Fecal Shedding of Multidrug Resistant Escherichia coli Isolates in Dogs Fed with Raw Meat-Based Diets in Brazil
Abstract
1. Introduction
2. Results
2.1. Phylogroups and Virulence Factors
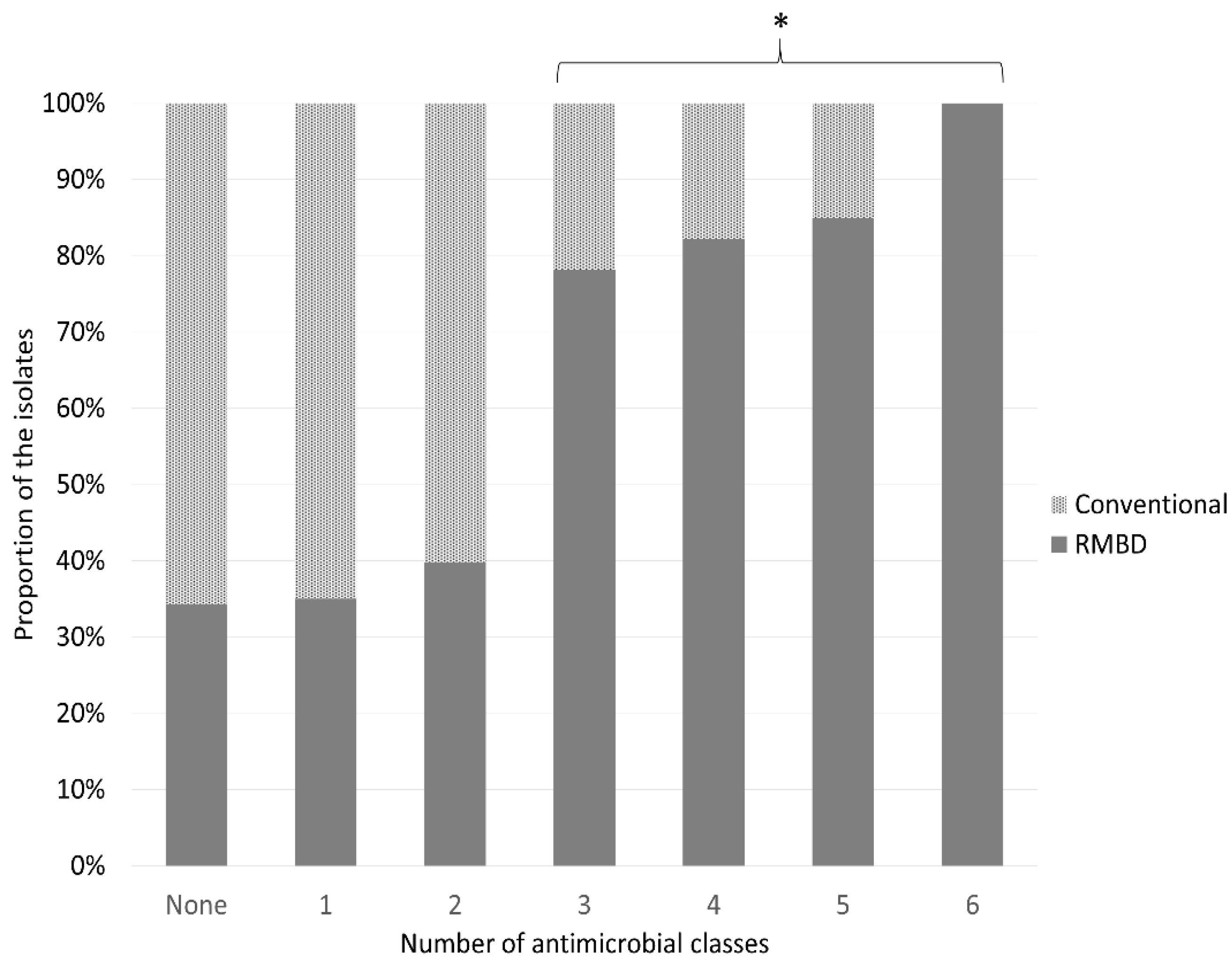
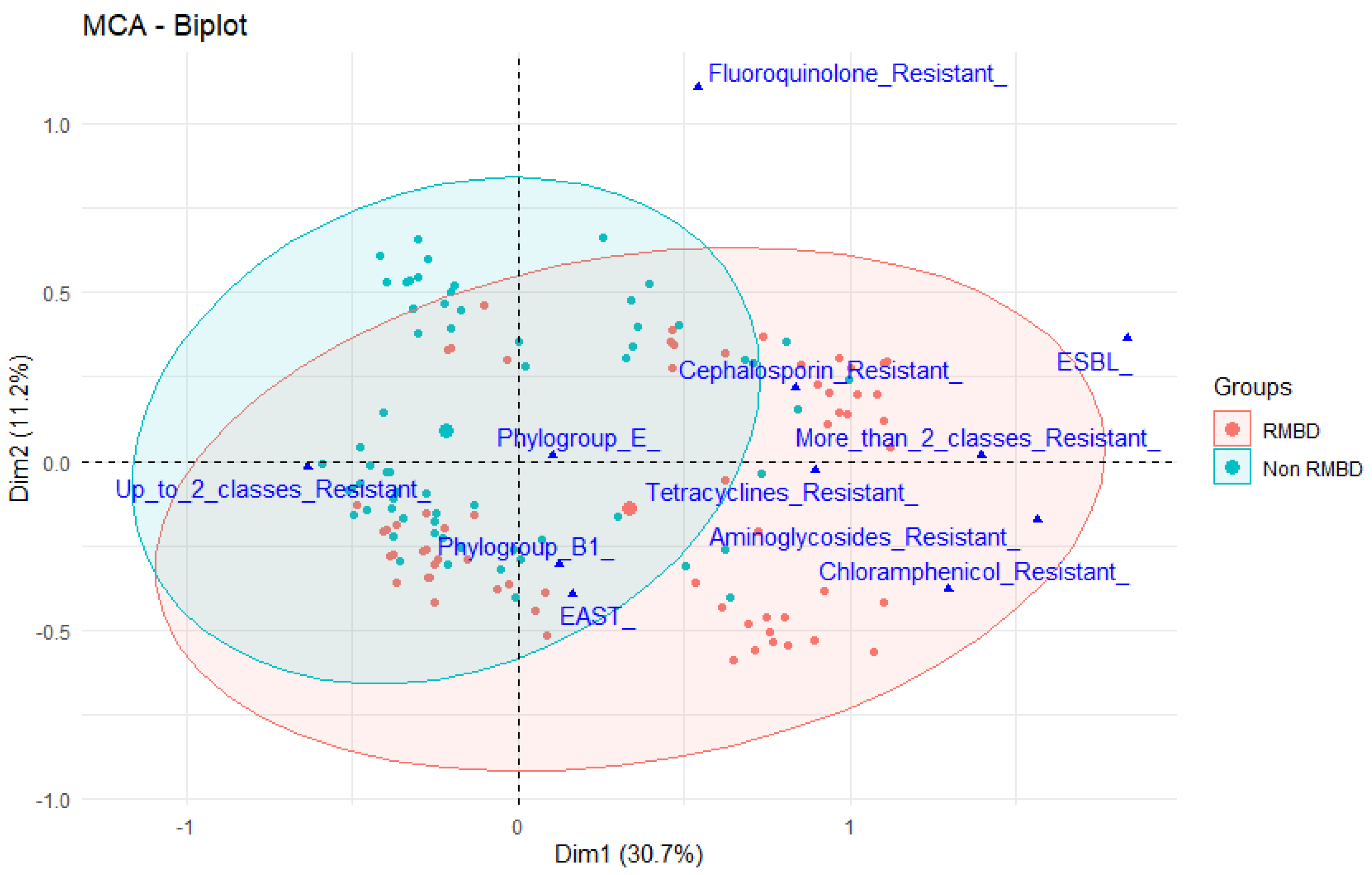
2.2. Antimicrobial Resistance
3. Discussion
4. Materials and Methods
4.1. Sampling
4.2. Isolation and Characterization of Escherichia coli Strains
4.3. Antimicrobial Susceptibility
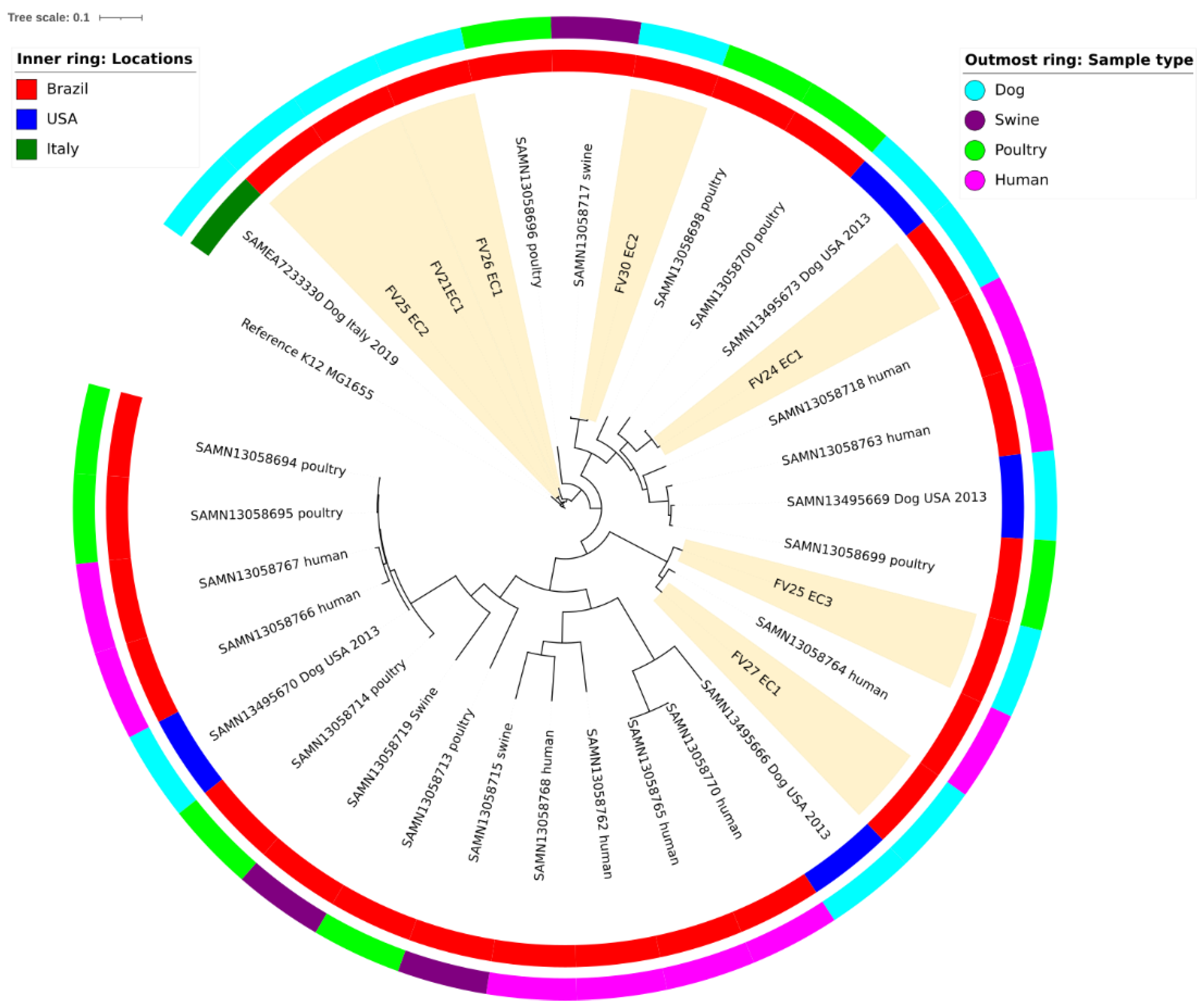
4.4. Whole-Genome Sequencing Analysis
4.5. Statistical Analysiss
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Davies, R.H.; Lawes, J.R.; Wales, A.D. Raw Diets for Dogs and Cats: A Review, with Particular Reference to Microbiological Hazards. J. Small Anim. Pract. 2019, 60, 329–339. [Google Scholar] [CrossRef]
- Freeman, L.M.; Chandler, M.L.; Hamper, B.A.; Weeth, L.P. Current Knowledge about the Risks and Benefits of Raw Meat–Based Diets for Dogs and Cats. J. Am. Vet. Med. Assoc. 2013, 243, 1549–1558. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; An, J.-U.; Kim, W.; Lee, S.; Cho, S. Differences in the Gut Microbiota of Dogs (Canis Lupus Familiaris) Fed a Natural Diet or a Commercial Feed Revealed by the Illumina MiSeq Platform. Gut Pathog. 2017, 9, 68. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, M.; Unterer, S.; Suchodolski, J.S.; Honneffer, J.B.; Guard, B.C.; Lidbury, J.A.; Steiner, J.M.; Fritz, J.; Kolle, P. The Fecal Microbiome and Metabolome Differs between Dogs Fed Bones and Raw Food (BARF) Diets and Dogs Fed Commercial Diets. PLoS ONE 2018, 13, e0201279. [Google Scholar] [CrossRef] [PubMed]
- Viegas, F.M.; Ramos, C.P.; Xavier, R.G.C.; Bagno, R.M.; Lopes, E.O.; Oliveira Junior, C.A.; Diniz, A.N.; Lobato, F.C.F.; Silva, R.O.S. Fecal Shedding of Salmonella Spp., Clostridium Perfringens, and Clostridioides difficile in Dogs Fed Raw Meat-Based Diets in Brazil and Their Owners’ Motivation. PLoS ONE 2020, 15, e0231275. [Google Scholar] [CrossRef]
- Lefebvre, S.L.; Reid-Smith, R.; Boerlin, P.; Weese, J.S. Evaluation of the Risks of Shedding Salmonellae and Other Potential Pathogens by Therapy Dogs Fed Raw Diets in Ontario and Alberta: The Risks of Shedding Salmonellae by Therapy Dogs Fed Raw Diets. Zoonoses Public Health 2008, 55, 470–480. [Google Scholar] [CrossRef] [PubMed]
- Morgan, S.K.; Willis, S.; Shepherd, M.L. Survey of Owner Motivations and Veterinary Input of Owners Feeding Diets Containing Raw Animal Products. PeerJ 2017, 5, e3031. [Google Scholar] [CrossRef]
- Van Bree, F.P.J.; Bokken, G.C.A.M.; Mineur, R.; Franssen, F.; Opsteegh, M.; van der Giessen, J.W.B.; Lipman, L.J.A.; Overgaauw, P.A.M. Zoonotic Bacteria and Parasites Found in Raw Meat-Based Diets for Cats and Dogs. Vet. Rec. 2018, 182, 50. [Google Scholar] [CrossRef]
- Baede, V.O.; Broens, E.M.; Spaninks, M.P.; Timmerman, A.J.; Graveland, H.; Wagenaar, J.A.; Duim, B.; Hordijk, J. Raw Pet Food as a Risk Factor for Shedding of Extended-Spectrum Beta-Lactamase-Producing Enterobacteriaceae in Household Cats. PLoS ONE 2017, 12, e0187239. [Google Scholar] [CrossRef]
- Wedley, A.L.; Dawson, S.; Maddox, T.W.; Coyne, K.P.; Pinchbeck, G.L.; Clegg, P.; Nuttall, T.; Kirchner, M.; Williams, N.J. Carriage of Antimicrobial Resistant Escherichia Coli in Dogs: Prevalence, Associated Risk Factors and Molecular Characteristics. Vet. Microbiol. 2017, 199, 23–30. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, V.M.; Pinchbeck, G.L.; Nuttall, T.; McEwan, N.; Dawson, S.; Williams, N.J. Antimicrobial Resistance Risk Factors and Characterisation of Faecal E. Coli Isolated from Healthy Labrador Retrievers in the United Kingdom. Prev. Vet. Med. 2015, 119, 31–40. [Google Scholar] [CrossRef] [PubMed]
- Baede, V.O.; Wagenaar, J.A.; Broens, E.M.; Duim, B.; Dohmen, W.; Nijsse, R.; Timmerman, A.J.; Hordijk, J. Longitudinal Study of Extended-Spectrum-β-Lactamase- and AmpC-Producing Enterobacteriaceae in Household Dogs. Antimicrob. Agents Chemother. 2015, 59, 3117–3124. [Google Scholar] [CrossRef] [PubMed]
- Johnson, J.R.; Stell, A.L.; Delavari, P. Canine Feces as a Reservoir of Extraintestinal Pathogenic Escherichia Coli. Infect. Immun. 2001, 69, 1306–1314. [Google Scholar] [CrossRef] [PubMed]
- Johnson, J.R.; Clabots, C. Sharing of Virulent Escherichia Coli Clones among Household Members of a Woman with Acute Cystitis. Clin. Infect. Dis. 2006, 43, e101–e108. [Google Scholar] [CrossRef] [PubMed]
- Johnson, J.R.; Johnston, B.; Clabots, C.R.; Kuskowski, M.A.; Roberts, E.; DebRoy, C. Virulence Genotypes and Phylogenetic Background of Escherichia Coli Serogroup O6 Isolates from Humans, Dogs, and Cats. J. Clin. Microbiol. 2008, 46, 417–422. [Google Scholar] [CrossRef] [PubMed]
