Microbial Community Structure among Honey Samples of Different Pollen Origin
Abstract
1. Introduction
2. Results and Discussion
3. Materials and Methods
3.1. Collection of Honey Samples
3.2. DNA Extraction from Honey Samples of Different Pollen Origin and Performance of Amplicon Sequencing
3.3. Bioinformatic Analysis and Analysis of Variance (ANOVA)
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Zhou, X.; Taylor, M.P.; Salouros, H.; Prasad, S. Authenticity and geographic origin of global honeys determined using carbon isotope ratios and trace elements. Sci. Rep. 2018, 8, 14639. [Google Scholar] [CrossRef] [PubMed]
- Potts, S.G.; Biesmeijer, J.C.; Kremen, C.; Neumann, P.; Schweiger, O.; Kunin, W.E. Global pollinator declines: Trends, impacts and drivers. Trends Ecol. Evol. 2010, 25, 345–353. [Google Scholar] [CrossRef]
- EU Overview. Detailed Information on Honey Production in the European Union. Agriculture and Rural Development, European Commission. 2022. Available online: https://agriculture.ec.europa.eu/farming/animal-products/honey_en (accessed on 9 December 2022).
- EU News Article. Beekeeping Sector: Results of the Pilot Study on Honey Bee Selection. Directorate-General for Agriculture and Rural Development, European Commission. 15 March 2022. Available online: https://agriculture.ec.europa.eu/news/beekeeping-sector-results-pilot-study-honey-bee-selection-2022-03-15_en (accessed on 9 December 2022).
- Eurostat. EU Imported €405.9 Million Worth in Honey in 2021. Products Eurostat News, Eurostat. 2022. Available online: https://ec.europa.eu/eurostat/web/products-eurostat-news/-/edn-20220819-2 (accessed on 9 December 2022).
- Alberoni, D.; Gaggìa, F.; Baffoni, L.; di Gioia, D. Beneficial microorganisms for honey bees: Problems and progresses. Appl. Microbiol. Biotechnol. 2016, 100, 9469–9482. [Google Scholar] [CrossRef]
- Da Silva, P.M.; Gauche, C.; Gonzaga, L.V.; Costa, A.C.O.; Fett, R. Honey: Chemical composition, stability and authenticity. Food Chem. 2016, 196, 309–323. [Google Scholar] [CrossRef] [PubMed]
- Bosancic, B.; Zabic, M.; Mihajlovic, D.; Samardzic, J.; Mirjanic, G. Comparative study of toxic heavy metal residues and other properties of honey from different environmental production systems. Environ. Sci. Pollut. Res. 2020, 27, 38200–38211. [Google Scholar] [CrossRef] [PubMed]
- Iurlina, M.O.; Fritz, R. Characterization of microorganisms in Argentinean honeys from different sources. Int. J. Food Microbiol. 2005, 105, 297–304. [Google Scholar] [CrossRef] [PubMed]
- Fangio, M.F.; Iurlina, M.O.; Fritz, R. Characterisation of Argentinean honeys and evaluation of its inhibitory action on Escherichia coli growth. Int. J. Food Sci. Technol. 2010, 45, 520–529. [Google Scholar] [CrossRef]
