Survey on Carbapenem-Resistant Bacteria in Pigs at Slaughter and Comparison with Human Clinical Isolates in Italy
Abstract
1. Introduction
2. Results
2.1. Phenotypical Antimicrobial Resistance in Pig Isolates
2.2. Phenotypical Antimicrobial Resistance in Human Isolates
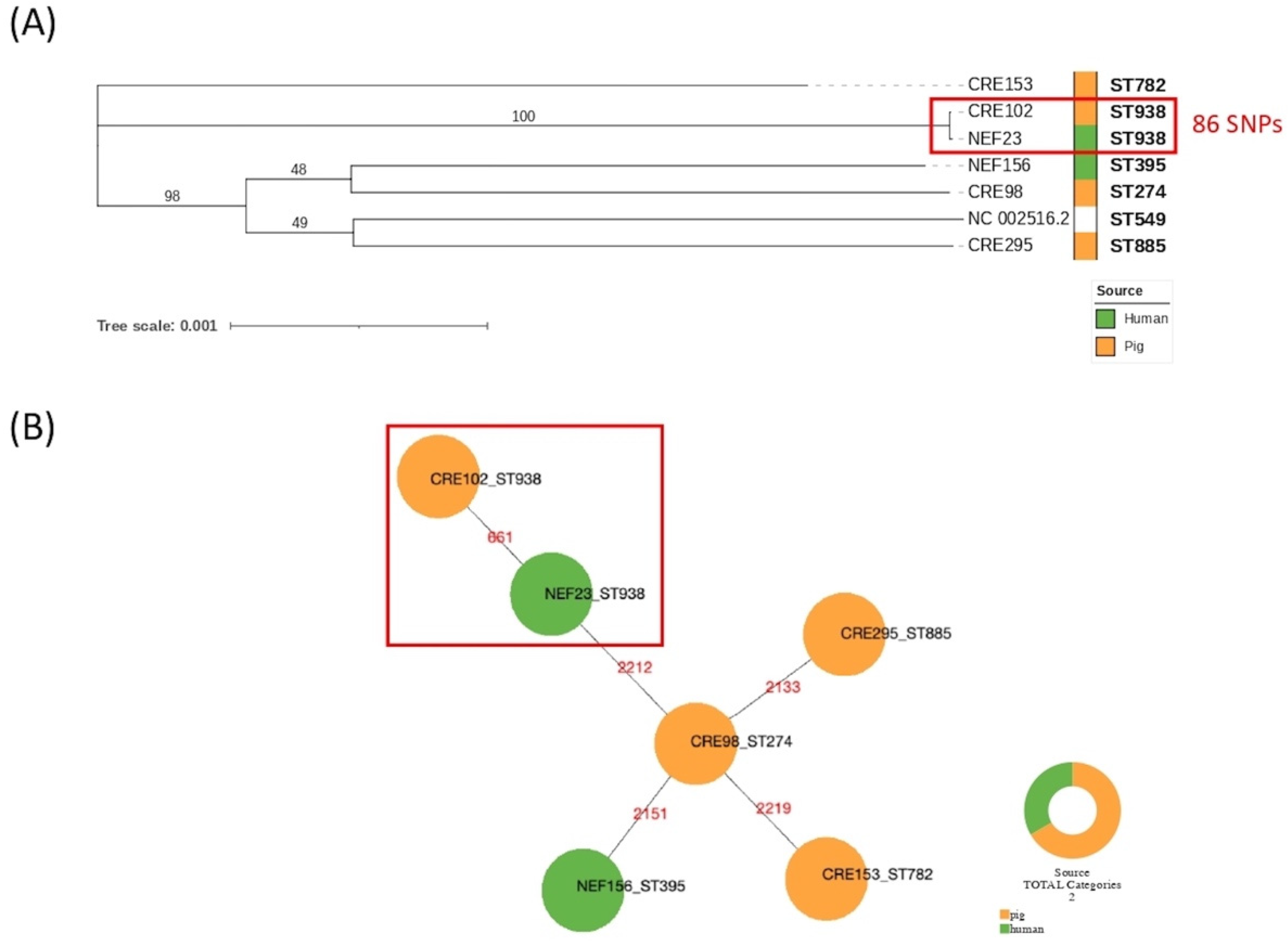
2.3. Whole-Genome Sequencing (WGS) of Porcine and Human Isolates
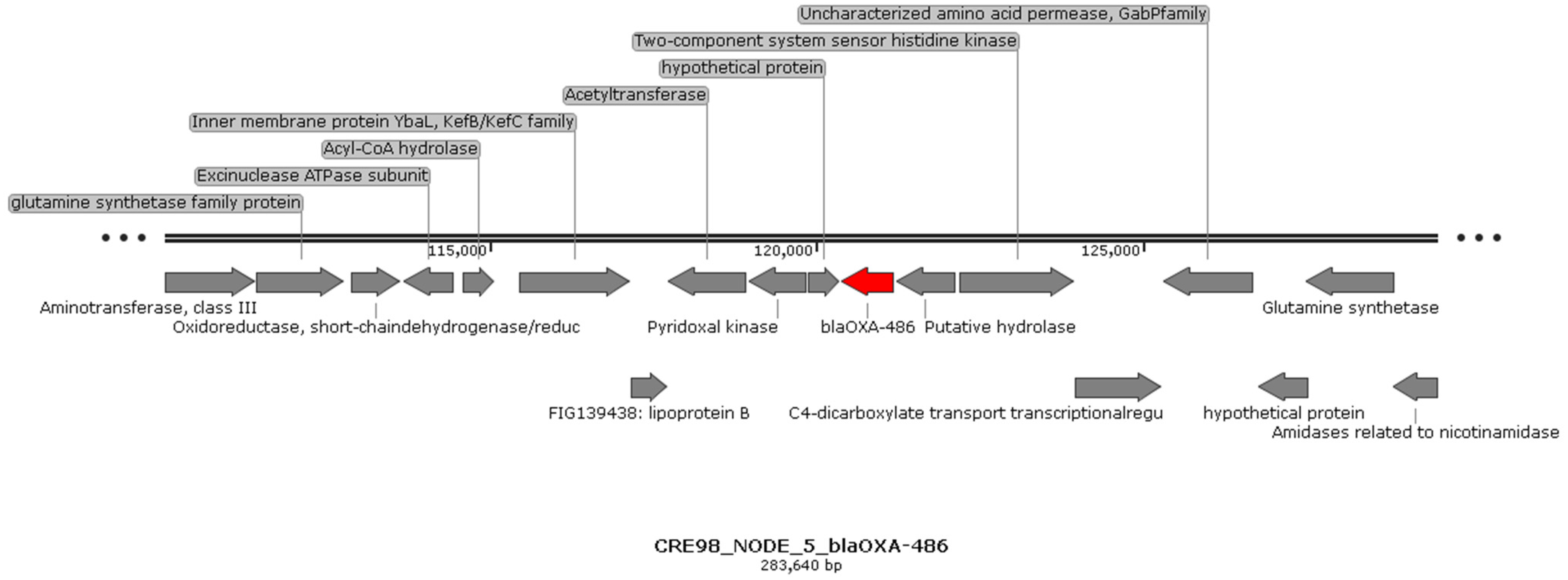
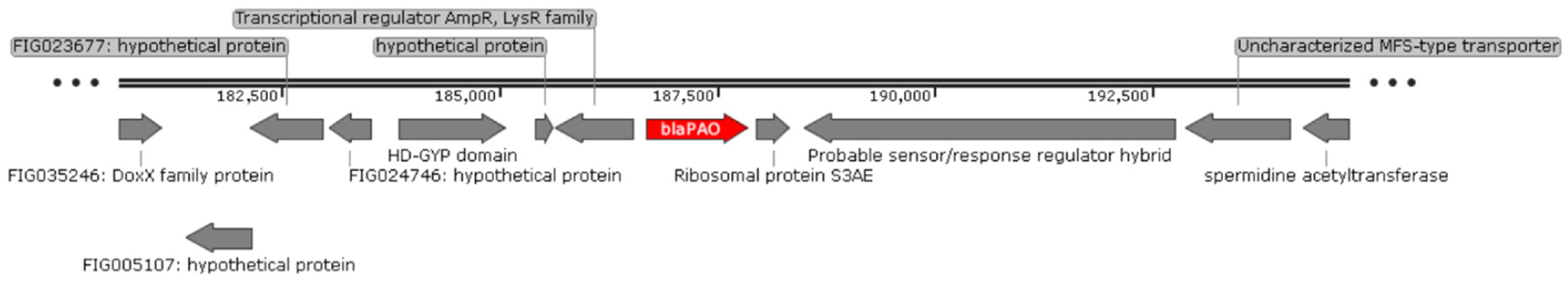
2.4. Localisation of Carbapenemase-Encoding Genes
2.5. Detection of Carbapenemases, β-Lactamases, and AmpC Genes in Other Pig Isolates
3. Discussion
3.1. CR P. aeruginosa in Pigs and Humans
3.2. ESBL/AmpC Genes Harbouring Porcine Isolates
4. Materials and Methods
4.1. Sample Collection
4.2. Sample Testing
4.3. Isolate Screening and Species Identification
4.4. MIC Testing
4.5. Comparison with CR Human Isolates
4.6. Genotypic Confirmation Test
4.7. Whole-Genome Sequencing
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Baquero, F.; Coque, T.M.; Martinez, J.-L.; Aracil-Gisbert, S.; Lanza, V.F. Gene transmission in the One Health microbiosphere and the channels of antimicrobial resistance. Front. Microbiol. 2019, 10, 2892. [Google Scholar] [CrossRef]
- Nordmann, P.; Naas, T.; Poirel, L. Global spread of carbapenemase producing Enterobacteriaceae. Emerg. Infect. Dis. 2011, 17, 1791–1798. [Google Scholar] [CrossRef]
