First Report of an Escherichia coli Strain Carrying the Colistin Resistance Determinant mcr-1 from a Dog in South Korea
Abstract
1. Introduction
2. Materials and Methods
2.1. Identification of Colistin-Resistant E. coli
2.2. Conjugation Assay
2.3. Molecular Characterization of mcr-1 Carrying E. coli
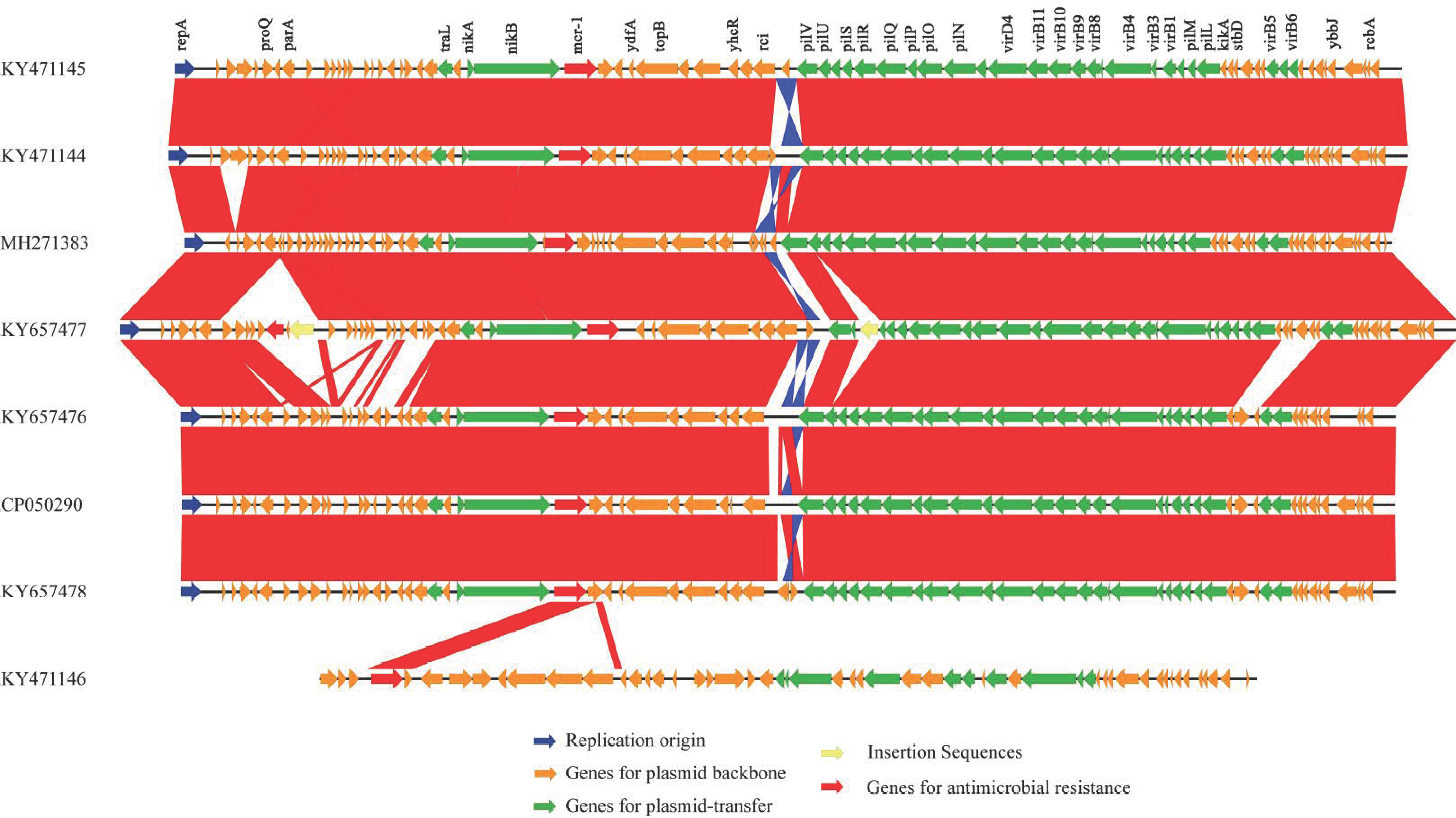
2.4. Comparative Analysis of mcr-1 Carrying Plasmids
3. Results and Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Liu, Y.Y.; Wang, Y.; Walsh, T.R.; Yi, L.X.; Zhang, R.; Spencer, J.; Doi, Y.; Tian, G.; Dong, B.; Huang, X.; et al. Emergence of plasmid-mediated colistin resistance mechanism mcr-1 in animals and human beings in China: A microbiological and molecular biological study. Lancet Infect. Dis. 2016, 16, 161–168. [Google Scholar] [CrossRef]
- Wang, C.; Feng, Y.; Sokerya, S.; Liu, L.; Wei, L.; Kang, M.; Zhong, Z. Identification of novel mobile colistin resistance gene mcr-10. Emerg. Microbes. Infect. 2020, 9, 508–516. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Zhang, H.; Liu, Y.H.; Feng, Y. Towards understanding mcr-like colistin resistance. Trends Microbiol. 2018, 26, 794–808. [Google Scholar] [CrossRef] [PubMed]
- Gao, R.; Hu, Y.; Li, Z.; Sun, J.; Wang, Q.; Lin, J.; Ye, H.; Liu, F.; Srinivas, S.; Li, D.; et al. Dissemination and mechanism for the mcr-1 colistin resistance. PLoS Pathog. 2016, 12, e1005957. [Google Scholar] [CrossRef]
- Poirel, L.; Nordmann, P. Emerging plasmid-encoded colistin resistance: The animal world as the culprit? J. Antimicrob. Chemother. 2016, 8, 2626–2627. [Google Scholar] [CrossRef] [PubMed]
- Doumith, M.; Godbole, G.; Ashton, P.; Larkin, L.; Dallman, T.; Day, M.; Day, M.; Muller-Peboy, B.; Ellington, M.J.; de Pinna, E.; et al. Detection of the plasmid-mediated mcr-1 gene conferring colistin resistance in human and food isolates of Salmonella enterica and Escherichia coli in England and Wales. J. Antimicrob. Chemother. 2016, 71, 2300–2305. [Google Scholar] [CrossRef] [PubMed]
- Stoesser, N.; Mathers, A.J.; Moore, C.E.; Nicholas, P.J.D.; Crook, D.W. Colistin resistance gene mcr-1 and pHNSHP45 plasmid in human isolates of Escherichia coli and Klebsiella pneumoniae. Lancet Infect. Dis. 2016, 16, 285–286. [Google Scholar] [CrossRef]
- Anjum, M.F.; Duggett, N.A.; AbuOun, M.; Randall, L.; Nunez-Gracia, J.; Ellis, R.J.; Rogers, J.; Horton, R.; Brena, C.; Williamson, S.; et al. Colistin resistance in Salmonella and Escherichia coli isolates from a pig farm in Great Britain. J. Antimicrob. Chemother. 2016, 71, 2306–2313. [Google Scholar] [CrossRef]
