Tracing the Evolutionary Pathways of Serogroup O78 Avian Pathogenic Escherichia coli
Abstract
1. Introduction
2. Results and Discussion
2.1. O78 APEC Strains Identified during 2012–2020
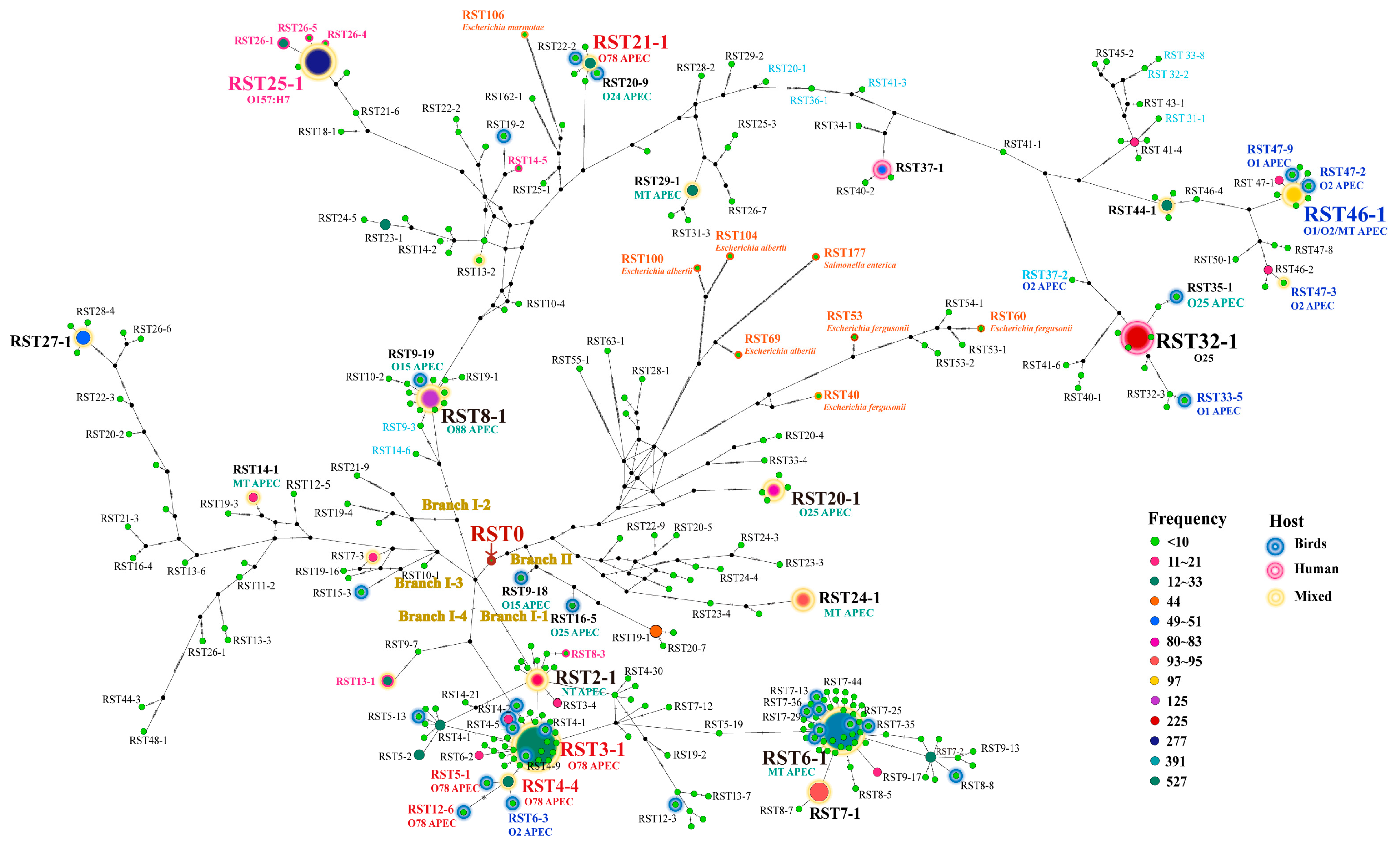
2.2. Modified rpoB Sequence Typing (RSTing) of E. coli Strains
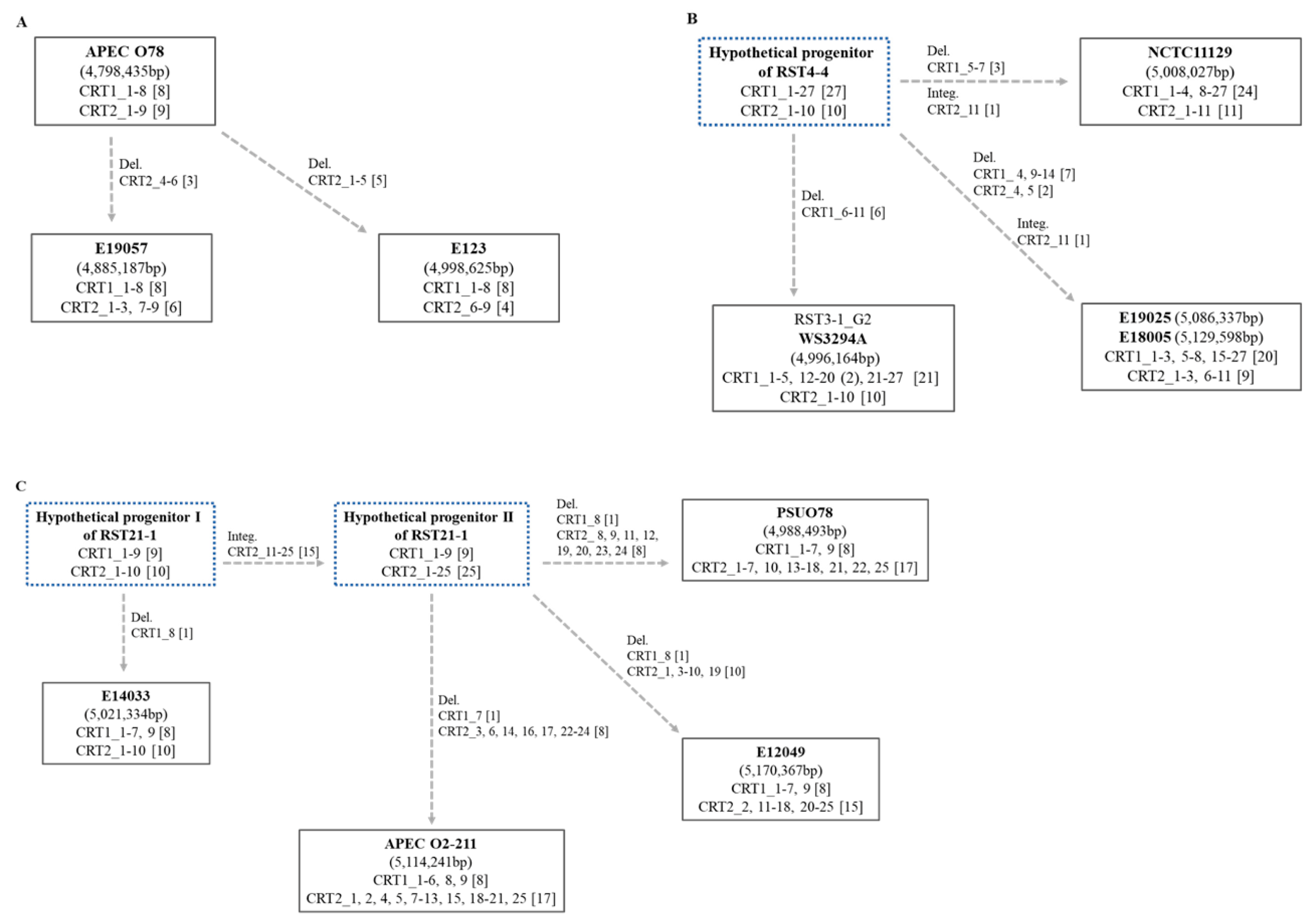
2.3. Hypothetical Evolutionary Pathways of E. coli and O78 APEC Strains
2.4. General Genomic Information of the Six Representative O78 APEC Strains from RST3-1, RST4-4, and RST21-1
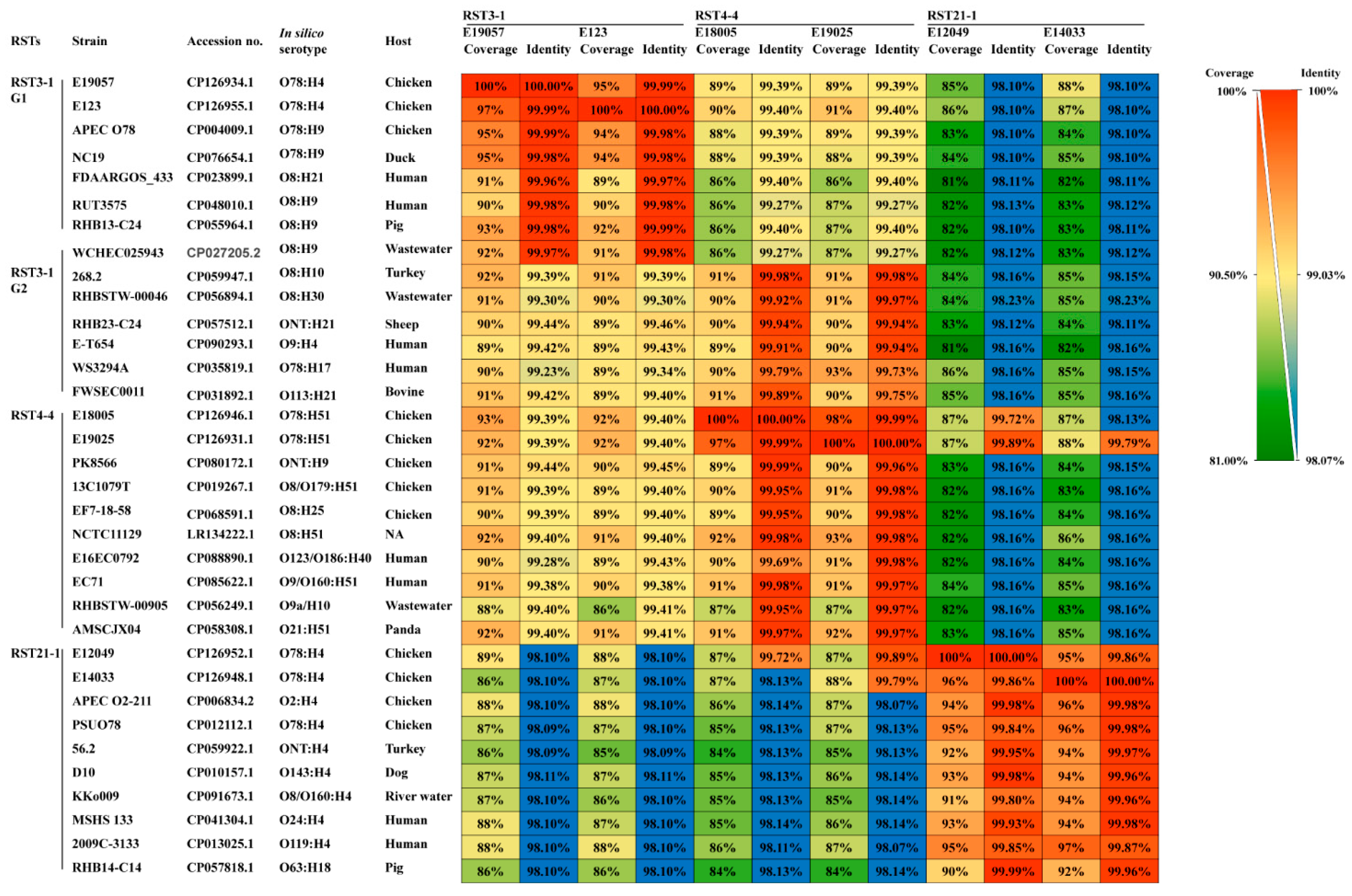
2.5. Correlation of Genomic Sequence Coverage/Identity and CRISPR Contents between the Same RSTs Containing Major O78 Strains
| E19057 | E123 | APEC O78 | E18005 | E19025 | NCTC11129 | E12049 | E14033 | PSUO78 | |
|---|---|---|---|---|---|---|---|---|---|
| Accession no. | CP126934.1 | CP126955.1 | CP004009.1 | CP126946.1 | CP126931.1 | LR134222.1 | CP126952.1 | CP126948.1 | CP012112.1 |
| Origin | Chicken | Chicken | Turkey | Chicken | Chicken | unknown | Chicken | Chicken | Chicken |
| RST | 3-1 | 3-1 | 3-1 | 4-4 | 4-4 | 4-4 | 21-1 | 21-1 | 21-1 |
| MLST (Achman) | ST23 | ST369 | ST23 | ST155 | ST155 | ST155 | ST11749 | ST48 | ST5940 |
| mPPT | 2-5 | 2-3-4 | 3-4 | 1-2-3-5(2)-6 | 1-2-3(2)-4-5-6 | 2-3-4-5 | 1-2-5 | 1-2-4-5 | 2-4-5-7 |
| Serotype | O78:H4 | O78:H4 | O78:H9 | O78:H51 | O78:H51 | O8, O60:H51 | O78:H4 | O78:H4 | O78:H4 |
| Total Length (bp) | 4,885,187 | 4,998,625 | 4,798,435 | 5,129,598 | 5,086,337 | 5,008,027 | 5,170,367 | 5,021,334 | 4,988,493 |
| GC Content (%) | 50.60% | 50.64% | 50.70% | 50.64% | 50.70% | 50.78% | 50.74% | 50.79% | 50.80% |
| No. of CDSs | 4586 | 4690 | 4504 | 4771 | 4751 | 4697 | 4788 | 4658 | 4791 |
| No. of rRNA | 22 | 22 | 19 | 21 | 22 | 22 | 22 | 22 | 22 |
| No. of tRNA | 86 | 82 | 88 | 92 | 95 | 89 | 97 | 90 | 88 |
| No. of CRISPRS | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| Coding ratio (%) | 87.30% | 87.10% | 87.60% | 87.00% | 87.20% | 87.10% | 86.80% | 87.20% | 86.60% |
| Plasmid 1 a | pEND_Eco19057-1 (CP126935.1) (IncI1-1 (alpha), 100%; 110,406 bp) | pEND_Eco 123-1 (CP126956.1) (IncFIB, 98.39%; 121,788 bp) | 217,830 bp | pEND_Eco 18005 (CP126947.1) (IncFIB, 98.39%; 204,838 bp) | pEND_Eco 19025-1 (CP126932.1) (IncFIB, 98.39%; 160,439 bp) | No data | pEND_Eco 12049-1 (CP126953.1) (IncFIB, 99.12%; 155,857 bp) | pEND_Eco 14033-1 (CP126949.1) (IncFIB, 98.39%; 145,212 bp) | pPSUO78_1 (CP012113.1) (132,464 bp) |
| Plasmid 2 | pEND_Eco 19057-2 (CP126936.1) (IncFIB, 98.39%; 104,962 bp) | pEND_Eco 123-2 (CP126957.1) (No hit; 68,426 bp) | 113,260 bp | pEND_Eco 19025-2 (CP126933.1) (IncY, 99.38%; 90,722 bp) | pEND_Eco 12049-2 (CP126954.1) (IncI1-1(alpha), 100%; 114,229 bp) | pEND_Eco 14033-2 (CP126950.1) (IncI1-1 (alpha), 100%; 115,173 bp) | pPSUO78_2 (CP012114.1) (109,613 bp) | ||
| Plasmid 3 | pEND_Eco 14033-3 (CP126951.1) (IncFIB (H89-PhagePlasmid), 98.7%; 108,542 bp) | ||||||||
| Plasmid 4 | pEND_Eco 14033-4 (CP130557.1) (IncI2(delta), 97.15%; 61,041 bp) | ||||||||
| No. Genomic islands (GI) b | 69 | 70 | 47 | 94 | 88 | 61 | 94 | 77 | 80 |
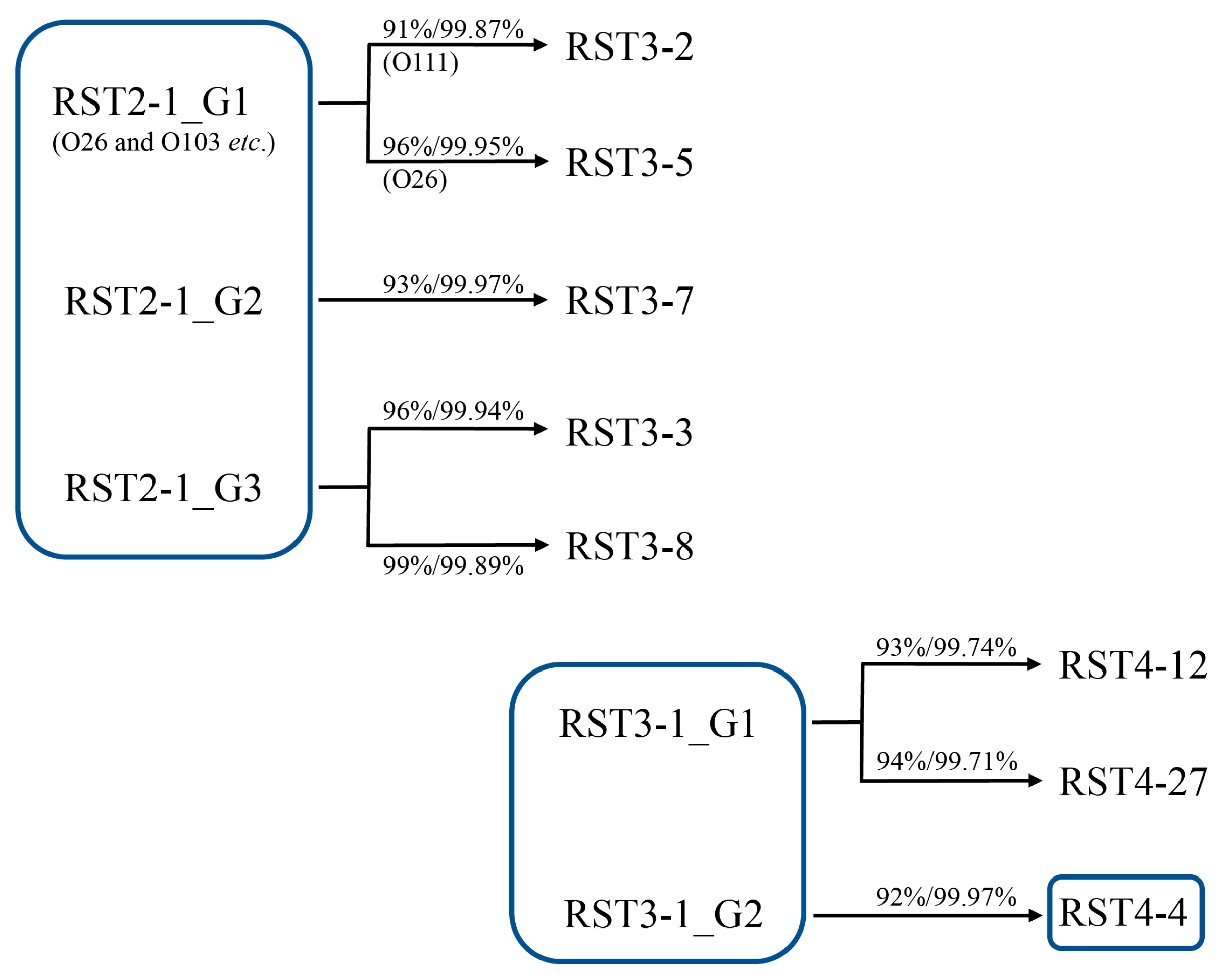
2.6. Evolutionary Linkage between Early-Appearing RSTs
2.7. Evolution of O Serogroups in Different RSTs
2.8. Profiling Virulence Genes of O78 APEC Strains
2.9. Antibiograms of O78 APEC Strains
2.10. Profiles of Microcins and Colicins of O78 APEC Strains
3. Materials and Methods
3.1. Bacterial Strains and Identification
3.2. Serotyping and Phylogrouping of E. coli
3.3. rpoB Sequence Typing (RSTing) and Molecular Prophage Typing (mPPTing)
3.4. Molecular Pathotyping
3.5. Antibiotic Susceptibility Testing and Molecular Profiling of Antibiotic Resistance Genes
3.6. Comparative Genomics
3.7. Identification of Bacteriocin Genes in O78 APEC
3.8. Antimicrobial Activity Test of Bacteriocin
3.9. Statistical Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Nataro, J.P.; Kaper, J.B. Diarrheagenic Escherichia coli. Clin. Microbiol. Rev. 1998, 11, 142–201. [Google Scholar] [CrossRef] [PubMed]