- Nordmann, P.; Dortet, L.; Poirel, L. Carbapenem resistance in Enterobacteriaceae: Here is the storm! Trends Mol. Med. 2012, 18, 263–272. [Google Scholar] [CrossRef] [PubMed]
- Patel, G.; Bonomo, R.A. “Stormy waters ahead”: Global emergence of carbapenemases. Front. Microbiol. 2013, 4, 48. [Google Scholar] [CrossRef]
- Zhanel, G.G.; Wiebe, R.; Dilay, L.; Thomson, K.; Rubinstein, E.; Hoban, D.J.; Noreddin, A.M.; Karlowsky, J.A. Comparative review of the carbapenems. Drugs 2007, 67, 1027–1052. [Google Scholar] [CrossRef]
- Roschanski, N.; Friese, A.; von Salviati-Claudius, C.; Hering, J.; Kaesbohrer, A.; Kreienbrock, L.; Roesler, U. Prevalence of carbapenemase producing Enterobacteriaceae isolated from German pig-fattening farms during the years 2011–2013. Vet. Microbiol. 2017, 200, 124–129. [Google Scholar] [CrossRef] [PubMed]
- Mollenkopf, D.F.; Stull, J.W.; Mathys, D.A.; Bowman, A.S.; Feicht, S.M.; Grooters, S.V.; Daniels, J.B.; Wittuma, T.E. Carbapenemase-Producing Enterobacteriaceae Recovered from the Environment of a Swine Farrow-to-Finish Operation in the United States. Antimicrob. Agents Chemother. 2017, 61, e01298-16. [Google Scholar] [CrossRef]
- FAO; OIE; WHO (Food and Agriculture Organization of the United Nations, World Organization for Animal Health, World Health Organization). Monitoring and Evaluation of the Global Action Plan on Antimicrobial Resistance: Framework and Recommended Indicators. 2019. Available online: http://www.who.int (accessed on 1 December 2021).
