Beyond Soil-Dwelling Actinobacteria: Fantastic Antibiotics and Where to Find Them
Abstract
1. Introduction
2. Bacterial Symbionts of Marine Invertebrates
2.1. Sponges
2.2. Tunicates
2.3. Other Groups of Marine Invertebrates
3. Insect-Associated Bacteria
4. Entomopathogenic Bacteria
5. Anaerobes
6. Myxobacteria
7. Cyanobacteria
8. Lichens
9. Pseudomonas
10. Burkholderia
11. Planctomycetes
12. The Mammalian Gut Microbiome
13. Culturing the Unculturable Treasures from the Soil
14. Extremophilic Bacteria
15. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Bentley, S.D.; Chater, K.F.; Cerdeño-Tárraga, A.-M.; Challis, G.L.; Thomson, N.R.; James, K.D.; Harris, D.E.; Quail, M.A.; Kieser, H.; Harper, D.; et al. Complete Genome Sequence of the Model Actinomycete Streptomyces coelicolor A3(2). Nature 2002, 417, 141–147. [Google Scholar] [CrossRef]
- Ikeda, H.; Ishikawa, J.; Hanamoto, A.; Shinose, M.; Kikuchi, H.; Shiba, T.; Sakaki, Y.; Hattori, M.; Ōmura, S. Complete Genome Sequence and Comparative Analysis of the Industrial Microorganism Streptomyces avermitilis. Nat. Biotechnol. 2003, 21, 526–531. [Google Scholar] [CrossRef]
- Oliynyk, M.; Samborskyy, M.; Lester, J.B.; Mironenko, T.; Scott, N.; Dickens, S.; Haydock, S.F.; Leadlay, P.F. Complete Genome Sequence of the Erythromycin-Producing Bacterium Saccharopolyspora erythraea NRRL23338. Nat. Biotechnol. 2007, 25, 447–453. [Google Scholar] [CrossRef]
- Schloss, P.D.; Handelsman, J. Status of the Microbial Census. Microbiol. Mol. Biol. Rev. 2004, 68, 686–691. [Google Scholar] [CrossRef] [PubMed]
- Amann, R.I.; Ludwig, W.; Schleifer, K.H. Phylogenetic Identification and in Situ Detection of Individual Microbial Cells without Cultivation. Microbiol. Rev. 1995, 59, 143–169. [Google Scholar] [CrossRef] [PubMed]
- Ventola, C.L. The Antibiotic Resistance Crisis: Part 1: Causes and Threats. P T 2015, 40, 277–283. [Google Scholar] [PubMed]
- Brown, E.D.; Wright, G.D. Antibacterial Drug Discovery in the Resistance Era. Nature 2016, 529, 336–343. [Google Scholar] [CrossRef] [PubMed]
- Santajit, S.; Indrawattana, N. Mechanisms of Antimicrobial Resistance in ESKAPE Pathogens. BioMed. Res. Int. 2016, 2016, 1–8. [Google Scholar] [CrossRef]
- Bachmann, B.O.; Van Lanen, S.G.; Baltz, R.H. Microbial Genome Mining for Accelerated Natural Products Discovery: Is a Renaissance in the Making? J. Ind. Microbiol. Biotechnol. 2014, 41, 175–184. [Google Scholar] [CrossRef]
- Zerikly, M.; Challis, G.L. Strategies for the Discovery of New Natural Products by Genome Mining. ChemBioChem 2009, 10, 625–633. [Google Scholar] [CrossRef]
- Ziemert, N.; Alanjary, M.; Weber, T. The Evolution of Genome Mining in Microbes—A Review. Nat. Prod. Rep. 2016, 33, 988–1005. [Google Scholar] [CrossRef]
- Lewis, W.H.; Tahon, G.; Geesink, P.; Sousa, D.Z.; Ettema, T.J.G. Innovations to Culturing the Uncultured Microbial Majority. Nat. Rev. Microbiol. 2021, 19, 225–240. [Google Scholar] [CrossRef]
- Mahler, L.; Niehs, S.P.; Martin, K.; Weber, T.; Scherlach, K.; Hertweck, C.; Roth, M.; Rosenbaum, M.A. Highly Parallelized Droplet Cultivation and Prioritization of Antibiotic Producers from Natural Microbial Communities. eLife 2021, 10, e64774. [Google Scholar] [CrossRef] [PubMed]
- Overmann, J.; Abt, B.; Sikorski, J. Present and Future of Culturing Bacteria. Annu. Rev. Microbiol. 2017, 71, 711–730. [Google Scholar] [CrossRef] [PubMed]
- Nichols, D.; Cahoon, N.; Trakhtenberg, E.M.; Pham, L.; Mehta, A.; Belanger, A.; Kanigan, T.; Lewis, K.; Epstein, S.S. Use of Ichip for High-Throughput In Situ Cultivation of “Uncultivable” Microbial Species. Appl. Environ. Microbiol. 2010, 76, 2445–2450. [Google Scholar] [CrossRef] [PubMed]
- Gomez-Escribano, J.; Alt, S.; Bibb, M. Next Generation Sequencing of Actinobacteria for the Discovery of Novel Natural Products. Mar. Drugs 2016, 14, 78. [Google Scholar] [CrossRef]
- Sukmarini, L. Recent Advances in Discovery of Lead Structures from Microbial Natural Products: Genomics- and Metabolomics-Guided Acceleration. Molecules 2021, 26, 2542. [Google Scholar] [CrossRef] [PubMed]
- Yamanaka, K.; Reynolds, K.A.; Kersten, R.D.; Ryan, K.S.; Gonzalez, D.J.; Nizet, V.; Dorrestein, P.C.; Moore, B.S. Direct Cloning and Refactoring of a Silent Lipopeptide Biosynthetic Gene Cluster Yields the Antibiotic Taromycin A. Proc. Natl. Acad. Sci. USA 2014, 111, 1957–1962. [Google Scholar] [CrossRef]
- Blin, K.; Shaw, S.; Kloosterman, A.M.; Charlop-Powers, Z.; van Wezel, G.P.; Medema, M.H.; Weber, T. AntiSMASH 6.0: Improving Cluster Detection and Comparison Capabilities. Nucleic Acids Res. 2021, 49, W29–W35. [Google Scholar] [CrossRef]
- Tietz, J.I.; Schwalen, C.J.; Patel, P.S.; Maxson, T.; Blair, P.M.; Tai, H.-C.; Zakai, U.I.; Mitchell, D.A. A New Genome-Mining Tool Redefines the Lasso Peptide Biosynthetic Landscape. Nat. Chem. Biol. 2017, 13, 470–478. [Google Scholar] [CrossRef]
- Skinnider, M.A.; Johnston, C.W.; Gunabalasingam, M.; Merwin, N.J.; Kieliszek, A.M.; MacLellan, R.J.; Li, H.; Ranieri, M.R.M.; Webster, A.L.H.; Cao, M.P.T.; et al. Comprehensive Prediction of Secondary Metabolite Structure and Biological Activity from Microbial Genome Sequences. Nat. Commun. 2020, 11, 6058. [Google Scholar] [CrossRef] [PubMed]
- Skinnider, M.A.; Merwin, N.J.; Johnston, C.W.; Magarvey, N.A. PRISM 3: Expanded Prediction of Natural Product Chemical Structures from Microbial Genomes. Nucleic Acids Res. 2017, 45, W49–W54. [Google Scholar] [CrossRef] [PubMed]
- Van Heel, A.J.; de Jong, A.; Montalbán-López, M.; Kok, J.; Kuipers, O.P. BAGEL3: Automated Identification of Genes Encoding Bacteriocins and (Non-)Bactericidal Posttranslationally Modified Peptides. Nucleic Acids Res. 2013, 41, W448–W453. [Google Scholar] [CrossRef] [PubMed]
- Santos-Aberturas, J.; Chandra, G.; Frattaruolo, L.; Lacret, R.; Pham, T.H.; Vior, N.M.; Eyles, T.H.; Truman, A.W. Uncovering the Unexplored Diversity of Thioamidated Ribosomal Peptides in Actinobacteria Using the RiPPER Genome Mining Tool. Nucleic Acids Res. 2019, 47, 4624–4637. [Google Scholar] [CrossRef] [PubMed]
- Benítez, X.; Gonzalez, E.G.; García, J.; Zúñiga, P.; Calle, F.; Cuevas, C. Detection of a Pederin-like Compound Using a Dilution-to-extinction-based Platform for the Isolation of Marine Bacteria in Drug Discovery Strategies. Microb. Biotechnol. 2021, 14, 241–250. [Google Scholar] [CrossRef]
- Torres, J.P.; Lin, Z.; Winter, J.M.; Krug, P.J.; Schmidt, E.W. Animal Biosynthesis of Complex Polyketides in a Photosynthetic Partnership. Nat. Commun. 2020, 11, 2882. [Google Scholar] [CrossRef]
- Morita, M.; Schmidt, E.W. Parallel Lives of Symbionts and Hosts: Chemical Mutualism in Marine Animals. Nat. Prod. Rep. 2018, 35, 357–378. [Google Scholar] [CrossRef]
- Haygood, M.G.; Schmidt, E.W.; Davidson, S.K.; Faulkner, D.J. Microbial Symbionts of Marine Invertebrates: Opportunities for Microbial Biotechnology. J. Mol. Microbiol. Biotechnol. 1999, 1, 33–43. [Google Scholar]
- Bakshani, C.R.; Morales-Garcia, A.L.; Althaus, M.; Wilcox, M.D.; Pearson, J.P.; Bythell, J.C.; Burgess, J.G. Evolutionary Conservation of the Antimicrobial Function of Mucus: A First Defence against Infection. Npj Biofilm. Microbiom. 2018, 4, 14. [Google Scholar] [CrossRef]
- Kennedy, J.; Codling, C.E.; Jones, B.V.; Dobson, A.D.W.; Marchesi, J.R. Diversity of Microbes Associated with the Marine Sponge, Haliclona Simulans, Isolated from Irish Waters and Identification of Polyketide Synthase Genes from the Sponge Metagenome. Environ. Microbiol. 2008, 10, 1888–1902. [Google Scholar] [CrossRef]
- Unson, M.D.; Holland, N.D.; Faulkner, D.J. A Brominated Secondary Metabolite Synthesized by the Cyanobacterial Symbiont of a Marine Sponge and Accumulation of the Crystalline Metabolite in the Sponge Tissue. Mar. Biol. 1994, 119, 1–11. [Google Scholar] [CrossRef]
- Siegl, A.; Kamke, J.; Hochmuth, T.; Piel, J.; Richter, M.; Liang, C.; Dandekar, T.; Hentschel, U. Single-Cell Genomics Reveals the Lifestyle of Poribacteria, a Candidate Phylum Symbiotically Associated with Marine Sponges. ISME J. 2011, 5, 61–70. [Google Scholar] [CrossRef] [PubMed]
- Wilson, M.C.; Mori, T.; Rückert, C.; Uria, A.R.; Helf, M.J.; Takada, K.; Gernert, C.; Steffens, U.A.E.; Heycke, N.; Schmitt, S.; et al. An Environmental Bacterial Taxon with a Large and Distinct Metabolic Repertoire. Nature 2014, 506, 58–62. [Google Scholar] [CrossRef] [PubMed]
- Hertweck, C. The Biosynthetic Logic of Polyketide Diversity. Angew. Chem. Int. Ed. 2009, 48, 4688–4716. [Google Scholar] [CrossRef] [PubMed]
- Fieseler, L.; Hentschel, U.; Grozdanov, L.; Schirmer, A.; Wen, G.; Platzer, M.; Hrvatin, S.; Butzke, D.; Zimmermann, K.; Piel, J. Widespread Occurrence and Genomic Context of Unusually Small Polyketide Synthase Genes in Microbial Consortia Associated with Marine Sponges. Appl. Environ. Microbiol. 2007, 73, 2144–2155. [Google Scholar] [CrossRef]
- Hamada, T.; Matsunaga, S.; Fujiwara, M.; Fujita, K.; Hirota, H.; Schmucki, R.; Güntert, P.; Fusetani, N. Solution Structure of Polytheonamide B, a Highly Cytotoxic Nonribosomal Polypeptide from Marine Sponge. J. Am. Chem. Soc. 2010, 132, 12941–12945. [Google Scholar] [CrossRef]
- Süssmuth, R.D.; Mainz, A. Nonribosomal Peptide Synthesis-Principles and Prospects. Angew. Chem. Int. Ed. 2017, 56, 3770–3821. [Google Scholar] [CrossRef]