- Bayram, N.E.; Kara, H.H.; Can, A.M.; Bozkurt, F.; Akman, P.K.; Vardar, S.U.; Çebi, N.; Yılmaz, M.T.; Sağdıç, O.; Dertli, E. Characterization of physicochemical and antioxidant properties of Bayburt honey from the North-east part of Turkey. J. Apic. Res. 2021, 60, 46–56. [Google Scholar] [CrossRef]
- Pătruică, S.; Alexa, E.; Obiștioiu, D.; Cocan, I.; Radulov, I.; Berbecea, A.; Lazăr, R.N.; Simiz, E.; Vicar, N.M.; Hulea, A.; et al. Chemical composition, antioxidant and antimicrobial activity of some types of honey from Banat region, Romania. Molecules 2022, 27, 4179. [Google Scholar] [CrossRef]
- Mandal, M.D.; Mandal, S. Honey: Its medicinal property and antibacterial activity. Asian Pac. J. Trop. Biomed. 2011, 1, 154–160. [Google Scholar] [CrossRef]
- Yupanqui Mieles, J.; Vyas, C.; Aslan, E.; Humphreys, G.; Diver, C.; Bartolo, P. Honey: An advanced antimicrobial and wound healing biomaterial for tissue engineering applications. Pharmaceutics 2022, 14, 1663. [Google Scholar] [CrossRef]
- Cheung, Y.; Meenu, M.; Yu, X.; Xu, B. Phenolic acids and flavonoids profiles of commercial honey from different floral sources and geographic sources. Int. J. Food Prop. 2019, 22, 290–308. [Google Scholar] [CrossRef]
- Mundo, M.A.; Padilla-Zakour, O.I.; Worobo, R.W. Growth inhibition of foodborne pathogens and food spoilage organisms by select raw honeys. Int. J. Food Microbiol. 2004, 97, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Stevenson, P.C.; Nicolson, S.W.; Wright, G.A. Plant secondary metabolites in nectar: Impacts on pollinators and ecological functions. Funct. Ecol. 2017, 31, 65–75. [Google Scholar] [CrossRef]
- Pajor, M.; Worobo, R.W.; Milewski, S.; Szweda, P. The antimicrobial potential of bacteria isolated from honey samples produced in the Apiaries located in Pomeranian Voivodeship in Northern Poland. Int. J. Environ. Res. Public Health 2018, 15, 2002. [Google Scholar] [CrossRef]
- Voidarou, C.; Antoniadou, M.; Rozos, G.; Alexopoulos, A.; Giorgi, E.; Tzora, A.; Giorgi, E.; Tzora, A.; Skoufos, I.; Varzakas, T.; et al. An in vitro study of different types of Greek honey as potential natural antimicrobials against dental caries and other oral pathogenic microorganisms. Case study simulation of oral cavity conditions. Appl. Sci. 2021, 11, 6318. [Google Scholar] [CrossRef]
- Masoura, M.; Gkatzionis, K. The antimicrobial mechanism of Greek thyme honeys against methicillin-resistant Staphylococcus aureus clinical isolates: A case study of comparison with Manuka honey. Int. J. Food Sci. Technol. 2022, 57, 7076–7084. [Google Scholar] [CrossRef]