- Zhang, X.F.; Doi, Y.; Huang, X.; Li, H.Y.; Zhong, L.L.; Zeng, K.J.; Zhang, Y.F.; Patil, S.; Tian, G.B. Possible transmission of mcr-1-harboring Escherichia coli between companion animals and human. Emerg. Infect. Dis. 2016, 22, 1679–1681. [Google Scholar] [CrossRef]
- Lei, L.; Wang, Y.; Schwarz, S.; Walsh, T.R.; Ou, Y.; Wu, Y.; Li, M.; Shen, Z. mcr-1 in Enterobacteriaceae from companion animals, Beijing, China, 2012–2016. Emerg. Infect. Dis. 2017, 23, 710–711. [Google Scholar] [CrossRef]
- Ortega-Paredes, D.; Haro, M.; Leoro-Garzón, P.; Barba, P.; Loaiza, K.; Mora, F.; Fors, M.; Vinueza-Burgos, C.; Fernández-Moreira, E. Multidrug-resistant Escherichia coli isolated from canine feces in a public park in Quito, Ecuador. J. Glob. Antimicrob. Resist. 2019, 18, 263–268. [Google Scholar] [CrossRef] [PubMed]
- Lim, S.K.; Kang, H.Y.; Lee, K.; Moon, D.C.; Lee, H.S.; Jung, S.C. First detection of the mcr-1 gene in Escherichia coli isolated from livestock between 2013 and 2015 in South Korea. Antimicrob. Agents Chemother. 2016, 60, 6991–6993. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Belaynehe, K.M.; Shin, S.W.; Park, K.Y.; Jang, J.Y.; Won, H.G.; Yoon, I.J.; Yoo, H.S. Emergence of mcr-1 and mcr-3 variants coding for plasmid-mediated colistin resistance in Escherichia coli isolates from food-producing animals in South Korea. Int. J. Infect. Dis. 2018, 72, 22–24. [Google Scholar] [CrossRef] [PubMed]
- Oh, S.S.; Song, J.; Kim, J.; Shin, J. Increasing prevalence of multidrug-resistant mcr-1-positive Escherichia coli isolates from fresh vegetables and healthy food animals in South Korea. Int. J. Infect. Dis. 2020, 92, 53–55. [Google Scholar] [CrossRef]
- Yoon, E.J.; Hong, J.S.; Yang, J.W.; Lee, K.J.; Lee, H.; Jeong, S.H. Detection of mcr-1 plasmids in Enterobacteriaceae isolates from human specimens: Comparisons with those of human isolates from livestock in Korea. Ann. Lab. Med. 2018, 38, 555–562. [Google Scholar] [CrossRef]
- Clinical and Laboratory Standards Institute (CLSI). Performance Standards for Antimicrobial Susceptibility Testing; Twentieth Informational Supplement; CLSI Document M100: Wayne, PA, USA, 2018. [Google Scholar]
- European Committee on Antimicrobial Susceptibility Testing. Breakpoint Tables for Interpretation of MICs and Zone Diameters. EUCAST, Version 8.1. 2018. Available online: http://www.eucast.org (accessed on 17 July 2020).