- Johnson, J.R.; O’Bryan, T.T.; Low, D.A.; Ling, G.; Delavari, P.; Fasching, C.; Russo, T.A.; Carlino, U.; Stell, A.L. Evidence of Commonality between Canine and Human Extraintestinal Pathogenic Escherichia coli Strains That Express papG Allele III. Infect. Immun. 2000, 68, 3327–3336. [Google Scholar] [CrossRef]
- Croxen, M.A.; Law, R.J.; Scholz, R.; Keeney, K.M.; Wlodarska, M.; Finlay, B.B. Recent Advances in Understanding Enteric Pathogenic Escherichia coli. Clin. Microbiol. Rev. 2013, 26, 822–880. [Google Scholar] [CrossRef] [PubMed]
- Clermont, O.; Christenson, J.K.; Denamur, E.; Gordon, D.M. The Clermont Escherichia coli Phylo-Typing Method Revisited: Improvement of Specificity and Detection of New Phylo-Groups. Environ. Microbiol. Rep. 2013, 5, 58–65. [Google Scholar] [CrossRef] [PubMed]
- Ramos, C.P.; Santana, J.A.; Morcatti Coura, F.; Xavier, R.G.C.; Leal, C.A.G.; Oliveira Junior, C.A.; Heinemann, M.B.; Lage, A.P.; Lobato, F.C.F.; Silva, R.O.S. Identification and Characterization of Escherichia coli, Salmonella Spp., Clostridium perfringens, and C. difficile Isolates from Reptiles in Brazil. BioMed Res. Int. 2019, 2019, 9530732. [Google Scholar] [CrossRef]
- Coura, F.M.; de Araújo Diniz, S.; Mussi, J.M.S.; Silva, M.X.; Lage, A.P.; Heinemann, M.B. Characterization of Virulence Factors and Phylogenetic Group Determination of Escherichia coli Isolated from Diarrheic and Non-Diarrheic Calves from Brazil. Folia Microbiol. 2017, 62, 139–144. [Google Scholar] [CrossRef] [PubMed]
- Coura, F.M.; Diniz, A.N.; Oliveira Junior, C.A.; Lage, A.P.; Lobato, F.C.F.; Heinemann, M.B.; Silva, R.O.S.; Coura, F.M.; Diniz, A.N.; Oliveira Junior, C.A.; et al. Detection of Virulence Genes and the Phylogenetic Groups of Escherichia coli Isolated from Dogs in Brazil. Ciênc. Rural 2018, 48, e20170478. [Google Scholar] [CrossRef]
- NandaKafle, G.; Seale, T.; Flint, T.; Nepal, M.; Venter, S.N.; Brözel, V.S. Distribution of Diverse Escherichia coli between Cattle and Pasture. Microbes Environ. 2017, 32, 226–233. [Google Scholar] [CrossRef]
- Coura, F.M.; de Araújo Diniz, S.; Silva, M.X.; Mussi, J.M.S.; Barbosa, S.M.; Lage, A.P.; Heinemann, M.B.; Heinemann, M.B. Phylogenetic Group Determination of Escherichia coli Isolated from Animals Samples. Sci. World J. 2015, 2015, 258424. [Google Scholar] [CrossRef]
- Tenaillon, O.; Skurnik, D.; Picard, B.; Denamur, E. The Population Genetics of Commensal Escherichia coli. Nat. Rev. Microbiol. 2010, 8, 207–217. [Google Scholar] [CrossRef] [PubMed]
- Escobar-Páramo, P.; Le Menac’h, A.; Le Gall, T.; Amorin, C.; Gouriou, S.; Picard, B.; Skurnik, D.; Denamur, E. Identification of Forces Shaping the Commensal Escherichia coli Genetic Structure by Comparing Animal and Human Isolates. Environ. Microbiol. 2006, 8, 1975–1984. [Google Scholar] [CrossRef] [PubMed]
- O’Brien, C.L.; Gordon, D.M. Effect of Diet and Gut Dynamics on the Establishment and Persistence of Escherichia coli. Microbiology 2011, 157, 1375–1384. [Google Scholar] [CrossRef]
- Runesvärd, E.; Wikström, C.; Fernström, L.-L.; Hansson, I. Presence of Pathogenic Bacteria in Faeces from Dogs Fed Raw Meat-Based Diets or Dry Kibble. Vet. Rec. 2020, 187, e71. [Google Scholar] [CrossRef] [PubMed]
- Mateus, L.; Henriques, S.; Merino, C. Virulence Genotypes of Escherichia coli Canine Isolates from Pyometra, Cystitis and Fecal Origin. Vet.-Microbiol. 2013, 166, 590–594. [Google Scholar] [CrossRef] [PubMed]
- Lyu, T.; Liu, G.; Zhang, H.; Wang, L.; Zhou, S.; Dou, H.; Pang, B.; Sha, W.; Zhang, H. Changes in Feeding Habits Promoted the Differentiation of the Composition and Function of Gut Microbiotas between Domestic Dogs (Canis lupus familiaris) and Gray Wolves (Canis lupus). AMB Express 2018, 8, 123. [Google Scholar] [CrossRef] [PubMed]
- Gordon, D.M.; Cowling, A. The Distribution and Genetic Structure of Escherichia coli in Australian Vertebrates: Host and Geographic Effects. Microbiology 2003, 149, 3575–3586. [Google Scholar] [CrossRef] [PubMed]
- Apostolakos, I.; Mughini-Gras, L.; Fasolato, L.; Piccirillo, A. Assessing the Occurrence and Transfer Dynamics of ESBL/PAmpC-Producing Escherichia coli across the Broiler Production Pyramid. PLoS ONE 2019, 14, e0217174. [Google Scholar] [CrossRef] [PubMed]
- Sary, K.; Fairbrother, J.M.; Arsenault, J.; De Lagarde, M.; Boulianne, M. Antimicrobial Resistance and Virulence Gene Profiles among Escherichia coli Isolates from Retail Chicken Carcasses in Vietnam. Foodborne Pathog. Dis. 2019, 16, 298–306. [Google Scholar] [CrossRef] [PubMed]