- Hoelle, J.; Johnson, J.R.; Johnston, B.D.; Kinkle, B.; Boczek, L.; Ryu, H.; Hayes, S. Survey of US wastewater for carbapenem-resistant Enterobacteriaceae. J. Water Health 2019, 17, 219–226. [Google Scholar] [CrossRef]
- Hu, Z.; Chen, W.; Guo, G.; Dong, C.; Shen, Y.; Qin, S.; Chen, L.; Zhang, W. An Escherichia coli isolate from hospital sewage carries bla(NDM-1) and bla(oxa-10). Arch. Microbiol. 2021, 203, 4427–4432. [Google Scholar] [CrossRef]
- Yao, Y.; Lazaro-Perona, F.; Falgenhauer, L.; Valverde, A.; Imirzalioglu, C.; Dominguez, L.; Cantón, R.; Mingorance, J.; Chakraborty, T. Insights into a Novel bla(KPC-2)-Encoding IncP-6 Plasmid Reveal Carbapenem-Resistance Circulation in Several Enterobacteriaceae Species from Wastewater and a Hospital Source in Spain. Front. Microbiol. 2017, 8, 1–7. [Google Scholar] [CrossRef]
- Jeong, J.H.; Lee, J.H.; Lee, J.J.; Park, K.S.; Karim, A.M.; Lee, C.R.; Jeong, B.C.; Lee, S.H. Structural basis for carbapenem-hydrolyzing mechanisms of carbapenemases conferring antibiotic resistance. Int. J. Mol. Sci. 2015, 16, 9654–9692. [Google Scholar] [CrossRef] [PubMed]
- Castanheira, M.; Sader, H.S.; Farrell, D.J.; Mendes, R.E.; Jones, R.N. Activity of ceftaroline-avibactam tested against Gram-negative organism populations, including strains expressing one or more β-lactamases and methicillin-resistant Staphylococcus aureus carrying various staphylococcal cassette chromosome mec types. Antimicrob. Agents Chemother. 2012, 56, 4779–4785. [Google Scholar] [CrossRef] [PubMed]
- Naas, T.; Oueslati, S.; Bonnin, R.A.; Dabos, M.L.; Zavala, A.; Dortet, L.; Retailleau, P.; Iorga, B.I. Beta-lactamase database (BLDB) -structure and function. J. Enzym. Inhib. Med. Chem. 2017, 32, 917–919. [Google Scholar] [CrossRef]
- Fischer, J.; Rodríguez, I.; Schmoger, S.; Friese, A.; Roesler, U.; Helmuth, R.; Guerra, B. Escherichia coli producing VIM-1 carbapenemase isolated on a pig farm. J. Antimicrob. Chemother. 2012, 67, 1793–1795. [Google Scholar] [CrossRef]
- Pulss, S.; Semmler, T.; Prenger-Berninghoff, E.; Bauerfeind, R.; Ewers, C. First report of an Escherichia coli strain from swine carrying and OXA-181 carbapenemase and the colisitin determinant MCR-1. Int. J. Antimicrob. Agents 2017, 50, 232–236. [Google Scholar] [CrossRef]
- Fischer, J.; Rodríguez, I.; Schmoger, S.; Friese, A.; Roesler, U.; Helmuth, R.; Guerra, B. Salmonella enterica subsp. enterica producing VIM-1 carbapenemase isolated from livestock farms. J. Antimicrob. Chemother. 2013, 68, 478–480. [Google Scholar] [CrossRef]
- Martínez- Martínez, L. Extended spectrum β-lactamase and the permeability barrier. Clin. Microbiol. Infect. 2008, 14 (Suppl. 1), 82–89. [Google Scholar] [CrossRef]
- Thomson, K.S. Extended-spectrum-β-lactamase, AmpC, and carbapenemase issues. J. Clin. Microbiol. 2010, 48, 1019–1025. [Google Scholar] [CrossRef] [PubMed]
- von Salviati, C.; Laube, H.; Guerra, B.; Roesler, U.; Friese, A. Emission of ESBL/AmpC-producing Escherichia coli from pig fattening farms to surrounding areas. Vet. Microbiol. 2015, 175, 77–84. [Google Scholar] [CrossRef]
- Bush, K.; Bradford, P.A. β-lactams and β-lactamase inhibitors: An overview. Cold Spring Harb. Perspect. Med. 2016, 6, a025247. [Google Scholar] [CrossRef]
- Jacoby, G.A. AmpC beta-lactamases. Clin. Microbiol. Rev. 2009, 22, 161–182. [Google Scholar] [CrossRef] [PubMed]
- Bergšpica, I.; Kaprou, G.; Alexa, E.A.; Prieto, M.; Alvarez-Ordóñez, A. Extended Spectrum β-Lactamase (ESBL) Producing Escherichia coli in Pigs and Pork Meat in the European Union. Antibiotics 2020, 9, 678. [Google Scholar] [CrossRef]
- Dohmen, W.; Bonten, M.J.M.; Bos, M.E.H.; van Marm, S.; Scharringa, J.; Wagenaar, J.A.; Heederik, D.J.J. Carriage of extended spectrum β-lactamases in pig farmers is associated with occurrence in pigs. Clin. Microbiol. Infect. 2015, 21, 917–923. [Google Scholar] [CrossRef]
- Hounmanou, Y.M.G.; Bortolaia, V.; Thanh Dang, S.T.; Truong, D.; Olsen, J.E.; Dalsgaard, A. ESBL and AmpC β-Lactamase Encoding Genes in E. coli From Pig and Pig Farm Workers in Vietnam and Their Association with Mobile Genetic Elements. Front. Microbiol. 2021, 12, 629139. [Google Scholar] [CrossRef]
- Commission Implementing Decision 2013/652/EU of 12 November 2013 on the monitoring and reporting of antimicrobial resistance in zoonotic and commensal bacteria. Off. J. Eur. Union 2013, L303, 26–39.