- Arnison, P.G.; Bibb, M.J.; Bierbaum, G.; Bowers, A.A.; Bugni, T.S.; Bulaj, G.; Camarero, J.A.; Campopiano, D.J.; Challis, G.L.; Clardy, J.; et al. Ribosomally Synthesized and Post-Translationally Modified Peptide Natural Products: Overview and Recommendations for a Universal Nomenclature. Nat. Prod. Rep. 2013, 30, 108–160. [Google Scholar] [CrossRef]
- Montalbán-López, M.; Scott, T.A.; Ramesh, S.; Rahman, I.R.; van Heel, A.J.; Viel, J.H.; Bandarian, V.; Dittmann, E.; Genilloud, O.; Goto, Y.; et al. New Developments in RiPP Discovery, Enzymology and Engineering. Nat. Prod. Rep. 2021, 38, 130–239. [Google Scholar] [CrossRef]
- Agarwal, V.; Blanton, J.M.; Podell, S.; Taton, A.; Schorn, M.A.; Busch, J.; Lin, Z.; Schmidt, E.W.; Jensen, P.R.; Paul, V.J.; et al. Metagenomic Discovery of Polybrominated Diphenyl Ether Biosynthesis by Marine Sponges. Nat. Chem. Biol. 2017, 13, 537–543. [Google Scholar] [CrossRef]
- Nguyen, N.A.; Lin, Z.; Mohanty, I.; Garg, N.; Schmidt, E.W.; Agarwal, V. An Obligate Peptidyl Brominase Underlies the Discovery of Highly Distributed Biosynthetic Gene Clusters in Marine Sponge Microbiomes. J. Am. Chem. Soc. 2021, 143, 10221–10231. [Google Scholar] [CrossRef]
- Dou, X.; Dong, B. Origins and Bioactivities of Natural Compounds Derived from Marine Ascidians and Their Symbionts. Mar. Drugs 2019, 17, 670. [Google Scholar] [CrossRef]
- Ramesh, C.; Tulasi, B.R.; Raju, M.; Thakur, N.; Dufossé, L. Marine Natural Products from Tunicates and Their Associated Microbes. Mar. Drugs 2021, 19, 308. [Google Scholar] [CrossRef] [PubMed]
- Cuevas, C.; Francesch, A. Development of Yondelis® (Trabectedin, ET-743). A Semisynthetic Process Solves the Supply Problem. Nat. Prod. Rep. 2009, 26, 322. [Google Scholar] [CrossRef] [PubMed]
- Rath, C.M.; Janto, B.; Earl, J.; Ahmed, A.; Hu, F.Z.; Hiller, L.; Dahlgren, M.; Kreft, R.; Yu, F.; Wolff, J.J.; et al. Meta-Omic Characterization of the Marine Invertebrate Microbial Consortium That Produces the Chemotherapeutic Natural Product ET-743. ACS Chem. Biol. 2011, 6, 1244–1256. [Google Scholar] [CrossRef] [PubMed]
- Schofield, M.M.; Jain, S.; Porat, D.; Dick, G.J.; Sherman, D.H. Identification and Analysis of the Bacterial Endosymbiont Specialized for Production of the Chemotherapeutic Natural Product ET-743: Metagenomic Analysis of the ET-743 Producer. Environ. Microbiol. 2015, 17, 3964–3975. [Google Scholar] [CrossRef]
- Patel, S.; Petty, W.J.; Sands, J.M. An Overview of Lurbinectedin as a New Second-Line Treatment Option for Small Cell Lung Cancer. Ther. Adv. Med. Oncol. 2021, 13, 175883592110205. [Google Scholar] [CrossRef]
- White, K.M.; Rosales, R.; Yildiz, S.; Kehrer, T.; Miorin, L.; Moreno, E.; Jangra, S.; Uccellini, M.B.; Rathnasinghe, R.; Coughlan, L.; et al. Plitidepsin Has Potent Preclinical Efficacy against SARS-CoV-2 by Targeting the Host Protein EEF1A. Science 2021, 371, 926–931. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Kersten, R.D.; Nam, S.-J.; Lu, L.; Al-Suwailem, A.M.; Zheng, H.; Fenical, W.; Dorrestein, P.C.; Moore, B.S.; Qian, P.-Y. Bacterial Biosynthesis and Maturation of the Didemnin Anti-Cancer Agents. J. Am. Chem. Soc. 2012, 134, 8625–8632. [Google Scholar] [CrossRef]
- Tsukimoto, M.; Nagaoka, M.; Shishido, Y.; Fujimoto, J.; Nishisaka, F.; Matsumoto, S.; Harunari, E.; Imada, C.; Matsuzaki, T. Bacterial Production of the Tunicate-Derived Antitumor Cyclic Depsipeptide Didemnin B. J. Nat. Prod. 2011, 74, 2329–2331. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, E.W.; Nelson, J.T.; Rasko, D.A.; Sudek, S.; Eisen, J.A.; Haygood, M.G.; Ravel, J. Patellamide A and C Biosynthesis by a Microcin-like Pathway in Prochloron Didemni, the Cyanobacterial Symbiont of Lissoclinum patella. Proc. Natl. Acad. Sci. USA 2005, 102, 7315–7320. [Google Scholar] [CrossRef]
- Schmidt, E.W.; Sudek, S.; Haygood, M.G. Genetic Evidence Supports Secondary Metabolic Diversity in Prochloron Spp., the Cyanobacterial Symbiont of a Tropical Ascidian. J. Nat. Prod. 2004, 67, 1341–1345. [Google Scholar] [CrossRef] [PubMed]
- Smith, T.E.; Pond, C.D.; Pierce, E.; Harmer, Z.P.; Kwan, J.; Zachariah, M.M.; Harper, M.K.; Wyche, T.P.; Matainaho, T.K.; Bugni, T.S.; et al. Accessing Chemical Diversity from the Uncultivated Symbionts of Small Marine Animals. Nat. Chem. Biol. 2018, 14, 179–185. [Google Scholar] [CrossRef]
- Donia, M.S.; Fricke, W.F.; Partensky, F.; Cox, J.; Elshahawi, S.I.; White, J.R.; Phillippy, A.M.; Schatz, M.C.; Piel, J.; Haygood, M.G.; et al. Complex Microbiome Underlying Secondary and Primary Metabolism in the Tunicate-Prochloron Symbiosis. Proc. Natl. Acad. Sci. USA 2011, 108, E1423–E1432. [Google Scholar] [CrossRef]
- Richardson, A.D.; Aalbersberg, W.; Ireland, C.M. The Patellazoles Inhibit Protein Synthesis at Nanomolar Concentrations in Human Colon Tumor Cells. Anti-Cancer Drugs 2005, 16, 533–541. [Google Scholar] [CrossRef]
- Helfrich, E.J.N.; Piel, J. Biosynthesis of Polyketides by trans-AT Polyketide Synthases. Nat. Prod. Rep. 2016, 33, 231–316. [Google Scholar] [CrossRef] [PubMed]
- Kwan, J.C.; Donia, M.S.; Han, A.W.; Hirose, E.; Haygood, M.G.; Schmidt, E.W. Genome Streamlining and Chemical Defense in a Coral Reef Symbiosis. Proc. Natl. Acad. Sci. USA 2012, 109, 20655–20660. [Google Scholar] [CrossRef]
- Ciavatta, M.L.; Lefranc, F.; Vieira, L.M.; Kiss, R.; Carbone, M.; van Otterlo, W.A.L.; Lopanik, N.B.; Waeschenbach, A. The Phylum Bryozoa: From Biology to Biomedical Potential. Mar. Drugs 2020, 18, 200. [Google Scholar] [CrossRef] [PubMed]
- Altamia, M.A.; Lin, Z.; Trindade-Silva, A.E.; Uy, I.D.; Shipway, J.R.; Wilke, D.V.; Concepcion, G.P.; Distel, D.L.; Schmidt, E.W.; Haygood, M.G. Secondary Metabolism in the Gill Microbiota of Shipworms (Teredinidae) as Revealed by Comparison of Metagenomes and Nearly Complete Symbiont Genomes. mSystems 2020, 5. [Google Scholar] [CrossRef] [PubMed]
- Coutinho, M.C.L.; Teixeira, V.L.; Santos, C.S.G. A Review of “Polychaeta” Chemicals and Their Possible Ecological Role. J. Chem. Ecol. 2018, 44, 72–94. [Google Scholar] [CrossRef] [PubMed]
- Trindade-Silva, A.E.; Lim-Fong, G.E.; Sharp, K.H.; Haygood, M.G. Bryostatins: Biological Context and Biotechnological Prospects. Curr. Opin. Biotechnol. 2010, 21, 834–842. [Google Scholar] [CrossRef] [PubMed]
- Sharp, K.H.; Davidson, S.K.; Haygood, M.G. Localization of ‘Candidatus Endobugula Sertula’ and the Bryostatins throughout the Life Cycle of the Bryozoan Bugula Neritina. ISME J. 2007, 1, 693–702. [Google Scholar] [CrossRef]
- Lim, G.E.; Haygood, M.G. “Candidatus Endobugula Glebosa,” a Specific Bacterial Symbiont of the Marine Bryozoan Bugula simplex. Appl. Environ. Microbiol. 2004, 70, 4921–4929. [Google Scholar] [CrossRef] [PubMed]
- Davidson, S.K.; Allen, S.W.; Lim, G.E.; Anderson, C.M.; Haygood, M.G. Evidence for the Biosynthesis of Bryostatins by the Bacterial Symbiont “Candidatus Endobugula Sertula” of the Bryozoan Bugula neritina. Appl. Environ. Microbiol. 2001, 67, 4531–4537. [Google Scholar] [CrossRef] [PubMed]
- Sudek, S.; Lopanik, N.B.; Waggoner, L.E.; Hildebrand, M.; Anderson, C.; Liu, H.; Patel, A.; Sherman, D.H.; Haygood, M.G. Identification of the Putative Bryostatin Polyketide Synthase Gene Cluster from “Candidatus Endobugula sertula”, the Uncultivated Microbial Symbiont of the Marine Bryozoan Bugula neritina. J. Nat. Prod. 2007, 70, 67–74. [Google Scholar] [CrossRef] [PubMed]
- Hildebrand, M.; Waggoner, L.E.; Liu, H.; Sudek, S.; Allen, S.; Anderson, C.; Sherman, D.H.; Haygood, M. bryA: An Unusual Modular Polyketide Synthase Gene from the Uncultivated Bacterial Symbiont of the Marine Bryozoan Bugula neritina. Chem. Biol. 2004, 11, 1543–1552. [Google Scholar] [CrossRef] [PubMed]
- Miller, B.W.; Lim, A.L.; Lin, Z.; Bailey, J.; Aoyagi, K.L.; Fisher, M.A.; Barrows, L.R.; Manoil, C.; Schmidt, E.W.; Haygood, M.G. Shipworm Symbiosis Ecology-Guided Discovery of an Antibiotic That Kills Colistin-Resistant Acinetobacter. Cell Chem. Biol. 2021, S2451945621002208. [Google Scholar] [CrossRef]
- Elshahawi, S.I.; Trindade-Silva, A.E.; Hanora, A.; Han, A.W.; Flores, M.S.; Vizzoni, V.; Schrago, C.G.; Soares, C.A.; Concepcion, G.P.; Distel, D.L.; et al. Boronated Tartrolon Antibiotic Produced by Symbiotic Cellulose-Degrading Bacteria in Shipworm Gills. Proc. Natl. Acad. Sci. USA 2013, 110, E295–E304. [Google Scholar] [CrossRef] [PubMed]
- Irschik, H.; Schummer, D.; Gerth, K.; Höfle, G.; Reichenbach, H. Antibiotics from Gliding Bacteria. No.60. The Tartrolons, New Boron-Containing Antibiotics from a Myxobacterium, Sorangium Cellulosum. J. Antibiot. 1995, 48, 26–30. [Google Scholar] [CrossRef] [PubMed]
- O’Connor, R.M.; Nepveux, V.F.J.; Abenoja, J.; Bowden, G.; Reis, P.; Beaushaw, J.; Bone Relat, R.M.; Driskell, I.; Gimenez, F.; Riggs, M.W.; et al. A Symbiotic Bacterium of Shipworms Produces a Compound with Broad Spectrum Anti-Apicomplexan Activity. PLoS Pathog. 2020, 16, e1008600. [Google Scholar] [CrossRef]
- Yang, J.C.; Madupu, R.; Durkin, A.S.; Ekborg, N.A.; Pedamallu, C.S.; Hostetler, J.B.; Radune, D.; Toms, B.S.; Henrissat, B.; Coutinho, P.M.; et al. The Complete Genome of Teredinibacter turnerae T7901: An Intracellular Endosymbiont of Marine Wood-Boring Bivalves (Shipworms). PLoS ONE 2009, 4, e6085. [Google Scholar] [CrossRef] [PubMed]
