Assessment of Antimicrobial Resistance and Virulence of Biofilm-Forming Uropathogenic Escherichia coli from Rio de Janeiro
Abstract
1. Introduction
2. Results
2.1. Bacterial Identification, Antimicrobial Susceptibility, and Biofilm Production
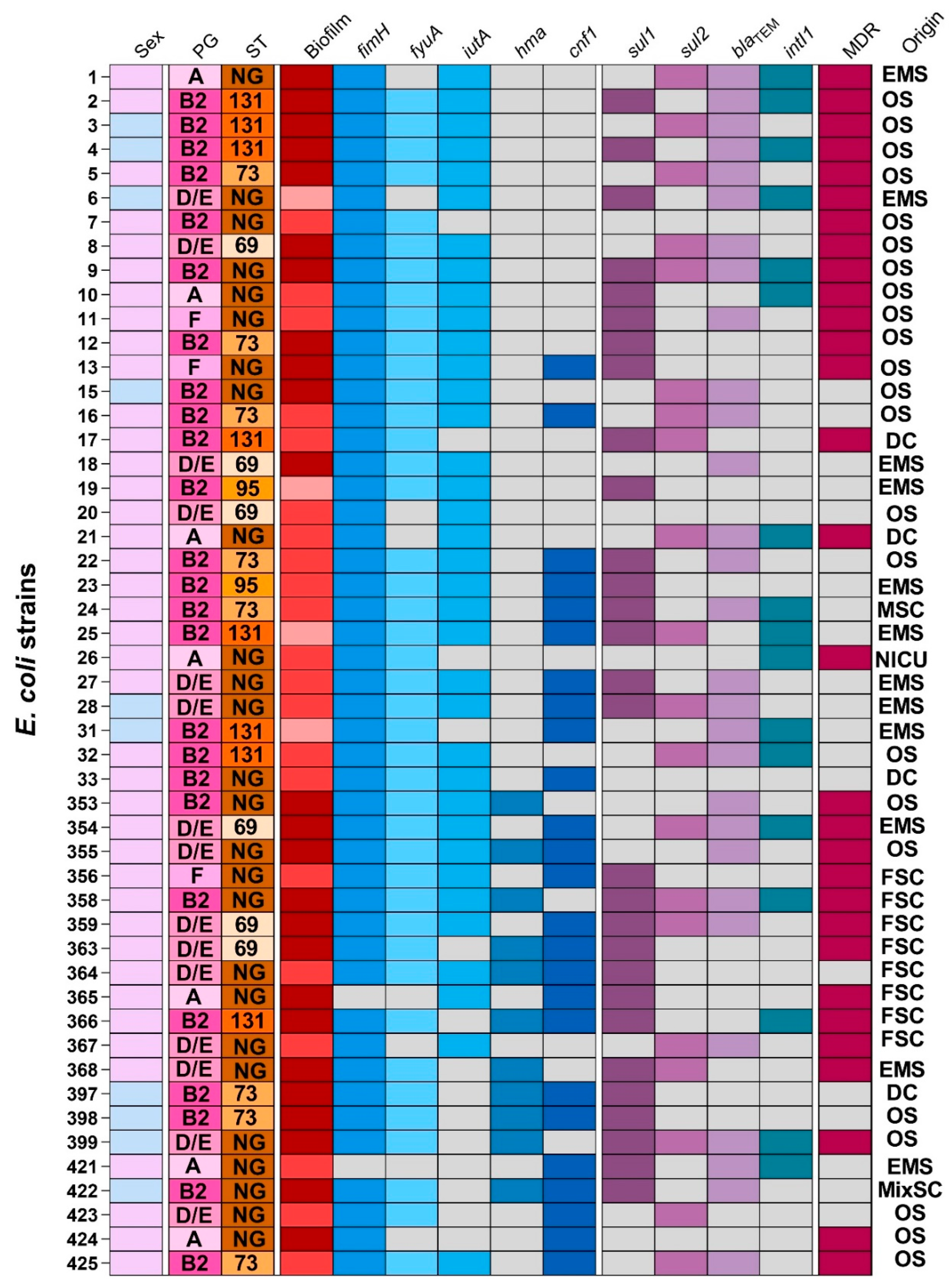
2.2. Resistance and Virulence Genes
2.3. Sequence Types and Phylogenetic Groups
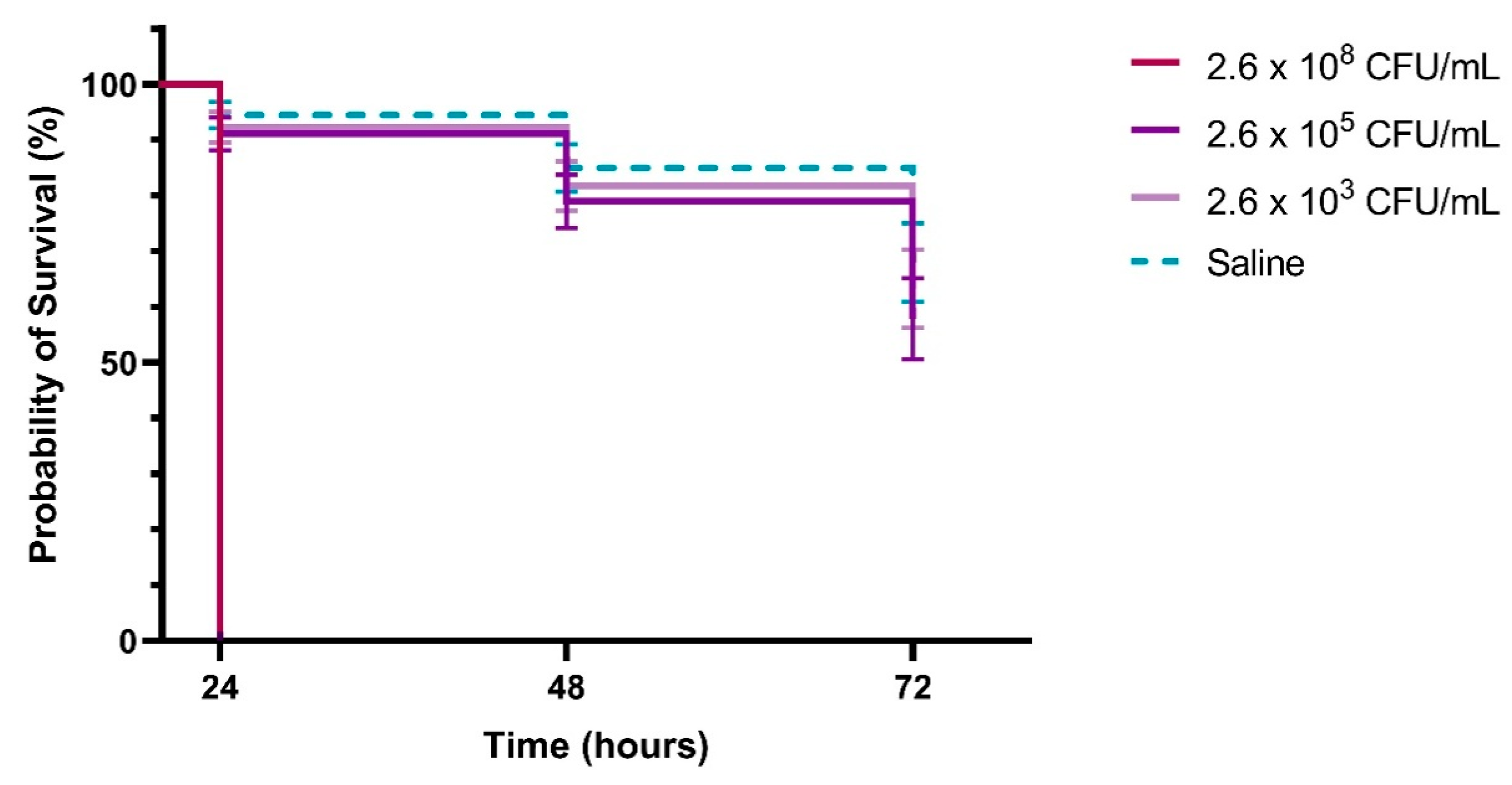
2.4. Larvae Infection
3. Discussion
4. Materials and Methods
4.1. Study Design, Patients, Bacterial Isolates and Ethical Approval
4.2. Bacterial Identification Confirmation
4.3. Evaluation of Biofilm Production
4.4. Antimicrobial Susceptibility Testing
4.5. Antimicrobial Resistance Genes Screening
4.6. Virulence Genes Investigation
4.7. Sequence-Type and Phylogenetic Groups Determination
4.8. Tenebrio Molitor Infection Model
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| UTI | Urinary tract infection |
| UPEC | Uropathogenic Escherichia coli |
| STs | sequence types |
| PCR | polymerase chain reaction |
| CFU | colony-forming units |
| mL | milliliter |
| NG | Not-grouped |
| MDR | multidrug-resistant isolate |
| EMS | Emergency service |
| OS | Outpatient service |
| DC | dialysis center |
| MSC | male surgical clinic |
| NICU | neonatal intensive care unit |
| FSC | female surgical clinic |
| MixSC | mixed surgical clinic |
| h | hour |
| CNF1 | cytotoxic necrotizing factor 1 |
| MALDI-TOF MS | Matrix-Assisted Laser Desorption/Ionization Time-of-Flight mass spectrometry |
| μL | microliter |
| ODc | cutoff value |
| OD | optical density |
| SD | standard deviation |
| CLSI | Clinical & Laboratory Standards Institute |
| LBB | Luria–Bertani broth |
References
- Campos, A.C.C.; Andrade, N.L.; Ferdous, M.; Chlebowicz, M.A.; Santos, C.C.; Correal, J.C.D.; Lo Ten Foe, J.R.; Rosa, A.C.P.; Damasco, P.V.; Friedrich, A.W.; et al. Comprehensive Molecular Characterization of Escherichia coli Isolates from Urine Samples of Hospitalized Patients in Rio de Janeiro, Brazil. Front. Microbiol. 2018, 9, 243, Erratum in Front. Microbiol. 2020, 11, 599031. https://doi.org/10.3389/fmicb.2020.599031. [Google Scholar] [CrossRef]
