Lysoquinone-TH1, a New Polyphenolic Tridecaketide Produced by Expressing the Lysolipin Minimal PKS II in Streptomyces albus
Abstract
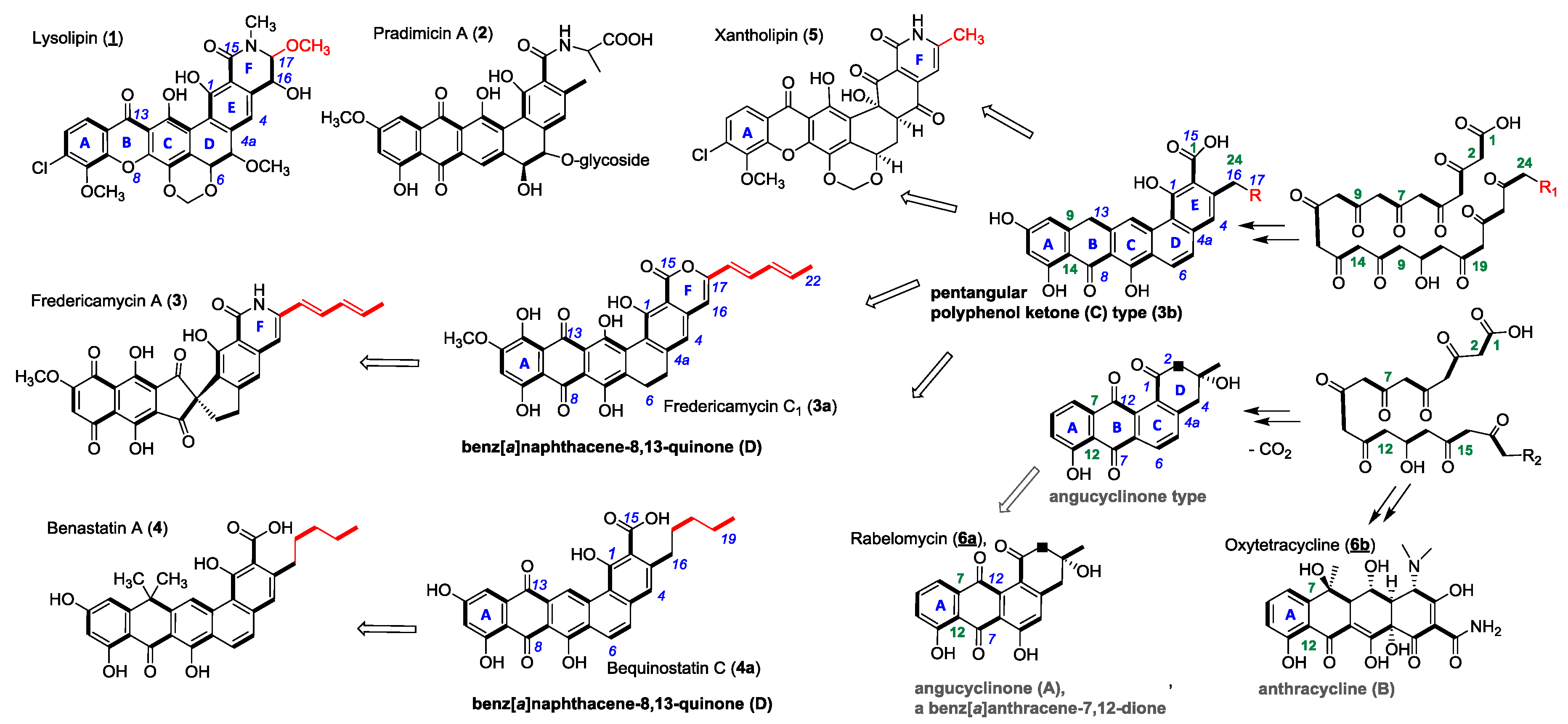
1. Introduction
2. Results and Discussion
2.1. Heterologous Production of Lysoquinone-TH 1 (7) in S. albus
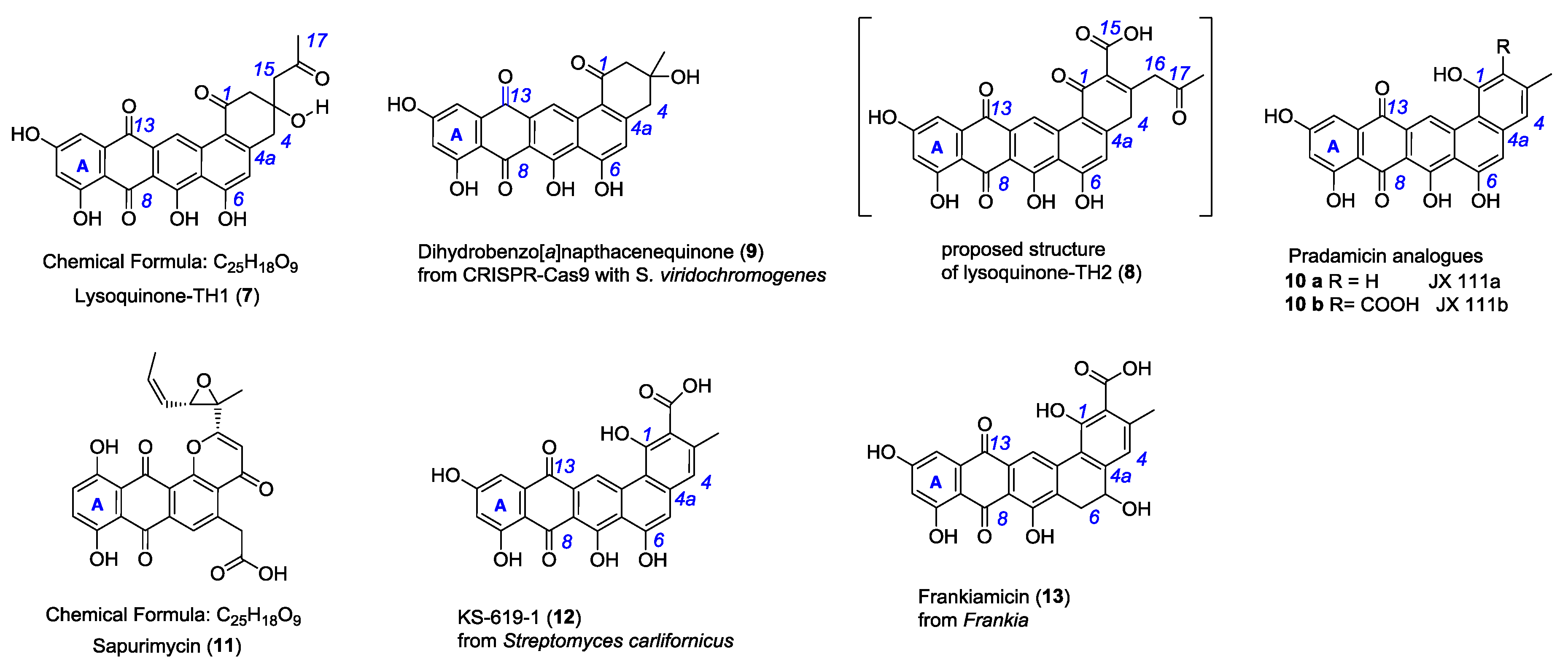
2.2. Isolation and Structure Elucidation of Lysoquinone-TH1 (7)
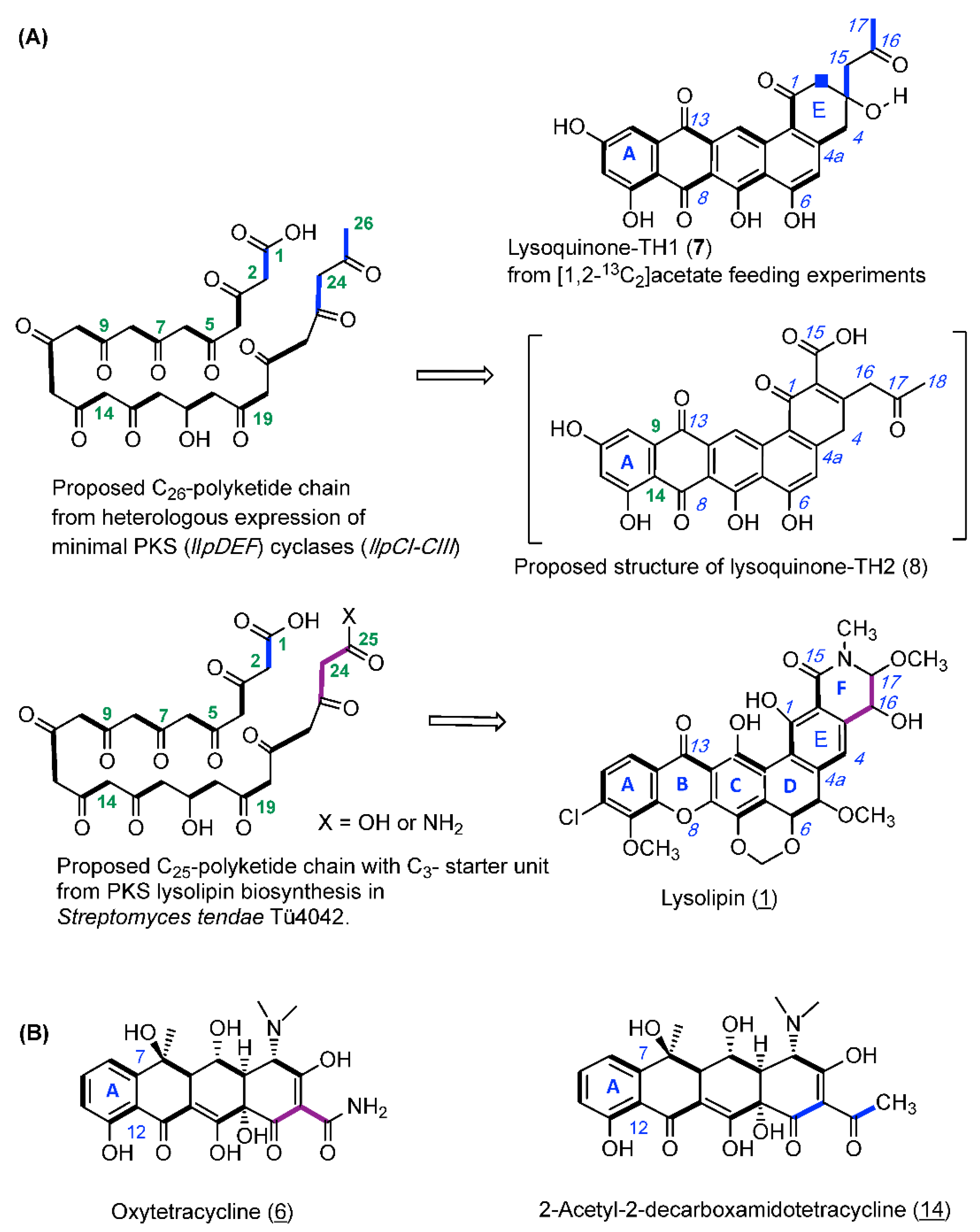
2.3. Feeding Experiment with [1,2-13C2]-Labeled Acetate
2.4. Biological Activity of Lysoquinone-TH1 (7)
3. Materials and Methods
3.1. Cloning of the Lysolipin Minimal PKS
3.2. Culture Conditions
3.3. Extraction and Isolation
3.4. Feeding Experiment with [1,2-13C2]-Labeled Acetate
3.5. Biological Activity Assays
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Hertweck, C. The biosynthetic logic of polyketide diversity. Angew. Chem. Int. Ed. 2009, 48, 4688–4716. [Google Scholar] [CrossRef] [PubMed]
- Kharel, M.K.; Pahari, P.; Shepherd, M.D.; Tibrewal, N.; Nybo, S.E.; Shaaban, K.A.; Rohr, J. Angucyclines: Biosynthesis, mode-of-action, new natural products, and synthesis. Nat. Prod. Rep. 2012, 29, 264–325. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Pan, H.-X.; Tang, G.-L. New insights into bacterial type II polyketide biosynthesis. F1000Research 2017, 6, 172. [Google Scholar] [CrossRef] [PubMed]
- Tang, Y.; Tsai, S.C.; Khosla, C. Polyketide chain length control by chain length factor. J. Am. Chem. Soc. 2003, 125, 12708–12709. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; Li, Y.; Tang, Y. Cyclization of aromatic polyketides from bacteria and fungi. Nat. Prod. Rep. 2010, 27, 839–868. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Watanabe, K.; Wang, C.C.C.; Tang, Y. Investigation of early tailoring reactions in the oxytetracycline biosynthetic pathway. J. Biol. Chem. 2007, 282, 25717–25725. [Google Scholar] [CrossRef] [PubMed]