- Pettit, G.R.; Kamano, Y.; Herald, C.L.; Tuinman, A.A.; Boettner, F.E.; Kizu, H.; Schmidt, J.M.; Baczynskyj, L.; Tomer, K.B.; Bontems, R.J. The Isolation and Structure of a Remarkable Marine Animal Antineoplastic Constituent: Dolastatin 10. J. Am. Chem. Soc. 1987, 109, 6883–6885. [Google Scholar] [CrossRef]
- Bai, R.; Petit, G.R.; Hamel, E. Dolastatin 10, a Powerful Cytostatic Peptide Derived from a Marine Animal. Biochem. Pharmacol. 1990, 39, 1941–1949. [Google Scholar] [CrossRef]
- Francisco, J.A.; Cerveny, C.G.; Meyer, D.L.; Mixan, B.J.; Klussman, K.; Chace, D.F.; Rejniak, S.X.; Gordon, K.A.; DeBlanc, R.; Toki, B.E.; et al. CAC10-VcMMAE, an Anti-CD30–Monomethyl Auristatin E Conjugate with Potent and Selective Antitumor Activity. Blood 2003, 102, 1458–1465. [Google Scholar] [CrossRef] [PubMed]
- Luesch, H.; Moore, R.E.; Paul, V.J.; Mooberry, S.L.; Corbett, T.H. Isolation of Dolastatin 10 from the Marine Cyanobacterium Symploca Species VP642 and Total Stereochemistry and Biological Evaluation of Its Analogue Symplostatin 1. J. Nat. Prod. 2001, 64, 907–910. [Google Scholar] [CrossRef]
- Harrigan, G.G.; Luesch, H.; Yoshida, W.Y.; Moore, R.E.; Nagle, D.G.; Paul, V.J.; Mooberry, S.L.; Corbett, T.H.; Valeriote, F.A. Symplostatin 1: A Dolastatin 10 Analogue from the Marine Cyanobacterium Symploca hydnoides. J. Nat. Prod. 1998, 61, 1075–1077. [Google Scholar] [CrossRef] [PubMed]
- Iwasaki, A.; Sumimoto, S.; Ohno, O.; Suda, S.; Suenaga, K. Kurahamide, a Cyclic Depsipeptide Analog of Dolastatin 13 from a Marine Cyanobacterial Assemblage of Lyngbya sp. BCSJ 2014, 87, 609–613. [Google Scholar] [CrossRef]
- Heming, B. Book Review: Grimaldi, D. & Engel, M. 2005: Evolution of the Insects. Eur. J. Entomol. 2006, 103, 273–275. [Google Scholar] [CrossRef]
- Mikheyev, A.S.; Vo, T.; Mueller, U.G. Phylogeography of Post-Pleistocene Population Expansion in a Fungus-Gardening Ant and Its Microbial Mutualists. Mol. Ecol. 2008, 17, 4480–4488. [Google Scholar] [CrossRef] [PubMed]
- Cafaro, M.J.; Currie, C.R. Phylogenetic Analysis of Mutualistic Filamentous Bacteria Associated with Fungus-Growing Ants. Can. J. Microbiol. 2005, 51, 441–446. [Google Scholar] [CrossRef]
- Haeder, S.; Wirth, R.; Herz, H.; Spiteller, D. Candicidin-Producing Streptomyces Support Leaf-Cutting Ants to Protect Their Fungus Garden against the Pathogenic Fungus Escovopsis. Proc. Natl. Acad. Sci. USA 2009, 106, 4742–4746. [Google Scholar] [CrossRef]
- Seipke, R.F.; Barke, J.; Brearley, C.; Hill, L.; Yu, D.W.; Goss, R.J.M.; Hutchings, M.I. A Single Streptomyces Symbiont Makes Multiple Antifungals to Support the Fungus Farming Ant Acromyrmex octospinosus. PLoS ONE 2011, 6, e22028. [Google Scholar] [CrossRef]
- Seipke, R.F.; Barke, J.; Heavens, D.; Yu, D.W.; Hutchings, M.I. Analysis of the Bacterial Communities Associated with Two Ant-Plant Symbioses. Microbiol. Open 2013, 2, 276–283. [Google Scholar] [CrossRef] [PubMed]
- Francoeur, C.B.; May, D.S.; Thairu, M.W.; Hoang, D.Q.; Panthofer, O.; Bugni, T.S.; Pupo, M.T.; Clardy, J.; Pinto-Tomás, A.A.; Currie, C.R. Burkholderia from Fungus Gardens of Fungus-Growing Ants Produces Antifungals That Inhibit the Specialized Parasite Escovopsis. Appl. Environ. Microbiol. 2021, 87. [Google Scholar] [CrossRef]
- Batey, S.F.D.; Greco, C.; Hutchings, M.I.; Wilkinson, B. Chemical Warfare between Fungus-Growing Ants and Their Pathogens. Curr. Opin. Chem. Biol. 2020, 59, 172–181. [Google Scholar] [CrossRef]
- Goldstein, S.L.; Klassen, J.L. Pseudonocardia Symbionts of Fungus-Growing Ants and the Evolution of Defensive Secondary Metabolism. Front. Microbiol. 2020, 11, 621041. [Google Scholar] [CrossRef]
- Oh, D.-C.; Poulsen, M.; Currie, C.R.; Clardy, J. Dentigerumycin: A Bacterial Mediator of an Ant-Fungus Symbiosis. Nat. Chem. Biol. 2009, 5, 391–393. [Google Scholar] [CrossRef] [PubMed]
- Prado-Alonso, L.; Pérez-Victoria, I.; Malmierca, M.G.; Montero, I.; Rioja-Blanco, E.; Martín, J.; Reyes, F.; Méndez, C.; Salas, J.A.; Olano, C. Colibrimycins, Novel Halogenated Hybrid PKS-NRPS Compounds Produced by Streptomyces sp. CS147. Appl. Environ. Microbiol. 2021, AEM0183921. [Google Scholar] [CrossRef]
- Garcia, I.; Vior, N.M.; Braña, A.F.; González-Sabin, J.; Rohr, J.; Moris, F.; Méndez, C.; Salas, J.A. Elucidating the Biosynthetic Pathway for the Polyketide-Nonribosomal Peptide Collismycin A: Mechanism for Formation of the 2,2′-Bipyridyl Ring. Chem. Biol. 2012, 19, 399–413. [Google Scholar] [CrossRef] [PubMed]
- Carr, G.; Derbyshire, E.R.; Caldera, E.; Currie, C.R.; Clardy, J. Antibiotic and Antimalarial Quinones from Fungus-Growing Ant-Associated Pseudonocardia sp. J. Nat. Prod. 2012, 75, 1806–1809. [Google Scholar] [CrossRef]
- Ortega, H.E.; Lourenzon, V.B.; Chevrette, M.G.; Ferreira, L.L.G.; Alvarenga, R.F.R.; Melo, W.G.P.; Venâncio, T.; Currie, C.R.; Andricopulo, A.D.; Bugni, T.S.; et al. Antileishmanial Macrolides from Ant-Associated Streptomyces sp. ISID311. Bioorganic Med. Chem. 2021, 32, 116016. [Google Scholar] [CrossRef]
- Fukuda, T.T.H.; Helfrich, E.J.N.; Mevers, E.; Melo, W.G.P.; Van Arnam, E.B.; Andes, D.R.; Currie, C.R.; Pupo, M.T.; Clardy, J. Specialized Metabolites Reveal Evolutionary History and Geographic Dispersion of a Multilateral Symbiosis. ACS Cent. Sci. 2021, 7, 292–299. [Google Scholar] [CrossRef] [PubMed]
- Holmes, N.A.; Devine, R.; Qin, Z.; Seipke, R.F.; Wilkinson, B.; Hutchings, M.I. Complete Genome Sequence of Streptomyces formicae KY5, the Formicamycin Producer. J. Biotechnol. 2018, 265, 116–118. [Google Scholar] [CrossRef]
- Qin, Z.; Munnoch, J.T.; Devine, R.; Holmes, N.A.; Seipke, R.F.; Wilkinson, K.A.; Wilkinson, B.; Hutchings, M.I. Formicamycins, Antibacterial Polyketides Produced by Streptomyces Formicae Isolated from African Tetraponera Plant-Ants. Chem. Sci. 2017, 8, 3218–3227. [Google Scholar] [CrossRef]
- Qin, Z.; Devine, R.; Hutchings, M.I.; Wilkinson, B. A Role for Antibiotic Biosynthesis Monooxygenase Domain Proteins in Fidelity Control during Aromatic Polyketide Biosynthesis. Nat. Commun. 2019, 10, 3611. [Google Scholar] [CrossRef]
- Qin, Z.; Devine, R.; Booth, T.J.; Farrar, E.H.E.; Grayson, M.N.; Hutchings, M.I.; Wilkinson, B. Formicamycin Biosynthesis Involves a Unique Reductive Ring Contraction. Chem. Sci. 2020, 11, 8125–8131. [Google Scholar] [CrossRef] [PubMed]
- Devine, R.; McDonald, H.P.; Qin, Z.; Arnold, C.J.; Noble, K.; Chandra, G.; Wilkinson, B.; Hutchings, M.I. Re-Wiring the Regulation of the Formicamycin Biosynthetic Gene Cluster to Enable the Development of Promising Antibacterial Compounds. Cell Chem. Biol. 2021, 28, 515–523.e5. [Google Scholar] [CrossRef]
- Yuan, J.; Wang, L.; Ren, J.; Huang, J.-P.; Yu, M.; Tang, J.; Yan, Y.; Yang, J.; Huang, S.-X. Antibacterial Pentacyclic Polyketides from a Soil-Derived Streptomyces. J. Nat. Prod. 2020, 83, 1919–1924. [Google Scholar] [CrossRef] [PubMed]
- Mueller, U.G.; Dash, D.; Rabeling, C.; Rodrigues, A. Coevolution betwwen Attine ants and actinomycetes bacteria: A reevaluation. Evolution 2008, 62, 2894–2912. [Google Scholar] [CrossRef] [PubMed]
- Sen, R.; Ishak, H.D.; Estrada, D.; Dowd, S.E.; Hong, E.; Mueller, U.G. Generalized Antifungal Activity and 454-Screening of Pseudonocardia and Amycolatopsis Bacteria in Nests of Fungus-Growing Ants. Proc. Natl. Acad. Sci. USA 2009, 106, 17805–17810. [Google Scholar] [CrossRef]
- Li, H.; Sosa-Calvo, J.; Horn, H.A.; Pupo, M.T.; Clardy, J.; Rabeling, C.; Schultz, T.R.; Currie, C.R. Convergent Evolution of Complex Structures for Ant–Bacterial Defensive Symbiosis in Fungus-Farming Ants. Proc. Natl. Acad. Sci. USA 2018, 115, 10720–10725. [Google Scholar] [CrossRef] [PubMed]
- Engl, T.; Kroiss, J.; Kai, M.; Nechitaylo, T.Y.; Svatoš, A.; Kaltenpoth, M. Evolutionary Stability of Antibiotic Protection in a Defensive Symbiosis. Proc. Natl. Acad. Sci. USA 2018, 115, E2020–E2029. [Google Scholar] [CrossRef] [PubMed]
- Kaltenpoth, M.; Göttler, W.; Herzner, G.; Strohm, E. Symbiotic Bacteria Protect Wasp Larvae from Fungal Infestation. Curr. Biol. 2005, 15, 475–479. [Google Scholar] [CrossRef]
- Nechitaylo, T.Y.; Sandoval-Calderón, M.; Engl, T.; Wielsch, N.; Dunn, D.M.; Goesmann, A.; Strohm, E.; Svatoš, A.; Dale, C.; Weiss, R.B.; et al. Incipient Genome Erosion and Metabolic Streamlining for Antibiotic Production in a Defensive Symbiont. Proc. Natl. Acad. Sci. USA 2021, 118, e2023047118. [Google Scholar] [CrossRef] [PubMed]
- Piel, J. A Polyketide Synthase-Peptide Synthetase Gene Cluster from an Uncultured Bacterial Symbiont of Paederus Beetles. Proc. Natl. Acad. Sci. USA 2002, 99, 14002–14007. [Google Scholar] [CrossRef] [PubMed]
- Piel, J.; Wen, G.; Platzer, M.; Hui, D. Unprecedented Diversity of Catalytic Domains in the First Four Modules of the Putative Pederin Polyketide Synthase. ChemBioChem 2004, 5, 93–98. [Google Scholar] [CrossRef]
- Kellner, R.L.L. Molecular Identification of an Endosymbiotic Bacterium Associated with Pederin Biosynthesis in Paederus sabaeus (Coleoptera: Staphylinidae). Insect Biochem. Mol. Biol. 2002, 32, 389–395. [Google Scholar] [CrossRef]
- Nakabachi, A.; Ueoka, R.; Oshima, K.; Teta, R.; Mangoni, A.; Gurgui, M.; Oldham, N.J.; van Echten-Deckert, G.; Okamura, K.; Yamamoto, K.; et al. Defensive Bacteriome Symbiont with a Drastically Reduced Genome. Curr. Biol. 2013, 23, 1478–1484. [Google Scholar] [CrossRef]