- Pereira, J.L.; Volcão, L.M.; Klafke, G.B.; Vieira, R.S.; Gonçalves, C.V.; Ramis, I.B.; da-Silva, P.E.A.; von Groll, A. Antimicrobial resistance and molecular characterization of extended-spectrum β-lactamases of Escherichia coli and Klebsiella spp. isolates from urinary tract infections in Southern Brasil. Microb. Drug Resist. 2019, 25, 173–181. [Google Scholar] [CrossRef]
- Nasrollahian, S.; Halaji, M.; Hosseini, A.; Teimourian, M.; Armaki, M.T.; Rajabnia, M.; Gholinia, H.; Pournajaf, A. Genetic Diversity, Carbapenem Resistance Genes, and Biofilm Formation in UPEC Isolated from Patients with Catheter-Associated Urinary Tract Infection in North of Iran. Int. J. Clin. Pract. 2022, 2022, 9520362. [Google Scholar] [CrossRef]
- Wagenlehner, F.M.E.; Bjerklund Johansen, T.E.; Cai, T.; Koves, B.; Kranz, J.; Pilatz, A.; Tandogdu, Z. Epidemiology, definition and treatment of complicated urinary tract infections. Nat. Rev. Urol. 2020, 17, 586–600. [Google Scholar] [CrossRef] [PubMed]
- Terlizzi, M.E.; Gribaudo, G.; Maffei, M.E. Uropathogenic Escherichia coli (UPEC) Infections: Virulence Factors, Bladder Responses, Antibiotic, and Non-Antibiotic Antimicrobial Strategies. Front. Microbiol. 2017, 8, 1566. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Chen, H.; Zheng, Y.; Qu, S.; Wang, H.; Yi, F. Disease burden and long-term trends of urinary tract infections: A worldwide report. Front. Public Health 2022, 10, 888205. [Google Scholar] [CrossRef]
- Reis, A.C.C.; Santos, S.R.S.; Souza, S.C.; Saldanha, M.G.; Pitanga, T.N.; Oliveira, R.R. Ciprofloxacin resistance pattern among bacteria isolated from patients with community-acquired urinary tract infection. Rev. Inst. Med. Trop. Sao Paulo 2016, 58, 53. [Google Scholar] [CrossRef]
- Kumar, S.; Dave, A.; Wolf, B.; Lerma, E.V. Urinary tract infections. Dis. Mon. 2015, 61, 45–59. [Google Scholar] [CrossRef]
- Galindo-Méndez, M. Molecular characterization and antimicrobial susceptibility pattern of extended-spectrum β-lactamase-producing Escherichia coli as cause of community acquired urinary tract infection. Rev. Chil. Infectol. 2018, 35, 29–35. [Google Scholar] [CrossRef]
- Antimicrobial Resistance Collaborators. Global burden of bacterial antimicrobial resistance in 2019: A systematic analysis. Lancet 2022, 399, 629–655. [Google Scholar] [CrossRef]