- Valentic, T.R.; Jackson, D.R.; Brady, S.F.; Tsai, S.C. Comprehensive Analysis of a Novel Ketoreductase for Pentangular Polyphenol Biosynthesis. ACS Chem. Biol. 2016, 11, 3421–3430. [Google Scholar] [CrossRef] [PubMed]
- Kong, L.; Zhang, W.; Chooi, Y.H.; Wang, L.; Cao, B.; Deng, Z.; Chu, Y.; You, D. A Multifunctional monooxygenase XanO4 catalyzes xanthone Formation in xantholipin biosynthesis via a cryptic demethoxylation. Cell Chem. Biol. 2016, 23, 508–516. [Google Scholar] [CrossRef] [PubMed]
- Gullon, S.; Olano, C.; Abdelfattah, M.S.; Brana, A.F.; Rohr, J.; Mendez, C.; Salas, J.A. Isolation, characterization, and heterologous expression of the biosynthesis gene cluster for the antitumor anthracycline steffimycin. Appl. Environ. Microbiol. 2006, 72, 4172–4183. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Ames, B.D.; Tsai, S.C.; Tang, Y. Engineered biosynthesis of a novel amidated polyketide, using the malonamyl-specific initiation module from the oxytetracycline polyketide synthase. Appl. Environ. Microbiol. 2006, 72, 2573–2580. [Google Scholar] [CrossRef] [PubMed]
- Luzhetskyy, A.; Vente, A.; Bechthold, A. Glycosyltransferases involved in the biosynthesis of biologically active natural products that contain oligosaccharides. Mol. Biosyst. 2005, 1, 117–126. [Google Scholar] [CrossRef] [PubMed]
- Drautz, H.; Keller-Schierlein, W.; Zähner, H. Metabolic products of microorganisms, 149. Lysolipin I, a new antibiotic from Streptomyces violaceoniger (author’s transl). Arch. Microbiol. 1975, 106, 175–190. [Google Scholar] [CrossRef] [PubMed]
- Oki, T.; Konishi, M.; Tomatsu, K.; Tomita, K.; Saitoh, K.; Tsunakawa, M.; Nishio, M.; Miyaki, T.; Kawaguchi, H. Pradimicin, a novel class of potent antifungal antibiotics. J. Antibiot. 1988, 41, 1701–1704. [Google Scholar] [CrossRef] [PubMed]
