Recent Developments in Methicillin-Resistant Staphylococcus aureus (MRSA) Treatment: A Review
Abstract
1. Introduction
1.1. Evolution of MRSA Treatment
1.2. Seriousness of MRSA
1.3. Current Threat
2. Current Therapeutic Strategies and Existing Drugs
2.1. Natural Extracts for MRSA Intervention
- ➢
- The increased penetrability of the cell wall membrane.
- ➢
- Prohibition of the efflux pump systems.
- ➢
- Modification of the active site.
- ➢
- Enzymatic ruin and alteration of bacterial enzymes.
2.2. Natural Drugs
2.3. Existing Synthetic Chemical Entities
2.4. Combining Synthetic and Natural Drugs
2.5. Multi-Drug Strategies
2.6. Bacteriophage Therapy
2.7. Bacteriophage-Antibiotic Therapy
2.8. Vaccines for MRSA Treatment
3. Current Treatment Approaches and Advances
3.1. Chemotherapeutics for MRSA
3.2. Nanomedicinal Platforms
3.3. Antimicrobial Activity of Surfactant Molecules
3.4. Radiation Therapy
4. In Vitro and In Vivo Studies
5. Clinical Analysis
6. Computational Analysis
Molecular Dynamics
7. Future Directions for MRSA Treatment
- Nanocarriers with good surface area for controlled release of antibiotic with minimum inhibitory concentrations.
- The design and development of antibody-based biologic agent treatments for the management of serious infections of MRSA.
- Multi-drug strategies for treating drug-resistant bacterial infections such as synthetic drugs, synthetic and natural drugs, and natural medicines.
- Breakage of biofilms formed by MRSA using a suitable targeting carrier system with the biotic drugs.
8. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Ogston, A. Report upon microorganisms in surgical diseases. Br. Med. J. 1881, 1, 369–375. [Google Scholar] [CrossRef] [PubMed]
- Ogston, A. Micrococcus poisoning. J. Anat. Physiol. 1882, 16, 526–567. [Google Scholar] [PubMed]
- Cowan, S.T.; Shaw, C.; Williams, R.E. Type strain for Staphylococcus aureus Rosenbach. J. Gen. Microbiol. 1954, 10, 174–176. [Google Scholar] [CrossRef] [PubMed]
- Cowan, S.T. Classification of staphylococci by slide agglutination. J. Pathol. Bacteriol. 1939, 48, 169–173. [Google Scholar] [CrossRef]
- Taylor, T.A.; Unakal, C.G. Staphylococcus aureus. StatPearls; 2020. Available online: https://www.ncbi.nlm.nih.gov/books/NBK441868/ (accessed on 27 May 2021). [PubMed]
- Turner, N.A.; Sharma-Kuinkel, B.K.; Maskarinec, S.A.; Eichenberger, E.M.; Shah, P.P.; Carugati, M.; Holland, T.L.; Fowler, V.G., Jr. Methicillin-resistant Staphylococcus: An overview of basic and clinical research. Nat. Rev. Microbiol. 2019, 17, 203–218. [Google Scholar] [CrossRef]
- Diekema, D.J.; Pfaller, M.A.; Schmitz, F.J.; Smayevsky, J.; Bell, J.; Jones, R.N.; Beach, M.; SENTRY Participants Group. Survey of infections due to Staphylococcus species: Frequency of occurrence and antimicrobial susceptibility of isolates collected in the United States, Canada, Latin America, Europe, and the Western Pacific region for the SENTRY Antimicrobial Surveillance Program, 1997–1999. Clin. Infect. Dis. 2001, 32 (Suppl. 2), S114–S132. [Google Scholar]
- Schito, G.C. The importance of the development of antibiotic resistance in Staphylococcus aureus. Clin. Microbiol. Infect. 2006, 12 (Suppl. 1), S3–S8. [Google Scholar] [CrossRef]
- Lindsay, J.A.; Holden, M.T. Staphylococcus aureus: Superbug, super genome? Trends Microbiol. 2004, 12, 378–385. [Google Scholar] [CrossRef]
- Lowy, F.D. Staphylococcus aureus infections. N. Engl. J. Med. 1998, 339, 520–532. [Google Scholar] [CrossRef]
- Wertheim, H.F.; Melles, D.C.; Vos, M.C.; van Leeuwen, W.; van Belkum, A.; Verbrugh, H.A.; Nouwen, J.L. The role of nasal carriage in Staphylococcus aureus infections. Lancet Infect. Dis. 2005, 5, 751–762. [Google Scholar] [CrossRef]
- Kirby, W.M. Extraction of a highly potent penicillin inactivator from penicillin resistant staphylococci. Science 1994, 99, 452–453. [Google Scholar] [CrossRef] [PubMed]
- Rammelkamp, C.H.; Maxon, T. Resistance of Staphylococcus aureus to the action of penicillin. Proc. Soc. Exp. Biol. Med. 1942, 51, 386–389. [Google Scholar] [CrossRef]
- Jevons, M.P. Celbenin-resistant staphylococci. Br. Med. J. 1961, 1, 124–125. [Google Scholar] [CrossRef]
- Hartman, B.J.; Tomasz, A. Low-affinity penicillin-binding protein associated with beta-lactam resistance in Staphylococcus aureus. J. Bacteriol. 1984, 158, 513–516. [Google Scholar] [CrossRef] [PubMed]
- Reynolds, P.E.; Brown, D.F. Penicillin-binding proteins of beta-lactam-resistant strains of Staphylococcus aureus. Effect of growth conditions. FEBS Lett. 1985, 192, 28–32. [Google Scholar] [CrossRef]
- Utsui, Y.; Yokota, T. Role of an altered penicillin-binding protein in methicillin and cephem-resistant Staphylococcus aureus. Antimicrob. Agents Chemother. 1985, 28, 397–403. [Google Scholar] [CrossRef] [PubMed]
- Matcuhashi, M.; Song, M.D.; Ishino, F.; Wachi, M.; Doi, M.; Inoue, M.; Ubukata, K.; Yamashita, N.; Konno, M. Molecular cloning of the gene of a penicillin-binding protein supposed to cause high resistance to beta-lactam antibiotics in Staphylococcus aureus. J. Bacteriol. 1986, 167, 975–980. [Google Scholar] [CrossRef]