- Ho, C.S.; Wong, C.T.H.; Aung, T.T.; Lakshminarayanan, R.; Mehta, J.S.; Rauz, S.; McNally, A.; Kintses, B.; Peacock, S.J.; de la Fuente-Nunez, C.; et al. Antimicrobial resistance: A concise update. Lancet Microbe 2025, 6, 100947. [Google Scholar] [CrossRef]
- Gillings, M.R. Integrons: Past, present, and future. Microbiol. Mol. Biol. Rev. 2014, 78, 257–277. [Google Scholar] [CrossRef] [PubMed]
- Gündoğdu, A.; Long, Y.B.; Vollmerhausen, T.L.; Katouli, M. Antimicrobial resistance and distribution of sul genes and integron-associated intI genes among uropathogenic Escherichia coli in Queensland, Australia. J. Med. Microbiol. 2011, 60, 1633–1642. [Google Scholar] [CrossRef] [PubMed]
- Keshavarz, M.; Jo, Y.H.; Patnaik, B.B.; Park, K.B.; Ko, H.J.; Kim, C.E.; Edosa, T.T.; Lee, Y.S.; Han, Y.S. TmRelish is required for regulating the antimicrobial responses to Escherichia coli and Staphylococcus aureus in Tenebrio molitor. Sci. Rep. 2020, 10, 4258. [Google Scholar] [CrossRef] [PubMed]
- Grape, M.; Sundström, L.; Kronvall, G. Sulphonamide Resistance Gene sul3 Found in Escherichia coli Isolates from Human Sources. J. Antimicrob. Chemother. 2003, 52, 1022–1024. [Google Scholar] [CrossRef] [PubMed]
- Muhammed Ali, R.A.; Alshara, J.M.R.; Tuwaij, S.S.; Al-Khilkhali, H.J.B. Study of Antibacterial Chemical Substances and Molecular Investigation among Sulfamethoxazole-Trimethoprim (SXT)-Resistant Escherichia coli Isolates. Rep. Biochem. Mol. Biol. 2022, 11, 166–175. [Google Scholar] [CrossRef]
- Rahman, M.M.; Hossain, M.M.K.; Rubaya, R.; Halder, J.; Karim, M.E.; Bhuiya, A.A.; Khatun, A.; Alam, J. Association of Antibiotic Resistance Traits in Uropathogenic Escherichia coli (UPEC) Isolates. Can. J. Infect. Dis. Med. Microbiol. 2022, 2022, 4251486. [Google Scholar] [CrossRef]
- Chin, J.J.; Lee, H.M.; Lee, S.Y.; Lee, Y.Y.; Chew, C.H. High Carriage of tetA, sul1, sul2 and blaTEM Resistance Genes among the Multidrug-resistant Uropathogenic Escherichia coli (UPEC) Strains from Malaysian Patients. Trop. Life Sci. Res. 2024, 35, 211–225. [Google Scholar] [CrossRef]
- Cardoso, A.M.; Flores, V.R.; do Rosario, G.G.; Succar, J.B.; Berbert, L.C.; Oliveira, M.C.d.F.; Canellas, A.L.B.; Laport, M.S.; Souza, C.R.V.M.; Chagas, T.P.G.; et al. Antimicrobial Susceptibility of Escherichia coli Isolates Causing Community-Acquired Urinary Tract Infections: Comparison of Methods. Microorganisms 2025, 13, 231. [Google Scholar] [CrossRef]