- Russo, T.A.; Johnson, J.R. Proposal for a new inclusive designation for extraintestinal pathogenic isolates of Escherichia coli: ExPEC. J. Infect. Dis. 2000, 181, 1753–1754. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; Furevi, A.; Perepelov, A.V.; Guo, X.; Cao, H.; Wang, Q.; Reeves, P.R.; Knirel, Y.A.; Wang, L.; Widmalm, G. Structure and genetics of Escherichia coli O antigens. FEMS Microbiol. Rev. 2020, 44, 655–683. [Google Scholar] [CrossRef]
- Kim, Y.B.; Yoon, M.Y.; Ha, J.S.; Seo, K.W.; Noh, E.B.; Son, S.H.; Lee, Y.J. Molecular characterization of avian pathogenic Escherichia coli from broiler chickens with colibacillosis. Poult. Sci. 2020, 99, 1088–1095. [Google Scholar] [CrossRef]
- Jeong, Y.W.; Kim, T.E.; Kim, J.H.; Kwon, H.J. Pathotyping avian pathogenic Escherichia coli strains in Korea. J. Vet. Sci. 2012, 13, 145–152. [Google Scholar] [CrossRef] [PubMed]
- Kwon, H.-J.; Seong, W.-J.; Kim, T.-E.; Won, Y.-J.; Kim, J.-H. rpoB gene sequencing for phylogenetic analysis of avian pathogenic Escherichia coli. Korean J. Vet. Res. 2015, 55, 31–39. [Google Scholar] [CrossRef]
- Kwon, H.J.; Seong, W.J.; Kim, J.H. Molecular prophage typing of avian pathogenic Escherichia coli. Vet. Microbiol. 2013, 162, 785–792. [Google Scholar] [CrossRef]
- Garrett, S.C. Pruning and Tending Immune Memories: Spacer Dynamics in the CRISPR Array. Front. Microbiol. 2021, 12, 664299. [Google Scholar] [CrossRef]
- Shin, H.; Kim, Y.; Unno, T.; Hur, H.G. Prevalence and Characterization of CRISPR Locus 2.1 Spacers in Escherichia coli Isolates Obtained from Feces of Animals and Humans. Microbiol. Spectr. 2023, 11, e0493422. [Google Scholar] [CrossRef]
- Zhou, Z.; Charlesworth, J.; Achtman, M. HierCC: A multi-level clustering scheme for population assignments based on core genome MLST. Bioinformatics 2021, 37, 3645–3646. [Google Scholar] [CrossRef]
- Ding, W.; Baumdicker, F.; Neher, R.A. panX: Pan-genome analysis and exploration. Nucleic Acids Res. 2018, 46, e5. [Google Scholar] [CrossRef]
- Ko, D.S.; Kim, D.; Kim, E.K.; Kim, J.H.; Kwon, H.J. Evolution of a major bovine mastitic genotype (rpoB sequence type 10-2) of Staphylococcus aureus in cows. J. Microbiol. 2019, 57, 587–596. [Google Scholar] [CrossRef]
- Ko, D.S.; Kim, N.H.; Kim, E.K.; Ha, E.J.; Ro, Y.H.; Kim, D.; Choi, K.S.; Kwon, H.J. Comparative genomics of bovine mastitis-origin Staphylococcus aureus strains classified into prevalent human genotypes. Res. Vet. Sci. 2021, 139, 67–77. [Google Scholar] [CrossRef]
- Guinane, C.M.; Ben Zakour, N.L.; Tormo-Mas, M.A.; Weinert, L.A.; Lowder, B.V.; Cartwright, R.A.; Smyth, D.S.; Smyth, C.J.; Lindsay, J.A.; Gould, K.A.; et al. Evolutionary genomics of Staphylococcus aureus reveals insights into the origin and molecular basis of ruminant host adaptation. Genome Biol. Evol. 2010, 2, 454–466. [Google Scholar] [CrossRef]
- Herron-Olson, L.; Fitzgerald, J.R.; Musser, J.M.; Kapur, V. Molecular correlates of host specialization in Staphylococcus aureus. PLoS ONE 2007, 2, e1120. [Google Scholar] [CrossRef] [PubMed]
- Lowder, B.V.; Guinane, C.M.; Ben Zakour, N.L.; Weinert, L.A.; Conway-Morris, A.; Cartwright, R.A.; Simpson, A.J.; Rambaut, A.; Nubel, U.; Fitzgerald, J.R. Recent human-to-poultry host jump, adaptation, and pandemic spread of Staphylococcus aureus. Proc. Natl. Acad. Sci. USA 2009, 106, 19545–19550. [Google Scholar] [CrossRef] [PubMed]
- Viana, D.; Comos, M.; McAdam, P.R.; Ward, M.J.; Selva, L.; Guinane, C.M.; Gonzalez-Munoz, B.M.; Tristan, A.; Foster, S.J.; Fitzgerald, J.R.; et al. A single natural nucleotide mutation alters bacterial pathogen host tropism. Nat. Genet. 2015, 47, 361–366. [Google Scholar] [CrossRef] [PubMed]
- Dahal, R.H.; Choi, Y.J.; Kim, S.; Kim, J. Differentiation of Escherichia fergusonii and Escherichia coli Isolated from Patients with Inflammatory Bowel Disease/Ischemic Colitis and Their Antimicrobial Susceptibility Patterns. Antibiotics 2023, 12, 154. [Google Scholar] [CrossRef]
- Gomes, T.A.T.; Ooka, T.; Hernandes, R.T.; Yamamoto, D.; Hayashi, T. Escherichia albertii Pathogenesis. EcoSal Plus 2020, 9. [Google Scholar] [CrossRef] [PubMed]
- Cobo-Simon, M.; Hart, R.; Ochman, H. Escherichia coli: What Is and Which Are? Mol. Biol. Evol. 2023, 40, msac273. [Google Scholar] [CrossRef] [PubMed]