- De Almeida, P.M.P.; Arais, L.R.; Andrade, J.R.C.; Prado, E.H.R.B.; Irino, K.; de Mello Figueiredo Cerqueira, A. Characterization of Atypical Enteropathogenic Escherichia coli (AEPEC) Isolated from Dogs. Vet. Microbiol. 2012, 158, 420–424. [Google Scholar] [CrossRef] [PubMed]
- Marks, S.L.; Kather, E.J. Bacterial-Associated Diarrhea in the Dog: A Critical Appraisal. Vet. Clin. N. Am.-Small Anim. Pract. 2003, 33, 1029–1060. [Google Scholar] [CrossRef]
- Nagy, B.; Fekete, P.Z. Enterotoxigenic Escherichia coli in Veterinary Medicine. Int. J. Med. Microbiol. 2005, 295, 443–454. [Google Scholar] [CrossRef]
- Nishikawa, Y.; Ogasawara, J.; Helander, A.; Haruki, K. An Outbreak of Gastroenteritis in Japan Due to Escherichia coli O166. Emerg. Infect. Dis. 1999, 5, 300. [Google Scholar] [CrossRef]
- Wang, L.; Nakamura, H.; Kage-Nakadai, E.; Hara-Kudo, Y.; Nishikawa, Y. Prevalence, Antimicrobial Resistance and Multiple-Locus Variable-Number Tandem-Repeat Analysis Profiles of Diarrheagenic Escherichia coli Isolated from Different Retail Foods. Int. J. Food Microbiol. 2017, 249, 44–52. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.; Ogasawara, J.; Nishikawa, Y.; Seto, Y.; Helander, A.; Hase, A.; Iritani, N.; Nakamura, H.; Arikawa, K.; Kai, A.; et al. An Outbreak of Gastroenteritis in Osaka, Japan Due to Escherichia coli Serogroup O166:H15 That Had a Coding Gene for Enteroaggregative E. coli Heat-Stable Enterotoxin 1 (EAST1). Epidemiol. Infect. 2002, 128, 363–371. [Google Scholar] [CrossRef] [PubMed]
- Sukkua, K.; Manothong, S.; Sukhumungoon, P. Seroprevalence and Molecular Epidemiology of EAST1 Gene-Carrying Escherichia coli from Diarrheal Patients and Raw Meats. J. Infect. Dev. Ctries. 2017, 11, 220–227. [Google Scholar] [CrossRef] [PubMed]
- Yasir, M.; Qureshi, A.K.; Kensarah, E.A.; Bibi, F.; Al-Zahrani, I.A.; Abd El Ghany, M.; Azhar, E.I. Draft Genome Sequence of Colistin-Resistant and Extended-Spectrum β-Lactamase (ESBL)-Producing Multidrug-Resistant Escherichia coli Isolated from Poultry Meat. J. Glob. Antimicrob. Resist. 2021, 27, 112–114. [Google Scholar] [CrossRef] [PubMed]
- Bacci, C.; Vismarra, A.; Dander, S.; Barilli, E.; Superchi, P. Occurrence and Antimicrobial Profile of Bacterial Pathogens in Former Foodstuff Meat Products Used for Pet Diets. J. Food Prot. 2019, 82, 316–324. [Google Scholar] [CrossRef] [PubMed]
- Naziri, Z.; Derakhshandeh, A.; Firouzi, R.; Motamedifar, M.; Shojaee Tabrizi, A. DNA Fingerprinting Approaches to Trace Escherichia coli Sharing between Dogs and Owners. J. Appl. Microbiol. 2016, 120, 460–468. [Google Scholar] [CrossRef] [PubMed]
- WHO. A European One Health Action Plan against Antimicrobial Resistance (AMR); WHO: Geneva, Switzerland, 2017; Volume 1, p. 24. [Google Scholar]
- WHO. Global Priority List of Antibiotic-Resistant Bacteria to Guide Research, Discovery, and Development of New Antibiotics; WHO: Geneva, Switzerland, 2013; pp. 1–7. [Google Scholar]
- WHO. Critically Important Antimicrobials for Human Medicine; 6th Revision; WHO: Geneva, Switzerland, 2018; pp. 1–52. [Google Scholar]
- OIE. OIE List of Antimicrobial Agents of Veterinary Importance; OIE: Paris, France, 2018; pp. 1–10. [Google Scholar]
- Wagner, S.; Gally, D.L.; Argyle, S.A. Multidrug-Resistant Escherichia coli from Canine Urinary Tract Infections Tend to Have Commensal Phylotypes, Lower Prevalence of Virulence Determinants and AmpC-Replicons. Vet. Microbiol. 2014, 169, 171–178. [Google Scholar] [CrossRef][Green Version]
- Johnson, J.R.; Kuskowski, M.A.; Gajewski, A.; Sahm, D.F.; Karlowsky, J.A. Virulence Characteristics and Phylogenetic Background of Multidrug-Resistant and Antimicrobial-Susceptible Clinical Isolates of Escherichia coli from across the United States, 2000–2001. J. Infect. Dis. 2004, 190, 1739–1744. [Google Scholar] [CrossRef][Green Version]
- Salgado-Caxito, M.; Benavides, J.A.; Adell, A.D.; Paes, A.C.; Moreno-Switt, A.I. Global Prevalence and Molecular Characterization of Extended-Spectrum β-Lactamase Producing-Escherichia coli in Dogs and Cats A Scoping Review and Meta-Analysis. One Health 2021, 12, 100236. [Google Scholar] [CrossRef]
- Karkaba, A.; Hill, K.; Benschop, J.; Pleydell, E.; Grinberg, A. Carriage and Population Genetics of Extended Spectrum β-Lactamase-Producing Escherichia coli in Cats and Dogs in New Zealand. Vet. Microbiol. 2019, 233, 61–67. [Google Scholar] [CrossRef] [PubMed]
- Meyer, E.; Gastmeier, P.; Kola, A.; Schwab, F. Pet Animals and Foreign Travel Are Risk Factors for Colonisation with Extended-Spectrum β-Lactamase-Producing Escherichia coli. Infection 2012, 40, 685–687. [Google Scholar] [CrossRef] [PubMed]
- Carvalho, A.C.; Barbosa, A.V.; Arais, L.R.; Ribeiro, P.F.; Carneiro, V.C.; Cerqueira, A.M.F. Resistance Patterns, ESBL Genes, and Genetic Relatedness of Escherichia coli from Dogs and Owners. Braz. J. Microbiol. 2016, 47, 150–158. [Google Scholar] [CrossRef]