- Soto, S.M.; Smithson, A.; Horcajada, J.P.; Martinez, J.A.; Mensa, J.P.; Vila, J. Implication of biofilm formation in the persistence of urinary tract infection caused by uropathogenic Escherichia coli. Clin. Microbiol. Infect. 2006, 12, 1034–1036. [Google Scholar] [CrossRef]
- Baldiris-Avila, R.; Montes-Robledo, A.; Buelvas-Montes, Y. Phylogenetic Classification, Biofilm-Forming Capacity, Virulence Factors, and Antimicrobial Resistance in Uropathogenic Escherichia coli (UPEC). Curr. Microbiol. 2020, 77, 3361–3370. [Google Scholar] [CrossRef] [PubMed]
- Yun, K.W.; Kim, H.Y.; Park, H.K.; Kim, W.; Lim, I.S. Virulence factors of uropathogenic Escherichia coli of urinary tract infections and asymptomatic bacteriuria in children. J. Microbiol. Immunol. Infect. 2014, 47, 455–461. [Google Scholar] [CrossRef] [PubMed]
- Flores, C.; Ling, J.; Loh, A.; Maset, R.G.; Aw, A.; White, I.J.; Fernando, R.; Rohn, J.L. A human urothelial microtissue model reveals shared colonization and survival strategies between uropathogens and commensals. Sci. Adv. 2023, 9, eadi9834. [Google Scholar] [CrossRef] [PubMed]
- Lindblom, A.; Kiszakiewicz, C.; Kristiansson, E.; Yazdanshenas, S.; Kamenska, N.; Karami, N.; Åhrén, C. The impact of the ST131 clone on recurrent ESBL-producing E. coli urinary tract infection: A prospective comparative study. Sci. Rep. 2022, 12, 10048. [Google Scholar] [CrossRef]
- Li, D.; Elankumaran, P.; Kudinha, T.; Kidsley, A.K.; Trott, D.J.; Jarocki, V.M.; Djordjevic, S.P. Dominance of Escherichia coli sequence types ST73, ST95, ST127 and ST131 in Australian urine isolates: A genomic analysis of antimicrobial resistance and virulence linked to F plasmids. Microb. Genom. 2023, 9, mgen001068. [Google Scholar] [CrossRef]
- Hagelueken, G.; Duthie, F.G.; Florin, N.; Schubert, E.; Schiemann, O. Expression, Purification and Spin Labelling of the Ferrous Iron Transporter FeoB from Escherichia coli BL21 for EPR Studies. Protein Expr. Purif. 2015, 114, 30–36. [Google Scholar] [CrossRef]
- Gatya Al-Mayahie, S.M.; Al-Guranie, D.R.T.; Hussein, A.A.; Bachai, Z.A. Prevalence of common carbapenemase genes and multidrug resistance among uropathogenic Escherichia coli phylogroup B2 isolates from outpatients in Wasit Province/Iraq. PLoS ONE 2022, 17, e0262984. [Google Scholar] [CrossRef]
