Comparative Genomics among Closely Related Streptomyces Strains Revealed Specialized Metabolite Biosynthetic Gene Cluster Diversity
Abstract
1. Introduction
2. Results and Discussion
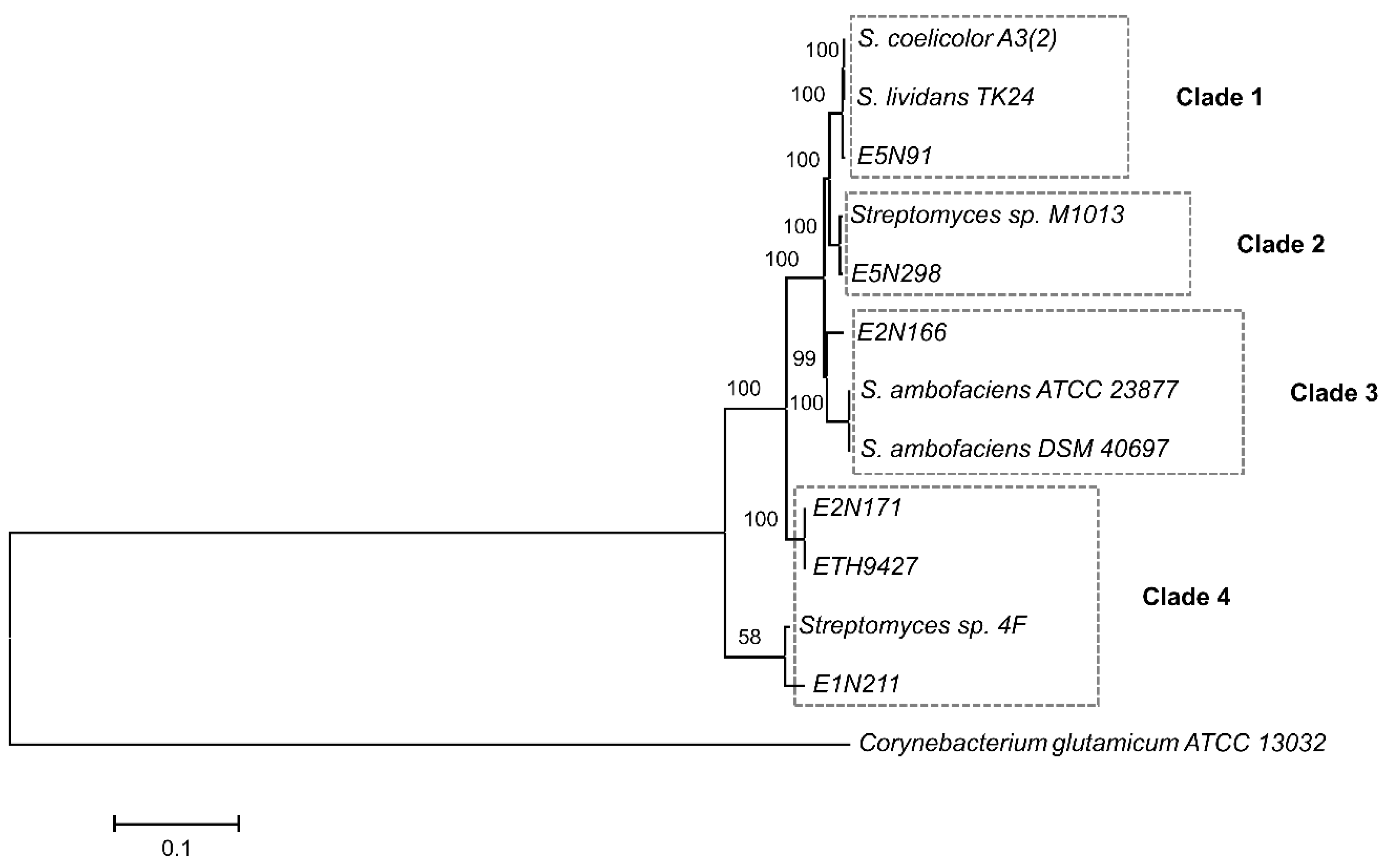
2.1. Analyzed Strains are Phylogenetically Diverse
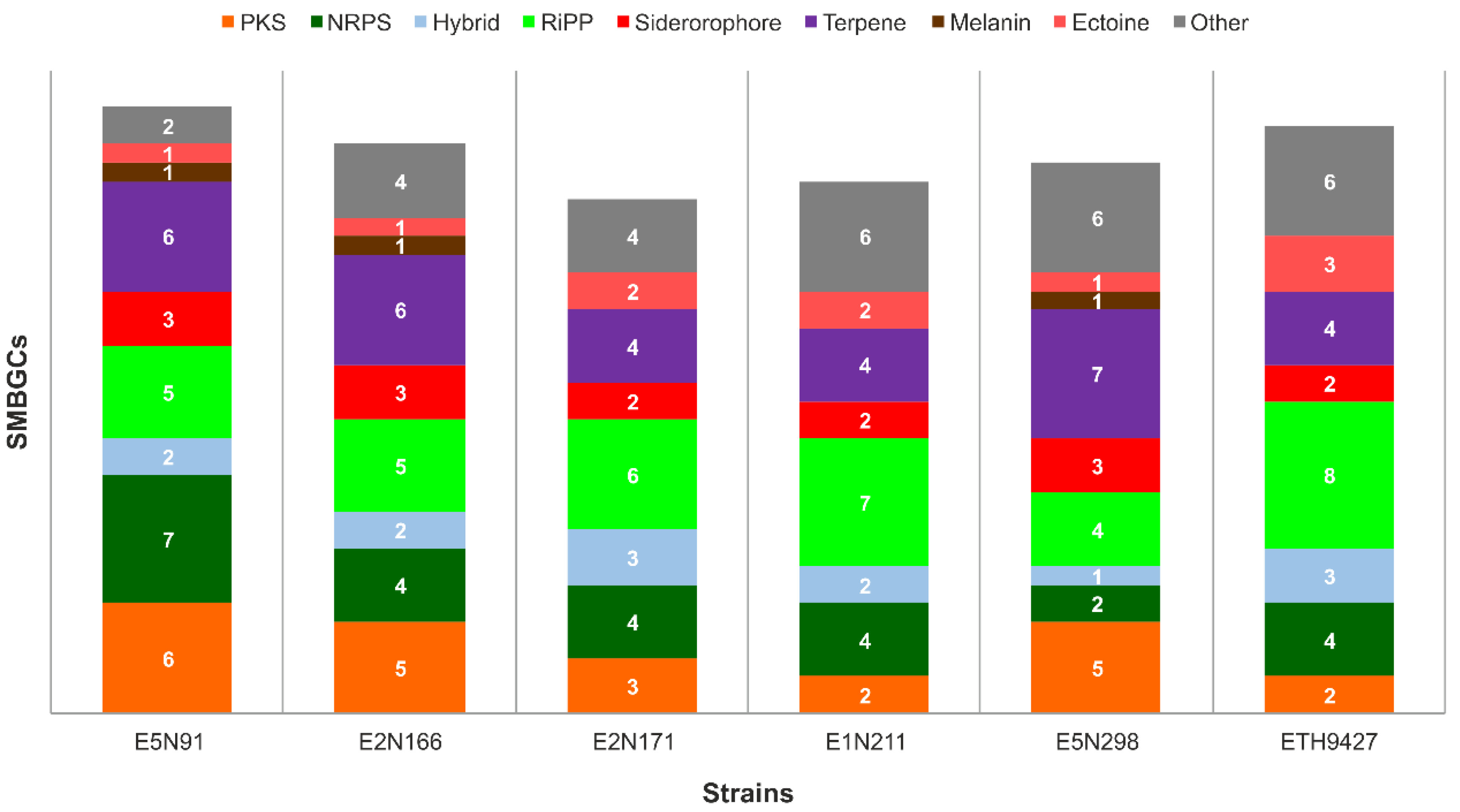
2.2. Specialized Metabolism Diversity
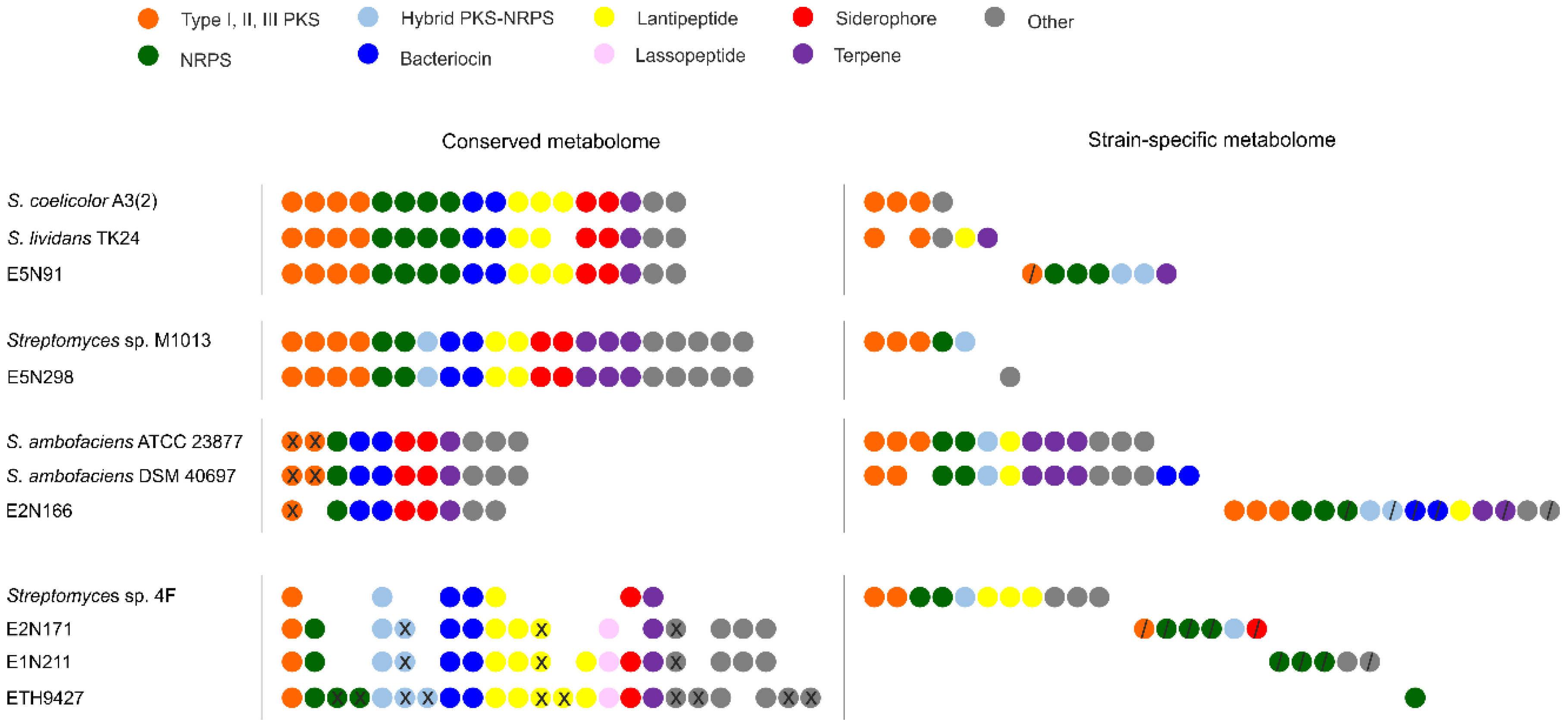
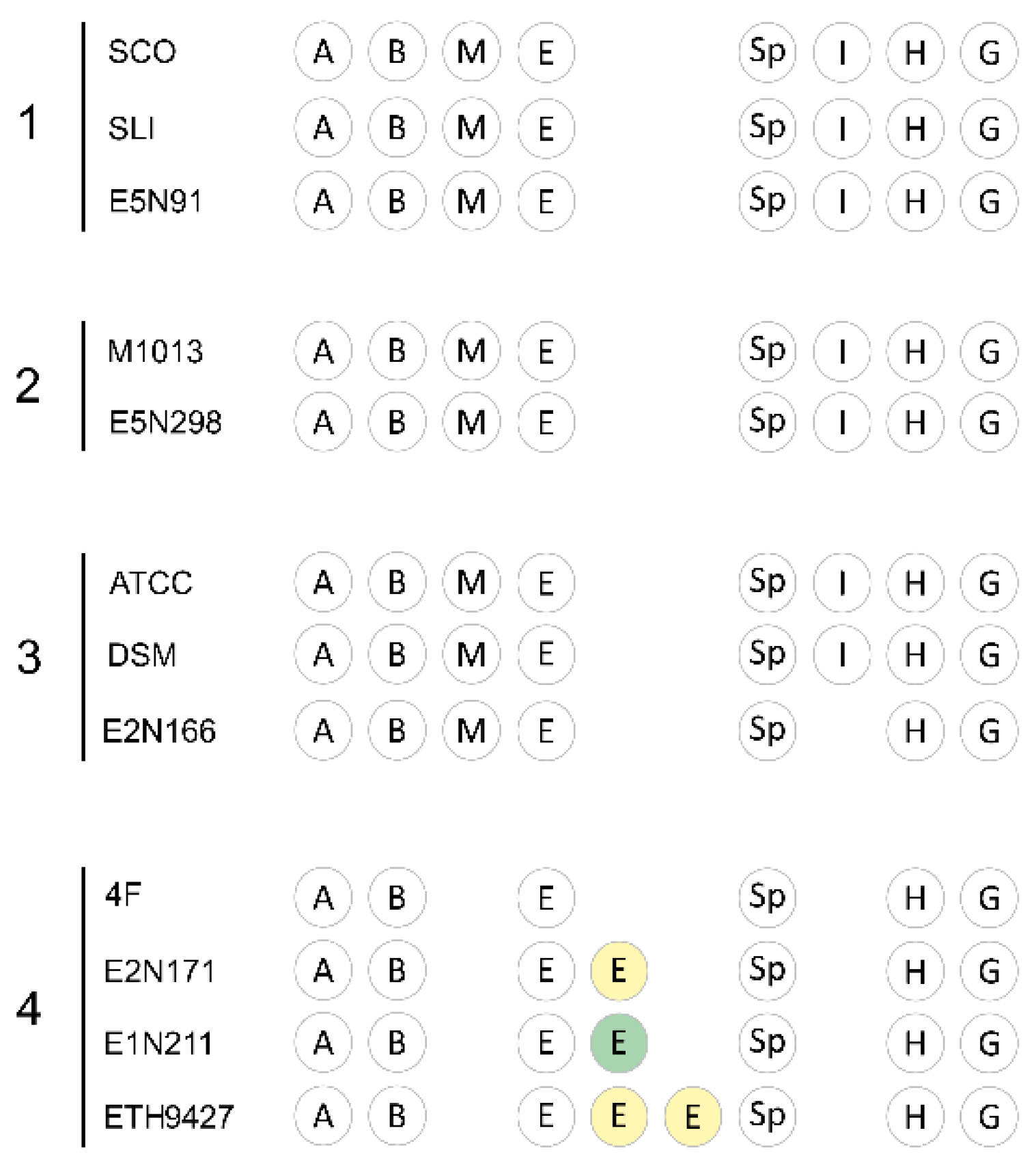
2.3. The Core and Conserved Specialized Metabolites
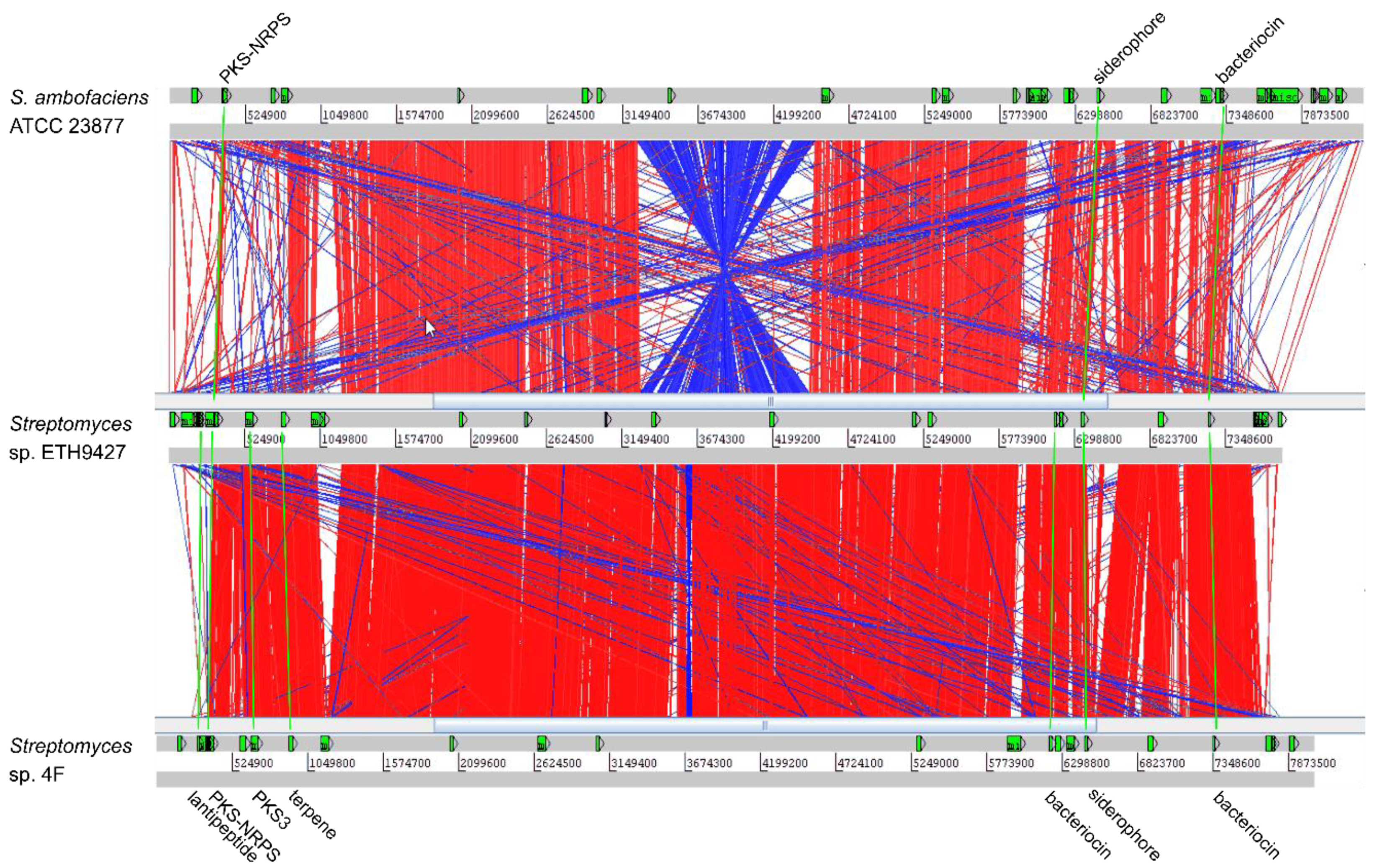
2.4. Strain-Specific Biosynthetic Gene Clusters
3. Materials and Methods
3.1. Strains and DNA Extraction
3.2. Whole-Genome Sequencing and Annotation
3.3. Phylogenetic Analysis and Biosynthetic Gene Cluster Identification
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Bérdy, J. Bioactive microbial metabolites. J. Antibiot. 2005, 58, 1–26. [Google Scholar] [CrossRef] [PubMed]
- Worrall, J.A.R.; Vijgenboom, E. Copper mining in Streptomyces: Enzymes, natural products and development. Nat. Prod. Rep. 2010, 27, 742–756. [Google Scholar] [CrossRef] [PubMed]
- Newman, D.J.; Cragg, G.M. Natural Products as Sources of New Drugs from 1981 to 2014. J. Nat. Prod. 2016, 79, 629–661. [Google Scholar] [CrossRef] [PubMed]
- Corre, C.; Challis, G.L. New natural product biosynthetic chemistry discovered by genome mining. Nat. Prod. Rep. 2009, 26, 977–986. [Google Scholar] [CrossRef] [PubMed]
- Lewis, K. Platforms for antibiotic discovery. Nat. Rev. Drug. Discov. 2013, 12, 371–387. [Google Scholar] [CrossRef] [PubMed]
- Rebets, Y.; Kormanec, J.; Luzhetskyy, A.; Bernaerts, K.; Anné, J. Cloning and Expression of Metagenomic DNA in Streptomyces lividans and Subsequent Fermentation for Optimized Production. Methods Mol. Biol. 2017, 1539, 99–144. [Google Scholar] [CrossRef] [PubMed]