- Gordienko, E.N.; Kazanov, M.D.; Gelfand, M.S. Evolution of pan-genomes of Escherichia coli, Shigella spp., and Salmonella enterica. J. Bacteriol. 2013, 195, 2786–2792. [Google Scholar] [CrossRef] [PubMed]
- Kim, N.H.; Ha, E.J.; Ko, D.S.; Lee, C.Y.; Kim, J.H.; Kwon, H.J. Molecular evolution of Salmonella enterica subsp. enterica serovar Gallinarum biovar Gallinarum in the field. Vet. Microbiol. 2019, 235, 63–70. [Google Scholar] [CrossRef] [PubMed]
- Krakauer, D.C.; Plotkin, J.B. Redundancy, antiredundancy, and the robustness of genomes. Proc. Natl. Acad. Sci. USA 2002, 99, 1405–1409. [Google Scholar] [CrossRef] [PubMed]
- Smith, H.W. Persistence of tetracycline resistance in pig E. coli. Nature 1975, 258, 628–630. [Google Scholar] [CrossRef] [PubMed]
- Crossman, L.C.; Chaudhuri, R.R.; Beatson, S.A.; Wells, T.J.; Desvaux, M.; Cunningham, A.F.; Petty, N.K.; Mahon, V.; Brinkley, C.; Hobman, J.L.; et al. A commensal gone bad: Complete genome sequence of the prototypical enterotoxigenic Escherichia coli strain H10407. J. Bacteriol. 2010, 192, 5822–5831. [Google Scholar] [CrossRef]
- Johnson, T.J.; Kariyawasam, S.; Wannemuehler, Y.; Mangiamele, P.; Johnson, S.J.; Doetkott, C.; Skyberg, J.A.; Lynne, A.M.; Johnson, J.R.; Nolan, L.K. The genome sequence of avian pathogenic Escherichia coli strain O1:K1:H7 shares strong similarities with human extraintestinal pathogenic E. coli genomes. J. Bacteriol. 2007, 189, 3228–3236. [Google Scholar] [CrossRef]
- Severinov, K.; Soushko, M.; Goldfarb, A.; Nikiforov, V. RifR mutations in the beginning of the Escherichia coli rpoB gene. Mol. Gen. Genet. 1994, 244, 120–126. [Google Scholar] [CrossRef]
- Doolittle, R.F.; Feng, D.F.; Tsang, S.; Cho, G.; Little, E. Determining divergence times of the major kingdoms of living organisms with a protein clock. Science 1996, 271, 470–477. [Google Scholar] [CrossRef]
- Bandelt, H.J.; Forster, P.; Rohl, A. Median-joining networks for inferring intraspecific phylogenies. Mol. Biol. Evol. 1999, 16, 37–48. [Google Scholar] [CrossRef]
- Leigh, J.W.; Bryant, D.; Nakagawa, S. popart: Full-feature software for haplotype network construction. Methods Ecol. Evol. 2015, 6, 1110–1116. [Google Scholar] [CrossRef]
- Rasko, D.A.; Rosovitz, M.J.; Myers, G.S.; Mongodin, E.F.; Fricke, W.F.; Gajer, P.; Crabtree, J.; Sebaihia, M.; Thomson, N.R.; Chaudhuri, R.; et al. The pangenome structure of Escherichia coli: Comparative genomic analysis of E. coli commensal and pathogenic isolates. J. Bacteriol. 2008, 190, 6881–6893. [Google Scholar] [CrossRef] [PubMed]
- Thomas, C.M.; Thomson, N.R.; Cerdeno-Tarraga, A.M.; Brown, C.J.; Top, E.M.; Frost, L.S. Annotation of plasmid genes. Plasmid 2017, 91, 61–67. [Google Scholar] [CrossRef] [PubMed]
- Redondo-Salvo, S.; Bartomeus-Penalver, R.; Vielva, L.; Tagg, K.A.; Webb, H.E.; Fernandez-Lopez, R.; de la Cruz, F. COPLA, a taxonomic classifier of plasmids. BMC Bioinform. 2021, 22, 390. [Google Scholar] [CrossRef]
- Touchon, M.; Charpentier, S.; Clermont, O.; Rocha, E.P.; Denamur, E.; Branger, C. CRISPR distribution within the Escherichia coli species is not suggestive of immunity-associated diversifying selection. J. Bacteriol. 2011, 193, 2460–2467. [Google Scholar] [CrossRef]
- Gupta, R.S. The natural evolutionary relationships among prokaryotes. Crit. Rev. Microbiol. 2000, 26, 111–131. [Google Scholar] [CrossRef] [PubMed]
- Lerouge, I.; Vanderleyden, J. O-antigen structural variation: Mechanisms and possible roles in animal/plant–microbe interactions. FEMS Microbiol. Rev. 2002, 26, 17–47. [Google Scholar] [CrossRef]
- DebRoy, C.; Fratamico, P.M.; Yan, X.; Baranzoni, G.; Liu, Y.; Needleman, D.S.; Tebbs, R.; O’Connell, C.D.; Allred, A.; Swimley, M.; et al. Comparison of O-Antigen Gene Clusters of All O-Serogroups of Escherichia coli and Proposal for Adopting a New Nomenclature for O-Typing. PLoS ONE 2016, 11, e0147434. [Google Scholar] [CrossRef]
- Ogura, Y.; Ooka, T.; Iguchi, A.; Toh, H.; Asadulghani, M.; Oshima, K.; Kodama, T.; Abe, H.; Nakayama, K.; Kurokawa, K.; et al. Comparative genomics reveal the mechanism of the parallel evolution of O157 and non-O157 enterohemorrhagic Escherichia coli. Proc. Natl. Acad. Sci. USA 2009, 106, 17939–17944. [Google Scholar] [CrossRef]