- Talebi, M.; Najar-Peerayeh, S.; Bakhshi, B. Hidden carbapenem resistance in the community-and hospital-associated OXA-48 gene-carrying uropathogenic Escherichia coli. Gene Rep. 2020, 21, 100897. [Google Scholar] [CrossRef]
- Lau, S.H.; Reddy, S.; Cheesbrough, J.; Bolton, F.J.; Willshaw, G.; Cheasty, T.; Fox, A.J.; Upton, M. Major uropathogenic Escherichia coli strain isolated in the northwest of England identified by multilocus sequence typing. J. Clin. Microbiol. 2008, 46, 1076–1080. [Google Scholar] [CrossRef]
- Johnson, J.R.; Clabots, C.; Porter, S.B.; Bender, T.; Johnston, B.D.; Thuras, P. Intestinal Persistence of Colonizing Escherichia coli Strains, Especially ST131-H30, in Relation to Bacterial and Host Factors. J. Infect. Dis. 2022, 225, 2197–2207. [Google Scholar] [CrossRef]
- Basu, S.; Mukherjee, S.K.; Hazra, A.; Mukherjee, M. Molecular Characterization of Uropathogenic Escherichia coli: Nalidixic Acid and Ciprofloxacin Resistance, Virulent Factors and Phylogenetic Background. J. Clin. Diagn. Res. 2013, 7, 2727–2731. [Google Scholar] [CrossRef]
- Alghoribi, M.F.; Gibreel, T.M.; Dodgson, A.R.; Beatson, S.A.; Upton, M. Galleria mellonella infection model demonstrates high lethality of ST69 and ST127 uropathogenic E. coli. PLoS ONE 2014, 9, e101547. [Google Scholar] [CrossRef]
- Canellas, A.L.B.; Lopes, I.R.; Mello, M.P.; Paranhos, R.; de Oliveira, B.F.R.; Laport, M.S. Vibrio Species in an Urban Tropical Estuary: Antimicrobial Susceptibility, Interaction with Environmental Parameters, and Possible Public Health Outcomes. Microorganisms 2021, 9, 1007. [Google Scholar] [CrossRef] [PubMed]
- Alves, G.D.S.O.; Canellas, A.L.B.; Gallo, M.N.; Vinzon, S.B.; Laport, M.S. In treacherous waters: Detection of colistin-resistant bacteria in water and plastic litter from a recreational estuary. Lett. Appl. Microbiol. 2024, 77, ovae082. [Google Scholar] [CrossRef] [PubMed]
- Stepanović, S.; Vuković, D.; Hola, V.; Di Bonaventura, G.; Djukić, S.; Cirković, I.; Ruzicka, F. Quantification of biofilm in microtiter plates: Overview of testing conditions and practical recommendations for assessment of biofilm production by staphylococci. APMIS 2007, 115, 891–899. [Google Scholar] [CrossRef] [PubMed]