- Jia, L.; Kosgey, J.C.; Wang, J.; Yang, J.; Nyamao, R.M.; Zhao, Y.; Teng, X.; Gao, L.; Wabo, M.T.C.; Vasilyeva, N.V.; et al. Antimicrobial and mechanism of antagonistic activity of Bacillus sp. A2 against pathogenic fungus and bacteria: The implication on honey’s regulatory mechanism on host’s microbiota. Food Sci. Nutr. 2020, 8, 4857–4867. [Google Scholar] [CrossRef]
- Wen, Y.; Wang, L.; Jin, Y.; Zhang, J.; Su, L.; Zhang, X.; Zhou, J.; Li, Y. The microbial community dynamics during the vitex honey ripening process in the honeycomb. Front. Microbiol. 2017, 8, 1649. [Google Scholar] [CrossRef]
- Yang, S.; Xu, W.; Tan, C.; Li, M.; Li, D.; Zhang, C.; Feng, L.; Chen, Q.; Jiang, J.; Li, Y.; et al. Heat stress weakens the skin barrier function in sturgeon by decreasing mucus secretion and disrupting the mucosal microbiota. Front. Microbiol. 2022, 13, 860079. [Google Scholar] [CrossRef]
- Kale, N.; Ashwini, M.; Jahagirdar, S.; Shirnalli, G. Potentials of lactic acid bacteria in enhancing nodulation of Bradyrhizobium daqingense and yield in soybean. Legume Res. 2022, 45, 507–513. [Google Scholar] [CrossRef]
- Disayathanoowat, T.; Li, H.; Supapimon, N.; Suwannarach, N.; Lumyong, S.; Chantawannakul, P.; Guo, J. Different dynamics of bacterial and fungal communities in hive-stored bee bread and their possible roles: A case study from two commercial honey bees in China. Microorganisms 2020, 8, 264. [Google Scholar] [CrossRef] [PubMed]
- Loncaric, I.; Ruppitsch, W.; Licek, E.; Moosbeckhofer, R.; Busse, H.J.; Rosengarten, R. Characterization of selected Gram-negative non-fermenting bacteria isolated from honey bees (Apis mellifera carnica). Apidologie 2011, 42, 312–325. [Google Scholar] [CrossRef]
- Anjum, S.I.; Aldakheel, F.; Shah, A.H.; Khan, S.; Ullah, A.; Hussain, R.; Khan, H.; Ansari, M.J.; Mahmoud, A.H.; Mohammed, O.B. Honey bee gut an unexpected niche of human pathogen. J. King Saud Univ. Sci. 2021, 33, 101247. [Google Scholar] [CrossRef]
- Belhadj, H.; Harzallah, D.; Khennouf, S.; Dahamna, S.; Ghadbane, M. Population dynamics of lactic acid bacteria during laboratory fermentation of honeybee-collected pollen. Der. Pharm. Lett. 2016, 8, 297–300. [Google Scholar]
- Kim, E.; Seo, J.; Yang, S.H.; Kim, I.S.; Koo, Y. Intestine bacterial microbiota of Asian hornet (Vespa velutina nigrithorax) and honey bee. Korean J. Environ. Agric. 2018, 37, 135–140. [Google Scholar] [CrossRef][Green Version]
- Li, Y.-H.; Huang, Y.-F.; Chen, T.-H.; Wu, S.-S.; Tang, H.-C.; Hsiao, C.-Y.; Huang, L.-C.; Chang, J.-C.; Chiu, K.-P.; Nai, Y.-S. Comparison of gut microbiota of healthy and diseased walking sticks, Phasmotaenia lanyuhensis. Arch. Insect Biochem. Physiol. 2020, 105, e21749. [Google Scholar] [CrossRef]