- Katayama, Y.; Ito, T.; Hiramatsu, K. A new class of genetic element, staphylococcus cassette chromosome mec, encodes methicillin resistance in Staphylococcus aureus. Antimicrob. Agents Chemother. 2000, 44, 1549–1555. [Google Scholar] [CrossRef]
- Hiramatsu, K.; Asada, K.; Suzuki, E.; Okonogi, K.; Yokota, T. Molecular cloning and nucleotide sequence determination of the regulator region of mecA gene in methicillin-resistant Staphylococcus aureus (MRSA). FEBS Lett. 1992, 298, 133–136. [Google Scholar] [CrossRef]
- Mickymaray, S.; Alturaiki, W.; Al-Aboody, M.S.; Mariappan, P.; Rajenderan, V.; Alsagaby, S.A.; Kalyanasundram, U.; Alarfajj, A.A. Antibacterial efficacy of bacteriocin produced by marine Bacillus subtilis against clinically important extended spectrum beta-lactamase strains and methicillin-resistant Staphylococcus aureus. IJMRHS 2018, 7, 75–83. [Google Scholar]
- Hetem, D.J.; Westh, H.; Boye, K.; Jarlov, J.O.; Bonten, M.J.; Bootsma, M.C. Nosocomail transmission of community-associated methicillin-resistant Staphylococcus aureus in Danish hospitals. J. Antimicrob. Chemother. 2012, 67, 1775–1780. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Moodley, A.; Oosthuysen, W.F.; Duse, A.G.; Marais, E.; South African MSG. Molecular characterization of clinical methicillin-resistant Staphylococcus aureus isolates in South Africa. J. Clin. Microbiol. 2010, 48, 4608–4611. [Google Scholar] [CrossRef] [PubMed][Green Version]
- King, M.D.; Humphrey, B.J.; Wang, Y.F.; Kourbatova, E.V.; Ray, S.M.; Blumberg, H.M. Emergence of community-acquired methicillin-resistant Staphylococcus aureus USA 300 clone as the predominant cause of skin and soft-tissue infections. Ann. Intern. Med. 2006, 144, 309–317. [Google Scholar] [CrossRef]
- Semple, S.J.; Reynolds, G.D.; O’Leary, M.C.; Flower, R.L. Screening of Australian medicinal plants for antiviral activity. J. Ethnopharmacol. 1998, 60, 163–172. [Google Scholar] [CrossRef]
- Bassett, I.B.; Pannowitz, D.L.; Barnetson, R.S. A comparative study of tea-tree oil versus benzylperoxide in the treatment of acne. Med. J. Aust. 1990, 153, 455–458. [Google Scholar] [CrossRef] [PubMed]
- Hemaiswarya, S.; Kruthiventi, A.K.; Doble, M. Synergism between natural products and antibiotics against infectious diseases. Phytomedicine 2008, 15, 639–652. [Google Scholar] [CrossRef]
- Nassan, M.A.; Mohamed, E.H.; Abdelhafez, S.; Ismail, T.A. Effect of clove and cinnamon extracts on experimental model of acute hematogenous pyelonephritis in albino rats: Immunopathological and antimicrobial study. Int. J. Immunopathol. Pharmacol. 2015, 28, 60–68. [Google Scholar] [CrossRef]
- Badei, A.; Faheld, S.; El-Akel, A.; Mahnoud, B. Application of some species in flavoring and preservation of cookies: 2-Antimicrobial and sensory properties of cardamom, cinnamon and clove. Dtsch. Lebensm. Rundsch. 2002, 98, 261–265. [Google Scholar]
- Hussain, M.B.; Hannan, A.; Absar, M.; Butt, N.S. In vitro susceptibility of methicillin-resistant Staphylococcus aureus to honey. Complement. Ther. Clin. Pract. 2017, 27, 57–60. [Google Scholar] [CrossRef]
- Sherlock, O.; Dolan, A.; Athman, R.; Power, A.; Gethin, G.; Cowman, S.; Humphreys, H. Comparison of the antimicrobial activity of Ulmo honey from Chile and Manuka honey against methicillin-resistant Staphylococcus aureus, Escherichia coli and Pseudomonas aeruginosa. BMC Complement. Altern. Med. 2010, 10, 47. [Google Scholar] [CrossRef]
- Tan, H.T.; Rahman, R.A.; Gan, S.H.; Halim, A.S.; Hassan, S.A.; Sulaiman, S.A.; Kaur, B.K. The antibacterial properties of Malaysian tualang honey against wound and enteric microorganisms in comparison to manuka honey. BMC Complement. Altern. Med. 2009, 9, 34. [Google Scholar] [CrossRef] [PubMed]
- Okwu, M.U.; Olley, M.; Akpoka, A.O.; Izevbuwa, O.E. Methicillin-resistant Staphylococcus aureus (MRSA) and anti-MRSA activities of extracts of some medicinal plants: A brief review. AIMS Microbiol. 2019, 5, 117–137. [Google Scholar] [CrossRef] [PubMed]
- Sahu, M.C.; Padhy, R.N. In vitro antibacterial potency of Butea monosperma Lam. against twelve clinically isolated multidrug resistant bacteria. Asian. Pac. J. Trop. Dis. 2013, 3, 217–226. [Google Scholar] [CrossRef]
- Voravuthikunchai, S.P.; Kitpipit, L. Activity of medicinal plant extracts against hospital isolates of Methicillin-resistant Staphylococcus aureus (MRSA). Clin. Microb. Infect. 2005, 11, 493–512. [Google Scholar] [CrossRef] [PubMed]
- Gomber, C.; Saxena, S. Anti-staphylococcal potential of Callistemon rigidus. Cent. Eur. J. Med. 2006, 2, 79–88. [Google Scholar] [CrossRef]
- Aliyu, A.B.; Musa, A.M.; Abdullahi, M.S. Activity of plant extracts used in Northern Nigerian traditional medicine against Methicillin-resistant Staphylococcus aureus (MRSA). Niger. J. Pharmaceu. Sci. 2008, 7, 1–8. [Google Scholar]
- Akinjogunla, O.J.; Yah, C.S.; Eghafona, N.O. Antibacterial activity of the leave extracts of Nymphaea lotus (Nymphaeceae) on Methicillin-resistant Staphylococcus aureus and Vancomycin-resistant S. aureus isolated from clinical samples. Annals. Biol. Res. 2010, 1, 174–184. [Google Scholar]
- Invasive Species Compendium-CABI. Available online: https://www.cabi.org/isc/datasheet/28765 (accessed on 27 May 2021).
