New Antimicrobial Bioactivity against Multidrug-Resistant Gram-Positive Bacteria of Kinase Inhibitor IMD0354
Abstract
1. Introduction
2. Results
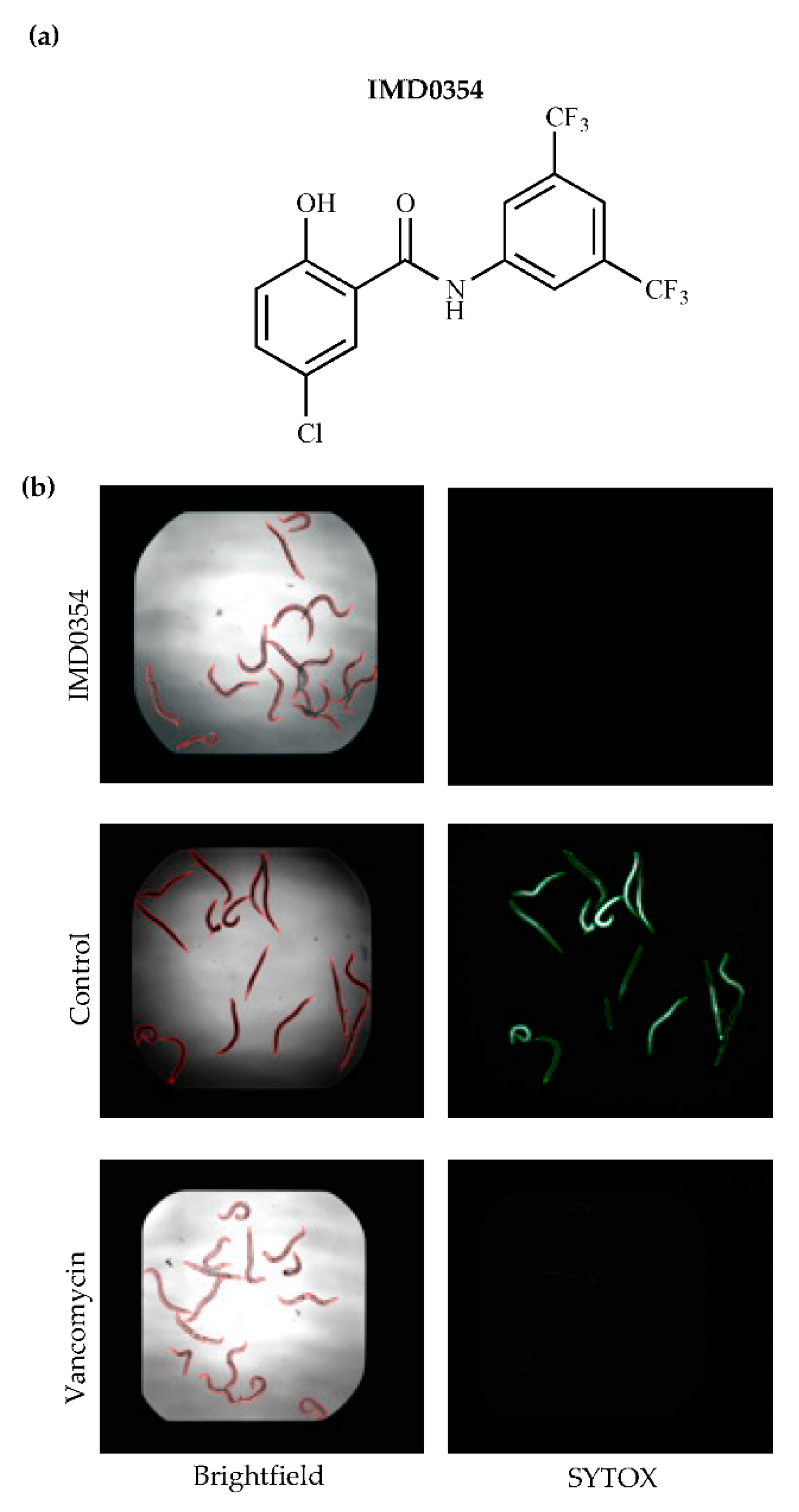
2.1. IMD0354 Exhibits Anti-Staphylococcal Activity In Vitro & in a Whole Animal C. elegans Infection Model
2.2. IMD0354 Shows Antimicrobial Activity against Multidrug-Resistant Gram-Positive Pathogens
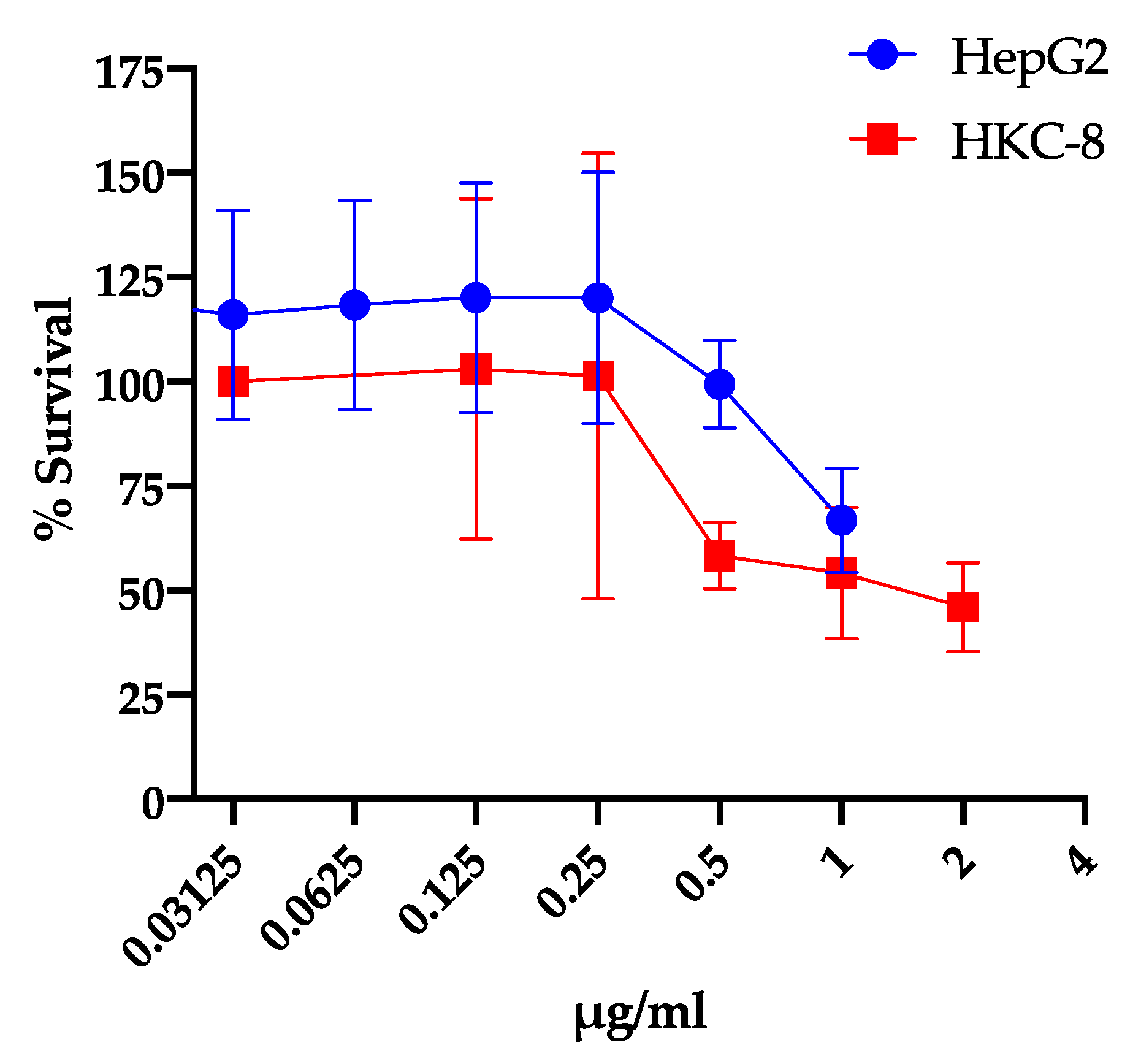
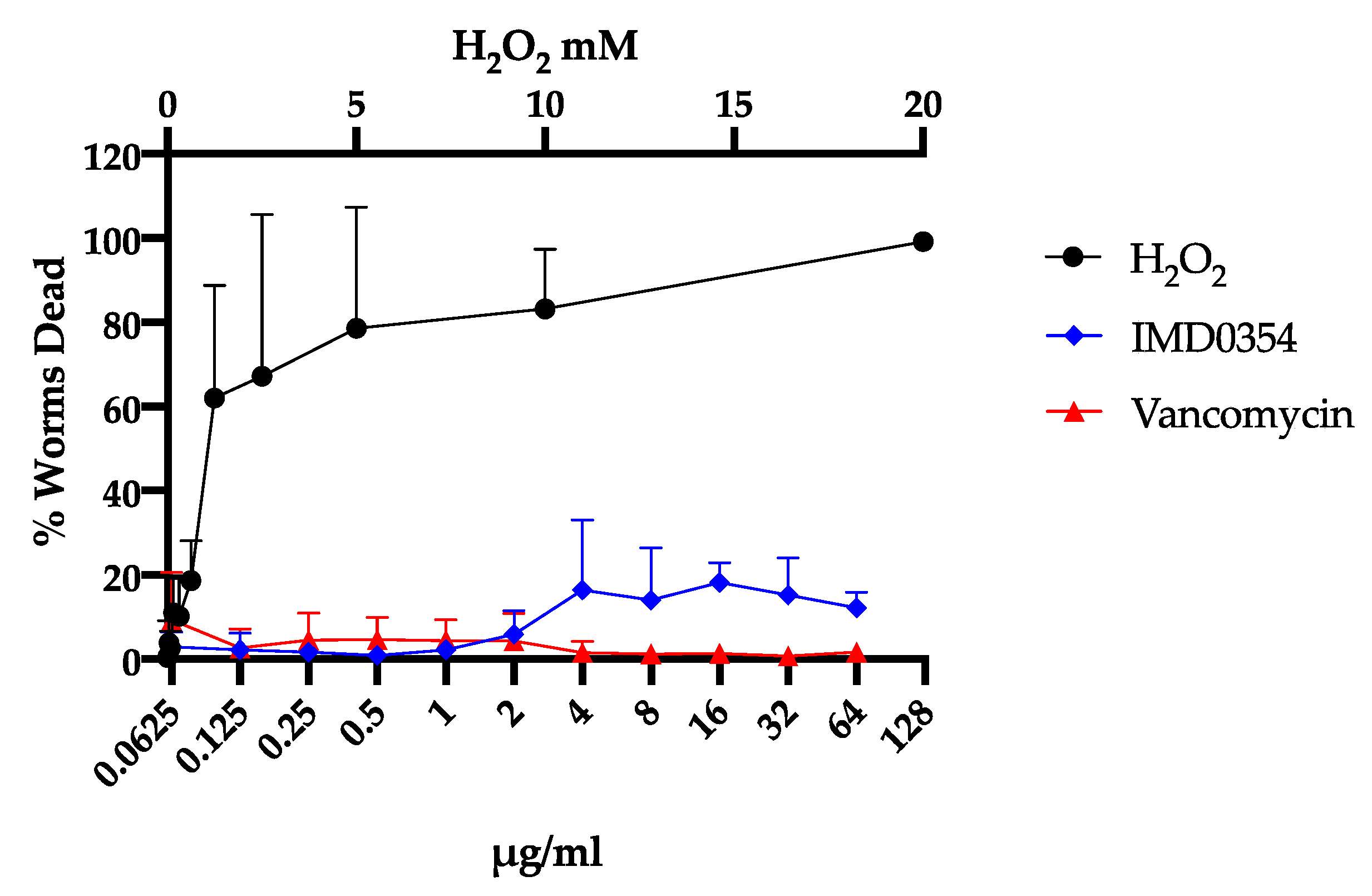
2.3. Cytotoxicity of IMD0354 on Human Cells and C. elegans
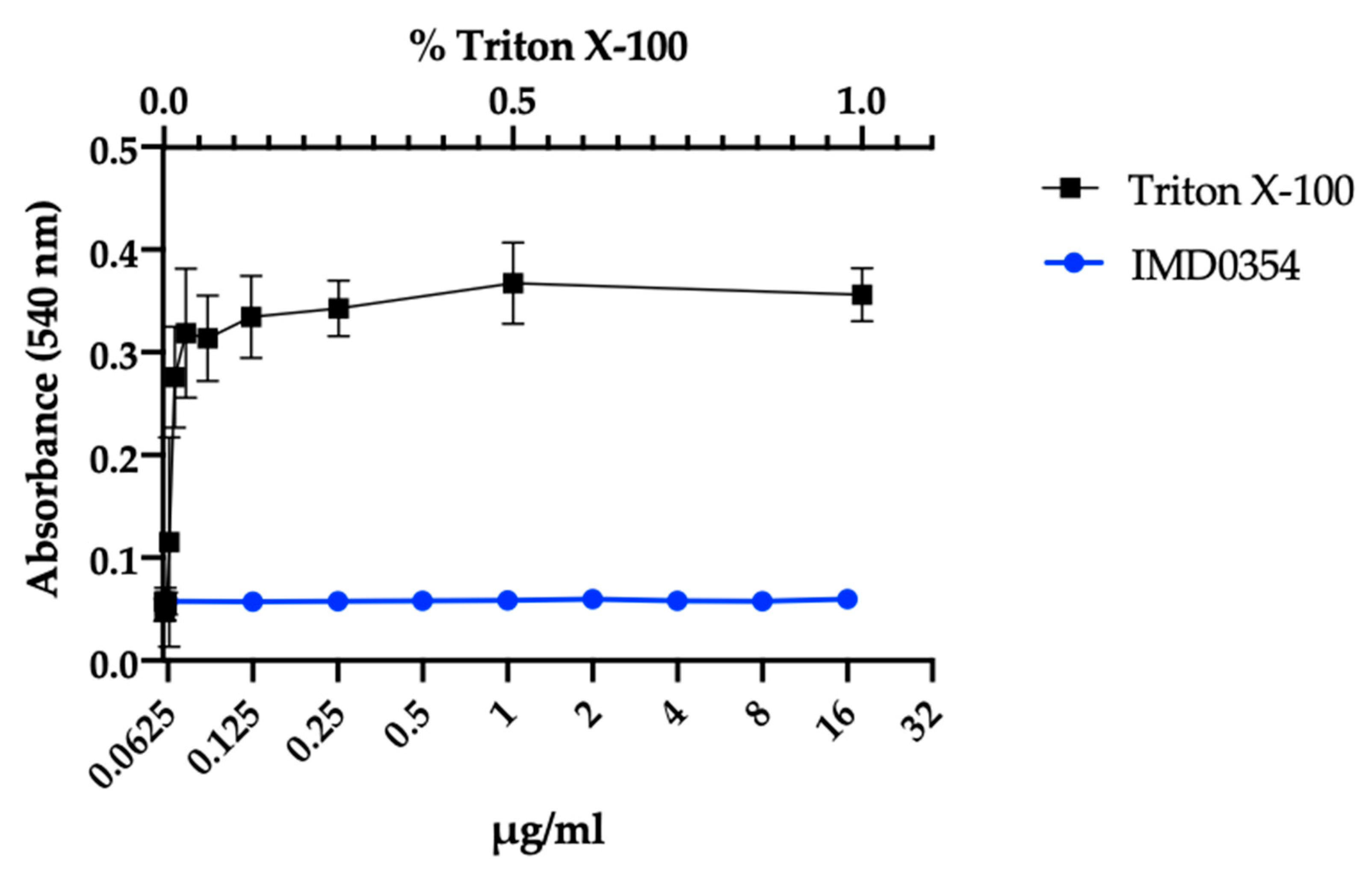
2.4. IMD0354 Shows Bacterial Membrane Permeability at High Concentrations
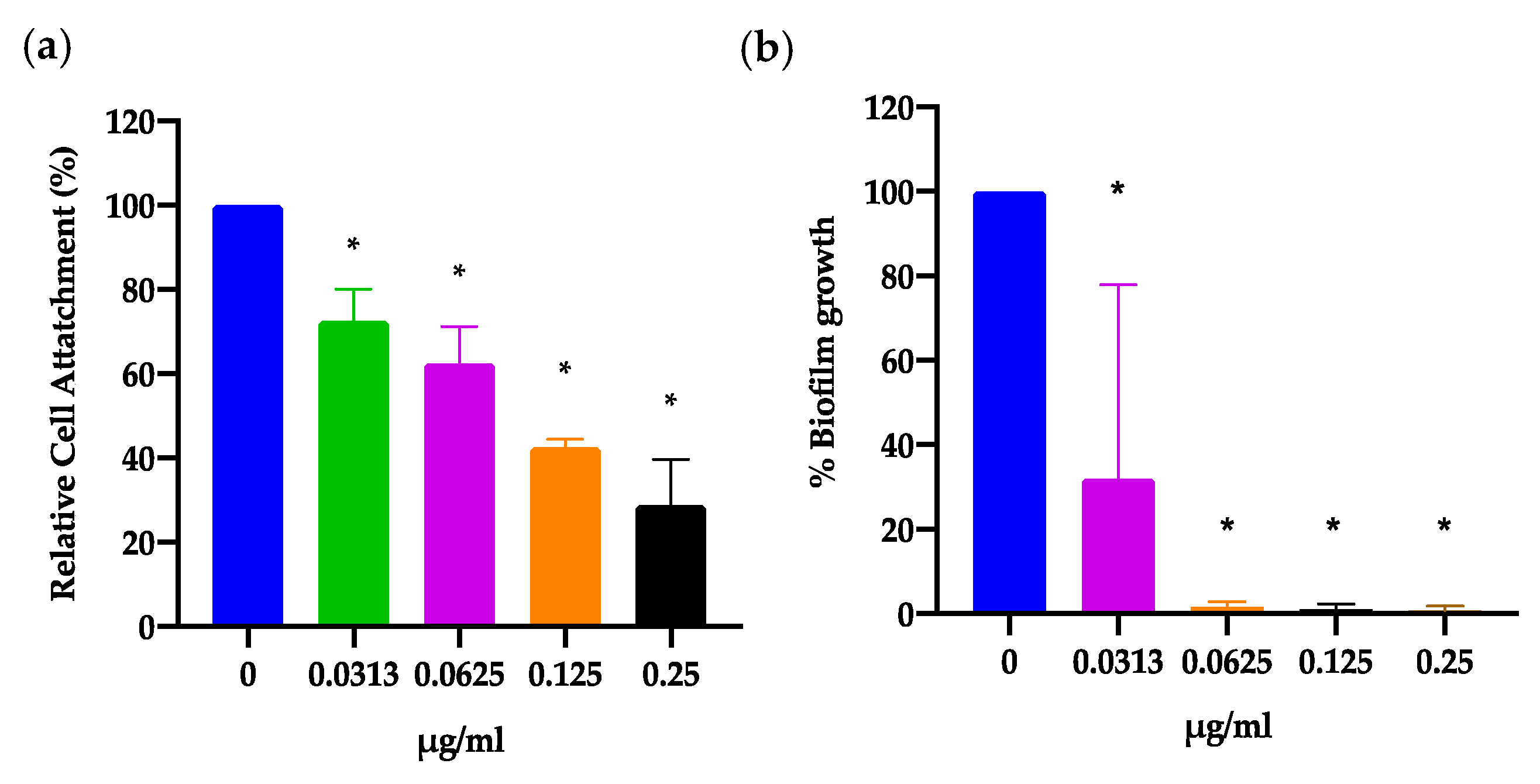
2.5. IMD0354 Inhibits Initial Cell Attachment in a Dose-Dependent Manner and Fully Inhibtis Biofilm Formation
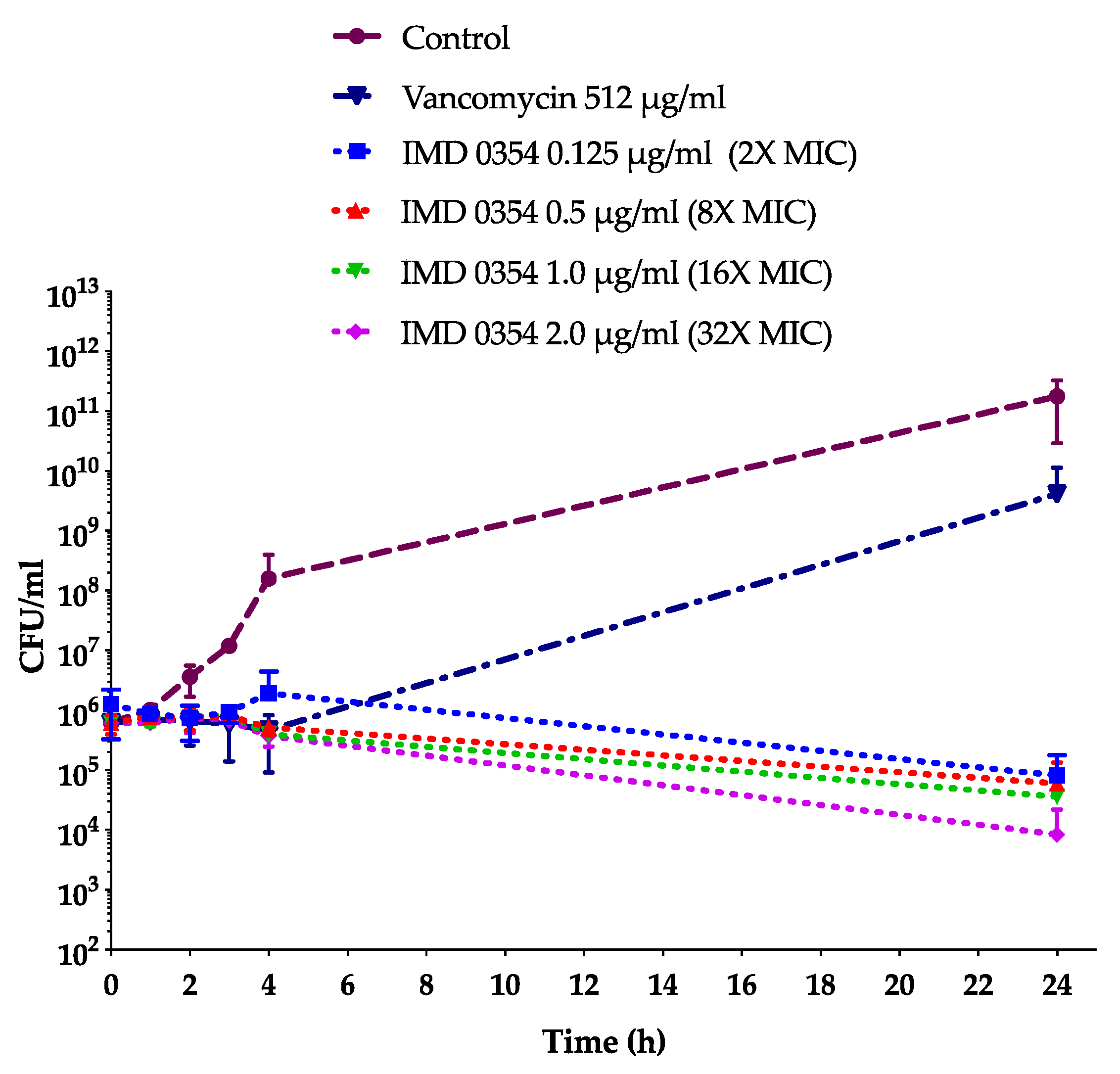
2.6. IMD0354 Does Not Affect the Viability of Antibiotic-Tolerant Cells nor Synergize with Conventional Antibiotics
3. Discussion
4. Materials and Methods
4.1. Bacterial Strains and Growth Conditions
4.2. Drugs and Antibiotics
4.3. Minimum Inhibitory Concentration Assay
4.4. C. elegans Infection Assay for Compound Screening
4.5. C. elegans Dose-Dependent Toxicity Assay
4.6. Bacterial Time-Course Killing Assay
4.7. Human Blood Hemolysis
4.8. Antibiotic-Tolerant Cell Killing
4.9. SYTOX Green Membrane Permeability Assay
4.10. Biofilm Initial Cell Attachment Assay
4.11. Biofilm Inhibition Assay
4.12. Mammalian Cancer Cell Viability Assay