- CLSI (Clinical and Laboratory Standards Institute). Performance Standards for Antimicrobial Susceptibility Testing. In CLSI Supplement M100, 30th ed.; CLSI: Wayne, PA, USA, 2020; ISBN 978-1-68440-067-6. [Google Scholar]
- Richter, L.; Plessis, E.M.; Duvenage, S.; Korsten, L. Occurrence, Phenotypic and Molecular Characterization of Extended-Spectrum- and AmpC- β-Lactamase Producing Enterobacteriaceae Isolated from Selected Commercial Spinach Supply Chains in South Africa. Front. Microbiol. 2020, 11, 638. [Google Scholar] [CrossRef]
- Karam, M.R.A.; Habibi, M.; Bouzari, S. Relationships between virulence factors and antimicrobial resistance among Escherichia coli isolated from urinary tract infections and commensal isolates in Tehran, Iran. Osong Public Health Res. Perspect. 2018, 9, 217–224. [Google Scholar] [CrossRef]
- Doumith, M.; Day, M.; Ciesielczuk, H.; Hope, R.; Underwood, A.; Reynolds, R.; Wain, J.; Livermore, D.M.; Woodford, N. Rapid identification of major Escherichia coli sequence types causing urinary tract and bloodstream infections. J. Clin. Microbiol. 2015, 53, 160–166. [Google Scholar] [CrossRef]
- Clermont, O.; Christenson, J.K.; Denamur, E.; Gordon, D.M. The Clermont Escherichia coli phylo-typing method revisited: Improvement of specificity and detection of new phylo-groups. Environ. Microbiol. Rep. 2013, 5, 58–65. [Google Scholar] [CrossRef]
| Primers Sequences | Annealing Temperature (°C) | Expected Fragment Size (bp) | |
|---|---|---|---|
| (1) ireaA | |||
| F | 5′-ATGAAGAACAAATATATC-3′ | 53 R1 | 2500 |
| R | 5′-GAAGGATACTCTTACATT-3′ | ||
| (2) fyuA | |||
| F | 5′-ATGAAAATGACACGGCT-3′ | 56 R1 | 2200 |
| R | 5′-GAAGAAATCAATTCGCG-3′ | ||
| (3) hma | |||
| F | 5′-ATGGTTAAAGATACAATC-3′ | 56 R1 | 1850 |
| R | 5′-CCACTGATAACGGGTAT-3′ | ||
| (4) iutA | |||
| F | 5′-ATGAAAATGACACGGCT-3′ | 60 R1 | 1750 |
| R | 5′-GAAGAAATCAATTCGCG-3′ | ||
| (5) fimH | |||
| F | 5′-TGCAGAACGGATAAGCCGTGG-3′ | 60 R2 | 508 |
| R | 5′-GCAGTCACCTGCCCTCCGGTA-3′ | ||
| (6) cnf1 | |||
| F | 5′-AAGATGGAGTTTCCTATGCAG-3′ | 54 R2 | 543 |
| R | 5′-TCAGAGTCCTGCCCTCATTAT-3′ | ||
| ST Target | Primer Name | Primers Sequences | Fragment Size (bp) |
|---|---|---|---|
| ST73 | ST73_for | TGGTTTTACCATTTTGTCGGA | 490 |
| ST73_rev | GGAAATCGTTGATGTTGGCT | ||
| ST131 | ST131_for | GACTGCATTTCGTCGCCATA | 310 |