- Jorgensen, J.H.; Maher, L.A.; Howell, A.W. Activity of meropenem against antibiotic-resistant or infrequently encountered Gram-negative bacilli. Antimicrob. Agents Chemother. 1991, 35, 2410–2414. [Google Scholar] [CrossRef][Green Version]
- Neu, H.C.; Novelli, A.; Chin, N.X. In vitro activity and beta-lactamase stability of a new carbapenem, SM-7338. Antimicrob. Agents Chemother. 1989, 33, 1009–1018. [Google Scholar] [CrossRef] [PubMed]
- Riera, E.; Cabot, G.; Mulet, X.; Garcıa-Castillo, M.; del Campo, R.; Juan, C.; Canton, R.; Oliver, A. Pseudomonas aeruginosa carbapenem resistance mechanisms in Spain: Impact on the activity of imipenem, meropenem and doripenem. J. Antimicrob. Chemother. 2011, 66, 2022–2027. [Google Scholar] [CrossRef] [PubMed]
- EUCAST (European Commettee on Antimicrobial Susceptibility Testing). Breakpoint Tables for Interpretations of MICs and Zone Diameters; Version 12.0; EUCAST: Växjö, Sweden, 2021; Available online: http://www.eucast.org (accessed on 3 March 2021).
- Klockgether, J.; Cramer, N.; Wiehlmann, L.; Davenport, C.; Tümmler, B. Pseudomonas aeruginosa Genomic Structure and Diversity. Front. Microbiol. 2011, 2, 150. [Google Scholar] [CrossRef]
- Molina-Mora, J.A.; Campos-Sánchez, R.; Rodríguez, C.; Shi, L.; García, F. High quality 3C de novo assembly and annotation of a multidrug resistant ST-111 Pseudomonas aeruginosa genome: Benchmark of hybrid and non-hybrid assemblers. Sci. Rep. 2020, 10, 1392. [Google Scholar] [CrossRef]
- Timme, R.E.; Wolfgang, W.J.; Balkey, M.; Gubbala Venkata, S.L.; Randolph, R.; Allard, M.; Strain, E. Optimizing Open Data to Support One Health: Best Practices to Ensure Interoperability of Genomic Data from Microbial Pathogens. One Health Outlook 2020, 2, 20. [Google Scholar] [CrossRef]
- Sims, D.; Sudbery, I.; Ilott, N.E.; Heger, A.; Ponting, C.P. Sequencing depth and coverage: Key considerations in genomic analyses. Nat. Rev. Genet. 2014, 15, 121–132. [Google Scholar] [CrossRef] [PubMed]
- Berends, B.R.; Van Knapen, F.; Snijeders, J.M.A.; Mossel, D.A.A. Identification and quantification of risk factors associated with Salmonella contamination on pork carcasses. Int. J. Food Microbiol. 1997, 36, 199–206. [Google Scholar] [CrossRef]
- Biasino, W.; De Zutter, L.; Mattheus, W.; Bertrand, S.; Uyttendaele, M.; Van Damme, I. Correlation between slaughter practices and the distribution of Salmonella and hygiene indicator bacteria on pig carcasses during slaughter. Food Microbiol. 2018, 70, 192–199. [Google Scholar] [CrossRef]
- Bonardi, S.; Bassi, L.; Brindani, F.; D’Incau, M.; Barco, L.; Carra, E.; Pongolini, S. Prevalence, characterization and antimicrobial susceptibility of Salmonella enterica and Yersinia enterocolitica in pigs at slaughter in Italy. Int. J. Food Microbiol. 2013, 163, 248–257. [Google Scholar] [CrossRef]
- EFSA (European Food Safety Authority). Report from the Task Force on Zoonoses Data Collection including guidance for harmonized monitoring and reporting of antimicrobial resistance in commensal Escherichia coli and Enterococcus spp. from food animals. EFSA J. 2008, 6, 141–144. [CrossRef]
- Livermore, D.M. Multiple mechanisms of antimicrobial resistance in Pseudomonas aeruginosa: Our worst nightmare? Clin. Infect. Dis. 2002, 34, 634–640. [Google Scholar] [CrossRef]
- Strateva, T.; Yordanov, D. Pseudomonas aeruginosa—A phenomenon of bacterial resistance. J. Med. Microbiol. 2009, 58, 1133–1148. [Google Scholar] [CrossRef]
- Wolter, D.J.; Lister, P.D. Mechanisms of β-lactam resistance among Pseudomonas aeruginosa. Curr. Pharm. Des. 2013, 19, 209–222. [Google Scholar] [CrossRef]
- Botelho, J.; Grosso, F.; Peixe, L. Antibiotic resistance in Pseudomonas aeruginosa—Mechanisms, epidemiology and evolution. Drug Resist. Updat. 2019, 44, 100640. [Google Scholar] [CrossRef] [PubMed]
- Micek, S.T.; Wunderink, R.G.; Kollef, M.H.; Chen, C.; Rello, J.; Chastre, J.; Antonelli, M.; Welte, T.; Clair, B.; Ostermann, H.; et al. An international multicenter retrospective study of Pseudomonas aeruginosa nosocomial pneumonia: Impact of multidrug resistance. Crit. Care 2015, 19, 219. [Google Scholar] [CrossRef]