- Brady, C.L.; Cleenwerck, I.; Denman, S.; Venter, S.N.; Rodríguez-Palenzuela, P.; Coutinho, T.A.; de Vos, P. Proposal to reclassify Brenneria quercina (Hildebrand and Schroth 1967) Hauben et al. 1999 into a new genus, Lonsdalea gen. nov., as Lonsdalea quercina comb. nov., descriptions of Lonsdalea quercina subsp. quercina comb. nov., Lonsdalea quercina subsp. iberica subsp. nov. and Lonsdalea quercina subsp. britannica subsp. nov., emendation of the description of the genus Brenneria, reclassification of Dickeya dieffenbachiae as Dickeya dadantii subsp. dieffenbachiae comb. nov., and emendation of the description of Dickeya dadantii. Int. J. Syst. Evol. Microbiol. 2012, 62, 1592–1602. [Google Scholar]
- Okamoto, T.; Taguchi, H.; Nakamura, K.; Ikenaga, H.; Kuraishi, H.; Yamasato, K. Zymobacter palmae gen. nov., sp. nov., a new ethanol-fermenting peritrichous bacterium isolated from palm sap. Arch. Microbiol. 1993, 160, 333–337. [Google Scholar] [CrossRef]
- Sitz, R.A.; Aquino, V.M.; Tisserat, N.A.; Cranshaw, W.S.; Stewart, J.E. Insects visiting drippy blight diseased red oak trees are contaminated with the pathogenic bacterium Lonsdalea quercina. Plant Dis. 2019, 103, 1940–1946. [Google Scholar] [CrossRef]
- Ntougias, S.; Zervakis, G.I.; Fasseas, C. Halotalea alkalilenta gen. nov., sp. nov., a novel osmotolerant and alkalitolerant bacterium from alkaline olive mill wastes, and emended description of the family Halomonadaceae Franzmann et al. 1989, emend. Dobson and Franzmann 1996. Int. J. Syst. Evol. Microbiol. 2007, 57, 1975–1983. [Google Scholar] [CrossRef] [PubMed]
- Suenami, S.; Konishi Nobu, M.; Miyazaki, R. Community analysis of gut microbiota in hornets, the largest eusocial wasps, Vespa mandarinia and V. simillima. Sci. Rep. 2019, 9, 9830. [Google Scholar] [CrossRef] [PubMed]
- Crotti, E.; Rizzi, A.; Chouaia, B.; Ricci, I.; Favia, G.; Alma, A.; Sacchi, L.; Bourtzis, K.; Mandrioli, M.; Cherif, A.; et al. Acetic acid bacteria, newly emerging symbionts of insects. Appl. Environ. Microbiol. 2010, 76, 6963–6970. [Google Scholar] [CrossRef]
- Boğ, E.Ş.; Ertürk, Ö.; Yaman, M. Pathogenicity of aerobic bacteria isolated from honeybees (Apis mellifera) in Ordu Province. Turk. J. Vet. Anim. Sci. 2020, 44, 714–719. [Google Scholar] [CrossRef]
- Martins, J.; Ares, A.; Costa, J.; Canhoto, J. A baseline of Arbutus unedo L. microbiome for future research: In vitro versus ex vitro. Sci. Hortic. 2022, 292, 110657. [Google Scholar] [CrossRef]
- Puopolo, G.; Tomada, S.; Pertot, I. The impact of the omics era on the knowledge and use of Lysobacter species to control phytopathogenic micro-organisms. J. Appl. Microbiol. 2018, 124, 15–27. [Google Scholar] [CrossRef]
- Yun, J.H.; Lee, J.Y.; Kim, P.S.; Jung, M.J.; Bae, J.W. Paenibacillus apis sp. nov. and Paenibacillus intestini sp. nov., isolated from the intestine of the honey bee Apis mellifera. Int. J. Syst. Evol. Microbiol. 2017, 67, 1918–1924. [Google Scholar] [CrossRef]