- Bentley, S.D.; Chater, K.F.; Cerdeño-Tárraga, A.-M.; Challis, G.L.; Thomson, N.R.; James, K.D.; Harris, D.E.; Quail, M.A.; Kieser, H.; Harper, D.; et al. Complete genome sequence of the model actinomycete Streptomyces coelicolor A3(2). Nature 2002, 417, 141–147. [Google Scholar] [CrossRef] [PubMed]
- Medema, M.H.; Fischbach, M.A. Computational approaches to natural product discovery. Nat. Chem. Biol. 2015, 11, 639–648. [Google Scholar] [CrossRef] [PubMed]
- Nett, M.; Ikeda, H.; Moore, B.S. Genomic basis for natural product biosynthetic diversity in the actinomycetes. Nat. Prod. Rep. 2009, 26, 1362–1384. [Google Scholar] [CrossRef] [PubMed]
- Seipke, R.F. Strain-level diversity of secondary metabolism in Streptomyces albus. PLoS ONE 2015, 10, e0116457. [Google Scholar] [CrossRef] [PubMed]
- Antony-Babu, S.; Stien, D.; Eparvier, V.; Parrot, D.; Tomasi, S.; Suzuki, M.T. Multiple Streptomyces species with distinct secondary metabolomes have identical 16S rRNA gene sequences. Sci. Rep. 2017, 7, 11089. [Google Scholar] [CrossRef] [PubMed]
- Ziemert, N.; Lechner, A.; Wietz, M.; Millán-Aguiñaga, N.; Chavarria, K.L.; Jensen, P.R. Diversity and evolution of secondary metabolism in the marine actinomycete genus Salinispora. Proc. Natl. Acad. Sci. USA 2014, 111, E1130–E1139. [Google Scholar] [CrossRef] [PubMed]
- Benhadj, M.; Gacemi-Kirane, D.; Toussaint, M.; Hotel, L.; Bontemps, C.; Duval, R.E.; Aigle, B.; Leblond, P. Diversity and antimicrobial activities of Streptomyces isolates from Fetzara Lake, north eastern Algeria. Ann. Biol. Clin. 2018, 76, 81–95. [Google Scholar] [CrossRef] [PubMed]
- Thibessard, A.; Vicente, C.M.; Bertrand, C.; Aigle, B.; Leblond, P. Draft genome of Streptomyces sp. ETH9427, whole genome shotgun sequence. Genome Announc. 2018, in press. [Google Scholar]
- Rückert, C.; Albersmeier, A.; Busche, T.; Jaenicke, S.; Winkler, A.; Friðjónsson, Ó.H.; Hreggviðsson, G.Ó.; Lambert, C.; Badcock, D.; Bernaerts, K.; et al. Complete genome sequence of Streptomyces lividans TK24. J. Biotechnol. 2015, 199, 21–22. [Google Scholar] [CrossRef] [PubMed]