- Schleissner, C.; Cañedo, L.M.; Rodríguez, P.; Crespo, C.; Zúñiga, P.; Peñalver, A.; de la Calle, F.; Cuevas, C. Bacterial Production of a Pederin Analogue by a Free-Living Marine Alphaproteobacterium. J. Nat. Prod. 2017, 80, 2170–2173. [Google Scholar] [CrossRef]
- Kust, A.; Mareš, J.; Jokela, J.; Urajová, P.; Hájek, J.; Saurav, K.; Voráčová, K.; Fewer, D.P.; Haapaniemi, E.; Permi, P.; et al. Discovery of a Pederin Family Compound in a Nonsymbiotic Bloom-Forming Cyanobacterium. ACS Chem. Biol. 2018, 13, 1123–1129. [Google Scholar] [CrossRef]
- Flórez, L.V.; Scherlach, K.; Miller, I.J.; Rodrigues, A.; Kwan, J.C.; Hertweck, C.; Kaltenpoth, M. An Antifungal Polyketide Associated with Horizontally Acquired Genes Supports Symbiont-Mediated Defense in Lagria villosa Beetles. Nat. Commun. 2018, 9, 2478. [Google Scholar] [CrossRef]
- Waterworth, S.C.; Flórez, L.V.; Rees, E.R.; Hertweck, C.; Kaltenpoth, M.; Kwan, J.C. Horizontal Gene Transfer to a Defensive Symbiont with a Reduced Genome in a Multipartite Beetle Microbiome. mBio 2020, 11. [Google Scholar] [CrossRef] [PubMed]
- Zalom, F.G. Pesticide Use Practices in Integrated Pest Management. In Hayes’ Handbook of Pesticide Toxicology; Elsevier: Amsterdam, The Netherlands, 2010; pp. 303–313. ISBN 978-0-12-374367-1. [Google Scholar]
- Kalha, C.S.; Singh, P.P.; Kang, S.S.; Hunjan, M.S.; Gupta, V.; Sharma, R. Entomopathogenic Viruses and Bacteria for Insect-Pest Control. In Integrated Pest Management; Elsevier: Amsterdam, The Netherlands, 2014; pp. 225–244. ISBN 978-0-12-398529-3. [Google Scholar]
- Lacey, L. Microbial Control of Insect and Mite Pests; Elsevier: Amsterdam, The Netherlands, 2017; ISBN 978-0-12-803527-6. [Google Scholar]
- Jouzani, G.S.; Valijanian, E.; Sharafi, R. Bacillus thuringiensis: A Successful Insecticide with New Environmental Features and Tidings. Appl. Microbiol. Biotechnol. 2017, 101, 2691–2711. [Google Scholar] [CrossRef]
- Ffrench-Constant, R.H.; Waterfield, N.; Daborn, P. Insecticidal Toxins from Photorhabdus and Xenorhabdus. In Comprehensive Molecular Insect Science; Elsevier: Amsterdam, The Netherlands, 2005; pp. 239–253. ISBN 978-0-444-51924-5. [Google Scholar]
- Booysen, E.; Dicks, L.M.T. Does the Future of Antibiotics Lie in Secondary Metabolites Produced by Xenorhabdus spp.? A Review. Probiotics Antimicro. Prot. 2020, 12, 1310–1320. [Google Scholar] [CrossRef]
- Tobias, N.J.; Wolff, H.; Djahanschiri, B.; Grundmann, F.; Kronenwerth, M.; Shi, Y.-M.; Simonyi, S.; Grün, P.; Shapiro-Ilan, D.; Pidot, S.J.; et al. Natural Product Diversity Associated with the Nematode Symbionts Photorhabdus and Xenorhabdus. Nat. Microbiol. 2017, 2, 1676–1685. [Google Scholar] [CrossRef] [PubMed]
- Bozhüyük, K.A.J.; Zhou, Q.; Engel, Y.; Heinrich, A.; Pérez, A.; Bode, H.B. Natural Products from Photorhabdus and Other Entomopathogenic Bacteria. In The Molecular Biology of Photorhabdus Bacteria; Ffrench-Constant, R.H., Ed.; Current Topics in Microbiology and Immunology; Springer International Publishing: Cham, Switzerland, 2016; Volume 402, ISBN 978-3-319-52714-7. [Google Scholar]
- Dreyer, J.; Malan, A.P.; Dicks, L.M.T. Bacteria of the Genus Xenorhabdus, a Novel Source of Bioactive Compounds. Front. Microbiol. 2018, 9, 3177. [Google Scholar] [CrossRef] [PubMed]
- Crawford, J.M.; Portmann, C.; Zhang, X.; Roeffaers, M.B.J.; Clardy, J. Small Molecule Perimeter Defense in Entomopathogenic Bacteria. Proc. Natl. Acad. Sci. USA 2012, 109, 10821–10826. [Google Scholar] [CrossRef] [PubMed]
- Seo, S.; Lee, S.; Hong, Y.; Kim, Y. Phospholipase A2 Inhibitors Synthesized by Two Entomopathogenic Bacteria, Xenorhabdus Nematophila and Photorhabdus Temperata Subsp. Temperata. Appl. Environ. Microbiol. 2012, 78, 3816–3823. [Google Scholar] [CrossRef]
- Ji, D.; Yi, Y.; Kang, G.-H.; Choi, Y.-H.; Kim, P.; Baek, N.-I.; Kim, Y. Identification of an Antibacterial Compound, Benzylideneacetone, from Xenorhabdus Nematophila against Major Plant-Pathogenic Bacteria. FEMS Microbiol. Lett. 2004, 239, 241–248. [Google Scholar] [CrossRef]
- Joyce, S.A.; Brachmann, A.O.; Glazer, I.; Lango, L.; Schwär, G.; Clarke, D.J.; Bode, H.B. Bacterial Biosynthesis of a Multipotent Stilbene. Angew. Chem. Int. Ed. 2008, 47, 1942–1945. [Google Scholar] [CrossRef]
- Lang, G.; Kalvelage, T.; Peters, A.; Wiese, J.; Imhoff, J.F. Linear and Cyclic Peptides from the Entomopathogenic Bacterium Xenorhabdus nematophilus. J. Nat. Prod. 2008, 71, 1074–1077. [Google Scholar] [CrossRef]
- Grundmann, F.; Kaiser, M.; Kurz, M.; Schiell, M.; Batzer, A.; Bode, H.B. Structure Determination of the Bioactive Depsipeptide Xenobactin from Xenorhabdus sp. PB30.3. RSC Adv. 2013, 3, 22072. [Google Scholar] [CrossRef]
- Ohlendorf, B.; Simon, S.; Wiese, J.; Imhoff, J.F. Szentiamide, an N-Formylated Cyclic Depsipeptide from Xenorhabdus Szentirmaii DSM 16338T. Nat. Prod. Commun. 2011, 6, 1934578X1100600. [Google Scholar] [CrossRef]
- Nollmann, F.I.; Dowling, A.; Kaiser, M.; Deckmann, K.; Grösch, S.; ffrench-Constant, R.; Bode, H.B. Synthesis of Szentiamide, a Depsipeptide from Entomopathogenic Xenorhabdus szentirmaii with Activity against Plasmodium falciparum. Beilstein J. Org. Chem. 2012, 8, 528–533. [Google Scholar] [CrossRef] [PubMed]
- Xi, X.; Lu, X.; Zhang, X.; Bi, Y.; Li, X.; Yu, Z. Two Novel Cyclic Depsipeptides Xenematides F and G from the Entomopathogenic Bacterium Xenorhabdus budapestensis. J. Antibiot. 2019, 72, 736–743. [Google Scholar] [CrossRef]
- Zhou, Q.; Dowling, A.; Heide, H.; Wöhnert, J.; Brandt, U.; Baum, J.; ffrench-Constant, R.; Bode, H.B. Xentrivalpeptides A–Q: Depsipeptide Diversification in Xenorhabdus. J. Nat. Prod. 2012, 75, 1717–1722. [Google Scholar] [CrossRef]
- Böszörményi, E.; Érsek, T.; Fodor, A.; Fodor, A.M.; Földes, L.S.; Hevesi, M.; Hogan, J.S.; Katona, Z.; Klein, M.G.; Kormány, A.; et al. Isolation and Activity of Xenorhabdus Antimicrobial Compounds against the Plant Pathogens Erwinia amylovora and Phytophthora nicotianae. J. Appl. Microbiol. 2009, 107, 746–759. [Google Scholar] [CrossRef] [PubMed]
- Reimer, D.; Cowles, K.N.; Proschak, A.; Nollmann, F.I.; Dowling, A.J.; Kaiser, M.; Constant, R.; Ffrench-Constant, R.; Goodrich-Blair, H.; Bode, H.B. Rhabdopeptides as Insect-Specific Virulence Factors from Entomopathogenic Bacteria. ChemBioChem 2013, 14, 1991–1997. [Google Scholar] [CrossRef]
- Cai, X.; Nowak, S.; Wesche, F.; Bischoff, I.; Kaiser, M.; Fürst, R.; Bode, H.B. Entomopathogenic Bacteria Use Multiple Mechanisms for Bioactive Peptide Library Design. Nat. Chem. 2017, 9, 379–386. [Google Scholar] [CrossRef]
- Zhao, L.; Kaiser, M.; Bode, H.B. Rhabdopeptide/Xenortide-like Peptides from Xenorhabdus innexi with Terminal Amines Showing Potent Antiprotozoal Activity. Org. Lett. 2018, 20, 5116–5120. [Google Scholar] [CrossRef]
- Fuchs, S.W.; Proschak, A.; Jaskolla, T.W.; Karas, M.; Bode, H.B. Structure Elucidation and Biosynthesis of Lysine-Rich Cyclic Peptides in Xenorhabdus nematophila. Org. Biomol. Chem. 2011, 9, 3130. [Google Scholar] [CrossRef] [PubMed]
- Gualtieri, M.; Aumelas, A.; Thaler, J.-O. Identification of a New Antimicrobial Lysine-Rich Cyclolipopeptide Family from Xenorhabdus nematophila. J. Antibiot. 2009, 62, 295–302. [Google Scholar] [CrossRef] [PubMed]
- Nollmann, F.I.; Dauth, C.; Mulley, G.; Kegler, C.; Kaiser, M.; Waterfield, N.R.; Bode, H.B. Insect-Specific Production of New GameXPeptides in Photorhabdus Luminescens TTO1, Widespread Natural Products in Entomopathogenic Bacteria. ChemBioChem 2015, 16, 205–208. [Google Scholar] [CrossRef]
- Bode, H.B.; Reimer, D.; Fuchs, S.W.; Kirchner, F.; Dauth, C.; Kegler, C.; Lorenzen, W.; Brachmann, A.O.; Grün, P. Determination of the Absolute Configuration of Peptide Natural Products by Using Stable Isotope Labeling and Mass Spectrometry. Chem. Eur. J. 2012, 18, 2342–2348. [Google Scholar] [CrossRef]
- Challinor, V.L.; Bode, H.B. Bioactive Natural Products from Novel Microbial Sources: Natural Products from Microbial Sources. Annu. N. Y. Acad. Sci. 2015, 1354, 82–97. [Google Scholar] [CrossRef]
- Wenski, S.L.; Cimen, H.; Berghaus, N.; Fuchs, S.W.; Hazir, S.; Bode, H.B. Fabclavine Diversity in Xenorhabdus Bacteria. Beilstein J. Org. Chem. 2020, 16, 956–965. [Google Scholar] [CrossRef]
- Pantel, L.; Florin, T.; Dobosz-Bartoszek, M.; Racine, E.; Sarciaux, M.; Serri, M.; Houard, J.; Campagne, J.-M.; de Figueiredo, R.M.; Midrier, C.; et al. Odilorhabdins, Antibacterial Agents That Cause Miscoding by Binding at a New Ribosomal Site. Mol. Cell 2018, 70, 83–94.e7. [Google Scholar] [CrossRef]
- Zhao, L.; Awori, R.M.; Kaiser, M.; Groß, J.; Opatz, T.; Bode, H.B. Structure, Biosynthesis, and Bioactivity of Photoditritide from Photorhabdus temperata Meg1. J. Nat. Prod. 2019, 82, 3499–3503. [Google Scholar] [CrossRef]
- Zhao, L.; Vo, T.D.; Kaiser, M.; Bode, H.B. Phototemtide A, a Cyclic Lipopeptide Heterologously Expressed from Photorhabdus temperata Meg1, Shows Selective Antiprotozoal Activity. ChemBioChem 2020, 21, 1288–1292. [Google Scholar] [CrossRef] [PubMed]
- Li, J.-H.; Cho, W.; Hamchand, R.; Oh, J.; Crawford, J.M. A Conserved Nonribosomal Peptide Synthetase in Xenorhabdus bovienii Produces Citrulline-Functionalized Lipopeptides. J. Nat. Prod. 2021, 84, 2692–2699. [Google Scholar] [CrossRef] [PubMed]
- Guo, S.; Zhang, S.; Fang, X.; Liu, Q.; Gao, J.; Bilal, M.; Wang, Y.; Zhang, X. Regulation of Antimicrobial Activity and Xenocoumacins Biosynthesis by pH in Xenorhabdus nematophila. Microb. Cell Fact. 2017, 16, 203. [Google Scholar] [CrossRef] [PubMed]