- Tendon, V.D.; Poirel, L.; Nordman, P. Transferability of the mcr-1 colistin resistance gene. Antimicrob. Resist. Infect. Control. 2017, 23, 813–815. [Google Scholar]
- Gautom, R.K. Rapid pulse-field gel electrophoresis protocol for typing of Escherichia coli O157:H7 and other gram-negative organisms in day 1. J. Clin. Microbiol. 1997, 35, 2977–2980. [Google Scholar] [CrossRef]
- Escherichia coli MLST Database. Available online: http://enterobase.warwick.ac.uk/species/ecoli/allele_st_search (accessed on 15 August 2020).
- Altermann, E.; Lu, J.; McCulloch, A. GAMOLA2, a comprehensive software package for the annotation and curation of draft and complete microbial genomes. Front. Microbiol. 2017, 8, 346. [Google Scholar] [CrossRef][Green Version]
- Pritchard, L.; Glover, R.H.; Humphris, S.; Elphinstone, J.G.; Toth, I.K. Genomics and taxonomy in diagnostics for food security: Soft-rotting enterobacterial plant pathogens. Anal. Methods. 2016, 8, 12–24. [Google Scholar] [CrossRef]
- Camacho, C.; Coulouris, G.; Avagyan, V.; Ma, N.; Papadopolus, J.; Bealer, K.; Madden, T.L. BLAST+: Architecture and applications. BMC Bioinform. 2009, 10, 421. [Google Scholar] [CrossRef]
- Sullivan, M.J.; Petty, N.K.; Beatson, S.A. Easyfig: A genome comparison visualizer. Bioinformatics 2011, 27, 1009–1010. [Google Scholar] [CrossRef]
- Rumi, M.V.; Mas, J.; Elena, A.; Cerdeira, L.; Muñoz, M.E.; Lincopan, N.; Gentilini, É.R.; di Conza, J.; Gutkind, G. Co-occurrence of clinically relevant β-lactamases and mcr-1 encoding genes in Escherichia coli from companion animals in Argentina. Vet. Microbiol. 2019, 230, 228–234. [Google Scholar] [CrossRef]
- Lee, J.Y.; Lim, S.K.; Moon, Y.C.; Shin, J.; Ko, K.S. Whole sequences and characteristics of mcr-1- harboring plasmids of Escherichia coli strains isolated from livestock in South Korea. Microb. Drug. Resist. 2018, 24, 489–492. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Liu, Z.; Zhang, Y.; Zhang, Z.; Lei, L.; Xia, Z. Increasing prevalence of ESBL-producing multidrug resistance Escherichia coli from diseased pets in Beijing, China From 2012 to 2017. Front. Microbiol. 2019, 10, 2852. [Google Scholar] [CrossRef]
- Wang, J.; Huang, X.Y.; Xia, Y.B., Guo; Guo, Z.W.; Ma, Z.B.; Yi, M.Y.; Lv, L.C.; Lu, P.L.; Yan, J.C.; Huang, J.W.; et al. Clonal spread of Escherichia coli ST93 carrying mcr-1-harboring IncN1-IncHI2/ST3 plasmid among companion animals, China. Front. Microbiol. 2018, 9, 2989. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Wang, J.; Bai, X.; Ma, J.; Xiong, Y.; Fanning, S.; Bai, L.; Yang, Z. Characterization of five Escherichia coli isolates co-expressing ESBL and mcr-1 resistance mechanisms from different origins in China. Front. Microbiol. 2019, 10, 1994. [Google Scholar] [CrossRef]
- Zhuge, X.; Jiang, M.; Tang, F.; Sun, Y.; Ji, Y.; Xue, F.; Ren, J.; Zhu, W.; Dai, J. Avian-source mcr-1-positive Escherichia coli is phylogenetically diverse and shares virulence characteristics with E. coli causing human extra-intestinal infections. Vet. Microbiol. 2019, 239, 108483. [Google Scholar] [CrossRef]
- Wong, M.H.; Liu, L.; Yan, M.; Chan, E.W.; Chen, S. Dissemination of IncI2 plasmids that harbor the blaCTX-M element among clinical Salmonella isolates. Antimicrob. Agents Chemother. 2015, 59, 5026. [Google Scholar] [CrossRef]