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- David, M.Z.; Daum, R.S. Community-Associated Methicillin-Resistant Staphylococcus aureus: Epidemiology and Clinical Consequences of an Emerging Epidemic. Clin. Microbiol. Rev. 2010, 23, 616–687. [Google Scholar] [CrossRef] [PubMed]
- Lowy, F.D. Staphylococcus aureus infections. N. Engl. J. Med. 1998, 339, 520–532. [Google Scholar] [CrossRef] [PubMed]
- Lakhundi, S.; Zhang, K. Methicillin-Resistant Staphylococcus aureus: Molecular Characterization, Evolution, and Epidemiology. Clin. Microbiol. Rev. 2018, 31. [Google Scholar] [CrossRef]
- Stogios, P.J.; Savchenko, A. Molecular mechanisms of vancomycin resistance. Protein Sci. 2020, 29, 654–669. [Google Scholar] [CrossRef] [PubMed]
- Hiramatsu, K.; Aritaka, N.; Hanaki, H.; Kawasaki, S.; Hosoda, Y.; Hori, S.; Fukuchi, Y.; Kobayashi, I. Dissemination in Japanese hospitals of strains of Staphylococcus aureus heterogeneously resistant to vancomycin. Lancet 1997, 350, 1670–1673. [Google Scholar] [CrossRef]
- Howe, R.A.; Bowker, K.E.; Walsh, T.R.; Feest, T.G.; MacGowan, A.P. Vancomycin-resistant Staphylococcus aureus. Lancet 1998, 351, 602. [Google Scholar] [CrossRef]
- Hidayat, L.K.; Hsu, D.I.; Quist, R.; Shriner, K.A.; Wong-Beringer, A. High-dose vancomycin therapy for methicillin-resistant Staphylococcus aureus infections: Efficacy and toxicity. Arch. Intern. Med. 2138, 166, 2006–2144. [Google Scholar] [CrossRef]
- Leclercq, R.; Derlot, E.; Duval, J.; Courvalin, P. Plasmid-mediated resistance to vancomycin and teicoplanin in Enterococcus faecium. N. Engl. J. Med. 1988, 319, 157–161. [Google Scholar] [CrossRef]