- McInerney, B.V.; Taylor, W.C.; Lacey, M.J.; Akhurst, R.J.; Gregson, R.P. Biologically Active Metabolites from Xenorhabdus Spp., Part 2. Benzopyran-1-One Derivatives with Gastroprotective Activity. J. Nat. Prod. 1991, 54, 785–795. [Google Scholar] [CrossRef] [PubMed]
- Park, H.; Perez, C.; Perry, E.; Crawford, J. Activating and Attenuating the Amicoumacin Antibiotics. Molecules 2016, 21, 824. [Google Scholar] [CrossRef]
- Fuchs, S.W.; Grundmann, F.; Kurz, M.; Kaiser, M.; Bode, H.B. Fabclavines: Bioactive Peptide-Polyketide-Polyamino Hybrids from Xenorhabdus. ChemBioChem 2014, 15, 512–516. [Google Scholar] [CrossRef] [PubMed]
- Brachmann, A.O.; Reimer, D.; Lorenzen, W.; Augusto Alonso, E.; Kopp, Y.; Piel, J.; Bode, H.B. Reciprocal Cross Talk between Fatty Acid and Antibiotic Biosynthesis in a Nematode Symbiont. Angew. Chem. Int. Ed. 2012, 51, 12086–12089. [Google Scholar] [CrossRef]
- Stein, M.L.; Beck, P.; Kaiser, M.; Dudler, R.; Becker, C.F.W.; Groll, M. One-Shot NMR Analysis of Microbial Secretions Identifies Highly Potent Proteasome Inhibitor. Proc. Natl. Acad. Sci. USA 2012, 109, 18367–18371. [Google Scholar] [CrossRef] [PubMed]
- Theodore, C.M.; King, J.B.; You, J.; Cichewicz, R.H. Production of Cytotoxic Glidobactins/Luminmycins by Photorhabdus asymbiotica in Liquid Media and Live Crickets. J. Nat. Prod. 2012, 75, 2007–2011. [Google Scholar] [CrossRef]
- Zhao, L.; Le Chapelain, C.; Brachmann, A.O.; Kaiser, M.; Groll, M.; Bode, H.B. Activation, Structure, Biosynthesis and Bioactivity of Glidobactin-like Proteasome Inhibitors from Photorhabdus laumondii. ChemBioChem 2021, 22, 1582–1588. [Google Scholar] [CrossRef]
- Bian, X.; Plaza, A.; Zhang, Y.; Müller, R. Luminmycins A–C, Cryptic Natural Products from Photorhabdus luminescens Identified by Heterologous Expression in Escherichia coli. J. Nat. Prod. 2012, 75, 1652–1655. [Google Scholar] [CrossRef]
- Singh, J.; Banerjee, N. Transcriptional Analysis and Functional Characterization of a Gene Pair Encoding Iron-Regulated Xenocin and Immunity Proteins of Xenorhabdus nematophila. J. Bacteriol. 2008, 190, 3877–3885. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Singh, P.; Park, D.; Forst, S.; Banerjee, N. Xenocin Export by the Flagellar Type III Pathway in Xenorhabdus nematophila. J. Bacteriol. 2013, 195, 1400–1410. [Google Scholar] [CrossRef] [PubMed]
- Morales-Soto, N.; Forst, S.A. The Xnp1 P2-Like Tail Synthesis Gene Cluster Encodes Xenorhabdicin and Is Required for Interspecies Competition. J. Bacteriol. 2011, 193, 3624–3632. [Google Scholar] [CrossRef][Green Version]
- Morales-Soto, N.; Gaudriault, S.; Ogier, J.-C.; Thappeta, K.R.V.; Forst, S. Comparative Analysis of P2-Type Remnant Prophage Loci in Xenorhabdus bovienii and Xenorhabdus nematophila Required for Xenorhabdicin Production. FEMS Microbiol. Lett. 2012, 333, 69–76. [Google Scholar] [CrossRef][Green Version]
- Thappeta, K.R.V.; Ciezki, K.; Morales-Soto, N.; Wesener, S.; Goodrich-Blair, H.; Stock, S.P.; Forst, S. R-Type Bacteriocins of Xenorhabdus bovienii Determine the Outcome of Interspecies Competition in a Natural Host Environment. Microbiology 2020, 166, 1074–1087. [Google Scholar] [CrossRef] [PubMed]
- Imai, Y.; Meyer, K.J.; Iinishi, A.; Favre-Godal, Q.; Green, R.; Manuse, S.; Caboni, M.; Mori, M.; Niles, S.; Ghiglieri, M.; et al. A New Antibiotic Selectively Kills Gram-Negative Pathogens. Nature 2019, 576, 459–464. [Google Scholar] [CrossRef] [PubMed]
- Booysen, E.; Rautenbach, M.; Stander, M.A.; Dicks, L.M.T. Profiling the Production of Antimicrobial Secondary Metabolites by Xenorhabdus khoisanae J194 Under Different Culturing Conditions. Front. Chem. 2021, 9, 626653. [Google Scholar] [CrossRef] [PubMed]
- Grady, E.N.; MacDonald, J.; Liu, L.; Richman, A.; Yuan, Z.-C. Current Knowledge and Perspectives of Paenibacillus: A Review. Microb. Cell Fact. 2016, 15, 203. [Google Scholar] [CrossRef]
- Cochrane, S.A.; Vederas, J.C. Lipopeptides from Bacillus and Paenibacillus spp.: A Gold Mine of Antibiotic Candidates: Bacillus and Paenibacillus Lipopeptides. Med. Res. Rev. 2016, 36, 4–31. [Google Scholar] [CrossRef] [PubMed]
- Choi, S.-K.; Park, S.-Y.; Kim, R.; Kim, S.-B.; Lee, C.-H.; Kim, J.F.; Park, S.-H. Identification of a Polymyxin Synthetase Gene Cluster of Paenibacillus polymyxa and Heterologous Expression of the Gene in Bacillus subtilis. J. Bacteriol. 2009, 191, 3350–3358. [Google Scholar] [CrossRef]
- Velkov, T.; Roberts, K.D.; Nation, R.L.; Thompson, P.E.; Li, J. Pharmacology of Polymyxins: New Insights into an ‘Old’ Class of Antibiotics. Future Microbiol. 2013, 8, 711–724. [Google Scholar] [CrossRef]
- Kajimura, Y.; Kaneda, M. Fusaricidin A, a New Depsipeptide Antibiotic Produced by Bacillus polymyxa KT-8 Taxonomy, Fermentation, Isolation, Structure Elucidation and Biological Activity. J. Antibiot. 1996, 49, 129–135. [Google Scholar] [CrossRef] [PubMed]
- Mülner, P.; Schwarz, E.; Dietel, K.; Herfort, S.; Jähne, J.; Lasch, P.; Cernava, T.; Berg, G.; Vater, J. Fusaricidins, Polymyxins and Volatiles Produced by Paenibacillus polymyxa Strains DSM 32871 and M1. Pathogens 2021, 10, 1485. [Google Scholar] [CrossRef]
- Menegatti, C.; Da Paixão Melo, W.G.; Carrão, D.B.; De Oliveira, A.R.M.; Do Nascimento, F.S.; Lopes, N.P.; Pupo, M.T. Paenibacillus polymyxa Associated with the Stingless Bee Melipona scutellaris Produces Antimicrobial Compounds against Entomopathogens. J. Chem. Ecol. 2018, 44, 1158–1169. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Cheng, H.; Qian, W.; Zhao, W.; Liang, C.; Liu, C.; Cui, G.; Liu, H.; Zhang, L. Comparative Genome Analysis and Mining of Secondary Metabolites of Paenibacillus polymyxa. Genes Genet. Syst. 2020, 95, 141–150. [Google Scholar] [CrossRef]
- Meng, J.; Zhong, Z.; Qian, P.-Y. Paenialvin A–D, Four Peptide Antibiotics Produced by Paenibacillus alvei DSM 29. J. Antibiot. 2018, 71, 769–777. [Google Scholar] [CrossRef] [PubMed]
- Sood, S.; Steinmetz, H.; Beims, H.; Mohr, K.I.; Stadler, M.; Djukic, M.; von der Ohe, W.; Steinert, M.; Daniel, R.; Müller, R. Paenilarvins: Iturin Family Lipopeptides from the Honey Bee Pathogen Paenibacillus larvae. ChemBioChem 2014, 15, 1947–1955. [Google Scholar] [CrossRef] [PubMed]
- Jangra, M.; Kaur, M.; Tambat, R.; Rana, R.; Maurya, S.K.; Khatri, N.; Ghafur, A.; Nandanwar, H. Tridecaptin M, a New Variant Discovered in Mud Bacterium, Shows Activity against Colistin- and Extremely Drug-Resistant Enterobacteriaceae. AntiMicrob. Agents Chemother. 2019, 63. [Google Scholar] [CrossRef] [PubMed]
- Pichard, B.; Larue, J.-P.; Thouvenot, D. Gavaserin and Saltavalin, New Peptide Antibiotics Produced by Bacillus polymyxa. FEMS Microbiol. Lett. 1995, 133, 215–218. [Google Scholar] [CrossRef] [PubMed]
- Qian, C.-D.; Wu, X.-C.; Teng, Y.; Zhao, W.-P.; Li, O.; Fang, S.-G.; Huang, Z.-H.; Gao, H.-C. Battacin (Octapeptin B5), a New Cyclic Lipopeptide Antibiotic from Paenibacillus tianmuensis Active against Multidrug-Resistant Gram-Negative Bacteria. Antimicrob. Agents Chemother. 2012, 56, 1458–1465. [Google Scholar] [CrossRef]
- Velkov, T.; Gallardo-Godoy, A.; Swarbrick, J.D.; Blaskovich, M.A.T.; Elliott, A.G.; Han, M.; Thompson, P.E.; Roberts, K.D.; Huang, J.X.; Becker, B.; et al. Structure, Function, and Biosynthetic Origin of Octapeptin Antibiotics Active against Extensively Drug-Resistant Gram-Negative Bacteria. Cell Chem. Biol. 2018, 25, 380–391.e5. [Google Scholar] [CrossRef] [PubMed]
- Qian, C.-D.; Liu, T.-Z.; Zhou, S.-L.; Ding, R.; Zhao, W.-P.; Li, O.; Wu, X.-C. Identification and Functional Analysis of Gene Cluster Involvement in Biosynthesis of the Cyclic Lipopeptide Antibiotic Pelgipeptin Produced by Paenibacillus elgii. BMC Microbiol. 2012, 12, 197. [Google Scholar] [CrossRef]
- Guo, Y.; Huang, E.; Yuan, C.; Zhang, L.; Yousef, A.E. Isolation of a Paenibacillus sp. Strain and Structural Elucidation of Its Broad-Spectrum Lipopeptide Antibiotic. Appl. Environ. Microbiol. 2012, 78, 3156–3165. [Google Scholar] [CrossRef]
- Huang, E.; Guo, Y.; Yousef, A.E. Biosynthesis of the New Broad-Spectrum Lipopeptide Antibiotic Paenibacterin in Paenibacillus thiaminolyticus OSY-SE. Res. Microbiol. 2014, 165, 243–251. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.-X.; Zhong, Z.; Zhang, W.-P.; Qian, P.-Y. Discovery of Cationic Nonribosomal Peptides as Gram-Negative Antibiotics through Global Genome Mining. Nat. Commun. 2018, 9, 3273. [Google Scholar] [CrossRef] [PubMed]
- Olishevska, S.; Nickzad, A.; Déziel, E. Bacillus and Paenibacillus Secreted Polyketides and Peptides Involved in Controlling Human and Plant Pathogens. Appl. Microbiol. Biotechnol. 2019, 103, 1189–1215. [Google Scholar] [CrossRef] [PubMed]
- He, Z.; Kisla, D.; Zhang, L.; Yuan, C.; Green-Church, K.B.; Yousef, A.E. Isolation and Identification of a Paenibacillus Polymyxa Strain That Coproduces a Novel Lantibiotic and Polymyxin. Appl. Environ. Microbiol. 2007, 73, 168–178. [Google Scholar] [CrossRef] [PubMed]
- Huang, E.; Yousef, A.E. Biosynthesis of Paenibacillin, a Lantibiotic with N-Terminal Acetylation, by Paenibacillus polymyxa. Microbiol. Res. 2015, 181, 15–21. [Google Scholar] [CrossRef] [PubMed]