- Melo, L.C.; Oresco, C.; Leigue, L.; Netto, H.M.; Melville, P.A.; Benites, N.R.; Saras, E.; Haenni, M.; Lincopan, N.; Madec, J.-Y. Prevalence and Molecular Features of ESBL/PAmpC-Producing Enterobacteriaceae in Healthy and Diseased Companion Animals in Brazil. Vet. Microbiol. 2018, 221, 59–66. [Google Scholar] [CrossRef]
- Sfaciotte, R.A.P.; Parussolo, L.; Melo, F.D.; Wildemann, P.; Bordignon, G.; Israel, N.D.; Leitzke, M.; Wosiacki, S.R.; Salbego, F.Z.; da Costa, U.M.; et al. Identification and Characterization of Multidrug-Resistant Extended-Spectrum Beta-Lactamase-Producing Bacteria from Healthy and Diseased Dogs and Cats Admitted to a Veterinary Hospital in Brazil. Microb. Drug Resist. 2021, 27, 855–864. [Google Scholar] [CrossRef] [PubMed]
- Carvalho, I.; Cunha, R.; Martins, C.; Martínez-Álvarez, S.; Safia Chenouf, N.; Pimenta, P.; Pereira, A.R.; Ramos, S.; Sadi, M.; Martins, Â.; et al. Antimicrobial Resistance Genes and Diversity of Clones among Faecal ESBL-Producing Escherichia coli Isolated from Healthy and Sick Dogs Living in Portugal. Antibiot. Basel Switz. 2021, 10, 1013. [Google Scholar] [CrossRef] [PubMed]
- Rocha-Gracia, R.C.; Cortés-Cortés, G.; Lozano-Zarain, P.; Bello, F.; Martínez-Laguna, Y.; Torres, C. Faecal Escherichia coli Isolates from Healthy Dogs Harbour CTX-M-15 and CMY-2 β-Lactamases. Vet. J. 2015, 203, 315–319. [Google Scholar] [CrossRef] [PubMed]
- Zogg, A.L.; Zurfluh, K.; Schmitt, S.; Nüesch-Inderbinen, M.; Stephan, R. Antimicrobial Resistance, Multilocus Sequence Types and Virulence Profiles of ESBL Producing and Non-ESBL Producing Uropathogenic Escherichia coli Isolated from Cats and Dogs in Switzerland. Vet. Microbiol. 2018, 216, 79–84. [Google Scholar] [CrossRef] [PubMed]
- Lee, W.; Chung, H.-S.; Lee, H.; Yum, J.H.; Yong, D.; Jeong, S.H.; Lee, K.; Chong, Y. CTX-M-55-Type Extended-Spectrum β-Lactamase-Producing Shigella Sonnei Isolated from a Korean Patient Who Had Travelled to China. Ann. Lab. Med. 2013, 33, 141–144. [Google Scholar] [CrossRef]
- Zeng, S.; Luo, J.; Li, X.; Zhuo, C.; Wu, A.; Chen, X.; Huang, L. Molecular Epidemiology and Characteristics of CTX-M-55 Extended-Spectrum β-Lactamase-Producing Escherichia coli From Guangzhou, China. Front. Microbiol. 2021, 12, 730012. [Google Scholar] [CrossRef]
- Mathers, A.J.; Peirano, G.; Pitout, J.D.D. The Role of Epidemic Resistance Plasmids and International High-Risk Clones in the Spread of Multidrug-Resistant Enterobacteriaceae. Clin. Microbiol. Rev. 2015, 28, 565–591. [Google Scholar] [CrossRef]
- Bortolami, A.; Zendri, F.; Maciuca, E.I.; Wattret, A.; Ellis, C.; Schmidt, V.; Pinchbeck, G.; Timofte, D. Diversity, Virulence, and Clinical Significance of Extended-Spectrum β-Lactamase- and PAmpC-Producing Escherichia coli From Companion Animals. Front. Microbiol. 2019, 10, 1260. [Google Scholar] [CrossRef]
- Huang, Y.-H.; Kuan, N.-L.; Yeh, K.-S. Characteristics of Extended-Spectrum β-Lactamase–Producing Escherichia coli From Dogs and Cats Admitted to a Veterinary Teaching Hospital in Taipei, Taiwan From 2014 to 2017. Front. Vet. Sci. 2020, 7, 395. [Google Scholar] [CrossRef]
- Saliu, E.-M.; Vahjen, W.; Zentek, J. Types and Prevalence of Extended-Spectrum Beta-Lactamase Producing Enterobacteriaceae in Poultry. Anim. Health Res. Rev. 2017, 18, 46–57. [Google Scholar] [CrossRef]
- De Souza Gazal, L.E.; Medeiros, L.P.; Dibo, M.; Nishio, E.K.; Koga, V.L.; Gonçalves, B.C.; Grassotti, T.T.; de Camargo, T.C.L.; Pinheiro, J.J.; Vespero, E.C.; et al. Detection of ESBL/AmpC-Producing and Fosfomycin-Resistant Escherichia coli from Different Sources in Poultry Production in Southern Brazil. Front. Microbiol. 2021, 11, 604544. [Google Scholar] [CrossRef]
- Cunha, M.P.V.; Lincopan, N.; Cerdeira, L.; Esposito, F.; Dropa, M.; Franco, L.S.; Moreno, A.M.; Knöbl, T. Coexistence of CTX-M-2, CTX-M-55, CMY-2, FosA3, and QnrB19 in Extraintestinal Pathogenic Escherichia coli from Poultry in Brazil. Antimicrob. Agents Chemother. 2017, 61, e02474-16. [Google Scholar] [CrossRef] [PubMed]
- Coppola, N.; Cordeiro, N.F.; Trenchi, G.; Esposito, F.; Fuga, B.; Fuentes-Castillo, D.; Lincopan, N.; Iriarte, A.; Bado, I.; Vignoli, R. Imported One-Day-Old Chicks as Trojan Horses for Multidrug-Resistant Priority Pathogens Harboring Mcr-9, RmtG and Extended-Spectrum β-Lactamase Genes. Appl. Environ. Microbiol. 2021, 88, e01675-21. [Google Scholar] [CrossRef] [PubMed]
- Nüesch-Inderbinen, M.; Treier, A.; Zurfluh, K.; Stephan, R. Raw Meat-Based Diets for Companion Animals: A Potential Source of Transmission of Pathogenic and Antimicrobial-Resistant Enterobacteriaceae. R. Soc. Open Sci. 2019, 6, 191170. [Google Scholar] [CrossRef] [PubMed]