- Wang, L.; Huskic, S.; Cisterne, A.; Rothemund, D.; Reeves, P.R. The O-antigen gene cluster of Escherichia coli O55:H7 and identification of a new UDP-GlcNAc C4 epimerase gene. J. Bacteriol. 2002, 184, 2620–2625. [Google Scholar] [CrossRef]
- Jeong, J.; Lee, J.Y.; Kang, M.S.; Lee, H.J.; Kang, S.I.; Lee, O.M.; Kwon, Y.K.; Kim, J.H. Comparative Characteristics and Zoonotic Potential of Avian Pathogenic Escherichia coli (APEC) Isolates from Chicken and Duck in South Korea. Microorganisms 2021, 9, 946. [Google Scholar] [CrossRef]
- Kim, T.-E.; Jeong, Y.-W.; Cho, S.-H.; Kim, S.-J.; Kwon, H.-J. Chronological study of antibiotic resistances and their relevant genes in Korean avian pathogenic Escherichia coli isolates. J. Clin. Microbiol. 2007, 45, 3309–3315. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Agunos, A.; Leger, D.F.; Carson, C.A.; Gow, S.P.; Bosman, A.; Irwin, R.J.; Reid-Smith, R.J. Antimicrobial use surveillance in broiler chicken flocks in Canada, 2013-2015. PLoS ONE 2017, 12, e0179384. [Google Scholar] [CrossRef] [PubMed]
- Verrette, L.; Fairbrother, J.M.; Boulianne, M. Effect of Cessation of Ceftiofur and Substitution with Lincomycin-Spectinomycin on Extended-Spectrum-beta-Lactamase/AmpC Genes and Multidrug Resistance in Escherichia coli from a Canadian Broiler Production Pyramid. Appl. Environ. Microbiol. 2019, 85, e00037-19. [Google Scholar] [CrossRef]
- de Los Santos, E.; Lavina, M.; Poey, M.E. Strict relationship between class 1 integrons and resistance to sulfamethoxazole in Escherichia coli. Microb. Pathog. 2021, 161, 105206. [Google Scholar] [CrossRef]
- Shis, D.L.; Bennett, M.R.; Igoshin, O.A. Dynamics of Bacterial Gene Regulatory Networks. Annu. Rev. Biophys. 2018, 47, 447–467. [Google Scholar] [CrossRef]
- Molstad, S.; Erntell, M.; Hanberger, H.; Melander, E.; Norman, C.; Skoog, G.; Lundborg, C.S.; Soderstrom, A.; Torell, E.; Cars, O. Sustained reduction of antibiotic use and low bacterial resistance: 10-year follow-up of the Swedish Strama programme. Lancet Infect. Dis. 2008, 8, 125–132. [Google Scholar] [CrossRef] [PubMed]
- Nikaido, H. Molecular Basis of Bacterial Outer Membrane Permeability Revisited. Microbiol. Mol. Biol. Rev. 2003, 67, 593–656. [Google Scholar] [CrossRef]
- Blair, J.M.; Webber, M.A.; Baylay, A.J.; Ogbolu, D.O.; Piddock, L.J. Molecular mechanisms of antibiotic resistance. Nat. Rev. Microbiol. 2015, 13, 42–51. [Google Scholar] [CrossRef]
- Hall, C.W.; Mah, T.F. Molecular mechanisms of biofilm-based antibiotic resistance and tolerance in pathogenic bacteria. FEMS Microbiol. Rev. 2017, 41, 276–301. [Google Scholar] [CrossRef]
- Gordon, D.M.; O’Brien, C.L. Bacteriocin diversity and the frequency of multiple bacteriocin production in Escherichia coli. Microbiology 2006, 152, 3239–3244. [Google Scholar] [CrossRef]
- Baquero, F.; Lanza, V.F.; Baquero, M.R.; Del Campo, R.; Bravo-Vazquez, D.A. Microcins in Enterobacteriaceae: Peptide Antimicrobials in the Eco-Active Intestinal Chemosphere. Front. Microbiol. 2019, 10, 2261. [Google Scholar] [CrossRef]
- Cameron, A.; Zaheer, R.; Adator, E.H.; Barbieri, R.; Reuter, T.; McAllister, T.A. Bacteriocin Occurrence and Activity in Escherichia coli Isolated from Bovines and Wastewater. Toxins 2019, 11, 475. [Google Scholar] [CrossRef] [PubMed]
- Weisburg, W.G.; Barns, S.M.; Pelletier, D.A.; Lane, D.J. 16S ribosomal DNA amplification for phylogenetic study. J. Bacteriol. 1991, 173, 697–703. [Google Scholar] [CrossRef] [PubMed]
- DebRoy, C.; Fratamico, P.M.; Roberts, E. Molecular serogrouping of Escherichia coli. Anim. Health Res. Rev. 2018, 19, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Joensen, K.G.; Tetzschner, A.M.; Iguchi, A.; Aarestrup, F.M.; Scheutz, F. Rapid and Easy In Silico Serotyping of Escherichia coli Isolates by Use of Whole-Genome Sequencing Data. J. Clin. Microbiol. 2015, 53, 2410–2426. [Google Scholar] [CrossRef] [PubMed]
- Dong, A.; Liu, C.; Hua, X.; Yu, Y.; Guo, Y.; Wang, D.; Liu, X.; Chen, H.; Wang, H.; Zhu, L. Bioinformatic analysis of structures and encoding genes of Escherichia coli surface polysaccharides sheds light on the heterologous biosynthesis of glycans. BMC Genom. 2023, 24, 168. [Google Scholar] [CrossRef]