- Lohans, C.T.; Huang, Z.; van Belkum, M.J.; Giroud, M.; Sit, C.S.; Steels, E.M.; Zheng, J.; Whittal, R.M.; McMullen, L.M.; Vederas, J.C. Structural Characterization of the Highly Cyclized Lantibiotic Paenicidin A via a Partial Desulfurization/Reduction Strategy. J. Am. Chem. Soc. 2012, 134, 19540–19543. [Google Scholar] [CrossRef]
- Lohans, C.T.; van Belkum, M.J.; Cochrane, S.A.; Huang, Z.; Sit, C.S.; McMullen, L.M.; Vederas, J.C. Biochemical, Structural, and Genetic Characterization of Tridecaptin A1, an Antagonist of Campylobacter jejuni. ChemBioChem 2014, 15, 243–249. [Google Scholar] [CrossRef]
- Baindara, P.; Chaudhry, V.; Mittal, G.; Liao, L.M.; Matos, C.O.; Khatri, N.; Franco, O.L.; Patil, P.B.; Korpole, S. Characterization of the Antimicrobial Peptide Penisin, a Class Ia Novel Lantibiotic from Paenibacillus sp. Strain A3. AntiMicrob. Agents Chemother. 2016, 60, 580–591. [Google Scholar] [CrossRef] [PubMed]
- Müller, S.; Garcia-Gonzalez, E.; Mainz, A.; Hertlein, G.; Heid, N.C.; Mösker, E.; van den Elst, H.; Overkleeft, H.S.; Genersch, E.; Süssmuth, R.D. Paenilamicin: Structure and Biosynthesis of a Hybrid Nonribosomal Peptide/Polyketide Antibiotic from the Bee Pathogen Paenibacillus larvae. Angew. Chem. Int. Ed. 2014, 53, 10821–10825. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Gonzalez, E.; Müller, S.; Hertlein, G.; Heid, N.; Süssmuth, R.D.; Genersch, E. Biological Effects of Paenilamicin, a Secondary Metabolite Antibiotic Produced by the Honey Bee Pathogenic Bacterium Paenibacillus larvae. Microbiol. Open 2014, 3, 642–656. [Google Scholar] [CrossRef]
- Pineda-Castellanos, M.; Rodríguez-Segura, Z.; Villalobos, F.; Hernández, L.; Lina, L.; Nuñez-Valdez, M. Pathogenicity of Isolates of Serratia Marcescens towards Larvae of the Scarab Phyllophaga blanchardi (Coleoptera). Pathogens 2015, 4, 210–228. [Google Scholar] [CrossRef] [PubMed]
- Clements, T.; Rautenbach, M.; Ndlovu, T.; Khan, S.; Khan, W. A Metabolomics and Molecular Networking Approach to Elucidate the Structures of Secondary Metabolites Produced by Serratia marcescens Strains. Front. Chem. 2021, 9, 633870. [Google Scholar] [CrossRef] [PubMed]
- Cleto, S.; Haslinger, K.; Prather, K.L.J.; Lu, T.K. Natural Combinatorial Genetics and Prolific Polyamine Production Enable Siderophore Diversification in Serratia Plymuthica. BMC Biol. 2021, 19, 46. [Google Scholar] [CrossRef]
- Nguyen, H.T.; Kim, H.-G.; Yu, N.H.; Hwang, I.M.; Kim, H.; Kim, Y.C.; Kim, J.-C. In Vitro and In Vivo Antibacterial Activity of Serratamid, a Novel Peptide–Polyketide Antibiotic Isolated from Serratia plymuthica C1, against Phytopathogenic Bacteria. J. Agric. Food Chem. 2021, 69, 5471–5480. [Google Scholar] [CrossRef]
- Maglangit, F.; Yu, Y.; Deng, H. Bacterial Pathogens: Threat or Treat (a Review on Bioactive Natural Products from Bacterial Pathogens). Nat. Prod. Rep. 2021, 38, 782–821. [Google Scholar] [CrossRef]
- Lincke, T.; Behnken, S.; Ishida, K.; Roth, M.; Hertweck, C. Closthioamide: An Unprecedented Polythioamide Antibiotic from the Strictly Anaerobic Bacterium Clostridium cellulolyticum. Angew. Chem. Int. Ed. 2010, 49, 2011–2013. [Google Scholar] [CrossRef]
- Chiriac, A.I.; Kloss, F.; Krämer, J.; Vuong, C.; Hertweck, C.; Sahl, H.-G. Mode of Action of Closthioamide: The First Member of the Polythioamide Class of Bacterial DNA Gyrase Inhibitors. J. Antimicrob. Chemother. 2015, 70, 2576–2588. [Google Scholar] [CrossRef] [PubMed]
- Dunbar, K.L.; Büttner, H.; Molloy, E.M.; Dell, M.; Kumpfmüller, J.; Hertweck, C. Genome Editing Reveals Novel Thiotemplated Assembly of Polythioamide Antibiotics in Anaerobic Bacteria. Angew. Chem. Int. Ed. 2018, 57, 14080–14084. [Google Scholar] [CrossRef] [PubMed]
- Dunbar, K.L.; Dell, M.; Gude, F.; Hertweck, C. Reconstitution of Polythioamide Antibiotic Backbone Formation Reveals Unusual Thiotemplated Assembly Strategy. Proc. Natl. Acad. Sci. USA 2020, 117, 8850–8858. [Google Scholar] [CrossRef] [PubMed]
- Dunbar, K.L.; Dell, M.; Molloy, E.M.; Kloss, F.; Hertweck, C. Reconstitution of Iterative Thioamidation in Closthioamide Biosynthesis Reveals Tailoring Strategy for Nonribosomal Peptide Backbones. Angew. Chem. Int. Ed. 2019, 58, 13014–13018. [Google Scholar] [CrossRef] [PubMed]
- Dunbar, K.L.; Dell, M.; Molloy, E.M.; Büttner, H.; Kumpfmüller, J.; Hertweck, C. An Unexpected Split-Merge Pathway in the Assembly of the Symmetric Nonribosomal Peptide Antibiotic Closthioamide. Angew. Chem. Int. Ed. 2021, 60, 4104–4109. [Google Scholar] [CrossRef] [PubMed]
- Letzel, A.-C.; Pidot, S.J.; Hertweck, C. Genome Mining for Ribosomally Synthesized and Post-Translationally Modified Peptides (RiPPs) in Anaerobic Bacteria. BMC Genom. 2014, 15, 983. [Google Scholar] [CrossRef] [PubMed]
- Behnken, S.; Hertweck, C. Anaerobic Bacteria as Producers of Antibiotics. Appl. Microbiol. Biotechnol. 2012, 96, 61–67. [Google Scholar] [CrossRef]
- Neuwirth, T.; Letzel, A.; Tank, C.; Ishida, K.; Cyrulies, M.; Schmölz, L.; Lorkowski, S.; Hertweck, C. Induced Production, Synthesis, and Immunomodulatory Action of Clostrisulfone, a Diarylsulfone from Clostridium acetobutylicum. Chem. Eur. J. 2020, 26, 15855–15858. [Google Scholar] [CrossRef]
- Pidot, S.; Ishida, K.; Cyrulies, M.; Hertweck, C. Discovery of Clostrubin, an Exceptional Polyphenolic Polyketide Antibiotic from a Strictly Anaerobic Bacterium. Angew. Chem. Int. Ed. 2014, 53, 7856–7859. [Google Scholar] [CrossRef]
- Hertweck, C.; Luzhetskyy, A.; Rebets, Y.; Bechthold, A. Type II Polyketide Synthases: Gaining a Deeper Insight into Enzymatic Teamwork. Nat. Prod. Rep. 2007, 24, 162–190. [Google Scholar] [CrossRef] [PubMed]
- Shabuer, G.; Ishida, K.; Pidot, S.J.; Roth, M.; Dahse, H.-M.; Hertweck, C. Plant Pathogenic Anaerobic Bacteria Use Aromatic Polyketides to Access Aerobic Territory. Science 2015, 350, 670–674. [Google Scholar] [CrossRef]
- Diez, J.; Martinez, J.P.; Mestres, J.; Sasse, F.; Frank, R.; Meyerhans, A. Myxobacteria: Natural Pharmaceutical Factories. Microb. Cell Fact. 2012, 11, 52. [Google Scholar] [CrossRef][Green Version]
- Landwehr, W.; Wolf, C.; Wink, J. Actinobacteria and Myxobacteria—Two of the Most Important Bacterial Resources for Novel Antibiotics. In How to Overcome the Antibiotic Crisis; Stadler, M., Dersch, P., Eds.; Current Topics in Microbiology and Immunology; Springer International Publishing: Cham, Switzerland, 2016; Volume 398, pp. 273–302. ISBN 978-3-319-49282-7. [Google Scholar]
- Gemperlein, K.; Zaburannyi, N.; Garcia, R.; La Clair, J.; Müller, R. Metabolic and Biosynthetic Diversity in Marine Myxobacteria. Mar. Drugs 2018, 16, 314. [Google Scholar] [CrossRef] [PubMed]
- Velicer, G.J.; Vos, M. Sociobiology of the Myxobacteria. Annu. Rev. Microbiol. 2009, 63, 599–623. [Google Scholar] [CrossRef] [PubMed]
- Han, K.; Li, Z.; Peng, R.; Zhu, L.; Zhou, T.; Wang, L.; Li, S.; Zhang, X.; Hu, W.; Wu, Z.; et al. Extraordinary Expansion of a Sorangium cellulosum Genome from an Alkaline Milieu. Sci. Rep. 2013, 3, 2101. [Google Scholar] [CrossRef]
- Schneiker, S.; Perlova, O.; Kaiser, O.; Gerth, K.; Alici, A.; Altmeyer, M.O.; Bartels, D.; Bekel, T.; Beyer, S.; Bode, E.; et al. Complete Genome Sequence of the Myxobacterium Sorangium cellulosum. Nat. Biotechnol. 2007, 25, 1281–1289. [Google Scholar] [CrossRef]
- Weissman, K.J.; Müller, R. A Brief Tour of Myxobacterial Secondary Metabolism. Bioorganic Med. Chem. 2009, 17, 2121–2136. [Google Scholar] [CrossRef] [PubMed]
- Bode, H.B.; Irschik, H.; Wenzel, S.C.; Reichenbach, H.; Müller, R.; Höfle, G. The Leupyrrins: A Structurally Unique Family of Secondary Metabolites from the Myxobacterium Sorangium cellulosum. J. Nat. Prod. 2003, 66, 1203–1206. [Google Scholar] [CrossRef]
- Bode, H.B.; Wenzel, S.C.; Irschik, H.; Höfle, G.; Müller, R. Unusual Biosynthesis of Leupyrrins in the Myxobacterium Sorangium cellulosum. Angew. Chem. Int. Ed. 2004, 43, 4163–4167. [Google Scholar] [CrossRef]
- Li, Y.; Zhuo, L.; Li, X.; Zhu, Y.; Wu, S.; Shen, T.; Hu, W.; Li, Y.; Wu, C. Myxadazoles, Myxobacterium-Derived Isoxazole–Benzimidazole Hybrids with Cardiovascular Activities. Angew. Chem. Int. Ed. 2021, 60, 21679–21684. [Google Scholar] [CrossRef]
- Wenzel, S.C.; Müller, R. Myxobacterial Natural Product Assembly Lines: Fascinating Examples of Curious Biochemistry. Nat. Prod. Rep. 2007, 24, 1211. [Google Scholar] [CrossRef]
- Popoff, A.; Hug, J.J.; Walesch, S.; Garcia, R.; Keller, L.; Müller, R. Structure and Biosynthesis of Myxofacyclines: Unique Myxobacterial Polyketides Featuring Varing and Rare Heterocycles. Chem. Eur. J. 2021, 27, 16654–16661. [Google Scholar] [CrossRef]
- Gerth, K.; Bedorf, N.; Irschik, H.; Höfle, G.; Reichenbach, H. The Soraphens: A Family of Novel Antifungal Compounds from Sorangium cellulosum (Myxobacteria). I. Soraphen A1.ALPHA.: Fermentation, Isolation, Biological Properties. J. Antibiot. 1994, 47, 23–31. [Google Scholar] [CrossRef]
- Ligon, J.; Hill, S.; Beck, J.; Zirkle, R.; Molnár, I.; Zawodny, J.; Money, S.; Schupp, T. Characterization of the Biosynthetic Gene Cluster for the Antifungal Polyketide Soraphen A from Sorangium cellulosum So Ce26. Gene 2002, 285, 257–267. [Google Scholar] [CrossRef]
- Bollag, D.M.; McQueney, P.A.; Zhu, J.; Hensens, O.; Koupal, L.; Liesch, J.; Goetz, M.; Lazarides, E.; Woods, C.M. Epothilones, a New Class of Microtubule-Stabilizing Agents with a Taxol-like Mechanism of Action. Cancer Res. 1995, 55, 2325–2333. [Google Scholar] [PubMed]