- Haas, D.; Gerbaud, C.; Sahin, N.; Pernodet, J.-L.; Lautru, S. Draft Genome Sequence of Streptomyces sp. M1013, a Close Relative of Streptomyces ambofaciens and Streptomyces coelicolor. Genome Announc. 2017, 5. [Google Scholar] [CrossRef] [PubMed]
- Thibessard, A.; Haas, D.; Gerbaud, C.; Aigle, B.; Lautru, S.; Pernodet, J.-L.; Leblond, P. Complete genome sequence of Streptomyces ambofaciens ATCC 23877, the spiramycin producer. J. Biotechnol. 2015, 214, 117–118. [Google Scholar] [CrossRef] [PubMed]
- Thibessard, A.; Leblond, P. Complete Genome Sequence of Streptomyces ambofaciens DSM 40697, a Paradigm for Genome Plasticity Studies. Genome Announc. 2016, 4. [Google Scholar] [CrossRef] [PubMed]
- Richter, M.; Rosselló-Móra, R.; Oliver Glöckner, F.; Peplies, J. JSpeciesWS: A web server for prokaryotic species circumscription based on pairwise genome comparison. Bioinformatics 2016, 32, 929–931. [Google Scholar] [CrossRef] [PubMed]
- Rosselló-Móra, R.; Amann, R. Past and future species definitions for Bacteria and Archaea. Syst. Appl. Microbiol. 2015, 38, 209–216. [Google Scholar] [CrossRef] [PubMed]
- Alam, M.T.; Merlo, M.E.; Takano, E.; Breitling, R. Genome-based phylogenetic analysis of Streptomyces and its relatives. Mol. Phylogenet. Evol. 2010, 54, 763–772. [Google Scholar] [CrossRef] [PubMed]
- Rong, X.; Liu, N.; Ruan, J.; Huang, Y. Multilocus sequence analysis of Streptomyces griseus isolates delineating intraspecific diversity in terms of both taxonomy and biosynthetic potential. Antonie. Van. Leeuwenhoek 2010, 98, 237–248. [Google Scholar] [CrossRef] [PubMed]
- Rong, X.; Huang, Y. Taxonomic evaluation of the Streptomyces hygroscopicus clade using multilocus sequence analysis and DNA-DNA hybridization, validating the MLSA scheme for systematics of the whole genus. Syst. Appl. Microbiol. 2012, 35, 7–18. [Google Scholar] [CrossRef] [PubMed]