- Clermont, O.; Christenson, J.K.; Denamur, E.; Gordon, D.M. The Clermont Escherichia coli phylo-typing method revisited: Improvement of specificity and detection of new phylo-groups. Environ. Microbiol. Rep. 2013, 5, 58–65. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Rozas, J.; Ferrer-Mata, A.; Sanchez-DelBarrio, J.C.; Guirao-Rico, S.; Librado, P.; Ramos-Onsins, S.E.; Sanchez-Gracia, A. DnaSP 6: DNA Sequence Polymorphism Analysis of Large Data Sets. Mol. Biol. Evol. 2017, 34, 3299–3302. [Google Scholar] [CrossRef]
- Huelsenbeck, J.P.; Ronquist, F. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 2001, 17, 754–755. [Google Scholar] [CrossRef]
- Ewers, C.; Li, G.; Wilking, H.; Kiessling, S.; Alt, K.; Antao, E.M.; Laturnus, C.; Diehl, I.; Glodde, S.; Homeier, T.; et al. Avian pathogenic, uropathogenic, and newborn meningitis-causing Escherichia coli: How closely related are they? Int. J. Med. Microbiol. 2007, 297, 163–176. [Google Scholar] [CrossRef]
- Janben, T.; Schwarz, C.; Preikschat, P.; Voss, M.; Philipp, H.C.; Wieler, L.H. Virulence-associated genes in avian pathogenic Escherichia coli (APEC) isolated from internal organs of poultry having died from colibacillosis. Int. J. Med. Microbiol. 2001, 291, 371–378. [Google Scholar] [CrossRef] [PubMed]
- Schubert, S.; Rakin, A.; Karch, H.; Carniel, E.; Heesemann, J. Prevalence of the High-Pathogenicity Island Yersinia Species among Escherichia coli Strains That Are Pathogenic to Humans. Infect. Immun. 1998, 66, 480–485. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Clark, C.G.; Rodgers, F.G. Detection in Escherichia coli of the genes encoding the major virulence factors, the genes defining the O157:H7 serotype, and components of the type 2 Shiga toxin family by multiplex PCR. J. Clin. Microbiol. 2002, 40, 3613–3619. [Google Scholar] [CrossRef] [PubMed]
- Johnson, T.J.; Wannemuehler, Y.; Doetkott, C.; Johnson, S.J.; Rosenberger, S.C.; Nolan, L.K. Identification of minimal predictors of avian pathogenic Escherichia coli virulence for use as a rapid diagnostic tool. J. Clin. Microbiol. 2008, 46, 3987–3996. [Google Scholar] [CrossRef] [PubMed]
- Toma, C.; Lu, Y.; Higa, N.; Nakasone, N.; Chinen, I.; Baschkier, A.; Rivas, M.; Iwanaga, M. Multiplex PCR assay for identification of human diarrheagenic Escherichia coli. J. Clin. Microbiol. 2003, 41, 2669–2671. [Google Scholar] [CrossRef] [PubMed]
- Hudzicki, J. Kirby-Bauer disk diffusion susceptibility test protocol. Am. Soc. Microbiol. 2009, 15, 55–63. [Google Scholar]
- CLSI. Performance Standards for Antimivrobial Susceptibility Testing. In CLSI Supplement M100, 30th ed.; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2020; ISBN 978-1-68440-066-9 (Print). ISBN 978-1-68440-067-6 (Electronic). [Google Scholar]
- Wiegand, I.; Hilpert, K.; Hancock, R.E. Agar and broth dilution methods to determine the minimal inhibitory concentration (MIC) of antimicrobial substances. Nat. Protoc. 2008, 3, 163–175. [Google Scholar] [CrossRef]
- Jayol, A.; Nordmann, P.; Andre, C.; Poirel, L.; Dubois, V. Evaluation of three broth microdilution systems to determine colistin susceptibility of Gram-negative bacilli. J. Antimicrob. Chemother. 2018, 73, 1272–1278. [Google Scholar] [CrossRef]
- Jing Yu, W.; Pan, T.; En Hui, C.; Li Qin, W.; Wan Hua, L.; Juan Juan, R.; Ning, W.; Yuan Hao, Q.; Hung Jen, L. Characterization of antimicrobial resistance and related resistance genes in Escherichia coli strains isolated from chickens in China during 2007–2012. Afr. J. Microbiol. Res. 2013, 7, 5238–5247. [Google Scholar] [CrossRef]
- Dallenne, C.; Da Costa, A.; Decre, D.; Favier, C.; Arlet, G. Development of a set of multiplex PCR assays for the detection of genes encoding important beta-lactamases in Enterobacteriaceae. J. Antimicrob. Chemother. 2010, 65, 490–495. [Google Scholar] [CrossRef]
- Chen, S.; Zhao, S.; White, D.G.; Schroeder, C.M.; Lu, R.; Yang, H.; McDermott, P.F.; Ayers, S.; Meng, J. Characterization of multiple-antimicrobial-resistant Salmonella serovars isolated from retail meats. Appl. Environ. Microbiol. 2004, 70, 1–7. [Google Scholar] [CrossRef]