- Ebeling, J.; Knispel, H.; Hertlein, G.; Fünfhaus, A.; Genersch, E. Biology of Paenibacillus larvae, a deadly pathogen of honey bee larvae. Appl. Microbiol. Biotechnol. 2016, 100, 7387–7395. [Google Scholar] [CrossRef]
- Dedej, S.; Delaplane, K.S.; Scherm, H. Effectiveness of honey bees in delivering the biocontrol agent Bacillus subtilis to blueberry flowers to suppress mummy berry disease. Biol. Control 2004, 31, 422–427. [Google Scholar] [CrossRef]
- Kačániová, M.; Terentjeva, M.; Žiarovská, J.; Kowalczewski, P.Ł. In vitro antagonistic effect of gut bacteriota isolated from indigenous honey bees and essential oils against Paenibacillus larvae. Int. J. Mol. Sci. 2020, 21, 6736. [Google Scholar] [CrossRef]
- Shehabeldine, A.M.; Hashem, A.H.; Hasaballah, A.I. Antagonistic effect of gut microbiota of the Egyptian honeybees, Apis mellifera L. against the etiological agent of Stonebrood disease. Int. J. Trop. Insect Sci. 2022, 42, 1357–1366. [Google Scholar] [CrossRef]
- Graystock, P.; Rehan, S.M.; McFrederick, Q.S. Hunting for healthy microbiomes: Determining the core microbiomes of Ceratina, Megalopta, and Apis bees and how they associate with microbes in bee collected pollen. Conserv. Genet. 2017, 18, 701–711. [Google Scholar] [CrossRef]
- Cianciosi, D.; Forbes-Hernández, T.Y.; Afrin, S.; Gasparrini, M.; Reboredo-Rodriguez, P.; Manna, P.P.; Zhang, J.; Lamas, L.B.; Flórez, S.M.; Toyos, P.A.; et al. Phenolic compounds in honey and their associated health benefits: A review. Molecules 2018, 23, 2322. [Google Scholar] [CrossRef] [PubMed]
- Dong, Z.X.; Tang, Q.H.; Li, W.L.; Wang, Z.W.; Li, X.J.; Fu, C.M.; Li, D.; Qian, K.; Tian, W.-L.; Guo, J. Honeybee (Apis mellifera) resistance to deltamethrin exposure by modulating the gut microbiota and improving immunity. Environ. Pollut. 2022, 314, 120340. [Google Scholar] [CrossRef]
- Deng, Y.; Yang, S.; Zhao, H.; Luo, J.; Yang, W.; Hou, C. Antibiotics-induced changes in intestinal bacteria result in the sensitivity of honey bee to virus. Environ. Pollut. 2022, 314, 120278. [Google Scholar] [CrossRef]
- Wu, J.; Han, B.; Zhao, S.; Zhong, Y.; Han, W.; Gao, J.; Wang, S. Bioactive characterization of multifloral honeys from Apis cerana cerana, Apis dorsata, and Lepidotrigona flavibasis. Food Res. Int. 2022, 161, 111808. [Google Scholar] [CrossRef]
- Al-Ghamdi, A.; Al-Abbadi, A.A.; Khan, K.A.; Ghramh, H.A.; Ahmed, A.M.; Ansari, M.J. In vitro antagonistic potential of gut bacteria isolated from indigenous honey bee race of Saudi Arabia against Paenibacillus larvae. J. Apic. Res. 2020, 59, 825–833. [Google Scholar] [CrossRef]