- Invasive Species Compendium-CABI. Available online: https://www.cabi.org/isc/datasheet/24882 (accessed on 27 May 2021).
- Invasive Species Compendium-CABI. Available online: https://www.cabi.org/isc/datasheet/39510 (accessed on 27 May 2021).
- Invasive Species Compendium-CABI. Available online: https://www.cabi.org/isc/datasheet/45141 (accessed on 27 May 2021).
- National Resources Conservation Service, United States Department of Agriculture (USDA). Available online: https://plants.usda.gov/core/profile?symbol=PUGR2 (accessed on 27 May 2021).
- Globinmed. Available online: https://www.globinmed.com/index.php?option (accessed on 27 May 2021).
- Arefin, K.; Rahman, M.; Uddin, M.Z.; Hassan, A. Angiosperm flora of Satchari Natural Park, Habiganj, Bangladesh. Bangl. J. Plant. Taxon. 2011, 18, 117–140. [Google Scholar] [CrossRef]
- Al-Alusi, N.T.; Kadir, F.A.; Ismail, S.; Abdullah, M. In vitro interaction of combined plants: Tinospora crispa and Swietenia mahagoni against Methicillin-resistant Staphylococcus aureus (MRSA). Afr. J. Microbiol. Res. 2010, 4, 2309–2312. [Google Scholar]
- Wikaningtyas, P.; Sukandar, E.Y. The antibacterial activity of selected plants towards resistant bacteria isolated from clinical specimens. Asian. Pac. J. Trop. Biomed. 2016, 6, 16–19. [Google Scholar] [CrossRef]
- Zuo, G.Y.; Zhang, X.J.; Yang, C.X.; Han, J.; Wang, G.C.; Bian, Z.Q. Evaluation of traditional Chinese medicinal plants for anti-MRSA activity with reference to the treatment record of infectious diseases. Molecules 2012, 17, 2955–2967. [Google Scholar] [CrossRef] [PubMed]
- Heyman, H.M.; Hussein, A.A.; Meyer, J.J.M.; Lall, N. Antibacterial activity of South African medicinal plants against Methicillin-resistant Staphylococcus aureus (MRSA). Pharm. Biol. 2009, 47, 67–71. [Google Scholar] [CrossRef]
- Uddin, Q.; Samiulla, L.; Singh, V.K.; Jamil, S.S. Phytochemical and Pharmacological Profile of Withania somnifera Dunal: A review. J. Appl. Pharm. Sci. 2012, 2, 170–175. [Google Scholar]
- Sucilathangam, G.; Gomatheswari, S.N.; Velvizhi, G.; Vincent, C.P.; Palaniappan, N. Detection of antibacterial activity of Medicinal plant Quercus infectoria against methicillin-resistant Staphylococcus aureus (MRSA) isolates in clinical samples. J. Pharm. Biomed. Sci. 2012, 14, 1–5. [Google Scholar]
- Armas, J.R.; Quiroz, J.R.; Roman, R.A.; Sanchez, J.O.; Pacheco, M.P.; Valdivia, L.H.; Rivera, E.C.; Asmat, R.C.; Anampa-Guzmán, A. Antibacterial activities of essential oils from three medicinal plants in combination with EDTA against MRSA. Br. Microbiol. Res. J. 2016, 17, 1–10. [Google Scholar] [CrossRef]
- Belley, A.; McKay, G.A.; Arhin, F.F.; Sarmiento, I.; Beaulieu, S.; Fadhil, I.; Parr, T.R., Jr.; Moeck, G. Oritavancin disrupts membrane integrity of Staphylococcus aureus and vancomycin-resistant enterococci to effect rapid bacterial killing. Antimicrob. Agents Chemother. 2010, 54, 5369–5371. [Google Scholar] [CrossRef]
- Carpenter, C.F.; Chambers, H.F. Daptomycin: Another novel agent for treating infections due to drug-resistant Gram-positive pathogens. Clin. Infect. Dis. 2004, 38, 994–1000. [Google Scholar] [CrossRef]
- Hughes, J.; Mellows, G. Inhibition of isoleucyl-transfer ribonucleic acid synthetase in Echerichia coli by pseudomonic acid. Biochem. J. 1978, 176, 305–318. [Google Scholar] [CrossRef]
- French, G.L. What’s new and not so new on the antimicrobial horizon? Clin. Microbiol. Infect. 2008, 14 (Suppl. 6), 19–29. [Google Scholar] [CrossRef]
- Kuok, C.F.; Hoi, S.O.; Hoi, C.F.; Chan, C.H.; Fong, I.H.; Ngok, C.K.; Meng, L.R.; Fong, P. Synergistic antibacterial effects of herbal extracts and antibiotics on methicillin-resistant Staphylococcus aureus: A computational and experimental study. Exp. Biol. Med. 2017, 242, 731–743. [Google Scholar] [CrossRef]
- Blesson, J.; Saji, C.V.; Nivya, R.M.; Kumar, R. Synergistic Antibacterial Activity of Natural Plant Extracts and Antibiotics against Methicillin Resistant Staphylococcus aureus (MRSA). WJPPS 2015, 4, 741–763. [Google Scholar]
- Jiang, Q.W.; Chen, M.W.; Cheng, K.J.; Yu, P.Z.; Wei, X.; Shi, Z. Therapeutic potential of steroidal alkaloids in cancer and other diseases. Med. Res. Rev. 2016, 36, 119–143. [Google Scholar] [CrossRef]