- Misra, R.; Pandey, R.C.; Silverton, J.V. Fredericamycin A, an Antitumor antibiotic of a novel skeletal type. J. Am. Chem. Soc. 1982, 104, 4478–4479. [Google Scholar] [CrossRef]
- Aoyagi, T.; Aoyama, T.; Kojima, F.; Matsuda, N.; Maruyama, M.; Hamada, M.; Takeuchi, T. Benastatins A and B, new inhibitors of glutathione S-transferase, produced by Streptomyces sp. MI384-DF12. I. Taxonomy, production, isolation, physico-chemical properties and biological activities. J. Antibiot. 1992, 45, 1385–1390. [Google Scholar] [CrossRef] [PubMed]
- Yamazaki, T.; Tatee, T.; Aoyama; Kojima, F.; Takeuchi, T.; Aoyagi, T. Bequinostatins C and D, new inhibitors of glutathione S-transferase, produced by Streptomyces sp. MI384-DF12. J. Antibiot. 1993, 46, 1309–1311. [Google Scholar] [CrossRef] [PubMed]
- Terui, Y.; Yiwen, C.; Jun-Ying, L.; Ando, T.; Yamamoto, H.; Kawamura, Y.; Tomishima, Y.; Uchida, S.; Okazaki, T.; Munetomo, E.; et al. Xantholipin, a novel inhibitor of HSP47 gene expression produced by Streptomyces sp. Tetrahedron Lett. 2003, 44, 5427–5430. [Google Scholar] [CrossRef]
- Chen, Y.; Wendt-Pienkowski, E.; Ju, J.; Lin, S.; Rajski, S.R.; Shen, B. Characterization of FdmV as an amide synthetase for fredericamycin A biosynthesis in Streptomyces griseus ATCC 43944. J. Biol. Chem. 2010, 285, 38853–38860. [Google Scholar] [CrossRef] [PubMed]
- Kim, B.C.; Lee, J.M.; Ahn, J.S.; Kim, B.S. Cloning, sequencing, and characterization of the pradimicin biosynthetic gene cluster of Actinomadura hibisca P157-2. J. Microbiol. Biotechnol. 2007, 17, 830–839. [Google Scholar] [PubMed]
- Das, A.; Szu, P.-H.; Fitzgerald, J.T.; Khosla, C. Mechanism and engineering of polyketide chain initiation in fredericamycin biosynthesis. J. Am. Chem. Soc. 2010, 132, 8831–8833. [Google Scholar] [CrossRef] [PubMed]