- Ibekwe, A.; Durso, L.; Ducey, T.F.; Oladeinde, A.; Jackson, C.R.; Frye, J.G.; Dungan, R.; Moorman, T.; Brooks, J.P.; Obayiuwana, A.; et al. Diversity of Plasmids and Genes Encoding Resistance to Extended-Spectrum β-Lactamase in Escherichia coli from Different Animal Sources. Microorganisms 2021, 9, 1057. [Google Scholar] [CrossRef]
- Salinas, L.; Loayza, F.; Cárdenas, P.; Saraiva, C.; Johnson, T.J.; Amato, H.; Graham, J.P.; Trueba, G. Environmental Spread of Extended Spectrum Beta-Lactamase (ESBL) Producing Escherichia coli and ESBL Genes among Children and Domestic Animals in Ecuador. Environ. Health Perspect. 2021, 129, 027007. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhou, J.; Li, X.; Ma, L.; Cao, X.; Hu, W.; Zhao, L.; Jing, W.; Lan, X.; Li, Y.; et al. Genetic Diversity, Antimicrobial Resistance and Extended-Spectrum β-Lactamase Type of Escherichia coli Isolates from Chicken, Dog, Pig and Yak in Gansu and Qinghai Provinces, China. J. Glob. Antimicrob. Resist. 2020, 22, 726–732. [Google Scholar] [CrossRef]
- Kawamura, K.; Nagano, N.; Suzuki, M.; Wachino, J.-I.; Kimura, K.; Arakawa, Y. ESBL-Producing Escherichia coli and Its Rapid Rise among Healthy People. Food Saf. Tokyo Jpn. 2017, 5, 122–150. [Google Scholar] [CrossRef]
- Li, S.; Liu, J.; Zhou, Y.; Miao, Z. Characterization of ESBL-Producing Escherichia coli Recovered from Companion Dogs in Tai’an, China. J. Infect. Dev. Ctries. 2017, 11, 282–286. [Google Scholar] [CrossRef] [PubMed]
- Alegría, Á.; Arias Temprano, M.; Fernandez-Natal, M.; Rodríguez-Calleja, J.; García-López, M.-L.; Santos, J. Molecular Diversity of ESBL-Producing Escherichia coli from Foods of Animal Origin and Human Patients. Int. J. Environ. Res. Public Health 2020, 17, 1312. [Google Scholar] [CrossRef] [PubMed]
- Ewers, C.; Bethe, A.; Semmler, T.; Guenther, S.; Wieler, L.H. Extended-Spectrum β-Lactamase-Producing and AmpC-Producing Escherichia coli from Livestock and Companion Animals, and Their Putative Impact on Public Health: A Global Perspective. Clin. Microbiol. Infect. 2012, 18, 646–655. [Google Scholar] [CrossRef]
- Moreira da Silva, R.C.R.; de Oliveira Martins Júnior, P.; Gonçalves, L.F.; de Paulo Martins, V.; de Melo, A.B.F.; Pitondo-Silva, A.; de Campos, T.A. Ciprofloxacin Resistance in Uropathogenic Escherichia coli Isolates Causing Community-Acquired Urinary Infections in Brasília, Brazil. J. Glob. Antimicrob. Resist. 2017, 9, 61–67. [Google Scholar] [CrossRef] [PubMed]
- Drali, R.; Berrazeg, M.; Zidouni, L.L.; Hamitouche, F.; Abbas, A.A.; Deriet, A.; Mouffok, F. Emergence of mcr-1 Plasmid-Mediated Colistin-Resistant Escherichia coli Isolates from Seawater. Sci. Total Environ. 2018, 642, 90–94. [Google Scholar] [CrossRef] [PubMed]
- Paiva, Y.; Nagano, D.S.; Cotia, A.L.F.; Guimarães, T.; Martins, R.C.R.; Perdigão Neto, L.V.; Côrtes, M.F.; Marchi, A.P.; Corscadden, L.; Machado, A.S.; et al. Colistin-Resistant Escherichia coli Belonging to Different Sequence Types: Genetic Characterization of Isolates Responsible for Colonization, Community- and Healthcare-Acquired Infections. Rev. Inst. Med. Trop. Sao Paulo 2021, 63, e38. [Google Scholar] [CrossRef]
- Fuga, B.; Sellera, F.P.; Cerdeira, L.; Esposito, F.; Cardoso, B.; Fontana, H.; Moura, Q.; Cardenas-Arias, A.; Sano, E.; Ribas, R.M.; et al. WHO Critical Priority Escherichia coli as One Health Challenge for a Post-Pandemic Scenario: Genomic Surveillance and Analysis of Current Trends in Brazil. Microbiol. Spectr. 2022, 10, e01256-21. [Google Scholar] [CrossRef] [PubMed]
- Falgenhauer, L.; Imirzalioglu, C.; Ghosh, H.; Gwozdzinski, K.; Schmiedel, J.; Gentil, K.; Bauerfeind, R.; Kämpfer, P.; Seifert, H.; Michael, G.B.; et al. Circulation of Clonal Populations of Fluoroquinolone-Resistant CTX-M-15-Producing Escherichia coli ST410 in Humans and Animals in Germany. Int. J. Antimicrob. Agents 2016, 47, 457–465. [Google Scholar] [CrossRef] [PubMed]
- Schaufler, K.; Semmler, T.; Wieler, L.H.; Wöhrmann, M.; Baddam, R.; Ahmed, N.; Müller, K.; Kola, A.; Fruth, A.; Ewers, C.; et al. Clonal Spread and Interspecies Transmission of Clinically Relevant ESBL-Producing Escherichia coli of ST410—Another Successful Pandemic Clone? FEMS Microbiol. Ecol. 2016, 92, fiv155. [Google Scholar] [CrossRef] [PubMed]
- Nadimpalli, M.L.; de Lauzanne, A.; Phe, T.; Borand, L.; Jacobs, J.; Fabre, L.; Naas, T.; Le Hello, S.; Stegger, M. Escherichia coli ST410 among Humans and the Environment in Southeast Asia. Int. J. Antimicrob. Agents 2019, 54, 228–232. [Google Scholar] [CrossRef]