- Khameneh, B.; Iranshahy, M.; Ghandadi, M.; Atashbeyk, D.G.; Bazzaz, B.S.F.; Iranshahi, M. Investigation of the antibacterial activity and efflux pump inhibitory effect of co-loaded piperine and gentamicin nanoliposomes in methicillin-resistant Staphylococcus aureus. Drug. Dev. Ind. Pharm. 2015, 41, 989–994. [Google Scholar] [CrossRef] [PubMed]
- Tillotson, J.; Tillotson, G.S. The Regulatory Pathway for Antifungal Drugs: A US Perspective. Clin. Infect. Dis. 2015, 61, S678–S683. [Google Scholar] [CrossRef] [PubMed]
- Gudiol, F.; Aguado, J.M.; Almirante, B.; Bouza, E.; Cercenado, E.; Dominguez, M.A.; Gasch, O.; Lora-Tamayo, J.; Miró, J.M.; Palomar, M.; et al. Executive summary of the diagnosis and treatment of bacteremia and endocarditis due to Staphylococcus aureus. A clinical guideline from the Spanish Society of Clinical Microbiology and Infectious Diseases (SEIMC). Enferm. Infecc. Microbiol. Clínica 2015, 33, 626–632. [Google Scholar] [CrossRef]
- Bal, A.M.; David, M.Z.; Garau, J.; Gottlieb, T.; Mazzei, T.; Scaglione, F.; Tattevin, P.; Gould, I.M. Future trends in the treatment of Methicillin-Resistant Staphylococcus aureus (MRSA) infection: An in-depth review of newer antibiotics active against an enduring pathogen. J. Glob. Antimicrob. Resist. 2017, 10, 295–303. [Google Scholar] [CrossRef]
- Ippolito, J.A.; Kanyo, Z.F.; Wang, D.; Franceschi, F.J.; Moore, P.B.; Steitz, T.A.; Duffy, E.M. Crystal structure of the oxazolidinone antibiotic linezolid bound to the 50S ribosomal subunit. J. Med. Chem. 2008, 51, 3353–3356. [Google Scholar] [CrossRef]
- Lee, Y.C.; Chen, P.Y.; Wang, J.T.; Chang, S.C. A study on combination of daptomycin with selected antimicrobial agents: In vitro synergistic effect of MIC value of 1 mg/L against MRSA strains. BMC Pharmacol. Toxicol. 2019, 20, 25. [Google Scholar] [CrossRef]
- Debbia, E.; Pesce, A.; Schito, G.C. In vitro activity of LY146032 alone and in combination with other antibiotics against gram-positive bacteria. Antimicrob. Agents Chemother. 1988, 32, 279–281. [Google Scholar] [CrossRef]
- Saleh-Mghir, A.; Muller-Serieys, C.; Dinh, A.; Massias, L.; Cremieux, A.C. Adjunctive rifampin is crucial to optimizing daptomycin efficacy against rabbit prosthetic joint infection due to methicillin-resistant Staphylococcus aureus. Antimicrob. Agents Chemother. 2011, 55, 4589–4593. [Google Scholar] [CrossRef]
- Trookman, N.S.; Rizer, R.L.; Weber, T. Treatment of minor wounds from dermatologic procedures: A comparison of three topical wound care ointments using a laser wound model. J. Am. Acad. Dermatol. 2011, 64, S8–S15. [Google Scholar] [CrossRef] [PubMed]
- Deresinski, S. Vancomycin in Combination with other Antibiotics for the Treatment of Serious Methicillin-Resistant Staphylococcus aureus Infections. Clin. Infect. Dis. 2009, 49, 1072–1079. [Google Scholar] [CrossRef] [PubMed]
- Dilworth, T.J.; Ibrahim, O.; Hall, P.; Sliwinski, J.; Walraven, C.; Mercier, R.C. β-Lactams enhance vancomycin activity against Methicillin-Resistant Staphylococcus aureus bacteremia compared to vancomycin alone. Antimicrob. Agents Chemother. 2014, 58, 102–109. [Google Scholar] [CrossRef] [PubMed]
- Ruddaraju, L.K.; Pammi, S.V.N.; Guntuku, G.S.; Padavala, V.S.; Kolapalli, V.R.M. A review on anti-bacterials to combat resistance: From ancient era of plants and metals to present and future perspectives of green nano technological combinations. Asian J. Pharm. Sci. 2020, 15, 42–59. [Google Scholar] [CrossRef] [PubMed]
- Gurunathan, S. Biologically synthesized silver nanoparticles enhances antibiotic activity against gram-negative bacteria. J. Ind. Eng. Chem. 2015, 29, 217–226. [Google Scholar] [CrossRef]
- Ghosh, S.; Patil, S.; Ahire, M.; Kitture, R.; Kale, S.; Pardesi, K.; Cameotra, S.S.; Bellare, J.; Dhavale, D.D.; Jabgunde, A.; et al. Synthesis of silver nanoparticles using Dioscorea bulbifera tuber extract and evaluation of its synergistic potential in combination with antimicrobial agents. Int. J. Nanomed. 2012, 7, 483–496. [Google Scholar]
- Patra, J.K.; Baek, K.H. Biosynthesis of silver nanoparticles using aqueous extract of silky hairs of corn and investigation of its antibacterial and anticandidal synergistic activity and antioxidant potential. IET Nanobiotechnol. 2016, 10, 326–333. [Google Scholar] [CrossRef]
- Basker, S. Synergistic efficacy of antibiotics and silver nanoparticles synthesized from Eichhornia crassipes. Res. Plant. Biol. 2016, 6, 1–5. [Google Scholar] [CrossRef]