- Iorizzo, M.; Ganassi, S.; Albanese, G.; Letizia, F.; Testa, B.; Tedino, C.; Petrarca, S.; Mutinelli, F.; Mazzeo, A.; de Cristofaro, A. Antimicrobial activity from putative probiotic lactic acid bacteria for the biological control of American and European foulbrood diseases. Vet. Sci. 2022, 9, 236. [Google Scholar] [CrossRef]
- Vásquez, A.; Forsgren, E.; Fries, I.; Paxton, R.J.; Flaberg, E.; Szekely, L.; Olofsson, T.C. Symbionts as major modulators of insect health: Lactic acid bacteria and honeybees. PLoS ONE 2012, 7, e33188. [Google Scholar] [CrossRef]
- Goh, L.P.W.; Molujin, A.M.; Muthu, K.; Abdulla, R.; Sabullah, M.K.; Mohd Faik, A.A.; Gansau, J.A.; Jawan, R. Isolation and characterization of lactic acid bacteria from Sabah (North Borneo) stingless bees for probiotic and food applications. Int. J. Food Prop. 2021, 24, 564–578. [Google Scholar] [CrossRef]
- Kaškonienė, V.; Adaškevičiūtė, V.; Kaškonas, P.; Mickienė, R.; Maruška, A. Antimicrobial and antioxidant activities of natural and fermented bee pollen. Food Biosci. 2020, 34, 100532. [Google Scholar] [CrossRef]
- Jaafar, N.A.I.; Mohamad, S.A.S.; Razak, W.R.W.A. Antimicrobial activity and antibiotic resistance of lactic acid bacteria isolated from Malaysian stingless bee’s gut. Malays. J. Microbiol. 2019, 15, 233–241. [Google Scholar]
- Bağci, E.; Diğrak, M. Antimicrobial activity of essential oils of some Abies (Fir) species from Turkey. Flavour Fragr. J. 1996, 11, 251–256. [Google Scholar] [CrossRef]
- Chandra, H.; Bishnoi, P.; Yadav, A.; Patni, B.; Mishra, A.P.; Nautiyal, A.R. Antimicrobial resistance and the alternative resources with special emphasis on plant-based antimicrobials—A review. Plants 2017, 6, 16. [Google Scholar] [CrossRef]
- Reichenbach, H. The Genus Lysobacter. In The Prokaryotes; Dworkin, M., Falkow, S., Rosenberg, E., Schleifer, K.H., Stackebrandt, E., Eds.; Springer: New York, NY, USA, 2006. [Google Scholar]
- Shao, Y.; Arias-Cordero, E.; Guo, H.; Bartram, S.; Boland, W. In vivo Pyro-SIP assessing active gut microbiota of the cotton leafworm, Spodoptera littoralis. PLoS ONE 2014, 9, e85948. [Google Scholar] [CrossRef] [PubMed]
- Agila, A.; Barringer, S. Application of selected ion flow tube mass spectrometry coupled with chemometrics to study the effect of location and botanical origin on volatile profile of unifloral American honeys. J. Food Sci. 2012, 77, C1103–C1108. [Google Scholar] [CrossRef] [PubMed]
- Ozcan-Sinir, G.; Copur, O.U.; Barringer, S.A. Botanical and geographical origin of Turkish honeys by selected-ion flow-tube mass spectrometry and chemometrics. J. Sci. Food Agric. 2020, 100, 2198–2207. [Google Scholar] [CrossRef]
