; Carullo, M.R.; Tela, I.L.; Capparelli, R.; Barco, L.; Pasquale, V. Occurrence and antimicrobial resistance of Salmonella strains from food of animal origin in southern Italy. Folia Microbiol. 2016, 21–27. [Google Scholar] [CrossRef]
- Silveira, L.; Pinto, M.; Isidro, J.; Pista, Â.; Themudo, P.; Vieira, L.; Machado, J.; Gomes, J.P. Multidrug-Resistant Salmonella enterica Serovar Rissen Clusters Detected in Azores Archipelago, Portugal. Int. J. Genom. 2019, 2019. [Google Scholar] [CrossRef]
- Pan, H.; Paudyal, N.; Li, X.; Fang, W.; Yue, M. Multiple food-animal-borne route in transmission of antibiotic-resistant salmonella newport to humans. Front. Microbiol. 2018, 9, 1–10. [Google Scholar] [CrossRef]
- Silveira, C.S.D.; Sousa, O.V.D.; Evangelista-Barreto, N.S. Propagation of antimicrobial resistant salmonella spp. in bivalve mollusks from estuary areas of Bahia, Brazil. Rev. Caatinga 2016, 29, 450–457. [Google Scholar] [CrossRef]
- Smaldone, G.; Marrone, R.; Cappiello, S.; Martin, G.A.; Oliva, G.; Cortesi, M.L.; Anastasio, A. Occurrence of antibiotic resistance in bacteria isolated from seawater organisms caught in Campania Region: Preliminary study. BMC Vet. Res. 2014, 10, 161. [Google Scholar] [CrossRef]
- Wolfensberger, A.; Kuster, S.P.; Marchesi, M.; Zbinden, R.; Hombach, M. The effect of varying multidrug-resistence (MDR) definitions on rates of MDR gram-negative rods. Antimicrob. Resist. Infect. Control 2019, 3, 1–9. [Google Scholar] [CrossRef]
- Proroga, Y.T.R.; Mancusi, A.; Peruzy, M.F.; Carullo, M.R.; Montone, A.M.I.; Fulgione, A.; Capuano, F. Characterization of Salmonella Typhimurium and its monophasic variant 1,4, [5],12:i:-isolated from different sources. Folia Microbiol. 2019, 64, 711–718. [Google Scholar] [CrossRef] [PubMed]

| Antibiotics | Poultry | Pork | MS | Bovine | Buffalo | Ovine | Snail | Vegetable | MMP |
|---|---|---|---|---|---|---|---|---|---|
| Ampicillin | 66a | 49ac | 21bd | 53ae | 20bce | 0ad | 33ad | 0bce | 0ad |
| Cefotaxime | 34a | 6b | 10b | 13ab | 0ab | 0ab | 0ab | 0ab | 0ab |
| Cephtazidime | 12a | 6ac | 2bc | 0ac | 0ac | 0ac | 0ac | 0ac | 0ac |
| Chloramphenicol | 14a | 9a | 12a | 0a | 0a | 0a | 0a | 0a | 0a |
| Colistin sulfate | 0a | 9a | 0a | 7a | 0a | 0a | 0a | 0a | 0a |
| Trimethoprim sulfamethoxazole | 77a | 17bc | 17bc | 27bc | 20bc | 0ac | 0ac | 0bc | 33ac |
| Ciprofloxacin | 80a | 23bc | 25bd | 47ace | 20bcd | 0acd | 0acd | 50bde | 100acd |
| Nalidixic acid | 89a | 14b | 10bc | 33a | 20ac | 0ac | 0ac | 0ac | 100ac |
| Gentamicin | 0a | 6a | 2a | 0a | 0a | 0a | 0a | 25a | 0a |
| Tetracycline | 86ad | 46bcd | 27bcd | 73ac | 0d | 0acd | 0acd | 0d | 67acd |
| Species and Subspecies | Serotype | 2015 | 2016 | 2017 | 2018 | 2019 | Total |
|---|---|---|---|---|---|---|---|
| S. enterica subsp. enterica | N.I.* | 1 | 1 | 2 | |||
| Infantis | 2 | 7 | 2 | 32 | 16 | 59 | |
| Derby | 5 | 1 | 2 | 9 | 1 | 18 | |
| Monophasic S. Typhimurium | 1 | 5 | 10 | 2 | 18 | ||
| Typhimurium | 3 | 1 | 1 | 4 | 2 | 11 | |
| Rissen | 3 | 1 | 1 | 2 | 1 | 8 | |
| Brandenburg | 1 | 2 | 2 | 5 | |||
| Anatum | 1 | 1 | 2 | 4 | |||
| Livingstone | 1 | 1 | 2 | 4 | |||
| Bredeney | 2 | 1 | 3 | ||||
| Enteritidis | 1 | 2 | 3 | ||||
| London | 3 | 3 | |||||
| Muenster | 1 | 1 | 1 | 3 | |||
| Nottingham | 1 | 2 | 3 | ||||
| Fischerhuette | 2 | 2 | |||||
| Give | 1 | 1 | 2 | ||||
| Muenchen | 1 | 1 | 2 | ||||
| Newport | 1 | 1 | 2 | ||||
| Stanleyville | 1 | 1 | 2 | ||||
| Agona | 1 | 1 | |||||
| Blockley | 1 | 1 | |||||
| Bovismorbificans | 1 | 1 | |||||