- Uttley, A.H.; Collins, C.H.; Naidoo, J.; George, R.C. Vancomycin-resistant enterococci. Lancet 1988, 2, 57–58. [Google Scholar] [CrossRef]
- Dougherty, T.J.; Pucci, M.J. Antibiotic Discovery and Development; Springer Science & Business Media: Boston, MA, USA, 2011. [Google Scholar]
- Munita, J.M.; Arias, C.A. Mechanisms of Antibiotic Resistance. Microbiol. Spectr. 2016, 4, 481–511. [Google Scholar] [CrossRef]
- Arthur, M.; Molinas, C.; Courvalin, P. The VanS-VanR two-component regulatory system controls synthesis of depsipeptide peptidoglycan precursors in Enterococcus faecium BM4147. J. Bacteriol. 1992, 174, 2582–2591. [Google Scholar] [CrossRef]
- Meziane-Cherif, D.; Saul, F.A.; Haouz, A.; Courvalin, P. Structural and functional characterization of VanG D-Ala:D-Ser ligase associated with vancomycin resistance in Enterococcus faecalis. J. Biol. Chem. 2012, 287, 37583–37592. [Google Scholar] [CrossRef] [PubMed]
- Tattevin, P.; Arvieux, C.; Michelet, C. Alternative agents for the treatment of invasive infections due to methicillin-resistant Staphylococcus aureus strains with reduced susceptibility to vancomycin. Arch. Intern. Med. 2007, 167, 1206. [Google Scholar] [CrossRef] [PubMed]
- Gomes, D.M.; Ward, K.E.; LaPlante, K.L. Clinical implications of vancomycin heteroresistant and intermediately susceptible Staphylococcus aureus. Pharmacotherapy 2015, 35, 424–432. [Google Scholar] [CrossRef] [PubMed]