- Chen, H.; O’Connor, S.; Cane, D.E.; Walsh, C.T. Epothilone Biosynthesis: Assembly of the Methylthiazolylcarboxy Starter Unit on the EpoB Subunit. Chem. Biol. 2001, 8, 899–912. [Google Scholar] [CrossRef][Green Version]
- Xiao, Y.; Wei, X.; Ebright, R.; Wall, D. Antibiotic Production by Myxobacteria Plays a Role in Predation. J. Bacteriol. 2011, 193, 4626–4633. [Google Scholar] [CrossRef]
- Rosenberg, E.; Dworkin, M. Autocides and a Paracide, Antibiotic TA, Produced By Myxococcus xanthus. J. Ind. Microbiol. Biotechnol. 1996, 17, 424–431. [Google Scholar] [CrossRef]
- Findlay, B.L. The Chemical Ecology of Predatory Soil Bacteria. ACS Chem. Biol. 2016, 11, 1502–1510. [Google Scholar] [CrossRef]
- Kegler, C.; Gerth, K.; Müller, R. Establishment of a Real-Time PCR Protocol for Expression Studies of Secondary Metabolite Biosynthetic Gene Clusters in the G/C-Rich Myxobacterium Sorangium cellulosum So Ce56. J. Biotechnol. 2006, 121, 201–212. [Google Scholar] [CrossRef]
- Bibb, M.J. Regulation of Secondary Metabolism in Streptomycetes. Curr. Opin. Microbiol. 2005, 8, 208–215. [Google Scholar] [CrossRef] [PubMed]
- Panter, F.; Bader, C.D.; Müller, R. The Sandarazols Are Cryptic and Structurally Unique Plasmid-Encoded Toxins from a Rare Myxobacterium. Angew. Chem. Int. Ed. 2021, 60, 8081–8088. [Google Scholar] [CrossRef]
- Wenzel, S.C.; Gross, F.; Zhang, Y.; Fu, J.; Stewart, A.F.; Müller, R. Heterologous Expression of a Myxobacterial Natural Products Assembly Line in Pseudomonads via Red/ET Recombineering. Chem. Biol. 2005, 12, 349–356. [Google Scholar] [CrossRef]
- Schäberle, T.F.; Lohr, F.; Schmitz, A.; König, G.M. Antibiotics from Myxobacteria. Nat. Prod. Rep. 2014, 31, 953. [Google Scholar] [CrossRef] [PubMed]
- Herrmann, J.; Fayad, A.A.; Müller, R. Natural Products from Myxobacteria: Novel Metabolites and Bioactivities. Nat. Prod. Rep. 2017, 34, 135–160. [Google Scholar] [CrossRef]
- Gorges, J.; Panter, F.; Kjaerulff, L.; Hoffmann, T.; Kazmaier, U.; Müller, R. Structure, Total Synthesis, and Biosynthesis of Chloromyxamides: Myxobacterial Tetrapeptides Featuring an Uncommon 6-Chloromethyl-5-Methoxypipecolic Acid Building Block. Angew. Chem. Int. Ed. 2018, 57, 14270–14275. [Google Scholar] [CrossRef] [PubMed]
- Hug, J.J.; Dastbaz, J.; Adam, S.; Revermann, O.; Koehnke, J.; Krug, D.; Müller, R. Biosynthesis of Cittilins, Unusual Ribosomally Synthesized and Post-Translationally Modified Peptides from Myxococcus xanthus. ACS Chem. Biol. 2020, 15, 2221–2231. [Google Scholar] [CrossRef]
- Demay, J.; Bernard, C.; Reinhardt, A.; Marie, B. Natural Products from Cyanobacteria: Focus on Beneficial Activities. Mar. Drugs 2019, 17, 320. [Google Scholar] [CrossRef] [PubMed]
- Dhakal, D.; Chen, M.; Luesch, H.; Ding, Y. Heterologous Production of Cyanobacterial Compounds. J. Ind. Microbiol. Biotechnol. 2021, 48, kuab003. [Google Scholar] [CrossRef]
- Metcalf, J.S.; Codd, G.A. Co-Occurrence of Cyanobacteria and Cyanotoxins with Other Environmental Health Hazards: Impacts and Implications. Toxins 2020, 12, 629. [Google Scholar] [CrossRef] [PubMed]
- Banerjee, S.; Maity, S.; Guchhait, R.; Chatterjee, A.; Biswas, C.; Adhikari, M.; Pramanick, K. Toxic Effects of Cyanotoxins in Teleost Fish: A Comprehensive Review. Aquat. Toxicol. 2021, 240, 105971. [Google Scholar] [CrossRef]
- Gorham, T.; Dowling Root, E.; Jia, Y.; Shum, C.K.; Lee, J. Relationship between Cyanobacterial Bloom Impacted Drinking Water Sources and Hepatocellular Carcinoma Incidence Rates. Harmful Algae 2020, 95, 101801. [Google Scholar] [CrossRef]
- Buratti, F.M.; Manganelli, M.; Vichi, S.; Stefanelli, M.; Scardala, S.; Testai, E.; Funari, E. Cyanotoxins: Producing Organisms, Occurrence, Toxicity, Mechanism of Action and Human Health Toxicological Risk Evaluation. Arch. Toxicol. 2017, 91, 1049–1130. [Google Scholar] [CrossRef] [PubMed]
- Wong, J.L.; Oesterlin, R.; Rapoport, H. Structure of Saxitoxin. J. Am. Chem. Soc. 1971, 93, 7344–7345. [Google Scholar] [CrossRef] [PubMed]
- Schantz, E.J.; Ghazarossian, V.E.; Schnoes, H.K.; Strong, F.M.; Springer, J.P.; Pezzanite, J.O.; Clardy, J. Structure of Saxitoxin. J. Am. Chem. Soc. 1975, 97, 1238–1239. [Google Scholar] [CrossRef]
- Dittmann, E.; Gugger, M.; Sivonen, K.; Fewer, D.P. Natural Product Biosynthetic Diversity and Comparative Genomics of the Cyanobacteria. Trends Microbiol. 2015, 23, 642–652. [Google Scholar] [CrossRef]
- Jones, M.R.; Pinto, E.; Torres, M.A.; Dörr, F.; Mazur-Marzec, H.; Szubert, K.; Tartaglione, L.; Dell’Aversano, C.; Miles, C.O.; Beach, D.G.; et al. CyanoMetDB, a Comprehensive Public Database of Secondary Metabolites from Cyanobacteria. Water Res. 2021, 196, 117017. [Google Scholar] [CrossRef]
- Pereira, A.R.; Etzbach, L.; Engene, N.; Müller, R.; Gerwick, W.H. Molluscicidal Metabolites from an Assemblage of Palmyra Atoll Cyanobacteria. J. Nat. Prod. 2011, 74, 1175–1181. [Google Scholar] [CrossRef]
- Soares, A.R.; Engene, N.; Gunasekera, S.P.; Sneed, J.M.; Paul, V.J. Carriebowlinol, an Antimicrobial Tetrahydroquinolinol from an Assemblage of Marine Cyanobacteria Containing a Novel Taxon. J. Nat. Prod. 2015, 78, 534–538. [Google Scholar] [CrossRef]
- Berube, P.M.; Biller, S.J.; Hackl, T.; Hogle, S.L.; Satinsky, B.M.; Becker, J.W.; Braakman, R.; Collins, S.B.; Kelly, L.; Berta-Thompson, J.; et al. Single Cell Genomes of Prochlorococcus, Synechococcus, and Sympatric Microbes from Diverse Marine Environments. Sci. Data 2018, 5, 180154. [Google Scholar] [CrossRef] [PubMed]
- Nakayama, T.; Nomura, M.; Takano, Y.; Tanifuji, G.; Shiba, K.; Inaba, K.; Inagaki, Y.; Kawata, M. Single-Cell Genomics Unveiled a Cryptic Cyanobacterial Lineage with a Worldwide Distribution Hidden by a Dinoflagellate Host. Proc. Natl. Acad. Sci. USA 2019, 116, 15973–15978. [Google Scholar] [CrossRef]
- Alvarenga, D.O.; Fiore, M.F.; Varani, A.M. A Metagenomic Approach to Cyanobacterial Genomics. Front. Microbiol. 2017, 8, 809. [Google Scholar] [CrossRef]
- Griese, M.; Lange, C.; Soppa, J. Ploidy in Cyanobacteria. FEMS Microbiol. Lett. 2011, 323, 124–131. [Google Scholar] [CrossRef]
- Kelly, C.L.; Taylor, G.M.; Hitchcock, A.; Torres-Méndez, A.; Heap, J.T. A Rhamnose-Inducible System for Precise and Temporal Control of Gene Expression in Cyanobacteria. ACS Synth. Biol. 2018, 7, 1056–1066. [Google Scholar] [CrossRef] [PubMed]
- Long, P.F.; Dunlap, W.C.; Battershill, C.N.; Jaspars, M. Shotgun Cloning and Heterologous Expression of the Patellamide Gene Cluster as a Strategy to Achieving Sustained Metabolite Production. ChemBioChem 2005, 6, 1760–1765. [Google Scholar] [CrossRef]
- Watanabe, K.; Hotta, K.; Praseuth, A.P.; Koketsu, K.; Migita, A.; Boddy, C.N.; Wang, C.C.C.; Oguri, H.; Oikawa, H. Total Biosynthesis of Antitumor Nonribosomal Peptides in Escherichia Coli. Nat. Chem. Biol. 2006, 2, 423–428. [Google Scholar] [CrossRef]
- Weiz, A.R.; Ishida, K.; Makower, K.; Ziemert, N.; Hertweck, C.; Dittmann, E. Leader Peptide and a Membrane Protein Scaffold Guide the Biosynthesis of the Tricyclic Peptide Microviridin. Chem. Biol. 2011, 18, 1413–1421. [Google Scholar] [CrossRef]
- Włodarczyk, A.; Selão, T.T.; Norling, B.; Nixon, P.J. Newly Discovered Synechococcus sp. PCC 11901 Is a Robust Cyanobacterial Strain for High Biomass Production. Commun. Biol. 2020, 3, 215. [Google Scholar] [CrossRef]
- Mueller, T.J.; Ungerer, J.L.; Pakrasi, H.B.; Maranas, C.D. Identifying the Metabolic Differences of a Fast-Growth Phenotype in Synechococcus UTEX 2973. Sci. Rep. 2017, 7, 41569. [Google Scholar] [CrossRef]
- Taton, A.; Ecker, A.; Diaz, B.; Moss, N.A.; Anderson, B.; Reher, R.; Leão, T.F.; Simkovsky, R.; Dorrestein, P.C.; Gerwick, L.; et al. Heterologous Expression of Cryptomaldamide in a Cyanobacterial Host. ACS Synth. Biol. 2020, 9, 3364–3376. [Google Scholar] [CrossRef]
- Videau, P.; Wells, K.N.; Singh, A.J.; Eiting, J.; Proteau, P.J.; Philmus, B. Expanding the Natural Products Heterologous Expression Repertoire in the Model Cyanobacterium Anabaena sp. Strain PCC 7120: Production of Pendolmycin and Teleocidin B-4. ACS Synth. Biol. 2020, 9, 63–75. [Google Scholar] [CrossRef]
- Kim, W.J.; Lee, S.-M.; Um, Y.; Sim, S.J.; Woo, H.M. Development of SyneBrick Vectors As a Synthetic Biology Platform for Gene Expression in Synechococcus elongatus PCC 7942. Front. Plant. Sci. 2017, 8. [Google Scholar] [CrossRef]
- Vasudevan, R.; Gale, G.A.R.; Schiavon, A.A.; Puzorjov, A.; Malin, J.; Gillespie, M.D.; Vavitsas, K.; Zulkower, V.; Wang, B.; Howe, C.J.; et al. CyanoGate: A Modular Cloning Suite for Engineering Cyanobacteria Based on the Plant MoClo Syntax. Plant. Physiol. 2019, 180, 39–55. [Google Scholar] [CrossRef]
- Behler, J.; Vijay, D.; Hess, W.R.; Akhtar, M.K. CRISPR-Based Technologies for Metabolic Engineering in Cyanobacteria. Trends Biotechnol. 2018, 36, 996–1010. [Google Scholar] [CrossRef] [PubMed]
- Santos-Merino, M.; Singh, A.K.; Ducat, D.C. New Applications of Synthetic Biology Tools for Cyanobacterial Metabolic Engineering. Front. Bioeng. Biotechnol. 2019, 7, 33. [Google Scholar] [CrossRef]
- Shih, P.M.; Wu, D.; Latifi, A.; Axen, S.D.; Fewer, D.P.; Talla, E.; Calteau, A.; Cai, F.; Tandeau de Marsac, N.; Rippka, R.; et al. Improving the Coverage of the Cyanobacterial Phylum Using Diversity-Driven Genome Sequencing. Proc. Natl. Acad. Sci. USA 2013, 110, 1053–1058. [Google Scholar] [CrossRef] [PubMed]