- Labeda, D.P.; Dunlap, C.A.; Rong, X.; Huang, Y.; Doroghazi, J.R.; Ju, K.-S.; Metcalf, W.W. Phylogenetic relationships in the family Streptomycetaceae using multi-locus sequence analysis. Antonie Van Leeuwenhoek 2017, 110, 563–583. [Google Scholar] [CrossRef] [PubMed]
- Jiménez, G.; Urdiain, M.; Cifuentes, A.; López-López, A.; Blanch, A.R.; Tamames, J.; Kämpfer, P.; Kolstø, A.-B.; Ramón, D.; Martínez, J.F.; et al. Description of Bacillus toyonensis sp. nov., a novel species of the Bacillus cereus group, and pairwise genome comparisons of the species of the group by means of ANI calculations. Syst. Appl. Microbiol. 2013, 36, 383–391. [Google Scholar] [CrossRef] [PubMed]
- Weber, T.; Blin, K.; Duddela, S.; Krug, D.; Kim, H.U.; Bruccoleri, R.; Lee, S.Y.; Fischbach, M.A.; Müller, R.; Wohlleben, W.; et al. AntiSMASH 3.0-a comprehensive resource for the genome mining of biosynthetic gene clusters. Nucleic Acids Res. 2015, 43, W237–W243. [Google Scholar] [CrossRef] [PubMed]
- Bernan, V.; Filpula, D.; Herber, W.; Bibb, M.; Katz, E. The nucleotide sequence of the tyrosinase gene from Streptomyces antibioticus and characterization of the gene product. Gene 1985, 37, 101–110. [Google Scholar] [CrossRef]
- Malin, G.; Lapidot, A. Induction of synthesis of tetrahydropyrimidine derivatives in Streptomyces strains and their effect on Escherichia coli in response to osmotic and heat stress. J. Bacteriol. 1996, 178, 385–395. [Google Scholar] [CrossRef] [PubMed]
- Davis, N.K.; Chater, K.F. Spore colour in Streptomyces coelicolor A3(2) involves the developmentally regulated synthesis of a compound biosynthetically related to polyketide antibiotics. Mol. Microbiol. 1990, 4, 1679–1691. [Google Scholar] [CrossRef] [PubMed]
- Neilands, J.B. Microbial iron compounds. Annu. Rev. Biochem. 1981, 50, 715–731. [Google Scholar] [CrossRef] [PubMed]