- Rybak, M.J.; Hershberger, E.; Moldovan, T.; Grucz, R.G. In vitro activities of daptomycin, vancomycin, linezolid, and quinupristin-dalfopristin against Staphylococci and Enterococci, including vancomycin- intermediate and -resistant strains. Antimicrob. Agents Chemother. 2000, 44, 1062–1066. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.-J.; Huang, Y.-C.; Shie, S.-S. Evolution of Multi-Resistance to Vancomycin, Daptomycin, and Linezolid in Methicillin-Resistant Staphylococcus aureus Causing Persistent Bacteremia. Front. Microbiol. 2020, 11, 1414. [Google Scholar] [CrossRef]
- Nannini, E.; Murray, B.E.; Arias, C.A. Resistance or decreased susceptibility to glycopeptides, daptomycin, and linezolid in methicillin-resistant Staphylococcus aureus. Curr. Opin. Pharmacol. 2010, 10, 516–521. [Google Scholar] [CrossRef]
- Kim, W.; Zhu, W.; Hendricks, G.L.; Van Tyne, D.; Steele, A.D.; Keohane, C.E.; Fricke, N.; Conery, A.L.; Shen, S.; Pan, W.; et al. A new class of synthetic retinoid antibiotics effective against bacterial persisters. Nature 2018, 556, 103–107. [Google Scholar] [CrossRef]
- Kim, S.M.; Escorbar, I.; Lee, K.; Fuchs, B.B.; Mylonakis, E.; Kim, W. Anti-MRSA agent discovery using Caenorhabditis elegans-based high-throughput screening. J. Microbiol. 2020, 58, 431–444. [Google Scholar] [CrossRef]
- Kim, W.; Steele, A.D.; Zhu, W.; Csatary, E.E.; Fricke, N.; Dekarske, M.M.; Jayamani, E.; Pan, W.; Kwon, B.; Sinitsa, I.F.; et al. Discovery and Optimization of nTZDpa as an Antibiotic Effective against Bacterial Persisters. ACS Infect. Dis. 2018, 4, 1540–1545. [Google Scholar] [CrossRef]