- Tada, T.; Nhung, P.H.; Shimada, K.; Tsuchiya, M.; Phuong, D.M.; Anh, N.Q.; Ohmagari, N.; Kirikae, T. Emergence of Colistin-Resistant Escherichia coli Clinical Isolates Harboring mcr-1 in Vietnam. Int. J. Infect. Dis. 2017, 63, 72–73. [Google Scholar] [CrossRef] [PubMed]
- Monte, D.F.; Mem, A.; Fernandes, M.R.; Cerdeira, L.; Esposito, F.; Galvão, J.A.; Franco, B.D.G.M.; Lincopan, N.; Landgraf, M. Chicken Meat as a Reservoir of Colistin-Resistant Escherichia coli Strains Carrying mcr-1 Genes in South America . Antimicrob. Agents Chemother. 2017, 61, e02718-16. [Google Scholar] [CrossRef]
- Quiroga, C.; Nastro, M.; Di Conza, J. Current Scenario of Plasmid-Mediated Colistin Resistance in Latin America. Rev. Argent. Microbiol. 2019, 51, 93–100. [Google Scholar] [CrossRef]
- Brasil Instrução Normativa No 45 de 22 de Novembro de 2016. 2016. Available online: https://www.in.gov.br/materia/-/asset_publisher/Kujrw0TZC2Mb/content/id/22078290/do1-2016-11-30-instrucao-normativa-n-45-de-22-de-novembro-de-2016-22078259 (accessed on 14 April 2022).
- Carattoli, A.; Bertini, A.; Villa, L.; Falbo, V.; Hopkins, K.L.; Threlfall, E.J. Identification of Plasmids by PCR-Based Replicon Typing. J. Microbiol. Methods 2005, 63, 219–228. [Google Scholar] [CrossRef]
- Richter, L.; du Plessis, E.M.; Duvenage, S.; Allam, M.; Ismail, A.; Korsten, L. Whole Genome Sequencing of Extended-Spectrum- and AmpC- β-Lactamase-Positive Enterobacterales Isolated From Spinach Production in Gauteng Province, South Africa. Front. Microbiol. 2021, 12, 734649. [Google Scholar] [CrossRef] [PubMed]
- Valentin, L.; Sharp, H.; Hille, K.; Seibt, U.; Fischer, J.; Pfeifer, Y.; Michael, G.B.; Nickel, S.; Schmiedel, J.; Falgenhauer, L.; et al. Subgrouping of ESBL-Producing Escherichia coli from Animal and Human Sources: An Approach to Quantify the Distribution of ESBL Types between Different Reservoirs. Int. J. Med. Microbiol. 2014, 304, 805–816. [Google Scholar] [CrossRef] [PubMed]
- Voets, G.M.; Fluit, A.C.; Scharringa, J.; Schapendonk, C.; van den Munckhof, T.; Leverstein-van Hall, M.A.; Stuart, J.C. Identical Plasmid AmpC Beta-Lactamase Genes and Plasmid Types in E. coli Isolates from Patients and Poultry Meat in the Netherlands. Int. J. Food Microbiol. 2013, 167, 359–362. [Google Scholar] [CrossRef]
- Nilsson, O. Hygiene Quality and Presence of ESBL-Producing Escherichia coli in Raw Food Diets for Dogs. Infect. Ecol. Epidemiol. 2015, 5, 28758. [Google Scholar] [CrossRef]
- Kaesbohrer, A.; Bakran-Lebl, K.; Irrgang, A.; Fischer, J.; Kämpf, P.; Schiffmann, A.; Werckenthin, C.; Busch, M.; Kreienbrock, L.; Hille, K. Diversity in Prevalence and Characteristics of ESBL/PAmpC Producing E. coli in Food in Germany. Vet. Microbiol. 2019, 233, 52–60. [Google Scholar] [CrossRef] [PubMed]
- Gouvea, R.; dos Santos, F.F.; Aquino, M.H.C.; de Pereira VL, A. Fluoroquinolones in Industrial Poultry Production, Bacterial Resistance and Food Residues: A Review. Braz. J. Poult. Sci. 2015, 17, 1–10. [Google Scholar] [CrossRef]
- Van Boeckel, T.P.; Pires, J.; Silvester, R.; Zhao, C.; Song, J.; Criscuolo, N.G.; Gilbert, M.; Bonhoeffer, S.; Laxminarayan, R. Global Trends in Antimicrobial Resistance in Animals in Low- And Middle-Income Countries. Science 2019, 365, eaaw1944. [Google Scholar] [CrossRef]
- Vidovic, N.; Vidovic, S. Antimicrobial Resistance and Food Animals: Influence of Livestock Environment on the Emergence and Dissemination of Antimicrobial Resistance. Antibiotics 2020, 9, 52. [Google Scholar] [CrossRef]
- Vich Vila, A.; Collij, V.; Sanna, S.; Sinha, T.; Imhann, F.; Bourgonje, A.R.; Mujagic, Z.; Jonkers, D.M.A.E.; Masclee, A.A.M.; Fu, J.; et al. Impact of Commonly Used Drugs on the Composition and Metabolic Function of the Gut Microbiota. Nat. Commun. 2020, 11, 362. [Google Scholar] [CrossRef] [PubMed]
- McDaniels, A.E.; Rice, E.W.; Reyes, A.L.; Johnson, C.H.; Haugland, R.A.; Stelma, G.N., Jr. Confirmational Identification of Escherichia coli, a Comparison of Genotypic and Phenotypic Assays for Glutamate Decarboxylase and Confirmational Identification of Escherichia coli, a Comparison of Genotypic and Phenotypic Assays for Glutamate Decarboxy. Appl. Environ. Microbiol. 1996, 62, 3350–3354. [Google Scholar] [CrossRef] [PubMed]
- Clermont, O.; Gordon, D.M.; Brisse, S.; Walk, S.T.; Denamur, E. Characterization of the Cryptic Escherichia Lineages: Rapid Identification and Prevalence. Environ. Microbiol. 2011, 13, 2468–2477. [Google Scholar] [CrossRef] [PubMed]
- Performance Standards for Antimicrobial Disk and Dilution Susceptibility Tests for Bacteria Isolated From Animals. CLSI Supplement VET01S. 2020. Available online: https://clsi.org/standards/products/veterinary-medicine/documents/vet01s/ (accessed on 15 April 2022).