- Zilberberg, M.D.; Shorr, A.F. Prevalence of multidrug-resistant Pseudomonas aeruginosa and carbapenem-resistant Enterobacteriaceae among specimens from hospitalized patients with pneumonia and bloodstream infections in the United States from 2000 to 2009. J. Hosp. Med. 2013, 8, 559–563. [Google Scholar] [CrossRef] [PubMed]
- WHO Regional Office for Europe and European Centre for Disease Prevention and Control. Surveillance of Antimicrobial Resistance in Europe, 2020 Data. Executive Summary; WHO Regional Office for Europe: Copenhagen, Denmark, 2021. [Google Scholar]
- Girlich, D.; Naas, T.; Nordmann, P. Biochemical characterization of the naturally occurring oxacillinase OXA-50 of Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 2004, 48, 2043–2048. [Google Scholar] [CrossRef]
- Overbeek, R.; Olson, R.; Pusch, G.D.; Olsen, G.J.; Davis, J.J.; Disz, T.; Edwards, R.A.; Gerdes, S.; Parrello, B.; Shukla, M.; et al. The SEED and the Rapid Annotation of microbial genomes using Subsystems Technology (RAST). Nucleic Acids Res. 2014, 42, D206–D214. [Google Scholar] [CrossRef]
- OIE (World Organization for Animal Health). OIE List of Antimicrobial Agents of Veterinary Importance. 2018. Available online: https://www.oie.int/app/uploads/2021/03/a-oie-list-antimicrobials-may2018.pdf (accessed on 21 March 2022).
- Leman, A.D.; Straw, B.E.; Mengeling, W.L.; Sylvie, D.; Taylor, D.J. Diseases of Swine, 7th ed.; Iowa State University Press: Ames, Iowa, 1992. [Google Scholar]
- Miniats, O.P.; Johnson, J.A. Experimental atrophic rhinitis in gnotobiotic pigs. Can. J. Comp. Med. 1980, 44, 358–365. Available online: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1320090/ (accessed on 10 April 2022).
- Baljer, G.; Barrett, J.T. Nachweis von enterotoxinbildenden Pseudomonas aeruginosa-Stämmen im Darmligaturtest beim Ferkel. J. Vet. Med. 2010, 26, 740–747. [Google Scholar] [CrossRef]
- ECDC (European Centre for Disease Prevention and Control). Surveillance of Antimicrobial Resistance in Europe, 2017 Data. Annual Report of the European Antimicrobial Resistance Surveillance Network (EARS-Net) 2017; ECDC: Stockholm, Sweden, 2018.
- Lautenbach, E.; Synnestvedt, M.; Weiner, M.G.; Bilker, W.B.; Vo, L.; Schein, J.; Kim, M. Imipenem resistance in Pseudomonas aeruginosa: Emergence, epidemiology, and impact on clinical and economic outcomes. Infect Control. Hosp. Epidemiol. 2010, 31, 47–53. [Google Scholar] [CrossRef]
- Venditti, C.; Butera, O.; Proia, A.; Rigacci, L.; Mariani, B.; Parisi, G.; Messina, F.; Capone, A.; Nisii, C.; Di Caro, A. Reduced susceptibility to carbapenems in a Klebsiella pneumoniae clinical isolate producing SCO-1 and CTX-M-15 β lactamases together with OmpK35 and OmpK36 porin deficiency. Antimicrob. Agents Chemother. 2020, 64, e00556-20. [Google Scholar] [CrossRef] [PubMed]
- Wise, M.G.; Horvath, E.; Young, K.; Sahm, D.F.; Kazmierczak, K.M. Global survey of Klebsiella pneumoniae major porins from ertapenem non-susceptible isolates lacking carbapenemases. J. Med. Microbiol. 2018, 67, 289–295. [Google Scholar] [CrossRef]
- Andersen, V.D.; Jensen, V.F.; Vigre, H.; Andreasen, M.; Agersø, Y. The use of third and fourth generation cephalosporins affects the occurrence of extended-spectrum cephalosporinase-producing Escherichia coli in Danish pig herds. Vet. J. 2015, 204, 345–350. [Google Scholar] [CrossRef] [PubMed]
- Pauly, N.; Hammerl, J.A.; Grobbel, M.; Käsbohrer, A.; Tenhagen, B.A.; Malorny, B.; Schwarz, S.; Meemken, D.; Irrgang, A. Identification of a bla(VIM-1)-Carrying IncA/C(2) Multiresistance Plasmid in an Escherichia coli Isolate Recovered from the German Food Chain. Microorganisms 2020, 9, 29. [Google Scholar] [CrossRef]
- Irrgang, A.; Tausch, S.H.; Pauly, N.; Grobbel, M.; Kaesbohrer, A.; Hammerl, J.A. First Detection of GES-5-Producing Escherichia coli from Livestock-An Increasing Diversity of Carbapenemases Recognized from German Pig Production. Microorganisms 2020, 8, 1593. [Google Scholar] [CrossRef]
- Irrgang, A.; Pauly, N.; Tenhagen, B.A.; Grobbel, M.; Kaesbohrer, A.; Hammerl, A.J.A. Spill-Over from Public Health? First Detection of an OXA-48-Producing Escherichia coli in a German Pig Farm. Microorganisms 2020, 8, 855. [Google Scholar] [CrossRef]