- Kim, W.; Zou, G.; Hari, T.P.A.; Wilt, I.K.; Zhu, W.; Galle, N.; Faizi, H.A.; Hendricks, G.L.; Tori, K.; Pan, W.; et al. A selective membrane-targeting repurposed antibiotic with activity against persistent methicillin-resistant Staphylococcus aureus. Proc. Natl. Acad. Sci. USA 2019, 116, 16529–16534. [Google Scholar] [CrossRef] [PubMed]
- Kim, W.; Zou, G.; Pan, W.; Fricke, N.; Faizi, H.A.; Kim, S.M.; Khader, R.; Li, S.; Lee, K.; Escorba, L.; et al. The Neutrally Charged Diarylurea Compound PQ401 Kills Antibiotic-Resistant and Antibiotic-Tolerant Staphylococcus aureus. mBio 2020, 11, 603. [Google Scholar] [CrossRef] [PubMed]
- Onai, Y.; Suzuki, J.-I.; Kakuta, T.; Maejima, Y.; Haraguchi, G.; Fukasawa, H.; Muto, S.; Itai, A.; Isobe, M. Author NotesInhibition of IkappaB phosphorylation in cardiomyocytes attenuates myocardial ischemia/reperfusion injury. Cardiovasc. Res. 2004, 63, 51–59. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Ko, D.; Lee, Y.; Jang, S.; Lee, Y.; Lee, I.Y.; Kim, S. Anti-cancer activity of the novel 2-hydroxydiarylamide derivatives IMD-0354 and KRT1853 through suppression of cancer cell invasion, proliferation, and survival mediated by TMPRSS4. Sci. Rep. 2019, 9, 10003. [Google Scholar] [CrossRef]
- Kinose, Y.; Sawada, K.; Makino, H.; Tomonori, O.; Tomoko, M.; Noriko, S.; Tomoyuki, F.; Eiichi, M.; Koji, N.; Ikuko, S.; et al. IKKβ Regulates VEGF Expression and Is a Potential Therapeutic Target for Ovarian Cancer as an Antiangiogenic Treatment. Mol. Cancer Ther. 2015, 14, 909–919. [Google Scholar] [CrossRef] [PubMed]