- European Committee on Antimicrobial Susceptibility Testing (EUCAST). Break. Tables Interpret. MICs Zone Diameters 2022, 12. Available online: https://www.eucast.org/mic_distributions_and_ecoffs/ (accessed on 15 April 2022).
- Magiorakos, A.-P.; Srinivasan, A.; Carey, R.B.; Carmeli, Y.; Falagas, M.E.; Giske, C.G.; Harbarth, S.; Hindler, J.F.; Kahlmeter, G.; Olsson-Liljequist, B.; et al. Multidrug-Resistant, Extensively Drug-Resistant and Pandrug-Resistant Bacteria: An International Expert Proposal for Interim Standard Definitions for Acquired Resistance. Clin. Microbiol. Infect. Off. Publ. Eur. Soc. Clin. Microbiol. Infect. Dis. 2012, 18, 268–281. [Google Scholar] [CrossRef] [PubMed]
- Nurk, S.; Bankevich, A.; Antipov, D.; Gurevich, A.A.; Korobeynikov, A.; Lapidus, A.; Prjibelski, A.D.; Pyshkin, A.; Sirotkin, A.; Sirotkin, Y.; et al. Assembling Single-Cell Genomes and Mini-Metagenomes from Chimeric MDA Products. J. Comput. Biol. J. Comput. Mol. Cell Biol. 2013, 20, 714–737. [Google Scholar] [CrossRef]
- Seemann, T. Prokka: Rapid Prokaryotic Genome Annotation. Bioinform. Oxf. Engl. 2014, 30, 2068–2069. [Google Scholar] [CrossRef] [PubMed]
- Bortolaia, V.; Kaas, R.S.; Ruppe, E.; Roberts, M.C.; Schwarz, S.; Cattoir, V.; Philippon, A.; Allesoe, R.L.; Rebelo, A.R.; Florensa, A.F.; et al. ResFinder 4.0 for Predictions of Phenotypes from Genotypes. J. Antimicrob. Chemother. 2020, 75, 3491–3500. [Google Scholar] [CrossRef] [PubMed]
- Zankari, E.; Allesøe, R.; Joensen, K.G.; Cavaco, L.M.; Lund, O.; Aarestrup, F.M. PointFinder: A Novel Web Tool for WGS-Based Detection of Antimicrobial Resistance Associated with Chromosomal Point Mutations in Bacterial Pathogens. J. Antimicrob. Chemother. 2017, 72, 2764–2768. [Google Scholar] [CrossRef]
- Camacho, C.; Coulouris, G.; Avagyan, V.; Ma, N.; Papadopoulos, J.; Bealer, K.; Madden, T.L. BLAST+: Architecture and Applications. BMC Bioinform. 2009, 10, 421. [Google Scholar] [CrossRef]
- Joensen, K.G.; Scheutz, F.; Lund, O.; Hasman, H.; Kaas, R.S.; Nielsen, E.M.; Aarestrup, F.M. Real-Time Whole-Genome Sequencing for Routine Typing, Surveillance, and Outbreak Detection of Verotoxigenic Escherichia coli. J. Clin. Microbiol. 2014, 52, 1501–1510. [Google Scholar] [CrossRef]
- Malberg Tetzschner, A.M.; Johnson, J.R.; Johnston, B.D.; Lund, O.; Scheutz, F. In Silico Genotyping of Escherichia coli Isolates for Extraintestinal Virulence Genes by Use of Whole-Genome Sequencing Data. J. Clin. Microbiol. 2020, 58, e01269-20. [Google Scholar] [CrossRef]
- Jaureguy, F.; Landraud, L.; Passet, V.; Diancourt, L.; Frapy, E.; Guigon, G.; Carbonnelle, E.; Lortholary, O.; Clermont, O.; Denamur, E.; et al. Phylogenetic and Genomic Diversity of Human Bacteremic Escherichia coli Strains. BMC Genom. 2008, 9, 560. [Google Scholar] [CrossRef] [PubMed]
- Wirth, T.; Falush, D.; Lan, R.; Colles, F.; Mensa, P.; Wieler, L.H.; Karch, H.; Reeves, P.R.; Maiden, M.C.J.; Ochman, H.; et al. Sex and Virulence in Escherichia coli: An Evolutionary Perspective. Mol. Microbiol. 2006, 60, 1136–1151. [Google Scholar] [CrossRef] [PubMed]
- Larsen, M.V.; Cosentino, S.; Rasmussen, S.; Friis, C.; Hasman, H.; Marvig, R.L.; Jelsbak, L.; Sicheritz-Pontén, T.; Ussery, D.W.; Aarestrup, F.M.; et al. Multilocus Sequence Typing of Total-Genome-Sequenced Bacteria. J. Clin. Microbiol. 2012, 50, 1355–1361. [Google Scholar] [CrossRef]
- Jünemann, S.; Sedlazeck, F.; Prior, K.; Albersmeier, A.; John, U.; Kalinowski, J.; Mellmann, A.; Goesmann, A.; von Haeseler, A.; Stoye, J.; et al. Updating Benchtop Sequencing Performance Comparison. Nat. Biotechnol. 2013, 31, 294–296. [Google Scholar] [CrossRef] [PubMed]
- Thomsen, M.C.F.; Ahrenfeldt, J.; Cisneros, J.L.B.; Jurtz, V.; Larsen, M.V.; Hasman, H.; Aarestrup, F.M.; Lund, O. A Bacterial Analysis Platform: An Integrated System for Analysing Bacterial Whole Genome Sequencing Data for Clinical Diagnostics and Surveillance. PLoS ONE 2016, 11, e0157718. [Google Scholar] [CrossRef] [PubMed]
- Alikhan, N.-F.; Petty, N.K.; Ben Zakour, N.L.; Beatson, S.A. BLAST Ring Image Generator (BRIG): Simple Prokaryote Genome Comparisons. BMC Genom. 2011, 12, 402. [Google Scholar] [CrossRef] [PubMed]
- Kaas, R.S.; Leekitcharoenphon, P.; Aarestrup, F.M.; Lund, O. Solving the Problem of Comparing Whole Bacterial Genomes across Different Sequencing Platforms. PLoS ONE 2014, 9, e104984. [Google Scholar] [CrossRef] [PubMed]
- Zar, J.H. Bioestatistical Analysis, 5th ed.; Hall, P., Ed.; Pearson: Upper Saddle River, NJ, USA, 2009. [Google Scholar]
- Hosmer, D.W.; Lemeshow, S.; Sturdivant, R.X. Applied Logistic Regression, 3rd ed.; Sons, J.W., Ed.; Wiley: New York, NY, USA, 2013. [Google Scholar]
- Greenacre, M.; Blasius, J. Multiple Correspondence Analysis and Related Methods Multiple Correspondence Analysis and Related Methods, 1st ed.; Chapman and Hall: New York, NY, USA, 2006. [Google Scholar]
| Type of Diet | Phylogenetic Groups (% Total) | Total | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| A | B1 | B2 | C | D | E | F | Clade I | Unassignable ¹ | ||
| RMBD | 6 (2.8) | 31 (14.6) | 9 (4.2) a | 10 (4.7) | 0 | 15 (7.0) a | 7 (3.3) | 0 | 7 (3.3) | 85 (40.0) |