- Lopez, P.; Hornung, A.; Welzel, K.; Unsin, C.; Wohlleben, W.; Weber, T.; Pelzer, S. Isolation of the lysolipin gene cluster of Streptomyces tendae Tü 4042. Gene 2010, 461, 5–14. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Wang, L.; Kong, L.; Wang, T.; Chu, Y.; Deng, Z.; You, D. Unveiling the post-PKS redox tailoring steps in biosynthesis of the type II polyketide antitumor antibiotic xantholipin. Chem. Biol. 2012, 19, 422–432. [Google Scholar] [CrossRef] [PubMed]
- Lackner, G.; Schenk, A.; Xu, Z.; Reinhardt, K.; Yunt, Z.S.; Piel, J.; Hertweck, C. Biosynthesis of pentangular polyphenols: Deductions from the benastatin and griseorhodin pathways. J. Am. Chem. Soc. 2007, 129, 9306–9312. [Google Scholar] [CrossRef] [PubMed]
- Menges, R.; Muth, G.; Wohlleben, W.; Stegmann, E. The ABC transporter Tba of Amycolatopsis balhimycina is required for efficient export of the glycopeptide antibiotic balhimycin. Appl. Microbiol. Biotechnol. 2007, 77, 125–134. [Google Scholar] [CrossRef] [PubMed]
- Uosaki, Y.; Yasuzawa, T.; Hara, M.; Saitoh, Y.; Sano, H. Sapurimycin, new antitumor antibiotic produced by Streptomyces. Structure determination. J. Antibiot. 1991, 44, 40–44. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.M.; Wong, F.T.; Wang, Y.; Luo, S.; Lim, Y.H.; Heng, E.; Yeo, W.L.; Cobb, R.E.; Enghiad, B.; Ang, E.L.; et al. CRISPR-Cas9 strategy for activation of silent Streptomyces biosynthetic gene clusters. Nat. Chem. Biol. 2017, 13, 607–609. [Google Scholar] [CrossRef] [PubMed]
- Zhan, J.; Watanabe, K.; Tang, Y. Synergistic actions of a monooxygenase and cyclases in aromatic polyketide biosynthesis. Chembiochem 2008, 9, 1710–1715. [Google Scholar] [CrossRef] [PubMed]
- Yasuzawa, T.; Yoshida, M.; Shirahata, K.; Sano, H. Structure of a novel Ca2+ and calmodulin-dependent cyclic nucleotide phosphodiesterase inhibitor KS-619-1. J. Antibiot. 1987, 40, 1111–1114. [Google Scholar] [CrossRef] [PubMed]