| Carno | 1 | 1 | |||||
| Eko | 1 | 1 | |||||
| Goldcoast | 1 | 1 | |||||
| Havana | 1 | 1 | |||||
| Hvittingfoss | 1 | 1 | |||||
| Kapemba | 1 | 1 | |||||
| Kasenyi | 1 | 1 | |||||
| Kentucky | 1 | 1 | |||||
| Litchfield | 1 | 1 | |||||
| Manchester | 1 | 1 | |||||
| Mbandaka | 1 | 1 | |||||
| Mishmarhaemek | 1 | 1 | |||||
| Panama | 1 | 1 | |||||
| Pomona | 1 | 1 | |||||
| Saintpaul | 1 | 1 | |||||
| Tennessee | 1 | 1 | |||||
| Winston | 1 | 1 | |||||
| Worthington | 1 | 1 | |||||
| S. enterica subsp. diarizonae | N.I.* | 2 | 1 | 3 | |||
| Total | 23 | 16 | 20 | 79 | 40 | 178 |
| Species and Subspecies | Serotype | Poultry | Bovine | Pork | Molluscan Shellfish | Buffalo | Ovine | Snail | Vegetable | Mixed Meat |
|---|---|---|---|---|---|---|---|---|---|---|
| S. enterica subsp. enterica | N.I.* | 1 | 1 | 1 | ||||||
| Infantis | 53 | 2 | 3 | |||||||
| monophasic S. Typhimurium | 2 | 5 | 8 | 3 | ||||||
| Newport | 2 | 1 | ||||||||
| Bredeney | 1 | 1 | 1 | |||||||
| Derby | 1 | 12 | 4 | |||||||
| Livingstone | 1 | 3 | ||||||||
| London | 1 | 2 | ||||||||
| Manchester | 1 | 1 | ||||||||
| Saintpaul | 1 | |||||||||
| Tennessee | 1 | 1 | ||||||||
| Agona | 1 | |||||||||
| Anatum | 4 | |||||||||
| Blockley | ||||||||||
| Bovismorbificans | 1 | |||||||||
| Brandenburg | 1 | 4 | 1 | |||||||
| Carno | 1 | 1 | ||||||||
| Eko | 1 | |||||||||
| Enteritidis | 2 | 1 | ||||||||
| Fischerhuette | 2 | |||||||||
| Give | 1 | 1 | ||||||||
| Goldcoast | 1 | |||||||||
| Havana | 1 | |||||||||
| Hvittingfoss | ||||||||||
| Kapemba | 1 | |||||||||
| Kasenyi | ||||||||||
| Species and Subspecies | Serotype | Poultry | Bovine | Pork | Molluscan shellfish | Buffalo | Ovine | Snail | Vegetable | Mixed Meat |
| S. enterica subsp. enterica | Kentucky | 1 | ||||||||
| Litchfield | 1 | |||||||||
| Mbandaka | 1 | |||||||||
| Mishmarhaemek | 1 | |||||||||
| Muenchen | 1 | 1 | 1 | |||||||
| Muenster | 1 | 2 | ||||||||
| Nottingham | 2 | |||||||||
| Panama | 1 | |||||||||
| Pomona | 1 | |||||||||
| Rissen | 1 | 4 | 3 | |||||||
| Stanleyville | 1 | 1 | ||||||||
| Typhimurium | 6 | 5 | ||||||||
| Winston | 1 | |||||||||
| Worthington | 1 | |||||||||
| S. enterica subsp. diarizonae | N.I.* | 2 | ||||||||
| Total | 64 | 15 | 35 | 48 | 5 | 1 | 3 | 4 | 3 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Peruzy, M.F.; Capuano, F.; Proroga, Y.T.R.; Cristiano, D.; Carullo, M.R.; Murru, N. Antimicrobial Susceptibility Testing for Salmonella Serovars Isolated from Food Samples: Five-Year Monitoring (2015–2019). Antibiotics 2020, 9, 365. https://doi.org/10.3390/antibiotics9070365
Peruzy MF, Capuano F, Proroga YTR, Cristiano D, Carullo MR, Murru N. Antimicrobial Susceptibility Testing for Salmonella Serovars Isolated from Food Samples: Five-Year Monitoring (2015–2019). Antibiotics. 2020; 9(7):365. https://doi.org/10.3390/antibiotics9070365
Chicago/Turabian StylePeruzy, Maria Francesca, Federico Capuano, Yolande Thérèse Rose Proroga, Daniela Cristiano, Maria Rosaria Carullo, and Nicoletta Murru. 2020. "Antimicrobial Susceptibility Testing for Salmonella Serovars Isolated from Food Samples: Five-Year Monitoring (2015–2019)" Antibiotics 9, no. 7: 365. https://doi.org/10.3390/antibiotics9070365
APA StylePeruzy, M. F., Capuano, F., Proroga, Y. T. R., Cristiano, D., Carullo, M. R., & Murru, N. (2020). Antimicrobial Susceptibility Testing for Salmonella Serovars Isolated from Food Samples: Five-Year Monitoring (2015–2019). Antibiotics, 9(7), 365. https://doi.org/10.3390/antibiotics9070365