- Xiang, T.; Chen, C.; Wen, J.; Liu, Y.; Zhang, Q.; Cheng, N.; Wu, X.; Zhang, W. Resistance of Klebsiella pneumoniae Strains Carrying bla (NDM-1) Gene and the Genetic Environment of bla (NDM-1). Front. Microbiol. 2020, 11, 700. [Google Scholar] [CrossRef] [PubMed]
- Ramadan, A.A.; Abdelaziz, N.A.; Amin, M.A.; Aziz, R.K. Novel blaCTX-M variants and genotype-phenotype correlations among clinical isolates of extended spectrum beta lactamase-producing Escherichia coli. Sci. Rep. 2019, 9, 4224. [Google Scholar] [CrossRef] [PubMed]
- Komp Lindgren, P.; Karlsson, A.; Hughes, D. Mutation rate and evolution of fluoroquinolone resistance in Escherichia coli isolates from patients with urinary tract infections. Antimicrob. Agents Chemother. 2003, 47, 3222–3232. [Google Scholar] [CrossRef] [PubMed]
- Giraud, E.; Brisabois, A.; Martel, J.-L.; Chaslus-Dancla, E. Comparative Studies of Mutations in Animal Isolates and Experimental In Vitro- and In Vivo-Selected Mutants of Salmonella spp. Suggest a Counterselection of Highly Fluoroquinolone-Resistant Strains in the Field. Antimicrob. Agents Chemother. 1999, 43, 2131–2137. [Google Scholar] [CrossRef]
- Shahrani, M.; Dehkordi, F.S.; Momtaz, H. Characterization of Escherichia coli virulence genes, pathotypes and antibiotic resistance properties in diarrheic calves in Iran. Biol. Res. 2014, 47, 28. [Google Scholar] [CrossRef] [PubMed]
- Kozak, G.K.; Boerlin, P.; Janecko, N.; Reid-Smith, R.J.; Jardine, C. Antimicrobial resistance in Escherichia coli isolates from swine and wild small mammals in the proximity of swine farms and in natural environments in Ontario, Canada. Appl. Environ. Microbiol. 2009, 75, 559–566. [Google Scholar] [CrossRef] [PubMed]
- Aarestrup, F.M.; Lertworapreecha, M.; Evans, M.C.; Bangtrakulnonth, A.; Chalermchaikit, T.; Hendriksen, R.S.; Wegener, H.C. Antimicrobial susceptibility and occurrence of resistance genes among Salmonella enterica serovar Weltevreden from different countries. J. Antimicrob. Chemother. 2003, 52, 715–718. [Google Scholar] [CrossRef]
- Ibrahim, R.A.; Cryer, T.L.; Lafi, S.Q.; Basha, E.A.; Good, L.; Tarazi, Y.H. Identification of Escherichia coli from broiler chickens in Jordan, their antimicrobial resistance, gene characterization and the associated risk factors. BMC Vet. Res. 2019, 15, 159. [Google Scholar] [CrossRef]
- Perreten, V.; Boerlin, P. A new sulfonamide resistance gene (sul3) in Escherichia coli is widespread in the pig population of Switzerland. Antimicrob. Agents Chemother. 2003, 47, 1169–1172. [Google Scholar] [CrossRef] [PubMed]
- Sunde, M. Prevalence and characterization of class 1 and class 2 integrons in Escherichia coli isolated from meat and meat products of Norwegian origin. J. Antimicrob. Chemother. 2005, 56, 1019–1024. [Google Scholar] [CrossRef]
- Liu, J.; Zhang, Z.; Feng, Y.; Hu, H.; Yu, Y.; Qiu, L.; Liu, H.; Guo, Z.; Huang, J.; Du, C.; et al. Molecular Detection of the mcr Genes by Multiplex PCR. Infect. Drug Resist. 2020, 13, 3463–3468. [Google Scholar] [CrossRef] [PubMed]
- Maynard, C.; Fairbrother, J.M.; Bekal, S.; Sanschagrin, F.; Levesque, R.C.; Brousseau, R.; Masson, L.; Lariviere, S.; Harel, J. Antimicrobial resistance genes in enterotoxigenic Escherichia coli O149:K91 isolates obtained over a 23-year period from pigs. Antimicrob. Agents Chemother. 2003, 47, 3214–3221. [Google Scholar] [CrossRef] [PubMed]
- Dolejska, M.; Cizek, A.; Literak, I. High prevalence of antimicrobial-resistant genes and integrons in Escherichia coli isolates from black-headed gulls in the Czech Republic. J. Appl. Microbiol. 2007, 103, 11–19. [Google Scholar] [CrossRef] [PubMed]
- Allen, V.G.; Farrell, D.J.; Rebbapragada, A.; Tan, J.; Tijet, N.; Perusini, S.J.; Towns, L.; Lo, S.; Low, D.E.; Melano, R.G. Molecular Analysis of Antimicrobial Resistance Mechanisms in Neisseria gonorrhoeae Isolates from Ontario, Canada. Antimicrob. Agents Chemother. 2011, 55, 703–712. [Google Scholar] [CrossRef] [PubMed]