- Patra, J.K.; Baek, K.H. Novel green synthesis of gold nanoparticles using Citrullus lanatus rind and investigation of proteasome inhibitory activity, antibacterial, and antioxidant potential. Int. J. Nanomed. 2015, 10, 7253–7264. [Google Scholar]
- Patra, J.K.; Baek, K.H. Antibacterial activity and synergistic antibacterial potential of biosynthesized silver nanoparticles against foodborne pathogenic bacteria along with its anticandidal and antioxidant effects. Front. Microbiol. 2017, 8, 167. [Google Scholar] [CrossRef]
- Padalia, H.; Moteriya, P.; Chanda, S. Green synthesis of silver nanoparticles from marigold flower and its synergistic antimicrobial potential. Arab. J. Chem. 2015, 8, 732–741. [Google Scholar] [CrossRef]
- Rastogi, L.; Kora, A.J.; Sashidhar, R.B. Antibacterial effects of gum kondagogu reduced/stabilized silver nanoparticles in combination with various antibiotics: A mechanistic approach. Appl. Nanosci. 2015, 5, 535–543. [Google Scholar] [CrossRef]
- Singh, R.; Wagh, P.; Wadhwani, S.; Gaidhani, S.; Kumbhar, A.; Bellare, J.; Chopade, B.A. Synthesis, optimization, and characterization of silver nanoparticles from Acinetobacter calcoaceticus and their enhanced antibacterial activity when combined with antibiotics. Int. J. Nanomed. 2013, 8, 4277–4290. [Google Scholar]
- Gandhi, H.; Khan, S. Biological synthesis of silver nanoparticles and its antibacterial activity. J. Nanomed. Nanotechnol. 2016, 7, 366. [Google Scholar] [CrossRef]
- Barapatre, A.; Ram Aadil, K.; Jha, H. Synergistic antibacterial and antibiofilm activity of silver nanoparticles biosynthesized by lignin degrading fungus. Bioresour. Bioprocess. 2016, 3, 8. [Google Scholar] [CrossRef]
- Naqvi, S.Z.H.; Kiran, U.; Ali, M.I.; Jamal, A.; Hameed, A.; Ahmed, S.; Ali, N. Combined efficacy of biologically synthesized silver nanoparticles and different antibiotics against multidrug-resistant bacteria. Int. J. Nanomed. 2013, 8, 3187–3195. [Google Scholar] [CrossRef]
- Kumar, N.; Das, S.; Jyoti, A.; Kaushik, S. Synergistic effect of silver nanoparticles with doxycycline against klebsiella pneumonia. Int. J. Pharm. Sci. 2016, 8, 183–186. [Google Scholar]
- Wypij, M.; Czarnecka, J.; Swiecimska, M.; Dahm, H.; Rai, M.; Golinska, P. Synthesis, characterization and evaluation of antimicrobial and cytotoxic activities of biogenic silver nanoparticles synthesized from Streptomyces xinghaiensis OF1 strain. World J. Microbiol. Biotechnol. 2018, 34, 23. [Google Scholar] [CrossRef]
- Fayaz, A.M.; Balaji, K.; Girilal, M.; Yadav, R.; Kalaichelvan, P.T.; Venketesan, R. Biogenic synthesis of silver nanoparticles and their synergistic effect with antibiotics: A study against gram-positive and gram-negative bacteria. Nanomedicine 2010, 6, 103–109. [Google Scholar] [CrossRef]
- Łubowska, N.; Grygorcewicz, B.; Kosznik-Kwaśnicka, K.; Zauszkiewicz-Pawlak, A.; Węgrzyn, A.; Dołęgowska, B.; Piechowicz, L. Characterization of the Three New Kayviruses and Their Lytic Activity against Multidrug-Resistant Staphylococcus aureus. Microorganisms 2019, 7, 471. [Google Scholar] [CrossRef]
- Grygorcewicz, B.; Wojciuk, B.; Roszak, M.; Łubowska, N.; Błażejczak, P.; Jursa-Kulesza, J.; Rakoczy, R.; Masiuk, H.; Dołęgowska, B. Environmental Phage-Based Cocktail and Antibiotic Combination Effects on Acinetobacter baumannii Biofilm in a Human Urine Model. Microb. Drug Resist. 2021, 27, 25–35. [Google Scholar] [CrossRef] [PubMed]
- Clegg, J.; Soldaini, E.; McLoughlin, R.M.; Rittenhouse, S.; Bagnoli, F.; Phogat, S. Staphylococcus aureus Vaccine Research and Development: The Past, Present and Future, Including Novel Therapeutic Strategies. Front. Immunol. 2021, 12, 705360. [Google Scholar] [CrossRef] [PubMed]
- Hiramatsu, K.; Katayama, Y.; Matsuo, M.; Sasaki, T.; Morimoto, Y.; Sekiguchi, A.; Baba, T. Multi-drug resistant Staphylococcus aureus and future chemotherapy. J. Infect. Chemother. 2014, 20, 593–601. [Google Scholar] [CrossRef] [PubMed]
- Jacqueline, C.; Amador, G.; Batard, E.; Mabecque, V.L.; Miegeville, A.F.; Biek, D.; Caillon, J.; Potel, G. Comparison of ceftaroline fosamil, daptomycin and tigecycline in an experimental rabbit endocarditis model caused by methicillin-susceptible, methicillin-resistant and glycopeptide-intermediate Staphylococcus aureus. J. Antimicrob. Chemother. 2011, 66, 863–866. [Google Scholar] [CrossRef][Green Version]