- Gürtler, H.; Pedersen, R.; Anthoni, U.; Christophersen, C.; Nielsen, P.H.; Wellington, E.M.; Pedersen, C.; Bock, K. Albaflavenone, a sesquiterpene ketone with a zizaene skeleton produced by a streptomycete with a new rope morphology. J. Antibiot. 1994, 47, 434–439. [Google Scholar] [CrossRef] [PubMed]
- Poralla, K.; Muth, G.; Härtner, T. Hopanoids are formed during transition from substrate to aerial hyphae in Streptomyces coelicolor A3(2). FEMS Microbiol. Lett. 2000, 189, 93–95. [Google Scholar] [CrossRef] [PubMed]
- Krügel, H.; Krubasik, P.; Weber, K.; Saluz, H.P.; Sandmann, G. Functional analysis of genes from Streptomyces griseus involved in the synthesis of isorenieratene, a carotenoid with aromatic end groups, revealed a novel type of carotenoid desaturase. Biochim. Biophys. Acta 1999, 1439, 57–64. [Google Scholar] [CrossRef]
- Gerber, N.N.; Lechevalier, H.A. Geosmin, an earthly-smelling substance isolated from actinomycetes. Appl. Microbiol. 1965, 13, 935–938. [Google Scholar] [PubMed]
- Choudoir, M.J.; Pepe-Ranney, C.; Buckley, D.H. Diversification of Secondary Metabolite Biosynthetic Gene Clusters Coincides with Lineage Divergence in Streptomyces. Antibiotics 2018, 7, 12. [Google Scholar] [CrossRef]
- Lautru, S.; Deeth, R.J.; Bailey, L.M.; Challis, G.L. Discovery of a new peptide natural product by Streptomyces coelicolor genome mining. Nat. Chem. Biol. 2005, 1, 265–269. [Google Scholar] [CrossRef] [PubMed]
- Hesketh, A.; Kock, H.; Mootien, S.; Bibb, M. The role of absC, a novel regulatory gene for secondary metabolism, in zinc-dependent antibiotic production in Streptomyces coelicolor A3(2). Mol. Microbiol. 2009, 74, 1427–1444. [Google Scholar] [CrossRef] [PubMed]
- Chong, P.P.; Podmore, S.M.; Kieser, H.M.; Redenbach, M.; Turgay, K.; Marahiel, M.; Hopwood, D.A.; Smith, C.P. Physical identification of a chromosomal locus encoding biosynthetic genes for the lipopeptide calcium-dependent antibiotic (CDA) of Streptomyces coelicolor A3(2). Microbiology 1998, 144 Pt 1, 193–199. [Google Scholar] [CrossRef] [PubMed]