- Diaconu, E.L.; Carfora, V.; Alba, P.; Di Matteo, P.; Stravino, F.; Buccella, C.; Dell’Aira, E.; Onorati, R.; Sorbara, L.; Battisti, A.; et al. Novel IncFII plasmid harbouring blaNDM-4 in a carbapenem-resistant Escherichia coli of pig origin, Italy. J. Antimicrob. Chemother. 2020, 75, 3475–3479. [Google Scholar] [CrossRef]
- Rega, M.; Carmosino, I.; Bonilauri, P.; Frascolla, V.; Vismarra, A.; Bacci, C. Prevalence of ESβL, AmpC and Colistin-Resistant, E. coli in Meat: A Comparison between Pork and Wild Boar. Microorganisms 2021, 9, 214. [Google Scholar] [CrossRef]
- Hassan, S.; Amer, S.; Mittal, C.; Sharma, R. Ewingella americana: An emerging true pathogen. Case Rep. Infect. Dis. 2012, 2012, 730720. [Google Scholar] [CrossRef]
- Meisler, S.; Kamity, R.; Noor, A.; Krilov, L.; Tiozzo, C. First Case of Ewingella americana Meningitis in a Term Newborn: A Rare but Real Pathogen. Front. Pediatr. 2020, 8, 308. [Google Scholar] [CrossRef]
- Esteban, J.; Martin, J.; Ortiz, A.; Santos-O’Connor, F.; Cabria, F.; Reyero, A. Pseudomonas oryzihabitans peritonitis in a patient on continuous ambulatory peritoneal dialysis. Clin. Microbiol. Infect. 2002, 8, 607–608. [Google Scholar] [CrossRef][Green Version]
- Nei, T.; Sonobe, K.; Onodera, A.; Itabashi, T.; Yamaguchi, H.; Maeda, M.; Saito, R. Two cases with bacteremia suspected to be due to relatively rare Pseudomonas (Flavimonas) oryzihabitans. J. Infect. Chemother. 2015, 21, 751–755. [Google Scholar] [CrossRef]
- Govan, J.R.; Hughes, J.E.; Vandamme, P. 19 Burkholderia cepacia: Medical, taxonomic and ecological isues. J. Med. Microbiol. 1996, 45, 395–407. [Google Scholar] [CrossRef]
- Mukerji, R.; Kakarala, R.; Smith, S.J.; Kusz, H.G. Chryseobacterium indologenes: An emerging infection in the USA. BMJ Case Rep. 2016, 2016, bcr2016214486. [Google Scholar] [CrossRef]
- Poole, T.L.; Schlosser, W.D.; Anderson, R.C.; Norman, K.N.; Beier, R.C.; Nisbet, D.J. Whole-Genome Sequence of Aeromonas hydrophila CVM861 Isolated from Diarrhetic Neonatal Swine. Microorganisms 2020, 8, 1648. [Google Scholar] [CrossRef]
- Igbinosa, I.H.; Igbinosa, E.O.; Okoh, A.I. Antibiogram characterization and putative virulence genes in Aeromonas species isolated from pig fecal samples. Environ. Sci. Pollut. Res. Int. 2016, 23, 12199–12205. [Google Scholar] [CrossRef] [PubMed]
- Janda, J.M.; Abbott, S.L. The Genus Aeromonas: Taxonomy, Pathogenicity, and Infection. Clin. Micro. Rev. 2010, 23, 35–73. [Google Scholar] [CrossRef]
- Manfredi, R.; Nanetti, A.; Ferri, M.; Mastroianni, A.; Coronado, O.V.; Chiodo, F. Flavobacterium spp. organisms as opportunistic bacterial pathogens during advanced HIV disease. J. Infect. 1999, 39, 146–152. [Google Scholar] [CrossRef]
- Brooke, J.S. Stenotrophomonas maltophilia: An emerging global opportunistic pathogen. Clin. Microbiol. Rev. 2012, 25, 2–41. [Google Scholar] [CrossRef]
- Abbott, I.J.; Slavin, M.A.; Turnidge, J.D.; Thursky, K.A.; Worth, L.J. Stenotrophomonas maltophilia: Emerging disease patterns and challenges for treatment. Expert Rev. Anti. Infect. Ther. 2011, 9, 471–488. [Google Scholar] [CrossRef] [PubMed]
- Ibn Saied, W.; Merceron, S.; Schwebel, C.; Le Monnier, A.; Oziel, J.; Garrouste-Orgeas, M.; Marcotte, G.; Ruckly, S.; Souweine, B.; Darmon, M.; et al. Ventilator-associated pneumonia due to Stenotrophomonas maltophilia: Risk factors and outcome. J. Infect. 2020, 80, 279–285. [Google Scholar] [CrossRef]
- Marchaim, D.; Chopra, T.; Perez, F.; Hayakawa, K.; Lephart, P.R.; Bheemreddy, S.; Blunden, C.; Hujer, A.M.; Rudin, S.; Shango, M.; et al. Outcomes and genetic relatedness of carbapenem-resistant Enterobacteriaceae at Detroit medical center. Infect. Control. Hosp. Epidemiol. 2011, 32, 861–871. [Google Scholar] [CrossRef][Green Version]
- Xu, L.; Sun, X.; Ma, X. Systematic review and meta-analysis of mortality of patients infected with carbapenem-resistant Klebsiella pneumoniae. Ann. Clin. Microbiol. Antimicrob. 2017, 16, 18. [Google Scholar] [CrossRef] [PubMed]
- DTU (Technical University of Denmark—National Food Institute). Laboratory Protocol—Isolation of ESBL, AmpC- and Carbapenemase-Producing E. coli from Caecal Samples; Version 5; DTU: Kongens Lyngby, Denmark, 2017; Available online: https://www.eurl-ar.eu/CustomerData/Files/Folders/21-protocols/530_esbl-ampc-cpeprotocol-version-caecal-v7-09-12-19.pdf (accessed on 1 March 2022).