- Wilcox, M.H.; Tack, K.J.; Bouza, E.; Herr, D.L.; Ruf, B.R.; Ijzerman, M.M.; Croos-Dabrera, R.V.; Kunkel, M.; Knirsch, C. Complicated skin and skin-structure infections and catheter-related bloodstream infections: Noninfereiority of linezolid in a phase 3 study. Clin. Infect. Dis. 2009, 48, 203–212. [Google Scholar] [CrossRef]
- Tsai, C.Y.; Lee, C.H.; Chien, C.C.; Chen, I.L. Impact of teicoplanin maintenance dose and MIC values on the clinical outcomes of patients treated for methicillin-resistant Staphylococcus aureus bacteremia. Infect. Drug Resist. 2018, 11, 1205–1217. [Google Scholar] [CrossRef]
- Stryjewski, M.E.; O’Riordan, W.D.; Lau, W.K.; Pien, F.D.; Dunbar, L.M.; Vallee, M.; Fowler, V.G., Jr.; Chu, V.H.; Spencer, E.; Barriere, S.L.; et al. FAST Investigator Group. Telavancin versus Standard Therapy for Treatment of Complicated Skin and Soft-Tissue Infections Due to Gram-Positive Bacteria. Clin. Infect. Dis. 2005, 40, 1601–1607. [Google Scholar] [CrossRef][Green Version]
- Huh, A.J.; Kwon, Y.J. Nanoantibiotics: A new paradigm for treating infectious diseases using nanomaterials in the antibiotics resistant era. J. Control. Release 2011, 156, 128–145. [Google Scholar] [CrossRef]
- Mamun, M.M.; Sorinolu, A.J.; Munir, M.; Vejerano, E.P. Nanoantibiotics: Functions and Properties at the Nanoscale to Combat Antibiotic Resistance. Front. Chem. 2021, 9, 687660. [Google Scholar] [CrossRef]
- Hassan, D.; Omolo, C.A.; Fasiku, V.O.; Elrashedy, A.A.; Mocktar, C.; Nkambule, B.; Soliman, M.E.S.; Govender, T. Formulation of pH-Responsive Quatsomes from Quaternary Bicephalic Surfactants and Cholesterol for Enhanced Delivery of Vancomycin against Methicillin Resistant Staphylococcus aureus. Pharmaceutics 2020, 12, 1093. [Google Scholar] [CrossRef]
- Giersing, B.K.; Dastgheyb, S.S.; Modjarrad, K.; Moorthy, V. Status of vaccine research and development of vaccines for Staphylococcus aureus. Vaccine 2016, 34, 2962–2966. [Google Scholar] [CrossRef] [PubMed]
- Knetsch, M.L.W.; Koole, L.H. New Strategies in the Development of Antimicrobial Coatings: The Example of Increasing Usage of Silver and Silver Nanoparticles. Polymers 2011, 3, 340–366. [Google Scholar] [CrossRef]
- Vimbela, G.V.; Ngo, S.M.; Fraze, C.; Yang, L.; Stout, D.A. Antibacterial properties and toxicity from metallic nanomaterials. Int. J. Nanomed. 2017, 12, 3941–3965. [Google Scholar] [CrossRef] [PubMed]
- Tancer, R.J.; Baynes, K.; Wiedman, G.R. Synergy among humimycins against methicillin-resistant Staphylococcus aureus. Pept. Sci. 2020, 113, e24197. [Google Scholar] [CrossRef]
- Dong, P.T.; Mohammad, H.; Hui, J.; Leanse, L.G.; Li, J.; Liang, L.; Dai, T.; Seleem, M.N.; Cheng, J.X. Photolysis of Staphyloxanthin in Methicillin-Resistant Staphylococcus aureus Potentiates Killing by Reactive Oxygen Species. Adv. Sci. 2019, 6, 1900030. [Google Scholar] [CrossRef] [PubMed]
- Eftekhar, F.; Dadaei, T. Biofilm formation and detection of IcaAB genes in clinical isolates of methicillin resistant Staphylococcus aureus. Iran. J. Basic Med. Sci. 2011, 14, 132–136. [Google Scholar]
- National Research Council (US) Committee on Methods of Producing Monoclonal Antibodies; National Academies Press (US): Washington, DC, USA. Available online: https://www.ncbi.nlm.nih.gov/books/NBK100199 (accessed on 27 May 2021).
- Mohammad, H.; Cushman, M.; Seleem, M.N. Antibacterial Evaluation of Synthetic Thiazole Compounds In vitro and In vivo in a Methicillin-Resistant Staphylococcus aureus (MRSA) Skin Infection Mouse Model. PLoS ONE 2015, 10, e0142321. [Google Scholar] [CrossRef]
- Coello, R.; Glynn, J.R.; Gaspar, C.; Picazo, J.J.; Fereres, J. Risk factors for developing clinical infection with methicillin-resistant Staphylococcus aureus (MRSA) amongst hospital patients initially only colonized with MRSA. J. Hosp. Infect. 1997, 37, 39–46. [Google Scholar] [CrossRef]
- Jang, H.C.; Kim, S.H.; Kim, K.H.; Kim, C.J.; Lee, S.; Song, K.H.; Jeon, J.H.; Park, W.B.; Kim, H.B.; Park, S.W.; et al. Salvage treatment for persistent methicillin-resistant Staphylococcus aureus bacteremia: Efficacy of linezolid with or without carbapenem. Clin. Infect. Dis. 2009, 49, 395–401. [Google Scholar] [CrossRef]