- Kargar, M.; Mohammadalipour, Z.; Doosti, A.; Lorzadeh, S.; Japoni-Nejad, A. High Prevalence of Class 1 to 3 Integrons among Multidrug-Resistant Diarrheagenic Escherichia coli in Southwest of Iran. Osong Public. Health Res. Perspect. 2014, 5, 193–198. [Google Scholar] [CrossRef] [PubMed]
- Tanizawa, Y.; Fujisawa, T.; Nakamura, Y. DFAST: A flexible prokaryotic genome annotation pipeline for faster genome publication. Bioinformatics 2018, 34, 1037–1039. [Google Scholar] [CrossRef]
- Larsen, M.V.; Cosentino, S.; Rasmussen, S.; Friis, C.; Hasman, H.; Marvig, R.L.; Jelsbak, L.; Sicheritz-Ponten, T.; Ussery, D.W.; Aarestrup, F.M.; et al. Multilocus sequence typing of total-genome-sequenced bacteria. J. Clin. Microbiol. 2012, 50, 1355–1361. [Google Scholar] [CrossRef]
- Jolley, K.A.; Bray, J.E.; Maiden, M.C.J. Open-access bacterial population genomics: BIGSdb software, the PubMLST.org website and their applications. Wellcome Open Res. 2018, 3, 124. [Google Scholar] [CrossRef]
- Couvin, D.; Bernheim, A.; Toffano-Nioche, C.; Touchon, M.; Michalik, J.; Neron, B.; Rocha, E.P.C.; Vergnaud, G.; Gautheret, D.; Pourcel, C. CRISPRCasFinder, an update of CRISRFinder, includes a portable version, enhanced performance and integrates search for Cas proteins. Nucleic Acids Res. 2018, 46, W246–W251. [Google Scholar] [CrossRef] [PubMed]
- Biswas, A.; Gagnon, J.N.; Brouns, S.J.; Fineran, P.C.; Brown, C.M. CRISPRTarget: Bioinformatic prediction and analysis of crRNA targets. RNA Biol. 2013, 10, 817–827. [Google Scholar] [CrossRef] [PubMed]
- Camacho, C.; Coulouris, G.; Avagyan, V.; Ma, N.; Papadopoulos, J.; Bealer, K.; Madden, T.L. BLAST+: Architecture and applications. BMC Bioinform. 2009, 10, 421. [Google Scholar] [CrossRef] [PubMed]
- Alikhan, N.F.; Petty, N.K.; Ben Zakour, N.L.; Beatson, S.A. BLAST Ring Image Generator (BRIG): Simple prokaryote genome comparisons. BMC Genom. 2011, 12, 402. [Google Scholar] [CrossRef]

| Strain | Year of Isolation | Chicken Type a | Age (Day-Old) | Phylo- Group | RST b | No. of Virulence Genes | Strain | Year of Isolation | Chicken Type a | Age (Day-Old) | Phylo- Group | RST | No. of Virulence Genes |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| E12049 | 2012 | KNC | 40 | F | 21-1 | 20 | E16015 | 2016 | B | 24 | B1 | 5-1 | 15 |
| E14033 | 2014 | L | 76 | F | 21-1 | 19 | E17039 | 2017 | BB | 263 | B1 | 12-6 | 15 |
| E12090 | 2012 | L | 124 | F | 21-1 | 18 | E19033 | 2019 | BB | 134 | B1 | 4-4 | 15 |
| E18005 | 2018 | BB | 209 | B1 | 4-4 | 17 | E19057 | 2019 | L | 148 | C | 3-1 | 14 |
| E16018 | 2016 | L | 451 | B1 | 4-4 | 16 | E16023 | 2016 | BB | 203 | E | 3-1 | 14 |
| E18016 | 2018 | BB | 163 | B1 | 4-4 | 16 | E16039 | 2016 | L | 252 | B1 | 4-4 | 14 |
| E18027 | 2018 | B | 2 | E | 21-1 | 16 | E19024 | 2019 | L | 315 | C | 3-1 | 14 |
| E19014 | 2019 | BB | 7 | B1 | 4-4 | 16 | E20019 | 2020 | L | 310 | E | 3-1 | 14 |
| E19025 | 2019 | B | 7 | B1 | 4-4 | 16 | E17001 | 2017 | BB | 264 | C | 3-1 | 13 |
| E19034 | 2019 | BB | 235 | B1 | 4-4 | 16 | E12050 | 2012 | B | 26 | C | 3-1 | 12 |
| E19045 | 2019 | L | 19 | B1 | 4-4 | 16 | E13028 | 2013 | BB | 29 | C | 3-1 | 12 |
| E19070 | 2019 | BB | 170 | B1 | 4-4 | 16 | E20042 | 2020 | L | 161 | C | 3-1 | 11 |
| E12091 | 2012 | BB | 236 | B1 | 4-4 | 15 | E19013 | 2019 | BB | 77 | C | 4-5 | 9 |
| E15016 | 2015 | B | 30 | B1 | 4-4 | 15 | E19040 | 2019 | BB | 195 | C | 3-1 | 9 |
| E15026 | 2015 | BB | 273 | B1 | 4-4 | 15 | E19056 | 2019 | B | 3 | A | 3-1 | 9 |
| E16011 | 2016 | L | 136 | B1 | 4-4 | 15 | E123 | 2003 | NA c | NA | C | 3-1 | 17 |
| Pathogenic Bacteria | Strain | Accession No. | Size of rpoB | RST | Identity (%) |
|---|---|---|---|---|---|
| Corynebacterium diphtheriae | 31A | CP003206.1 | 3531 | ND | ND |
| Bacillus anthracis | A2084 | NC_007530.2 | 3534 | ND | ND |
| Staphylococcus aureus | PMB81 | CP03444.1 | 3552 | ND | ND |
| Listeria monocytogenes | EGD-e | NC_003210.1 | 3555 | ND | ND |
| Streptococcus pneumoniae | Hu17 | NZ_CP020549.1 | 3612 | ND | ND |
| Enterococcus faecalis | T5 | NZ_KB944666.1 | 3615 | ND | ND |
| Clostridium perfringens | CPI 18-6 | NZ_CP075979.1 | 3705 | ND | ND |
| Clostridium difficile | s-0253 | NZ_CP076401.1 | 3717 | ND | ND |
| Vibrio cholerae | RFB16 | CP043554.1 | 4026 | ND | ND |