- Ogasawara, Y.; Yackley, B.J.; Greenberg, J.A.; Rogelj, S.; Melançon, C.E. Expanding our understanding of sequence-function relationships of Type II polyketide biosynthetic gene clusters: Bioinformatics-guided identification of frankiamicin a from Frankia sp. EAN1pec. PLoS ONE 2015, 10, 1–25. [Google Scholar] [CrossRef] [PubMed]
- Scott, A.I.; Townsend, C.A.; Okada, K.; Kajiwara, M.; Cushley, R.J.; Whitman, P.J. Biosynthesis of corrins. II. Incorporation of 13C-labeled substrate in vitamins B12. J. Am. Chem. Soc. 1974, 96, 8069–8080. [Google Scholar] [CrossRef] [PubMed]
- Bockholt, H.; Udvarnoki, G.; Rohr, J.; Mocek, U.; Beale, J.M.; Floss, H.G. Biosynthetic studies on the xanthone antibiotics lysolipins X and I. J. Org. Chem. 1994, 59, 2064–2069. [Google Scholar] [CrossRef]
- Fu, H.; Ebert-Khosla, S.; Khosla, C.; Hopwood, D.A. Relaxed Specificity of the oxytetracycline polyketide synthase for an acetate primer in the absence of a malonamyl primer. J. Am. Chem. Soc. 1994, 116, 6443–6444. [Google Scholar] [CrossRef]
- Xu, Z.; Schenk, A.; Hertweck, C. Molecular analysis of the benastatin biosynthetic pathway and genetic engineering of altered fatty acid-polyketide hybrids. J. Am. Chem. Soc. 2007, 129, 6022–6030. [Google Scholar] [CrossRef] [PubMed]
- Winter, D.K.; Sloman, D.L.; Porco, J., Jr. A. Polycyclic xanthone natural products: Structure, biological activity and chemical synthesis. Nat. Prod. Rep. 2013, 30, 382–391. [Google Scholar] [CrossRef] [PubMed]
- Silber, J.; Ohlendorf, B.; Labes, A.; Erhard, A.; Imhoff, J.F. Calcarides A–E, antibacterial macrocyclic and linear polyesters from a calcarisporium strain. Mar. Drugs 2013, 11, 3309–3323. [Google Scholar] [CrossRef] [PubMed]
- Matsuda, Y.; Kase, H. KS-619-1, a new inhibitor of Ca2+ and calmodulin-dependent cyclic nucleotide phosphodiesterase from Streptomyces californicus. J. Antibiot. 1987, 40, 1104–1110. [Google Scholar] [CrossRef] [PubMed]