- Choi, Y.J.; Gringorten, J.L.; Bélanger, L.; Morel, L.; Bourque, D.; Masson, L.; Groleau, D.; Míguez, C.B. Production of an insecticidal crystal protein from Bacillus thuringiensis by the methylotroph Methylobacterium extorquens. Appl. Environ. Microbiol. 2008, 74, 5178–5182. [Google Scholar] [CrossRef]
- Galbally, I.E.; Kirstine, W. The production of methanol by flowering plants and the global cycle of methanol. J. Atmos. Chem. 2002, 43, 195–229. [Google Scholar] [CrossRef]
- Sharma, P.; Kumar, T.; Yadav, M.; Gill, S.S.; Chauhan, N.S. Plant-microbe interactions for the sustainable agriculture and food security. Plant Gene 2021, 28, 100325. [Google Scholar] [CrossRef]
- Stavropoulou, E.; Ieronymaki, E.; Dimitroulia, E.; Constantinidis, T.C.; Vrioni, G.; Tsatsanis, C.; Tsakris, A. Anti-Inflammatory and antibacterial effects and mode of action of Greek arbutus, chestnut, and fir honey in mouse models of inflammation and sepsis. Microorganisms 2022, 10, 2374. [Google Scholar] [CrossRef] [PubMed]
- Stavropoulou, E.; Voidarou, C.; Rozos, G.; Vaou, N.; Bardanis, M.; Konstantinidis, T.; Vrioni, G.; Tsakris, A. Antimicrobial evaluation of various honey types against carbapenemase-producing Gram-negative clinical isolates. Antibiotics 2022, 11, 422. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. UNOISE2: Improved error-correction for Illumina 16S and ITS amplicon sequencing. bioRxiv 2016. [Google Scholar] [CrossRef]
- Edgar, R.C. Accuracy of microbial community diversity estimated by closed-and open-reference OTUs. Peer J. 2017, 5, e3889. [Google Scholar] [CrossRef]
- Dhariwal, A.; Chong, J.; Habib, S.; King, I.L.; Agellon, L.B.; Xia, J. MicrobiomeAnalyst: A web-based tool for comprehensive statistical, visual and meta-analysis of microbiome data. Nucleic Acids Res. 2017, 45, W180–W188. [Google Scholar] [CrossRef] [PubMed]
- Hammer, Ø.; Harper, D.A.; Ryan, P.D. PAST: Paleontological statistics software package for education and data analysis. Palaeontol. Electron. 2001, 4, 9. [Google Scholar]
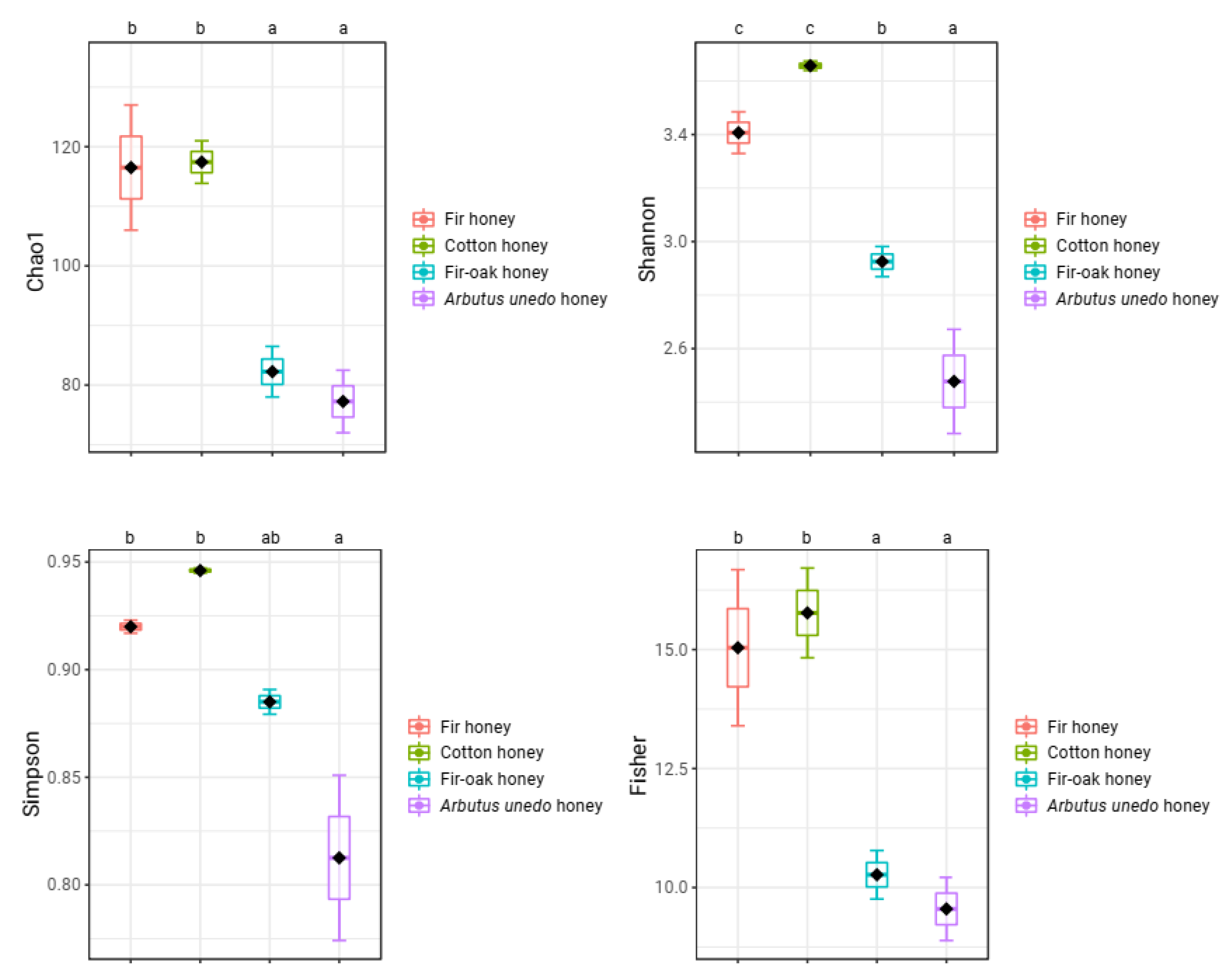

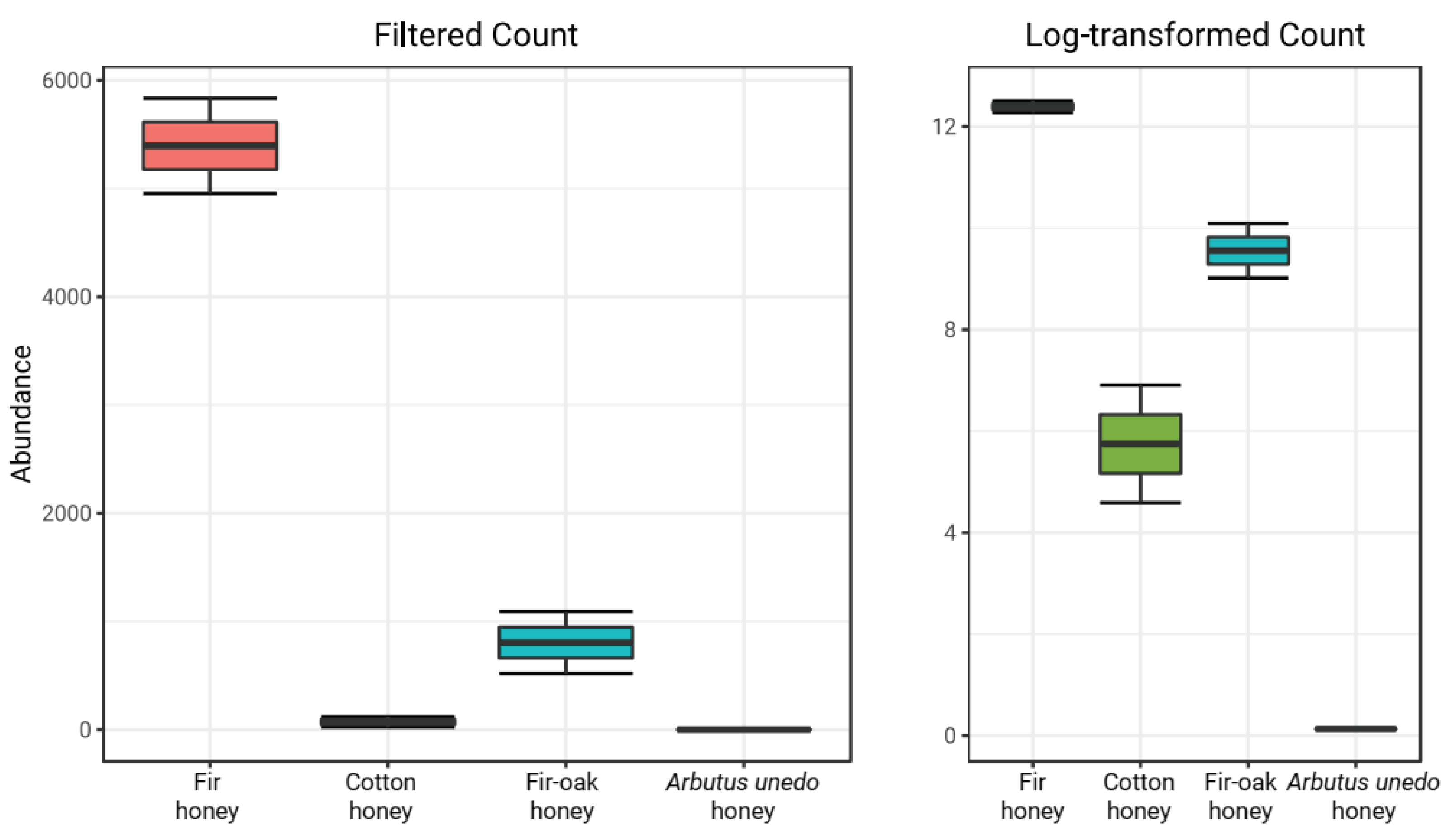
| Taxon | Relative Abundance (%) |
|---|---|
| Lactobacillus kunkeei | 19.34 ± 0.54 |
| Lactobacillus johnsonii | 0.31 ± 0.03 |
| Lactobacillus alvei | Marginally detected |
| Lactobacillus sakei | Marginally detected |
| Lactobacillus mellis | Marginally detected |
| Lactobacillus melliventris | Marginally detected |
| Taxon 1 | Fir Honey | Cotton Honey | Fir-Oak Honey | A. unedo Honey |
|---|---|---|---|---|
| Pseudomonas | 11.42 ± 1.68 (b) | 14.35 ± 2.10 (b) | 2.02 ± 1.23 (a) | 0.32 ± 0.03 (a) |
| Lysobacter | 2.67 ± 1.79 (a) | 7.07 ± 1.75 (a) | 14.93 ± 10.05 (ab) | 37.31 ± 5.58 (b) |
| Anoxybacillus | 2.24 ± 0.72 (a) | 5.47 ± 1.80 (a) | 1.78 ± 0.07 (a) | 4.60 ± 2.97 (a) |
| Bacillus | 0.30 ± 0.30 (a) | 3.35 ± 1.47 (a) | 3.61 ± 0.76 (a) | 3.69 ± 2.66 (a) |