- Takano, E.; Kinoshita, H.; Mersinias, V.; Bucca, G.; Hotchkiss, G.; Nihira, T.; Smith, C.P.; Bibb, M.; Wohlleben, W.; Chater, K. A bacterial hormone (the SCB1) directly controls the expression of a pathway-specific regulatory gene in the cryptic type I polyketide biosynthetic gene cluster of Streptomyces coelicolor. Mol. Microbiol. 2005, 56, 465–479. [Google Scholar] [CrossRef] [PubMed]
- Wright, L.F.; Hopwood, D.A. Actinorhodin is a chromosomally-determined antibiotic in Streptomyces coelicolar A3(2). J. Gen. Microbiol. 1976, 96, 289–297. [Google Scholar] [CrossRef] [PubMed]
- Rudd, B.A.; Hopwood, D.A. A pigmented mycelial antibiotic in Streptomyces coelicolor: Control by a chromosomal gene cluster. J. Gen. Microbiol. 1980, 119, 333–340. [Google Scholar] [CrossRef] [PubMed]
- Aigle, B.; Pang, X.; Decaris, B.; Leblond, P. Involvement of AlpV, a new member of the Streptomyces antibiotic regulatory protein family, in regulation of the duplicated type II polyketide synthase alp gene cluster in Streptomyces ambofaciens. J. Bacteriol. 2005, 187, 2491–2500. [Google Scholar] [CrossRef] [PubMed]
- Aigle, B.; Lautru, S.; Spiteller, D.; Dickschat, J.S.; Challis, G.L.; Leblond, P.; Pernodet, J.-L. Genome mining of Streptomyces ambofaciens. J. Ind. Microbiol. Biotechnol. 2014, 41, 251–263. [Google Scholar] [CrossRef] [PubMed]
- Jensen, P.R.; Williams, P.G.; Oh, D.-C.; Zeigler, L.; Fenical, W. Species-specific secondary metabolite production in marine actinomycetes of the genus Salinispora. Appl. Environ. Microbiol. 2007, 73, 1146–1152. [Google Scholar] [CrossRef] [PubMed]
- Penn, K.; Jenkins, C.; Nett, M.; Udwary, D.W.; Gontang, E.A.; McGlinchey, R.P.; Foster, B.; Lapidus, A.; Podell, S.; Allen, E.E.; et al. Genomic islands link secondary metabolism to functional adaptation in marine actinobacteria. ISME J. 2009, 3, 1193–1203. [Google Scholar] [CrossRef] [PubMed]