- Matsuda, Y.; Asano, K.; Kawamoto, I.; Kase, H. K-259-2, a new inhibitor of Ca2+ and calmodulin-dependent cyclic nucleotide phosphodiesterase from Micromonospora olivasterospora. J. Antibiot. 1987, 40, 1092–1100. [Google Scholar] [CrossRef] [PubMed]
- Yasuzawa, T.; Yoshida, M.; Shirahata, K.; Sano, H. Structure of a novel Ca2+ and calmodulin-dependent cyclic nucleotide phosphodiesterase inhibitor K-259-2. J. Antibiot. 1987, 40, 1101–1103. [Google Scholar] [CrossRef] [PubMed]
- Schulz, D.; Beese, P.; Ohlendorf, B.; Erhard, A.; Zinecker, H.; Dorador, C.; Imhoff, J.F. Abenquines A–D: Aminoquinone derivatives produced by Streptomyces sp. strain DB634. J. Antibiot. 2011, 64, 763–768. [Google Scholar] [CrossRef] [PubMed]
- Houslay, M.D.; Schafer, P.; Zhang, K.Y.J. Keynote review: Phosphodiesterase-4 as a therapeutic target. Drug Discov. Today 2005, 10, 1503–1519. [Google Scholar] [CrossRef]
- Boswell-Smith, V.; Spina, D.; Page, C.P. Phosphodiesterase inhibitors. Br. J. Pharmacol. 2009, 147, S252–S257. [Google Scholar] [CrossRef] [PubMed]
- Millar, J.K.; Pickard, B.S.; Mackie, S.; James, R.; Christie, S.; Buchanan, S.R.; Malloy, M.P.; Chubb, J.E.; Huston, E.; Baillie, G.S.; et al. DISC1 and PDE4B are interacting genetic factors in schizophrenia that regulate cAMP signaling. Science 2005, 310, 1187–1191. [Google Scholar] [CrossRef] [PubMed]
- Park, S.-J.; Ahmad, F.; Philp, A.; Baar, K.; Williams, T.; Luo, H.; Ke, H.; Rehmann, H.; Taussig, R.; Brown, A.L.; et al. Resveratrol ameliorates aging-related metabolic phenotypes by inhibiting cAMP phosphodiesterases. Cell 2012, 148, 421–433. [Google Scholar] [CrossRef] [PubMed]
- Kanes, S.J.; Tokarczyk, J.; Siegel, S.J.; Bilker, W.; Abel, T.; Kelly, M.P. Rolipram: A specific phosphodiesterase 4 inhibitor with potential antipsychotic activity. Neuroscience 2007, 144, 239–246. [Google Scholar] [CrossRef] [PubMed]