| Meiothermus | 1.91 ± 0.29 (a) | 6.53 ± 3.93 (a) | 0.61 ± 0.07 (a) | 1.65 ± 0.26 (a) |
| Sphingomonas | 1.14 ± 0.03 (a) | 0.63 ± 0.49 (a) | 2.86 ± 0.05 (b) | 3.33 ± 0.43 (b) |
| Cupriavidus | 0.97 ± 0.42 (a) | 1.94 ± 0.72 (ab) | 1.52 ± 0.18 (ab) | 3.41 ± 0.59 (b) |
| Phenylobacterium | 0.20 ± 0.06 (a) | 0.58 ± 0.23 (a) | 1.21 ± 0.03 (a) | 4.61 ± 0.87 (b) |
| Paracoccus | 0.44 ± 0.05 (a) | 1.04 ± 0.43 (a) | 0.79 ± 0.54 (a) | 0.48 ± 0.23 (a) |
| Staphylococcus | 0.35 ± 0.28 (a) | 1.23 ± 0.64 (a) | 0.23 ± 0.09 (a) | 0.57 ± 0.32 (a) |
| Methylibium | 1.02 ± 0.48 (a) | 0.22 ± 0.06 (a) | 0.69 ± 0.12 (a) | 0.29 ± 0.03 (a) |
| Propionibacterium | 0.65 ± 0.26 (a) | 0.61 ± 0.21 (a) | 0.27 ± 0.17 (a) | 0.17 ± 0.01 (a) |
| Methylobacterium | 0.50 ± 0.12 (b) | 0.67 ± 0.02 (b) | 0.07 ± 0.05 (a) | 0.05 ± 0.04 (a) |
| Microbacterium | 0.31 ± 0.03 (a) | 0.26 ± 0.10 (a) | 0.23 ± 0.02 (a) | 0.18 ± 0.04 (a) |
| Taxon | Fir Honey | Cotton Honey | Fir-Oak Honey | Arbutus unedo Honey |
|---|---|---|---|---|
| Methylibium | 1.02 ± 0.48 | 0.22 ± 0.06 | 0.69 ± 0.12 | 0.29 ± 0.03 |
| Methylobacterium | 0.50 ± 0.12 | 0.67 ± 0.02 | 0.07 ± 0.05 | 0.05 ± 0.04 |
| Methylocapsa | 0.18 ± 0.18 | n.d. | n.d. | n.d. |
| Methylopila | 0.02 ± 0.02 | n.d. | 0.16 ± 0.16 | n.d. |
| Methylosinus | 0.25 ± 0.16 | n.d. | 0.12 ± 0.08 | n.d. |
| Methylotenera | 0.40 ± 0.01 | 0.85 ± 0.26 | n.d. | n.d. |
| Methyloversatilis | 0.60 ± 0.60 | 1.26 ± 0.99 | 0.61 ± 0.53 | 1.00 ± 0.58 |
| Total relative abundance 1 | 2.97 ± 1.20 (a) | 3.00 ± 1.18 (a) | 1.65 ± 0.44 (a) | 1.34 ± 0.59 (a) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Stavropoulou, E.; Remmas, N.; Voidarou, C.; Vrioni, G.; Konstantinidis, T.; Ntougias, S.; Tsakris, A. Microbial Community Structure among Honey Samples of Different Pollen Origin. Antibiotics 2023, 12, 101. https://doi.org/10.3390/antibiotics12010101
Stavropoulou E, Remmas N, Voidarou C, Vrioni G, Konstantinidis T, Ntougias S, Tsakris A. Microbial Community Structure among Honey Samples of Different Pollen Origin. Antibiotics. 2023; 12(1):101. https://doi.org/10.3390/antibiotics12010101
Chicago/Turabian StyleStavropoulou, Elisavet, Nikolaos Remmas, Chrysoula (Chrysa) Voidarou, Georgia Vrioni, Theodoros Konstantinidis, Spyridon Ntougias, and Athanasios Tsakris. 2023. "Microbial Community Structure among Honey Samples of Different Pollen Origin" Antibiotics 12, no. 1: 101. https://doi.org/10.3390/antibiotics12010101
APA StyleStavropoulou, E., Remmas, N., Voidarou, C., Vrioni, G., Konstantinidis, T., Ntougias, S., & Tsakris, A. (2023). Microbial Community Structure among Honey Samples of Different Pollen Origin. Antibiotics, 12(1), 101. https://doi.org/10.3390/antibiotics12010101