- EUCAST (European Committee on Antimicrobial Susceptibility Testing). EUCAST Disc Diffusion Test Methodology. 2017. Available online: http://www.eucast.org (accessed on 30 November 2017).
- EUCAST (European Committee on Antimicrobial Susceptibility Testing). Breakpoint Tables for Interpretations of MICs and Zone Diameters, Version 8.0. 2017. Available online: http://www.eucast.org (accessed on 30 November 2017).
- EUCAST (European Committee on Antimicrobial Susceptibility Testing). EUCAST Reading Guide for Broth Microdilution. 2021. Available online: http://www.eucast.org (accessed on 3 March 2021).
- Adeolu, M.; Alnajar, S.; Naushad, S.; Gupta, R.S. Genome-based phylogeny and taxonomy of the ‘Enterobacteriales’: Proposal for Enterobacterales ord. nov. divided into the families Enterobacteriaceae, Erwiniaceae fam. Nov., Pectobacteriaceae fam. Nov., Yersiniaceae fam. Nov., Hafniaceae fam. Nov., Morganellaceae fam. Nov., and Budviciaceae fam Nov. Int. J. Syst. Evol. Microbiol. 2016, 65, 5575–5599. [Google Scholar] [CrossRef]
- CDC (Center for Disease Control and Prevention). Laboratory Protocol for Detection of Carbapenem-Resistant or Carbapenemase-Producing Klebsiella spp. and E. coli from Rectal Swabs; CDC: Atlanta, GA, USA, 2009.
- Doyle, D.; Peirano, G.; Lascols, C.; Lloyd, T.; Church, D.L.; Pitout, J.D. Laboratory detection of Enterobacteriaceae that produce carbapenemases. J. Clin. Microbiol. 2012, 50, 3877–3880. [Google Scholar] [CrossRef]
- Roschansky, N.; Fischer, J.; Guerra, B.; Roesler, U. Development of a Multiplex Real-Time PCR for the rapid detection of the predominant Beta-Lactamase genes CTX-M, SHV, TEM and CIT type AmpCs in Enterobacteriaceae. PLoS ONE 2014, 9, e100956. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Pérez, F.J.; Hanson, N.D. Detection of plasmid-mediated AmpC—Lactamase genes in clinical isolates by using Multiplex PCR. J. Clin. Microbiol. 2002, 40, 2153–2162. [Google Scholar] [CrossRef] [PubMed]
- INNUca. 2021. Available online: https://github.com/B-UMMI/INNUca (accessed on 1 December 2021).
- Contig_info. 2022. Available online: https://gitlab.pasteur.fr/GIPhy/contig_info (accessed on 15 May 2022).
- Snippy. 2021. Available online: https://github.com/tseemann/snippy (accessed on 1 December 2021).
- FastANI. 2022. Available online: https://github.com/ParBLiSS/FastANI (accessed on 15 May 2022).
- RAxML. 2022. Available online: https://github.com/stamatak/standard-RAxML (accessed on 15 May 2022).
- Itol. 2021. Available online: https://itol.embl.de/ (accessed on 1 December 2021).
- ChewBBACA. 2022. Available online: https://github.com/B-UMMI/chewBBACA (accessed on 15 May 2022).
- de Sales, R.O.; Migliorini, L.B.; Puga, R.; Kocsis, B.; Severino, P. A core genome multilocus sequence typing scheme for Pseudomonas aeruginosa. Front. Microbiol. 2020, 11, 1049. [Google Scholar] [CrossRef] [PubMed]
- Phyloviz. 2022. Available online: http://www.phyloviz.net/ (accessed on 15 May 2022).
- ABRicate. 2021. Available online: https://github.com/tseemann/abricate/ (accessed on 1 December 2021).
- Center for Genomic Epidemiology. 2021. Available online: https://cge.cbs.dtu.dk/services/ResFinder/ (accessed on 9 March 2021).
- Prokka. 2022. Available online: https://github.com/tseemann/prokka (accessed on 15 May 2022).
- SnapGene. 2021. Available online: https://www.snapgene.com/ (accessed on 1 December 2021).
- National Center for Biotechnology Information. 2021. Available online: http://www.ncbi.nlm.nih.gov (accessed on 1 December 2021).