- Gomez-Cabrero, A.; Wrasidlo, W.; Reisfeld, R.A. IMD-0354 targets breast cancer stem cells: A novel approach for an adjuvant to chemotherapy to prevent multidrug resistance in a murine model. PLoS ONE 2013, 8, e73607. [Google Scholar] [CrossRef][Green Version]
- Sugita, A.; Ogawa, H.; Azuma, M.; Muto, S.; Honjo, A.; Yanagawa, H.; Nishioka, Y.; Tani, K.; Itai, A.; Sone, S. Antiallergic and anti-inflammatory effects of a novel I kappaB kinase beta inhibitor, IMD-0354, in a mouse model of allergic inflammation. Int. Arch. Allergy Immunol. 2009, 148, 186–198. [Google Scholar] [CrossRef]
- Barker, W.T.; Nemeth, A.M.; Brackett, S.M.; Basak, A.K.; Chandler, C.E.; Jania, L.A.; Zuercher, W.J.; Melander, R.J.; Koller, B.H.; Ernst, R.K.; et al. Repurposing Eukaryotic Kinase Inhibitors as Colistin Adjuvants in Gram-Negative Bacteria. ACS Infect. Dis. 2019, 5, 1764–1771. [Google Scholar] [CrossRef]
- Rice, L.B. Federal funding for the study of antimicrobial resistance in nosocomial pathogens: No ESKAPE. J. Infect. Dis. 2008, 197, 1079–1081. [Google Scholar] [CrossRef]
- Pendleton, J.N.; Gorman, S.P.; Gilmore, B.F. Clinical relevance of the ESKAPE pathogens. Expert Rev. Anti-Infect. Ther. 2013, 11, 297–308. [Google Scholar] [CrossRef]
- Weigel, L.M.; Clewell, D.B.; Gill, S.R.; Clark, N.C.; McDougal, L.K.; Flannagan, S.E.; Kolonay, J.F.; Shetty, J.; Killgore, G.E.; Tenover, F.C. Genetic analysis of a high-level vancomycin-resistant isolate of Staphylococcus aureus. Science 2003, 302, 1569–1571. [Google Scholar] [CrossRef] [PubMed]
- Centers for Disease Control and Prevention. CDC & FDA Antibiotic Resistance (AR) Isolate Bank; CDC: Atlanta, GA, USA, 2019.