- Lai, C.C.; Sheng, W.H.; Wang, J.T.; Cheng, A.; Chuang, Y.C.; Chen, Y.C.; Chang, S.C. Safety and efficacy of high-dose daptomycin as salvage therapy for severe gram-positive bacterial sepsis in hospitalized adult patients. BMC Infect. Dis. 2013, 13, 66. [Google Scholar] [CrossRef]
- Caelli, M.; Porteous, J.; Carson, C.F.; Heller, R.; Riley, T.V. Tea tree oil as an alternative topical decolonization agent for methicillin-resistant Staphylococcus aureus. J. Hosp. Infect. 2000, 46, 236–237. [Google Scholar] [CrossRef]
- Huiskes, R.; Chao, E.Y.S. A survey of finite element analysis in orthopedic biomechanics: The first decade. J. Biomech. 1983, 16, 385–409. [Google Scholar] [CrossRef]
- Sangeetha, M.; Saranya, T.S.; Sathianarayanan, S.; Vyshnavi, A.M.H.; Namboori, P.K.K. Design and Development of Potential Flavonoid Moiety for Pbp2a Inhibition for MRSA Therapy—A Computational Technique. Biomed. Pharma. J. 2020, 13, 687–692. [Google Scholar] [CrossRef]
- Skariyachan, S.; Krishnan, R.S.; Siddapa, S.B.; Salian, C.; Bora, P.; Sebastian, D. Computer aided screening and evaluation of herbal therapeutics against MRSA infections. Bioinformation 2011, 7, 222–233. [Google Scholar] [CrossRef] [PubMed]
- Gioia, D.; Bertazzo, M.; Recanatini, M.; Masetti, M.; Cavalli, A. Dynamic Docking: A Paradigm Shift in Computational Drug Discovery. Molecules 2017, 22, 2029. [Google Scholar] [CrossRef] [PubMed]
- Cavalli, A.; Bottegoni, G.; Raco, C.; De Vivo, M.; Recanatini, M. A computational study of the binding of propidium to the peripheral anionic site of human acetylcholinesterase. J. Med. Chem. 2004, 47, 3991–3999. [Google Scholar] [CrossRef]
- Alonso, H.; Bliznyuk, A.A.; Gready, J.E. Combining docking and molecular dynamic simulations in drug design. Med. Res. Rev. 2006, 26, 531–568. [Google Scholar] [CrossRef]
- Decherchi, S.; Masetti, M.; Vyalov, I.; Rocchia, W. Implicit solvent methods for free energy estimation. Eur. J. Med. Chem. 2015, 91, 27–42. [Google Scholar] [CrossRef]
- Amaro, R.E.; Baron, R.; McCammon, J.A. An improved relaxed complex scheme for receptor flexibility in computer-aided drug design. J. Comput. Aided Mol. Des. 2008, 22, 693–705. [Google Scholar] [CrossRef]
- Lin, J.H.; Perryman, A.L.; Schames, J.R.; McCammon, J.A. Computational drug design accommodating receptor flexibility: The relaxed complex scheme. J. Am. Chem. Soc. 2002, 124, 5632–5633. [Google Scholar] [CrossRef]
- Buonfiglio, R.; Ferraro, M.; Falchi, F.; Cavalli, A.; Masetti, M.; Recanatini, M. Collecting and Assessing Human Lactate Dehydrogenase-A Conformations for Structure-Based Virtual Screening. J. Chem. Inf. Model. 2013, 53, 2792–2797. [Google Scholar] [CrossRef] [PubMed]
- Rasmussen, R.V.; Fowler Jr, V.G.; Skov, R.; Bruun, N.E. Future challenges and treatment of Staphylococcus aureus bacteremia with emphasis on MRSA. Future Microbiol. 2011, 6, 43–56. [Google Scholar] [CrossRef] [PubMed]
| S. No | Botanical Name | Common Name | Plant Part Used | Extracting Solvent | MIC/MBC (mg/mL) MRSA | Ref. |
|---|---|---|---|---|---|---|
| 1. | Butea monosperma Lam. | Flame-of-the-forest | Leaf | Ethanol | 5.91/13.30 | [34] |
| 2. | Acacia catechu (L. f.) Willd | Cutch tree, black catechu | Wood | Ethanol | 1.6–3.2/25 | [35] |
| 3. | Callistemon rigidus R.Br. | Stiff bottlebrush | Leaf | Methanol | 0.00125–0.08 | [36] |
| 4. | Acacia albida Del. | Gawo | Stem bark | Methanol | 3.0/4.0 | [37] |
| 5. | Nymphaea lotus Linn. | White lotus | Leaf | Ethanol | 5.0–10.0/10.0–30.0 | [38] |
| 6. | Impatiens balsamina | Garden balsam | Leaf | Ethanol | 6.3/25 | [39] |
| 7. | Garcinia mangostana L. | Mangosteen | Fruit shell | Ethanol | 0.05–0.4/0.1–0.4 | [40] |
| 8. | Peltophorum pterocarpum (DC.) | Yellow flame tree | Bark | Ethanol | 0.1–0.8/6.3 | [41] |
| 9. | Psidium guajava L. | Guava | Leaf | Ethanol | 0.2–1.6/6.3 | [42] |
| 10. | Punica granatum L. | Pomegranate | Fruit shell | Ethanol | 0.2–0.4/1.6–3.2 | [43] |
| 11. | Uncaria gambir (Hunter) Roxb. | Gambier, White cutch | Leaf, stem | Ethanol | 0.4–0.8/3.2 | [44] |
| 12. | Walsura robusta | Bonlichu | Wood | Ethanol | 1.6–3.2/25 | [45] |
| 13. | Swietenia mahagoni | Mahagoni | Seed | Ethanol | 0.2–0.78/0.78–1.56 | [46] |
| 14. | Senna alata | Candle bush | Leaf | Ethanol | 0.5/ND | [47] |