| Species | MIC Values (Lg/mL) | bla Genes | ampC Genes | ||
|---|---|---|---|---|---|
| MEM | CAZ | CTX | |||
| Enterobacterales | |||||
| C. freundii | 0.016 | 2 | 8 | TEM-1 | CIT |
| E. agglomerans | 0.032 | 128 | 512 | CTX-M-1, TEM-1 | |
| E. coli | 4 | ≤0.5 | ≤0.5 | TEM-1 | |
| E. coli | 0.016 | 0.5 | 0.5 | TEM-1 | |
| E. coli | 0.016 | 8 | 4 | CTX-M-1, TEM-1 | |
| E. coli | 0.016 | 32 | 512 | TEM-1, SHV | |
| E. americana | 4 | 32 | 16 | CTX-M-1, TEM-1 | |
| Pseudomonas (no breakpoints for CTX) | |||||
| P. oryzihabitans | 256 | 256 | 32 | TEM-1 | |
| Acinetobacter (no breakpoints for CAZ and CTX) | |||||
| A. lwoffii | 0.016 | 0.5 | 8 | TEM-1 | |
| Species with no breakpoints | |||||
| A. hydrophila | 4 | 32 | 64 | CTX-M-1, TEM-1 | |
| A. hydrophila | 4 | 2 | 0.5 | TEM-1 | |
| B. cepacia | 4 | 8 | 16 | TEM-1 | |
| B. cepacia | 8 | 16 | 64 | CTX-M-1, TEM-1, SHV | |
| B. cepacia | 8 | 16 | 64 | CTX-M-1, TEM-1 | |
| C. indologenes | 16 | 512 | 256 | TEM-1 | |
| F. odoratum | 16 | 16 | 64 | TEM-1 | |
| S. maltophilia | 128 | 64 | 256 | TEM-1 | |
| S. maltophilia | 64 | 32 | 256 | TEM-1 | EBC |
| S. maltophilia | 64 | 8 | 64 | TEM-1 | EBC |
| S. maltophilia | 64 | 64 | 256 | TEM-1 | ACC |
| S. maltophilia | 64 | 64 | 64 | TEM-1 | CIT |
| S. maltophilia | 64 | 128 | 128 | CTX-M-1, TEM-1 | CIT |
| S. maltophilia | 64 | 128 | 256 | TEM-1 | CIT |
| S. maltophilia | 32 | 16 | 256 | TEM-1 | CIT |
| S. maltophilia | 32 | 64 | 256 | CTX-M-1, TEM-1 | CIT |
| S. maltophilia | 32 | 128 | 256 | CTX-M-1, TEM-1 | CIT |
| S. maltophilia | 32 | 64 | 256 | CTX-M-1, TEM-1 | CIT |
| S. maltophilia | 128 | 64 | 256 | TEM-1 | CIT |
| S. maltophilia | 128 | 4 | 64 | TEM-1 | CIT |
| S. maltophilia | 16 | 16 | 64 | TEM-1 | CIT |
| S. maltophilia | 16 | 128 | 512 | TEM-1 | CIT |
| S. maltophilia | 8 | 64 | 128 | TEM-1 | |
| S. maltophilia | 4 | 2 | 16 | TEM-1 | CIT |
| S. maltophilia | 1 | 2 | 128 | TEM-1 | |
| Source | ID Code | MIC Values (µg/mL) | Multi Locus Sequence Typing | bla Genes | Additional Resistance Genes |
|---|---|---|---|---|---|
| MEM | |||||
| pig | CRE 98 | 2 | 274 | OXA-486, PAO | aph(3′)-IIb, crpP, fosA4, catB7 |
| pig | CRE 102 | 16 | 938 | OXA-396, PAO | aph(3′)-IIb, fosA4, catB7 |
| pig | CRE 153 | 16 | 782 | OXA-50, PAO | aph(3′)-IIb, crpP, fosA4, catB7 |
| pig | CRE 295 | 2 | 885 | OXA-50, PAO | aph(3′)-IIb, crpP, fosA4, catB7 |
| human | NEF 23 | 16 | 938 | OXA-396, PAO | aph(3′)-IIb, fosA4, catB7 |
| human | NEF 156 | 8 | 395 | OXA-488, PAO | aph(3′)-IIb, crpP, fosA4, catB7 |
| Assembly ID | Source | Genome Size | GC% | No. of Contig | Coverage | N50 | Maximum Contig Length |
|---|---|---|---|---|---|---|---|
| NEF 23 | human | 6,436,450 | 66.40 | 201 | 92.5 | 79,626 | 377,949 |
| CRE 98 | pig | 6,415,664 | 66.40 | 124 | 99.8 | 126,411 | 561,477 |
| CRE 102 | pig | 6,438,088 | 66.40 | 225 | 76.6 | 75,398 | 238,923 |
| CRE 153 | pig | 6,396,234 | 66.43 | 317 | 79.2 | 38,674 | 223,278 |
| CRE 295 | pig | 6,426,083 | 66.35 | 141 | 105.2 | 102,638 | 328,392 |
| NEF 156 | human | 7,099,705 | 65.85 | 172 | 85.9 | 123,256 | 340,617 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bonardi, S.; Cabassi, C.S.; Manfreda, G.; Parisi, A.; Fiaccadori, E.; Sabatino, A.; Cavirani, S.; Bacci, C.; Rega, M.; Spadini, C.; et al. Survey on Carbapenem-Resistant Bacteria in Pigs at Slaughter and Comparison with Human Clinical Isolates in Italy. Antibiotics 2022, 11, 777. https://doi.org/10.3390/antibiotics11060777
Bonardi S, Cabassi CS, Manfreda G, Parisi A, Fiaccadori E, Sabatino A, Cavirani S, Bacci C, Rega M, Spadini C, et al. Survey on Carbapenem-Resistant Bacteria in Pigs at Slaughter and Comparison with Human Clinical Isolates in Italy. Antibiotics. 2022; 11(6):777. https://doi.org/10.3390/antibiotics11060777
Chicago/Turabian StyleBonardi, Silvia, Clotilde Silvia Cabassi, Gerardo Manfreda, Antonio Parisi, Enrico Fiaccadori, Alice Sabatino, Sandro Cavirani, Cristina Bacci, Martina Rega, Costanza Spadini, and et al. 2022. "Survey on Carbapenem-Resistant Bacteria in Pigs at Slaughter and Comparison with Human Clinical Isolates in Italy" Antibiotics 11, no. 6: 777. https://doi.org/10.3390/antibiotics11060777
APA StyleBonardi, S., Cabassi, C. S., Manfreda, G., Parisi, A., Fiaccadori, E., Sabatino, A., Cavirani, S., Bacci, C., Rega, M., Spadini, C., Iannarelli, M., Crippa, C., Ruocco, F., & Pasquali, F. (2022). Survey on Carbapenem-Resistant Bacteria in Pigs at Slaughter and Comparison with Human Clinical Isolates in Italy. Antibiotics, 11(6), 777. https://doi.org/10.3390/antibiotics11060777