- Houslay, M.D.; Baillie, G.S.; Maurice, D.H. cAMP-Specific phosphodiesterase-4 enzymes in the cardiovascular system: A molecular toolbox for generating compartmentalized cAMP signaling. Circ. Res. 2007, 100, 950–966. [Google Scholar] [CrossRef] [PubMed]
- Musiol, E.M.; Härtner, T.; Kulik, A.; Moldenhauer, J.; Piel, J.; Wohlleben, W.; Weber, T. Supramolecular templating in kirromycin biosynthesis: The acyltransferase KirCII loads ethylmalonyl-CoA extender onto a specific ACP of the trans-AT PKS. Chem. Biol. 2011, 18, 438–444. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hofeditz, T.; Unsin, C.E.-M.; Wiese, J.; Imhoff, J.F.; Wohlleben, W.; Grond, S.; Weber, T. Lysoquinone-TH1, a New Polyphenolic Tridecaketide Produced by Expressing the Lysolipin Minimal PKS II in Streptomyces albus. Antibiotics 2018, 7, 53. https://doi.org/10.3390/antibiotics7030053
Hofeditz T, Unsin CE-M, Wiese J, Imhoff JF, Wohlleben W, Grond S, Weber T. Lysoquinone-TH1, a New Polyphenolic Tridecaketide Produced by Expressing the Lysolipin Minimal PKS II in Streptomyces albus. Antibiotics. 2018; 7(3):53. https://doi.org/10.3390/antibiotics7030053
Chicago/Turabian StyleHofeditz, Torben, Claudia Eva-Maria Unsin, Jutta Wiese, Johannes F. Imhoff, Wolfgang Wohlleben, Stephanie Grond, and Tilmann Weber. 2018. "Lysoquinone-TH1, a New Polyphenolic Tridecaketide Produced by Expressing the Lysolipin Minimal PKS II in Streptomyces albus" Antibiotics 7, no. 3: 53. https://doi.org/10.3390/antibiotics7030053
APA StyleHofeditz, T., Unsin, C. E.-M., Wiese, J., Imhoff, J. F., Wohlleben, W., Grond, S., & Weber, T. (2018). Lysoquinone-TH1, a New Polyphenolic Tridecaketide Produced by Expressing the Lysolipin Minimal PKS II in Streptomyces albus. Antibiotics, 7(3), 53. https://doi.org/10.3390/antibiotics7030053