- Lv, L.; Partridge, S.R.; He, L.; Zeng, Z.; He, D.; Ye, J.; Liu, J.H. Genetic characterization of IncI2 plasmids carrying blaCTX-M-55 spreading in both pets and food animals in China. Antimicrob. Agents Chemother. 2013, 57, 2824–2827. [Google Scholar] [CrossRef]
- Han, H.; Liu, W.; Cui, X.; Cheng, X.; Jiang, X. Co-Existence of mcr-1 and blaNDM-5 in an Escherichia coli strain isolated from the pharmaceutical industry, WWTP. Infect. Drug Resist. 2020, 13, 851–854. [Google Scholar] [CrossRef] [PubMed]
- Islam, A.; Rahman, Z.; Monira, S.; Rahman, M.A.; Camilli, A.; George, C.M.; Ahmed, N.; Alam, M. Colistin-resistant Escherichia coli carrying mcr-1 in urban sludge samples: Dhaka, Bangladesh. Gut Pathog. 2017, 9, 77. [Google Scholar] [CrossRef]
- Sun, P.; Bi, Z.; Nilsson, M.; Zheng, B.; Berglund, B.; Lundborg, C.S.; Borjesson, S.; Li, X.; Chen, B.; Yin, H.; et al. Occurrence and genomic characterization of ESBL-producing, mcr-1-harboring Escherichia coli in farming soil. Antimicrob. Agents Chemother. 2017, 61, e02569-16. [Google Scholar] [CrossRef] [PubMed]
- Guenther, S.; Falgenhauer, L.; Semmler, T.; Imirzalioglu, C.; Chakraborty, T.; Roesler, U.; Roschanski, N. Environmental emission of multiresistant Escherichia coli carrying the colistin resistance gene mcr-1 from German swine farms. J. Antimicrob. Chemother. 2017, 72, 1289–1292. [Google Scholar]


| Plasmids | GenBank Accession Number | Bacterial Species | Host | Country | Reference |
|---|---|---|---|---|---|
| pV80 | MH271383 | E. coli | Dog | Argentina | [25] |
| pEC006 | KY471144 | E. coli | Chicken | Korea | [26] |
| pEC019 | KY471145 | E. coli | Chicken | Korea | [26] |
| pEC111 | KY471146 | E. coli | Pig | Korea | [26] |
| pCREC-527-4 | KY657476 | E. coli | Human | Korea | [15] |
| pCRENT-301-1 | KY657477 | Enterobacter aerogenes | Human | Korea | [15] |
| pUSU-ECO-12704-4 | KY657478 | E. coli | Human | Korea | [15] |
| pK19EC149 | CP050290 | E. coli | Dog | Korea | This study |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Moon, D.C.; Mechesso, A.F.; Kang, H.Y.; Kim, S.-J.; Choi, J.-H.; Kim, M.H.; Song, H.-J.; Yoon, S.-S.; Lim, S.-K. First Report of an Escherichia coli Strain Carrying the Colistin Resistance Determinant mcr-1 from a Dog in South Korea. Antibiotics 2020, 9, 768. https://doi.org/10.3390/antibiotics9110768
Moon DC, Mechesso AF, Kang HY, Kim S-J, Choi J-H, Kim MH, Song H-J, Yoon S-S, Lim S-K. First Report of an Escherichia coli Strain Carrying the Colistin Resistance Determinant mcr-1 from a Dog in South Korea. Antibiotics. 2020; 9(11):768. https://doi.org/10.3390/antibiotics9110768
Chicago/Turabian StyleMoon, Dong Chan, Abraham Fikru Mechesso, Hee Young Kang, Su-Jeong Kim, Ji-Hyun Choi, Mi Hyun Kim, Hyun-Ju Song, Soon-Seek Yoon, and Suk-Kyung Lim. 2020. "First Report of an Escherichia coli Strain Carrying the Colistin Resistance Determinant mcr-1 from a Dog in South Korea" Antibiotics 9, no. 11: 768. https://doi.org/10.3390/antibiotics9110768
APA StyleMoon, D. C., Mechesso, A. F., Kang, H. Y., Kim, S.-J., Choi, J.-H., Kim, M. H., Song, H.-J., Yoon, S.-S., & Lim, S.-K. (2020). First Report of an Escherichia coli Strain Carrying the Colistin Resistance Determinant mcr-1 from a Dog in South Korea. Antibiotics, 9(11), 768. https://doi.org/10.3390/antibiotics9110768