- Gupta, V.; Singla, N.; Behl, P.; Sahoo, T.; Chander, J. Antimicrobial susceptibility pattern of vancomycin resistant enterococci to newer antimicrobial agents. Indian J. Med. Res. 2015, 141, 483–486. [Google Scholar] [CrossRef] [PubMed]
- Carias, L.L.; Rudin, S.D.; Donskey, C.J.; Rice, L.B. Genetic linkage and cotransfer of a novel, vanB-containing transposon Tn5382) and a low-affinity penicillin-binding protein 5 gene in a clinical vancomycin-resistant Enterococcus faecium isolate. J. Bacteriol. 1998, 180, 4426–4434. [Google Scholar] [CrossRef] [PubMed]
- Williamson, R.; Al-Obeid, S.; Shlaes, J.H.; Goldstein, F.W.; Shlaes, D.M. Inducible resistance to vancomycin in Enterococcus faecium D366. J. Infect. Dis. 1989, 159, 1095–1104. [Google Scholar] [CrossRef]
- Thorisdottir, A.S.; Carias, L.L.; Marshall, S.H.; Green, M.; Zervos, M.J.; Giorgio, C.; Mermel, L.A.; Boyce, J.M.; Medeiros, A.A.; Fraimowet, H.; et al. IS6770, an enterococcal insertion-like sequence useful for determining the clonal relationship of clinical enterococcal isolates. J. Infect. Dis. 1994, 70, 1539–1548. [Google Scholar] [CrossRef]
- Evers, S.; Sahm, D.F.; Courvalin, P. The vanB gene of vancomycin-resistant Enterococcus faecalis V583 is structurally related to genes encoding D-Ala:D-Ala ligases and glycopeptide-resistance proteins VanA and VanC. Gene 1993, 24, 143–144. [Google Scholar] [CrossRef]
- Racusen, L.C.; Monteil, C.; Sgrignoli, A.; Lucskay, M.; Marouillat, S.; Rhim, J.G.; Morin, J.P. Cell lines with extended in vitro growth potential from human renal proximal tubule: Characterization, response to inducers, and comparison with established cell lines. J. Lab. Clin. Med. 1997, 129, 318–329. [Google Scholar] [CrossRef]
- Kim, W.; Conery, A.L.; Rajamuthiah, R.; Fuchs, B.B.; Ausubel, F.M.; Mylonakis, E. Identification of an antimicrobial agent effective against methicillin-resistant Staphylococcus aureus persisters using a fluorescence-based screening strategy. PLoS ONE 2015, 10, e0127640. [Google Scholar] [CrossRef]
- Percival, S.L.; Suleman, L.; Vuotto, C.; Donelli, G. Healthcare-associated infections, medical devices and biofilms: Risk, tolerance and control. J. Med. Microbiol. 2015, 64 Pt 4, 64323–64334. [Google Scholar] [CrossRef]
- Moormeier, D.E.; Bayles, K.W. Staphylococcus aureus biofilm: A complex developmental organism. Mol. Microbiol. 2017, 104, 365–376. [Google Scholar] [CrossRef]
- Chung, P.Y.; Toh, Y.S. Anti-biofilm agents: Recent breakthrough against multi-drug resistant Staphylococcus aureus. Pathog. Dis. 2014, 70, 231–239. [Google Scholar] [CrossRef] [PubMed]
- Bjarnsholt, T.; Ciofu, O.; Molin, S.; Givskov, M.; Høiby, N. Applying insights from biofilm biology to drug development—Can a new approach be developed? Nat. Rev. Drug Discov. 2013, 12, 791–808. [Google Scholar] [CrossRef] [PubMed]
- Hurdle, J.G.; O’Neill, A.J.; Chopra, I.; Lee, R.E. Targeting bacterial membrane function: An underexploited mechanism for treating persistent infections. Nat. Rev. Microbiol. 2010, 9, 62–75. [Google Scholar] [CrossRef] [PubMed]
- Kim, W.; Fricke, N.; Conery, A.L.; Fuchs, B.B.; Rajamuthiah, R.; Jayamani, E.; Vlahovska, P.M.; Ausubel, F.M.; Mylonakis, E. NH125 kills methicillin-resistant Staphylococcus aureus persisters by lipid bilayer disruption. Future Med. Chem. 2016, 8, 257–269. [Google Scholar] [CrossRef]
- Raetz, C.R.H.; Whitfield, C. Lipopolysaccharide endotoxins. Annu. Rev. Biochem. 2002, 71, 635–700. [Google Scholar] [CrossRef]
- Tanaka, A.; Konno, M.; Muto, S.; Kambe, N.; Morii, E.; Nakahata, T.; Itai, A.; Matsuda, H. A novel NF-kappaB inhibitor, IMD-0354, suppresses neoplastic proliferation of human mast cells with constitutively activated c-kit receptors. Blood 2005, 105, 2324–2331. [Google Scholar] [CrossRef]
- Kwan, B.W.; Chowdhury, N.; Wood, T.K. Combatting bacterial infections by killing persister cells with mitomycin C. Environ. Microbiol. 2015, 17, 4406–4414. [Google Scholar] [CrossRef]
- Chowdhury, N.; Wood, T.L.; Martinez-Vazquez, M.; García-Contreras, R.; Wood, T.K. DNA-crosslinker cisplatin eradicates bacterial persister cells. Biotechnol. Bioeng. 2016, 113, 1984–1992. [Google Scholar] [CrossRef]
- Yang, E.B.; Tang, W.Y.; Zhang, K.; Cheng, L.Y.; Mack, P.O. Norcantharidin inhibits growth of human HepG2 cell-transplanted tumor in nude mice and prolongs host survival. Cancer Lett. 1997, 117, 93–98. [Google Scholar] [CrossRef]