- Joynt, R.; Seipke, R.F. A phylogenetic and evolutionary analysis of antimycin biosynthesis. Microbiology 2018, 164, 28–39. [Google Scholar] [CrossRef] [PubMed]
- Kieser, T.; Bibb, M.; Buttner, M.; Chater, K.; Hopwood, D.A. Practical Streptomyces Genetics; The John Innes Foundation: Norwich, UK, 2000. [Google Scholar]
- Tatusova, T.; DiCuccio, M.; Badretdin, A.; Chetvernin, V.; Nawrocki, E.P.; Zaslavsky, L.; Lomsadze, A.; Pruitt, K.D.; Borodovsky, M.; Ostell, J. NCBI prokaryotic genome annotation pipeline. Nucleic. Acids. Res. 2016, 44, 6614–6624. [Google Scholar] [CrossRef] [PubMed]
- Carver, T.; Berriman, M.; Tivey, A.; Patel, C.; Böhme, U.; Barrell, B.G.; Parkhill, J.; Rajandream, M.-A. Artemis and ACT: Viewing, annotating and comparing sequences stored in a relational database. Bioinformatics 2008, 24, 2672–2676. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.; Zheng, W.; Rong, X.; Huang, Y. A multilocus phylogeny of the Streptomyces griseus 16S rRNA gene clade: Use of multilocus sequence analysis for streptomycete systematics. Int. J. Syst. Evol. Microbiol. 2008, 58, 149–159. [Google Scholar] [CrossRef] [PubMed]
- Gouy, M.; Guindon, S.; Gascuel, O. SeaView version 4: A multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Mol. Biol. Evol. 2010, 27, 221–224. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef] [PubMed]

| Strain | G + C Content (%) | Genome Size (Mb) |
|---|---|---|
| E5N91 | 71.9 | 8.51 |
| E2N166 | 70.2 | 7.91 |
| E2N171 | 72.3 | 7.00 |
| E1N211 | 72.1 | 7.32 |
| E5N298 | 71.9 | 7.87 |
| Streptomyces sp. ETH9427 | 72.1 | 7.75 |
| SLI | E5N91 | M1013 | E5N298 | ATCC | DSM | E2N166 | E2N171 | E1N211 | ETH9427 | 4F | |
| SCO | 99.0 | 97.3 | 91.5 | 91.3 | 86.9 | 86.9 | 87.5 | 81.7 | 81.8 | 81.9 | 81.8 |
| SLI | 97.7 | 91.8 | 91.6 | 87.0 | 87.0 | 87.7 | 81.6 | 81.7 | 81.8 | 81.7 | |