| Escherichia coli | RST0 | ND a | 4029 | 0 | 100 |
| Escherichia fergusonii | RHB10-C04 | CP057918.1 | 4029 | 40 | 99.1 |
| Escherichia fergusonii | ATCC 35471 | CP042945.1 | 4029 | 53 | 98.7 |
| Escherichia fergusonii | RHB02-C14 | CP055872.1 | 4029 | 60 | 98.5 |
| Escherichia albertii | RM10507 | CP043258.1 | 4029 | 69 | 98.3 |
| Escherichia albertii | ChinaSP140150 | CP025676.1 | 4029 | 100 | 97.5 |
| Escherichia albertii | BIA_36 | CP117590.1 | 4029 | 104 | 97.4 |
| Escherichia marmotae | W49-2 | CP093239.1 | 4029 | 106 | 97.4 |
| Salmonella enterica serovar Indiana | SI67 | CP050783.1 | 4029 | 177 | 95.6 |
| Salmonella bongori | N268-08 | CP006608.1 | 4029 | 251 | 93.8 |
| Klebsiella pneumoniae | XH210 | CP052761.1 | 4029 | 259 | 93.6 |
| Enterobacter cloacae | colR/S | CP010512.1 | 4029 | 264 | 93.4 |
| Serratia marcescens | SmUNAM836 | CP012685.1 | 4029 | 491 | 87.8 |
| Yersinia pseudotuberculosis | IP2666pIB1 | CP032566.1 | 4029 | 588 | 85.4 |
| Yersinia pestis | FDAARGOS_603 | CP033690.1 | 4029 | 590 | 85.4 |
| Proteus mirabilis | 1035 | CP072779.1 | 4029 | 758 | 81.1 |
| Pasteurella multocida | FDAARGOS_218 | CP020405.2 | 4029 | 1027 | 74.5 |
| Haemophilus influenzae | Hi375 | CP009610.1 | 4032 | ND | ND |
| Pseudomonas aeruginosa | PAO1 | AE004091.2 | 4074 | ND | ND |
| Acinetobacter baumannii | Ab421_GEIH-2010 | CP014266.1 | 4074 | ND | ND |
| Legionella pneumophila | C9 | CP015941.1 | 4107 | ND | ND |
| Bordetella bronchiseptica | NCTC10543 | LR134326.1 | 4113 | ND | ND |
| Neisseria gonorrheae | AT159 | CP097846.1 | 4179 | ND | ND |
| Moraxella catarrhalis | CCRI-195ME | CP018059.1 | 4239 | ND | ND |
| RST3-1 | RST4-4 | RST5-1 | RST12-6 | RST21-1 | |
|---|---|---|---|---|---|
| Frequency out of MDR strains (%) a | 3/18 | 11/18 | 1/18 | 1/18 | 2/18 |
| (16.7%) | (61.1%) b | (5.6%) | (5.6%) | (11.1%) | |
| MDR Frequency out of each RST (%) | 3/11 | 11/14 | 1/1 | 1/1 | 2/4 |
| (27.3%) | (78.6%) | (100%) | (100%) | (50%) |
| Antibiotics Resistance- Related Genes | gyrA | tetB | sul2 | blaTEM | blaCTX-M -G1 | strA/strB | tetA | aadA1 /A2 | aac(3)II | sul1 | dfrA1 | blaCTX-M -G9 | ampC | floR | blaSHV |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Frequency (%) | 29 | 16 | 12 | 10 | 9 | 7 | 6 | 6 | 6 | 5 | 4 | 3 | 2 | 2 | 1 |
| (90.6) | (50.0) | (37.5) | (31.3) | (28.1) | (21.9) | (18.8) | (18.8) | (18.8) | (15.6) | (12.5) | (9.4) | (6.3) | (6.3) | (3.1) |
| Strain | RST | Chromosome | Plasmid | No. of Microcin/Colicin a | |||
|---|---|---|---|---|---|---|---|
| 1 | 2 | 3 | 4 | ||||
| E123 | 3-1 | None | Microcin V (Colicin V) | None | 1 | ||
| E19057 | 3-1 | None | Colicin Ia | Microcin V (Colicin V) | 2 | ||
| E18005 | microcin H47 | Microcin J25 | 4 | ||||
| Colicin M | |||||||
| 4-4 | Microcin V (Colicin V) | ||||||
| Colicin B (pseudo) | |||||||
| E19025 | None | Microcin J25 | None | 3 | |||
| Colicin M | |||||||
| 4-4 | Microcin V (Colicin V) | ||||||
| Colicin B (pseudo) | |||||||
| E12049 | 21-1 | None | Colicin M | Colicin Ib | 2 | ||
| E14033 | 21-1 | None | None | ColIcin Ib | None | None | 1 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ha, E.-J.; Hong, S.-M.; Kim, S.-J.; Ahn, S.-M.; Kim, H.-W.; Choi, K.-S.; Kwon, H.-J. Tracing the Evolutionary Pathways of Serogroup O78 Avian Pathogenic Escherichia coli. Antibiotics 2023, 12, 1714. https://doi.org/10.3390/antibiotics12121714
Ha E-J, Hong S-M, Kim S-J, Ahn S-M, Kim H-W, Choi K-S, Kwon H-J. Tracing the Evolutionary Pathways of Serogroup O78 Avian Pathogenic Escherichia coli. Antibiotics. 2023; 12(12):1714. https://doi.org/10.3390/antibiotics12121714
Chicago/Turabian StyleHa, Eun-Jin, Seung-Min Hong, Seung-Ji Kim, Sun-Min Ahn, Ho-Won Kim, Kang-Seuk Choi, and Hyuk-Joon Kwon. 2023. "Tracing the Evolutionary Pathways of Serogroup O78 Avian Pathogenic Escherichia coli" Antibiotics 12, no. 12: 1714. https://doi.org/10.3390/antibiotics12121714
APA StyleHa, E.-J., Hong, S.-M., Kim, S.-J., Ahn, S.-M., Kim, H.-W., Choi, K.-S., & Kwon, H.-J. (2023). Tracing the Evolutionary Pathways of Serogroup O78 Avian Pathogenic Escherichia coli. Antibiotics, 12(12), 1714. https://doi.org/10.3390/antibiotics12121714