| 15. | Garcinia Morella Desr. | Gamboge | Whole plant | Ethanol | 0.016–0.064/0.064–0.26 | [48] |
| 16. | Melianthus major L. | Giant honey flower | Leaves | Ethanol | 0.78/3.12 | [49] |
| 17. | Melianthus comosus Vahl | Honey flower | Leaves | Ethanol | 0.39/1.56 | [49] |
| 18. | Withania somnifera L. | Ashwagandha | Roots & leaves | Ethanol | 1.56/>6.25 | [50] |
| 19. | Quercus infectoria Olivier | Oak galls | Nutgalls | Ethanol | 0.4–3.2/3.2–6.3 | [51] |
| 20. | Thymus vulgaris L. | Thyme | Leaves | Essential oil | 0.057/ND | [52] |
| Nano Particles | Source of Reducing Agent | Combination of Antibiotic | Targeted Bacteria | Ref. |
|---|---|---|---|---|
| Ag NP | Leaf extract of Typha angustifolia | Gentamicin, cefotaxime, meropenem | E. coli, K. pneumonia | [72] |
| Ag NP | Dioscorea bulbifera tuber extract | β-lactam (piperacillin) and macrolide (erythromycin) | A. baumannii | [73] |
| Ag NP | Silky hairs of aqueous corn extract | Kanamycin and rifampicin | E. coli, S. aureus | [74] |
| Ag NP | Eichhornia crassipes | Vancomycin, penicillin, streptomycin, and tetracycline | S. aureus, E. coli, K. pneumonia, Enterococcus | [75] |
| Au NP | Citrullus lanatus rind extract | Kanamycin, rifampicin | Bacillus cereus, E. coli, Listeria monocytogenes, S. aureus, S. typhi | [76] |
| Ag NP | Corn leaf waste of Zea mays extract | Kanamycin and rifampicin | E. coli ATCC 42890, Bacillus cereus ATCC 13061 19115, and S. aureus ATCC 49444 | [77] |
| Ag NP | Flower broth of Tagetes erecta | Commercial antibiotics | Gram-positive (S. aureus and B. cereus), Gram-negative (E. coli and P. aeruginosa) bacteria | [78] |
| Ag NP | Gum kondagogu | Ciprofloxacin, streptomycin, and gentamycin | Gram-positive (S. aureus 25923, S. aureus 49834) and Gram-negative (E. coli 25922, P. aeruginosa 27853) | [79] |
| Ag NP | Acinetobacter calcoaceticus | Aminoglycosides, β-lactams, cephalosporins, tetracyclines | A. baumannii, P. aeruginosa, S. aureus, S. typhi | [80] |
| Ag NP | E. coli | Bacitracin, ampicillin, kanamycin | Corynebacterium diphtheria, E. coli, K. pneumonia | [81] |
| Ag NP | Aspergillus flavus and Emericella nidulans | Amikacin, kanamycin and streptomycin | E. coli, S. aureus and P. aeruginosa | [82] |
| Ag NP | Aspergillus flavus | Ciprofloxacin, vancomycin, gentamycin | Gram-positive and Gram-negative bacteria | [83] |
| Ag NP | Bacteria from petroleum soil | Doxycycline | Klebsiella pneumonia | [84] |
| Ag NP | Streptomyces xinghaiensis OF1 strain | Ampicillin, kanamycin, and tetracycline | E. coli, P. aeruginosa, S. aureus and Klebsiella pneumonia | [85] |
| Ag NP | Trichoderma viride | Ampicillin, kanamycin, and erythromycin | Gram-positive and Gram-negative bacteria | [86] |
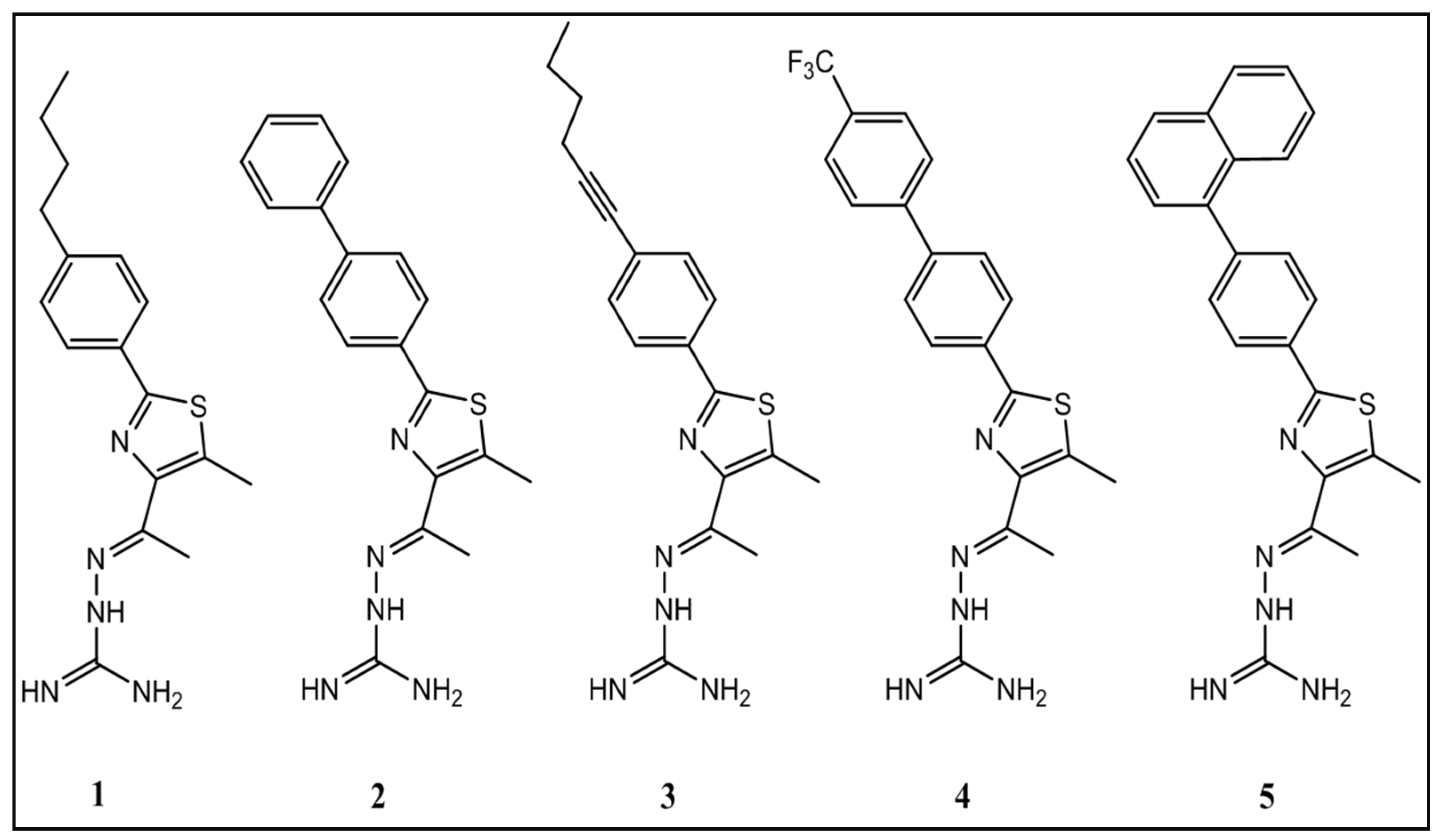
| Compound | S. aureus Strain Number | |||||
|---|---|---|---|---|---|---|
| NRS107 | USA300 | USA400 | USA800 | USA1000 | USA1100 | |
| 1 | 1.3 | 1.3 | 1.3 | 1.3 | 1.3 | 1.3 |
| 2 | 2.8 | 2.8 | 2.8 | 2.8 | 2.8 | 2.8 |
| 3 | 2.8 | 5.6 | 5.6 | 5.6 | 2.8 | 5.6 |
| 4 | 13.3 | 13.3 | 13.3 | 13.3 | 13.3 | 13.3 |
| 5 | 6.4 | 6.4 | 12.8 | 6.4 | 12.8 | 6.4 |
| Mupirocin | 1024.0 | 1.0 | 1.0 | 4.0 | 4.0 | 4.0 |
| Clindamycin | 0.1 | 1.8 | 0.1 | 0.1 | 0.1 | 0.1 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nandhini, P.; Kumar, P.; Mickymaray, S.; Alothaim, A.S.; Somasundaram, J.; Rajan, M. Recent Developments in Methicillin-Resistant Staphylococcus aureus (MRSA) Treatment: A Review. Antibiotics 2022, 11, 606. https://doi.org/10.3390/antibiotics11050606
Nandhini P, Kumar P, Mickymaray S, Alothaim AS, Somasundaram J, Rajan M. Recent Developments in Methicillin-Resistant Staphylococcus aureus (MRSA) Treatment: A Review. Antibiotics. 2022; 11(5):606. https://doi.org/10.3390/antibiotics11050606
Chicago/Turabian StyleNandhini, Palanichamy, Pradeep Kumar, Suresh Mickymaray, Abdulaziz S. Alothaim, Jayaprakash Somasundaram, and Mariappan Rajan. 2022. "Recent Developments in Methicillin-Resistant Staphylococcus aureus (MRSA) Treatment: A Review" Antibiotics 11, no. 5: 606. https://doi.org/10.3390/antibiotics11050606
APA StyleNandhini, P., Kumar, P., Mickymaray, S., Alothaim, A. S., Somasundaram, J., & Rajan, M. (2022). Recent Developments in Methicillin-Resistant Staphylococcus aureus (MRSA) Treatment: A Review. Antibiotics, 11(5), 606. https://doi.org/10.3390/antibiotics11050606