| E5N91 | 91.2 | 91.0 | 86.6 | 86.6 | 87.4 | 81.4 | 81.4 | 81.6 | 81.3 | ||
| M1013 | 98.2 | 87.1 | 87.1 | 87.9 | 81.5 | 81.5 | 81.7 | 81.4 | |||
| E5N298 | 87.2 | 87.2 | 87.9 | 81.7 | 81.7 | 81.8 | 81.6 | ||||
| ATCC | 99.0 | 88.9 | 81.8 | 81.7 | 81.8 | 81.9 | |||||
| DSM | 88.8 | 81.7 | 81.6 | 81.7 | 81.9 | ||||||
| E2N166 | 82.1 | 82.1 | 82.2 | 82.0 | |||||||
| E2N171 | 99.0 | 99.1 | 94.6 | ||||||||
| E1N211 | 99.9 | 94.4 | |||||||||
| ETH9427 | 94.1 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vicente, C.M.; Thibessard, A.; Lorenzi, J.-N.; Benhadj, M.; Hôtel, L.; Gacemi-Kirane, D.; Lespinet, O.; Leblond, P.; Aigle, B. Comparative Genomics among Closely Related Streptomyces Strains Revealed Specialized Metabolite Biosynthetic Gene Cluster Diversity. Antibiotics 2018, 7, 86. https://doi.org/10.3390/antibiotics7040086
Vicente CM, Thibessard A, Lorenzi J-N, Benhadj M, Hôtel L, Gacemi-Kirane D, Lespinet O, Leblond P, Aigle B. Comparative Genomics among Closely Related Streptomyces Strains Revealed Specialized Metabolite Biosynthetic Gene Cluster Diversity. Antibiotics. 2018; 7(4):86. https://doi.org/10.3390/antibiotics7040086
Chicago/Turabian StyleVicente, Cláudia M., Annabelle Thibessard, Jean-Noël Lorenzi, Mabrouka Benhadj, Laurence Hôtel, Djamila Gacemi-Kirane, Olivier Lespinet, Pierre Leblond, and Bertrand Aigle. 2018. "Comparative Genomics among Closely Related Streptomyces Strains Revealed Specialized Metabolite Biosynthetic Gene Cluster Diversity" Antibiotics 7, no. 4: 86. https://doi.org/10.3390/antibiotics7040086
APA StyleVicente, C. M., Thibessard, A., Lorenzi, J.-N., Benhadj, M., Hôtel, L., Gacemi-Kirane, D., Lespinet, O., Leblond, P., & Aigle, B. (2018). Comparative Genomics among Closely Related Streptomyces Strains Revealed Specialized Metabolite Biosynthetic Gene Cluster Diversity. Antibiotics, 7(4), 86. https://doi.org/10.3390/antibiotics7040086