- Li, Y.; Wang, Z.; Ma, X.; Shao, B.; Gao, X.; Zhang, B.; Xu, G.; Wei, Y. Low-dose cisplatin administration to septic mice improves bacterial clearance and programs peritoneal macrophage polarization to M1 phenotype. Pathog. Dis. 2014, 72, 111–123. [Google Scholar] [CrossRef]
- CLSI. M07-A9: Methods for Dilution Antimicrobial Susceptibility Tests for Bacteria That Grow Aerobically, 9th ed.; Approved Standard; CLSI: Sydney NSW, Australia, 2012; pp. 1–88. [Google Scholar]
- Fuchs, S.; Pané-Farré, J.; Kohler, C.; Hecker, M.; Engelmann, S. Anaerobic gene expression in Staphylococcus aureus. J. Bacteriol. 2007, 189, 4275–4289. [Google Scholar] [CrossRef] [PubMed]
- Yuen, G.J.; Ausubel, F.M. Both live and dead Enterococci activate Caenorhabditis elegans host defense via immune and stress pathways. Virulence 2018, 9, 683–699. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Lee, A.-R.; Kawasaki, I.; Strome, S.; Shim, Y.-H. A mutation of cdc-25.1 causes defects in germ cells but not in somatic tissues in C. elegans. Mol. Cells 2009, 28, 43–48. [Google Scholar] [CrossRef] [PubMed]
- Rajamuthiah, R.; Jayamani, E.; Conery, A.L.; Fuchs, B.B.; Kim, W.; Johnston, T.; Vilcinskas, A.; Ausubel, F.M.; Mylonakis, E. A Defensin from the Model Beetle Tribolium castaneum Acts Synergistically with Telavancin and Daptomycin against Multidrug Resistant Staphylococcus aureus. PLoS ONE 2015, 10, e0128576. [Google Scholar] [CrossRef]
- Mishra, B.; Golla, R.M.; Lau, K.; Lushnikova, T.; Wang, G. Anti-Staphylococcal Biofilm Effects of Human Cathelicidin Peptides. ACS Med. Chem. Lett. 2016, 7, 117–121. [Google Scholar] [CrossRef]
- O’Toole, G.A. Microtiter dish biofilm formation assay. J. Vis. Exp. 2011, 47, e2437. [Google Scholar] [CrossRef]

| Strain | MIC |
|---|---|
| Pseudomonas aeruginosa PA14 | >64 |
| Klebsiella pneumoniae WGLW2 | >64 |
| Acinetobacter baumannii ATCC 17978 | 16 |
| Enterobacter aerogenes EAE 2625 | >64 |
| Enterococcus faecalis MMH 594 | 0.25 |
| Enterococcus faecium E007 | 0.125 |
| IMD0354 | Vancomycin | |||
|---|---|---|---|---|
| Strain | MIC | MBC | MIC | MBC |
| VRS1 | 0.06 | 2 | >64 | >64 |
| VISA 215 | 0.25 | 0.5 | 8 | 8 |
| VISA 216 | 0.06 | 2 | 8 | 16 |
| VISA 217 | 0.125 | 0.5 | 8 | 8 |
| VISA 218 | 0.06 | 0.5 | 8 | 8 |
| VISA 219 | 0.06 | 1 | 8 | 16 |
| VISA 220 | 0.25 | 2.0 | 8 | 16 |
| VISA 221 | 0.06 | 1.0 | 8 | 8 |
| VISA 222 | 0.125 | 2.0 | 8 | 8 |
| VISA 223 | 0.125 | 1.0 | 4 | 8 |
| VISA 224 | 0.25 | 2 | 4 | 16 |
| VISA 225 | 0.125 | 0.25 | 4 | 8 |
| VISA 226 | 0.06 | 0.125 | 4 | 4 |
| VISA 227 | 0.25 | 0.5 | 8 | 8 |
| VISA 228 | 0.125 | 0.5 | 4 | 8 |
| IMD0354 | Vancomycin | |||
|---|---|---|---|---|
| Strain | MIC | MBC | MIC | MBC |
| EM C68 [35] | 0.25 | 8 | 64 | >64 |
| EM D366 [36] | 0.25 | 8 | 64 | >64 |
| EM WB312 [37] | 0.25 | 8 | 64 | >64 |
| EM WC176 [37] | 0.25 | 8 | 64 | >64 |
| EL V583 [38] | 0.25 | 8 | 64 | >64 |
| Bacterial Name | Strain |
|---|---|
| vancomycin-intermediate Staphylococcus aureus | VRS1 [32] |
| vancomycin-intermediate Staphylococcus aureus (clinical isolates) | VISA 215, 216, 217, 218, 219, 220, 221, 222, 223, 224, 225, 226, 227, 228 |
| vancomycin-resistant Enterococcus faecium (clinical isolates) | C68 [35], D366 [36], WB312 [37], WC176 [37] |
| vancomycin-resistant Enterococcus faecalis (clinical isolate) | V583 [38] |
| methicillin-resistant Staphylococcus aureus | MW2 |
| Klebsiella pneumoniae | WGLW2 |
| Acinetobacter baumannii | ATCC 17978 |
| Enterococcus faecalis | MMH 594 |
| Enterococcus faecium | E007 |
| Pseudomonas aeruginosa | PA14 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Escobar, I.E.; White, A.; Kim, W.; Mylonakis, E. New Antimicrobial Bioactivity against Multidrug-Resistant Gram-Positive Bacteria of Kinase Inhibitor IMD0354. Antibiotics 2020, 9, 665. https://doi.org/10.3390/antibiotics9100665
Escobar IE, White A, Kim W, Mylonakis E. New Antimicrobial Bioactivity against Multidrug-Resistant Gram-Positive Bacteria of Kinase Inhibitor IMD0354. Antibiotics. 2020; 9(10):665. https://doi.org/10.3390/antibiotics9100665
Chicago/Turabian StyleEscobar, Iliana E, Alexis White, Wooseong Kim, and Eleftherios Mylonakis. 2020. "New Antimicrobial Bioactivity against Multidrug-Resistant Gram-Positive Bacteria of Kinase Inhibitor IMD0354" Antibiotics 9, no. 10: 665. https://doi.org/10.3390/antibiotics9100665
APA StyleEscobar, I. E., White, A., Kim, W., & Mylonakis, E. (2020). New Antimicrobial Bioactivity against Multidrug-Resistant Gram-Positive Bacteria of Kinase Inhibitor IMD0354. Antibiotics, 9(10), 665. https://doi.org/10.3390/antibiotics9100665