| Conventional | 3 (1.4) | 37 (17.4) | 39 (18.3) b | 15 (7.0) | 1 (0.4) | 8 (3.7) b | 10 (4.7) | 3 (1.4) | 11 (5.1) | 127 (59.9) |
| Total | 9 (4.2) | 68 (32.0) | 48 (22.6) | 25 (11.7) | 1 (0.4) | 23 (10.8) | 17 (8.1) | 3 (1.4) | 18 (8.4) | 212 (100) |
| Antimicrobial Drug | Type of Diet (% Total) | p Value | |
|---|---|---|---|
| RMBD (n = 85) | Conventional (n = 127) | ||
| amoxicillin/clavulanic acid | 5 (5.8) | 3 (2.3) | 0.2 |
| ampicillin * | 46 (54.1) | 28 (22.0) | 0.0004 |
| ceftiofur * | 30 (35.2) | 25 (19.6) | 0.01 |
| enrofloxacin * | 20 (23.5) | 10 (7.8) | 0.002 |
| ciprofloxacin | 29 (34.1) | 45 (35.4) | 0.7 |
| trimethoprim/sulfamethoxazole * | 52 (61.1) | 44 (33.5) | 0.0004 |
| doxycycline * | 31 (36.4) | 22 (17.3) | 0.003 |
| oxytetracycline * | 41 (48.2) | 30 (23.6) | 0.0001 |
| florfenicol | 9 (10.5) | 6 (4.7) | 0.09 |
| gentamicin * | 15 (17.6) | 2 (1.5) | 0.0004 |
| neomycin * | 4 (4.7) | 0 (0) | 0.03 |
| Animal | Isolate | MLST 1 | Antimicrobial Resistance Genes | Virulence Factors | |
|---|---|---|---|---|---|
| ESBL 2 | Other | ||||
| FV21 | 1 | ST10 | blaCTX-M-55 | aph(3’)-Ia aadA22 mdf(A) lnu(F) gyrA* sul3 floR aadA22 | cif cma cvaC eae espA espB espF hlyF iucC iutA nleB ompT sitA tccP terC tir traT |
| FV24 | 1 | ST224 | blaCTX-M-55 blaTEM-1B | gyrA* fosA3 mdf(A) | cma cvaC gad hlyF iroN iss lpfA ompT sitA terC traT |
| FV25 | 2 | ST10 | blaCTX-M-55 | aph(3′)-Ia mdf(A) mdf(A) aadA22 lnu(F) | cif cma cvaC eae espA espB gad hlyF iucC iutA ompTb sitA terC tir traT |
| 3 | ST57 | blaCTX-M-55 blaCTX-M-2 | aph(3′)-Ia sul1 dfrA7 mdf(A) floR gyrA* sul3 mdf(A) aadA1 mcr-1.1 fosA3 tet(A) | astA cea chuA gad hra iha iss iucC iutA ompT sitA terC traT | |
| FV26 | 1 | ST744 | blaCTX-M-55 blaTEM-1B | aph(3′)-Ia sul1 catA1 gyrA dfrA17 fosA3 aph(3′)-Ia mph(A) aadA5 tet(B) aph(6)-Id | terC traT |
| FV27 | 1 | ST57 | blaCTX-M-2 | ant(2″)-Ia sul2 dfrA1 mdf(A) aadA1 aadA1 gyrA* | chuA cma etsC fyuA gad hlyF hra iroN irp2 iss iucC iutA ompT sitA terC traT tsh |
| FV30 | 2 | ST410 | blaSHV-12 blaTEM-1B | aac(3)-Iid sul1 dfrA1 mdf(A) aadA1 gyrA* mcr-1.1 | astA cib cma cvaC etsC hlyF hra iroN iss iucC iutA lpfA ompT papC sitA terC traT |
| Animal | Isolate | Relevant AMR Genes | Contig | Closest BLAST 1 Match Source, Country | Conjugative Plasmid Replicons |
|---|---|---|---|---|---|
| FV21 | 1 | blaCTX-M-55 | 64 | E. coli plasmid pRHB02-C09_2 (CP058073) Pig, UK | IncFIB; IncFIC; IncFII |
| FV24 | 1 | blaCTX-M-55 | 168 | E. coli plasmid pAH01-3 (CP055254) Poultry, China | IncFIB; IncFII; IncFII (pRSB107) |
| blaTEM-1B | |||||
| FV25 | 2 | blaCTX-M-55 | 70 | E. coli plasmid pTREC1 (MN158989) Wetland sediment, USA | IncFIB; IncFIC (FII); IncFII; IncI2 |
| 3 | blaCTX-M-55 | 429 | E. coli plasmid pAH01-3 (CP055254) Chicken, China | Col (MG828); Col156; IncFIB; IncFII; IncHI2; IncHI2A; IncI2; IncY | |
| blaCTX-M-2 | 74 | E. coli Integron in117 (DQ125241) Human, Spain | |||
| mcr-1 | 334 | E. coli mcr-1 cassette (LT159973) Cattle, Germany. | |||
| FV26 | 1 | blaCTX-M-55 | 87 | Proteus mirabilis genomic island PGI2C55 (MK847915) Chicken, China | IncFII; IncN; IncQ1 |
| blaTEM-1B | |||||
| FV27 | 1 | blaCTX-M-2 | 284 | E. coli plasmid RCS78_p (LT985296) Human, Brazil | ColpVC; IncFIB; IncFIC; IncI2 |
| FV30 | 2 | blaSHV-12 | 236 | E. coli plasmid pMCR_1525_C2 (MT929281) Turkey, Brazil | ColpVC; IncFIA; IncFIB; IncFII; IncI1-I; IncX4 |
| blaTEM-1B | 155 | E. coli plasmid pSHE-CTX-M (CP022359) Human, France | |||
| mcr-1 | 183 | E. coli plasmid pIncFIB_IncFII (CP066837) Chicken, USA |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ramos, C.P.; Kamei, C.Y.I.; Viegas, F.M.; de Melo Barbieri, J.; Cunha, J.L.R.; Hounmanou, Y.M.G.; Coura, F.M.; Santana, J.A.; Lobato, F.C.F.; Bojesen, A.M.; et al. Fecal Shedding of Multidrug Resistant Escherichia coli Isolates in Dogs Fed with Raw Meat-Based Diets in Brazil. Antibiotics 2022, 11, 534. https://doi.org/10.3390/antibiotics11040534
Ramos CP, Kamei CYI, Viegas FM, de Melo Barbieri J, Cunha JLR, Hounmanou YMG, Coura FM, Santana JA, Lobato FCF, Bojesen AM, et al. Fecal Shedding of Multidrug Resistant Escherichia coli Isolates in Dogs Fed with Raw Meat-Based Diets in Brazil. Antibiotics. 2022; 11(4):534. https://doi.org/10.3390/antibiotics11040534
Chicago/Turabian StyleRamos, Carolina Pantuzza, Carolina Yumi Iceri Kamei, Flávia Mello Viegas, Jonata de Melo Barbieri, João Luís Reis Cunha, Yaovi Mahuton Gildas Hounmanou, Fernanda Morcatti Coura, Jordana Almeida Santana, Francisco Carlos Faria Lobato, Anders Miki Bojesen, and et al. 2022. "Fecal Shedding of Multidrug Resistant Escherichia coli Isolates in Dogs Fed with Raw Meat-Based Diets in Brazil" Antibiotics 11, no. 4: 534. https://doi.org/10.3390/antibiotics11040534
APA StyleRamos, C. P., Kamei, C. Y. I., Viegas, F. M., de Melo Barbieri, J., Cunha, J. L. R., Hounmanou, Y. M. G., Coura, F. M., Santana, J. A., Lobato, F. C. F., Bojesen, A. M., & Silva, R. O. S. (2022). Fecal Shedding of Multidrug Resistant Escherichia coli Isolates in Dogs Fed with Raw Meat-Based Diets in Brazil. Antibiotics, 11(4), 534. https://doi.org/10.3390/antibiotics11040534