| ST131_rev | CCGGCGGCATCATAATGAAA | ||
| ST95 | ST95_for | ACTAATCAGGATGGCGAGAC | 200 |
| ST95_rev | ATCACGCCCATTAATCCAGT | ||
| ST69 | ST69_for | ATCTGGAGGCAACAAGCATA | 104 |
| ST69_rev | AGAGAAAGGGCGTTCAGAAT |
| Phylogroups of E. coli sensu stricto | Presence of PCR-Generated Fragments | |||
|---|---|---|---|---|
| arpA | chuA | yjaA | TspE4.C2 | |
| A | + | − | − | − |
| + | − | + | − | |
| B1 | + | − | + | + |
| B2 | − | + | + | + |
| − | + | + | − | |
| − | + | − | + | |
| C | + | − | + | − |
| D | + | + | − | − |
| + | + | − | + | |
| E | + | + | − | − |
| + | + | − | + | |
| + | + | + | − | |
| F | − | + | − | − |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Oliveira, M.C.F.; Canellas, A.L.B.; Berbert, L.C.; Cardoso, A.M.; Silva, V.A.; Garutti, S.S.T.; Rangel, D.H.F.; Dias, R.C.S.; Perini, J.A.; Souza, C.R.V.M.; et al. Assessment of Antimicrobial Resistance and Virulence of Biofilm-Forming Uropathogenic Escherichia coli from Rio de Janeiro. Antibiotics 2025, 14, 869. https://doi.org/10.3390/antibiotics14090869
Oliveira MCF, Canellas ALB, Berbert LC, Cardoso AM, Silva VA, Garutti SST, Rangel DHF, Dias RCS, Perini JA, Souza CRVM, et al. Assessment of Antimicrobial Resistance and Virulence of Biofilm-Forming Uropathogenic Escherichia coli from Rio de Janeiro. Antibiotics. 2025; 14(9):869. https://doi.org/10.3390/antibiotics14090869
Chicago/Turabian StyleOliveira, Maria Clara F., Anna Luiza B. Canellas, Lidiane C. Berbert, Alexander M. Cardoso, Vitoria A. Silva, Samantha S. T. Garutti, Débora Hosana F. Rangel, Rubens Clayton S. Dias, Jamila Alessandra Perini, Claudia R. V. M. Souza, and et al. 2025. "Assessment of Antimicrobial Resistance and Virulence of Biofilm-Forming Uropathogenic Escherichia coli from Rio de Janeiro" Antibiotics 14, no. 9: 869. https://doi.org/10.3390/antibiotics14090869
APA StyleOliveira, M. C. F., Canellas, A. L. B., Berbert, L. C., Cardoso, A. M., Silva, V. A., Garutti, S. S. T., Rangel, D. H. F., Dias, R. C. S., Perini, J. A., Souza, C. R. V. M., Chagas, T. P. G., Laport, M. S., & Pellegrino, F. L. P. C. (2025). Assessment of Antimicrobial Resistance and Virulence of Biofilm-Forming Uropathogenic Escherichia coli from Rio de Janeiro. Antibiotics, 14(9), 869. https://doi.org/10.3390/antibiotics14090869