- Leao, T.; Castelão, G.; Korobeynikov, A.; Monroe, E.A.; Podell, S.; Glukhov, E.; Allen, E.E.; Gerwick, W.H.; Gerwick, L. Comparative Genomics Uncovers the Prolific and Distinctive Metabolic Potential of the Cyanobacterial Genus Moorea. Proc. Natl. Acad. Sci. USA 2017, 114, 3198–3203. [Google Scholar] [CrossRef] [PubMed]
- Gu, W.; Dong, S.-H.; Sarkar, S.; Nair, S.K.; Schmidt, E.W. The Biochemistry and Structural Biology of Cyanobactin Pathways: Enabling Combinatorial Biosynthesis. In Methods in Enzymology; Elsevier: Amsterdam, The Netherlands, 2018; Volume 604, pp. 113–163. ISBN 978-0-12-813959-2. [Google Scholar]
- Tillett, D.; Dittmann, E.; Erhard, M.; von Döhren, H.; Börner, T.; Neilan, B.A. Structural Organization of Microcystin Biosynthesis in Microcystis aeruginosa PCC7806: An Integrated Peptide–Polyketide Synthetase System. Chem. Biol. 2000, 7, 753–764. [Google Scholar] [CrossRef]
- Moffitt, M.C.; Neilan, B.A. Characterization of the Nodularin Synthetase Gene Cluster and Proposed Theory of the Evolution of Cyanobacterial Hepatotoxins. Appl. Environ. Microbiol. 2004, 70, 6353–6362. [Google Scholar] [CrossRef]
- Ballot, A.; Swe, T.; Mjelde, M.; Cerasino, L.; Hostyeva, V.; Miles, C.O. Cylindrospermopsin- and Deoxycylindrospermopsin-Producing Raphidiopsis aaciborskii and Microcystin-Producing Microcystis spp. in Meiktila Lake, Myanmar. Toxins 2020, 12, 232. [Google Scholar] [CrossRef] [PubMed]
- Mihali, T.K.; Kellmann, R.; Muenchhoff, J.; Barrow, K.D.; Neilan, B.A. Characterization of the Gene Cluster Responsible for Cylindrospermopsin Biosynthesis. Appl. Environ. Microbiol. 2008, 74, 716–722. [Google Scholar] [CrossRef]
- Chang, Z.; Flatt, P.; Gerwick, W.H.; Nguyen, V.-A.; Willis, C.L.; Sherman, D.H. The Barbamide Biosynthetic Gene Cluster: A Novel Marine Cyanobacterial System of Mixed Polyketide Synthase (PKS)-Non-Ribosomal Peptide Synthetase (NRPS) Origin Involving an Unusual Trichloroleucyl Starter Unit. Gene 2002, 296, 235–247. [Google Scholar] [CrossRef]
- Hoffmann, D.; Hevel, J.M.; Moore, R.E.; Moore, B.S. Sequence Analysis and Biochemical Characterization of the Nostopeptolide A Biosynthetic Gene Cluster from Nostoc sp. GSV224. Gene 2003, 311, 171–180. [Google Scholar] [CrossRef]
- Lodin-Friedman, A.; Carmeli, S. Microginins from a Microcystis sp. Bloom Material Collected from the Kishon Reservoir, Israel. Mar. Drugs 2018, 16, 78. [Google Scholar] [CrossRef]
- Zervou, S.-K.; Gkelis, S.; Kaloudis, T.; Hiskia, A.; Mazur-Marzec, H. New Microginins from Cyanobacteria of Greek Freshwaters. Chemosphere 2020, 248, 125961. [Google Scholar] [CrossRef]
- Bornancin, L.; Alonso, E.; Alvariño, R.; Inguimbert, N.; Bonnard, I.; Botana, L.M.; Banaigs, B. Structure and Biological Evaluation of New Cyclic and Acyclic Laxaphycin-A Type Peptides. Bioorganic Med. Chem. 2019, 27, 1966–1980. [Google Scholar] [CrossRef]
- Heinilä, L.M.P.; Fewer, D.P.; Jokela, J.K.; Wahlsten, M.; Jortikka, A.; Sivonen, K. Shared PKS Module in Biosynthesis of Synergistic Laxaphycins. Front. Microbiol. 2020, 11, 578878. [Google Scholar] [CrossRef]
- Kinnel, R.B.; Esquenazi, E.; Leao, T.; Moss, N.; Mevers, E.; Pereira, A.R.; Monroe, E.A.; Korobeynikov, A.; Murray, T.F.; Sherman, D.; et al. A Maldiisotopic Approach to Discover Natural Products: Cryptomaldamide, a Hybrid Tripeptide from the Marine Cyanobacterium Moorea producens. J. Nat. Prod. 2017, 80, 1514–1521. [Google Scholar] [CrossRef]
- Becker, J.E.; Moore, R.E.; Moore, B.S. Cloning, Sequencing, and Biochemical Characterization of the Nostocyclopeptide Biosynthetic Gene Cluster: Molecular Basis for Imine Macrocyclization. Gene 2004, 325, 35–42. [Google Scholar] [CrossRef]
- Fidor, A.; Grabski, M.; Gawor, J.; Gromadka, R.; Węgrzyn, G.; Mazur-Marzec, H. Nostoc Edaphicum CCNP1411 from the Baltic Sea—A New Producer of Nostocyclopeptides. Mar. Drugs 2020, 18, 442. [Google Scholar] [CrossRef]
- Pancrace, C.; Jokela, J.; Sassoon, N.; Ganneau, C.; Desnos-Ollivier, M.; Wahlsten, M.; Humisto, A.; Calteau, A.; Bay, S.; Fewer, D.P.; et al. Rearranged Biosynthetic Gene Cluster and Synthesis of Hassallidin E in Planktothrix serta PCC 8927. ACS Chem. Biol. 2017, 12, 1796–1804. [Google Scholar] [CrossRef]
- Vestola, J.; Shishido, T.K.; Jokela, J.; Fewer, D.P.; Aitio, O.; Permi, P.; Wahlsten, M.; Wang, H.; Rouhiainen, L.; Sivonen, K. Hassallidins, Antifungal Glycolipopeptides, Are Widespread among Cyanobacteria and Are the End-Product of a Nonribosomal Pathway. Proc. Natl. Acad. Sci. USA 2014, 111, E1909–E1917. [Google Scholar] [CrossRef]
- Rantala-Ylinen, A.; Känä, S.; Wang, H.; Rouhiainen, L.; Wahlsten, M.; Rizzi, E.; Berg, K.; Gugger, M.; Sivonen, K. Anatoxin-a Synthetase Gene Cluster of the Cyanobacterium Anabaena sp. Strain 37 and Molecular Methods To Detect Potential Producers. Appl. Environ. Microbiol. 2011, 77, 7271–7278. [Google Scholar] [CrossRef]
- Kellmann, R.; Mihali, T.K.; Jeon, Y.J.; Pickford, R.; Pomati, F.; Neilan, B.A. Biosynthetic Intermediate Analysis and Functional Homology Reveal a Saxitoxin Gene Cluster in Cyanobacteria. Appl. Environ. Microbiol. 2008, 74, 4044–4053. [Google Scholar] [CrossRef] [PubMed]
- Minowa, T.; Cho, Y.; Oshima, Y.; Konoki, K.; Yotsu-Yamashita, M. Identification of a Novel Saxitoxin Analogue, 12β-Deoxygonyautoxin 3, in the Cyanobacterium, Anabaena circinalis (TA04). Toxins 2019, 11, 539. [Google Scholar] [CrossRef] [PubMed]
- D’Agostino, P.M.; Boundy, M.J.; Harwood, T.D.; Carmichael, W.W.; Neilan, B.A.; Wood, S.A. Re-Evaluation of Paralytic Shellfish Toxin Profiles in Cyanobacteria Using Hydrophilic Interaction Liquid Chromatography-Tandem Mass Spectrometry. Toxicon 2019, 158, 1–7. [Google Scholar] [CrossRef]
- Tao, Y.; Li, P.; Zhang, D.; Glukhov, E.; Gerwick, L.; Zhang, C.; Murray, T.F.; Gerwick, W.H. Samholides, Swinholide-Related Metabolites from a Marine Cyanobacterium Cf. Phormidium sp. J. Org. Chem. 2018, 83, 3034–3046. [Google Scholar] [CrossRef] [PubMed]
- Andrianasolo, E.H.; Gross, H.; Goeger, D.; Musafija-Girt, M.; McPhail, K.; Leal, R.M.; Mooberry, S.L.; Gerwick, W.H. Isolation of Swinholide A and Related Glycosylated Derivatives from Two Field Collections of Marine Cyanobacteria. Org. Lett. 2005, 7, 1375–1378. [Google Scholar] [CrossRef] [PubMed]
- Humisto, A.; Jokela, J.; Liu, L.; Wahlsten, M.; Wang, H.; Permi, P.; Machado, J.P.; Antunes, A.; Fewer, D.P.; Sivonen, K. The Swinholide Biosynthesis Gene Cluster from a Terrestrial Cyanobacterium, Nostoc sp. Strain UHCC 0450. Appl. Environ. Microbiol. 2018, 84. [Google Scholar] [CrossRef]
- Leikoski, N.; Liu, L.; Jokela, J.; Wahlsten, M.; Gugger, M.; Calteau, A.; Permi, P.; Kerfeld, C.A.; Sivonen, K.; Fewer, D.P. Genome Mining Expands the Chemical Diversity of the Cyanobactin Family to Include Highly Modified Linear Peptides. Chem. Biol. 2013, 20, 1033–1043. [Google Scholar] [CrossRef]
- Cegłowska, M.; Szubert, K.; Wieczerzak, E.; Kosakowska, A.; Mazur-Marzec, H. Eighteen New Aeruginosamide Variants Produced by the Baltic Cyanobacterium Limnoraphis CCNP1324. Mar. Drugs 2020, 18, 446. [Google Scholar] [CrossRef]
- Pearson, L.A.; Crosbie, N.D.; Neilan, B.A. Distribution and Conservation of Known Secondary Metabolite Biosynthesis Gene Clusters in the Genomes of Geographically Diverse Microcystis aeruginosa Strains. Mar. Freshw. Res. 2020, 71, 701. [Google Scholar] [CrossRef]
- Ziemert, N.; Ishida, K.; Liaimer, A.; Hertweck, C.; Dittmann, E. Ribosomal Synthesis of Tricyclic Depsipeptides in Bloom-Forming Cyanobacteria. Angew. Chem. Int. Ed. 2008, 47, 7756–7759. [Google Scholar] [CrossRef]
- Do Amaral, S.C.; Monteiro, P.R.; Neto, J.d.S.P.; Serra, G.M.; Gonçalves, E.C.; Xavier, L.P.; Santos, A.V. Current Knowledge on Microviridin from Cyanobacteria. Mar. Drugs 2021, 19, 17. [Google Scholar] [CrossRef]
- Cubillos-Ruiz, A.; Berta-Thompson, J.W.; Becker, J.W.; van der Donk, W.A.; Chisholm, S.W. Evolutionary Radiation of Lanthipeptides in Marine Cyanobacteria. Proc. Natl. Acad. Sci. USA 2017, 114, E5424–E5433. [Google Scholar] [CrossRef]
- Li, B.; Sher, D.; Kelly, L.; Shi, Y.; Huang, K.; Knerr, P.J.; Joewono, I.; Rusch, D.; Chisholm, S.W.; van der Donk, W.A. Catalytic Promiscuity in the Biosynthesis of Cyclic Peptide Secondary Metabolites in Planktonic Marine Cyanobacteria. Proc. Natl. Acad. Sci. USA 2010, 107, 10430–10435. [Google Scholar] [CrossRef]
- Sieber, S.; Grendelmeier, S.M.; Harris, L.A.; Mitchell, D.A.; Gademann, K. Microviridin 1777: A Toxic Chymotrypsin Inhibitor Discovered by a Metabologenomic Approach. J. Nat. Prod. 2020, 83, 438–446. [Google Scholar] [CrossRef]
- Bösch, N.M.; Borsa, M.; Greczmiel, U.; Morinaka, B.I.; Gugger, M.; Oxenius, A.; Vagstad, A.L.; Piel, J. Landornamides: Antiviral Ornithine-Containing Ribosomal Peptides Discovered through Genome Mining. Angew. Chem. 2020, 132, 11861–11866. [Google Scholar] [CrossRef]
- Morinaka, B.I.; Lakis, E.; Verest, M.; Helf, M.J.; Scalvenzi, T.; Vagstad, A.L.; Sims, J.; Sunagawa, S.; Gugger, M.; Piel, J. Natural Noncanonical Protein Splicing Yields Products with Diverse β-Amino Acid Residues. Science 2018, 359, 779–782. [Google Scholar] [CrossRef]
- Eyles, T.H.; Vior, N.M.; Lacret, R.; Truman, A.W. Understanding Thioamitide Biosynthesis Using Pathway Engineering and Untargeted Metabolomics. Chem. Sci. 2021, 12, 7138–7150. [Google Scholar] [CrossRef]
- Kampa, A.; Gagunashvili, A.N.; Gulder, T.A.M.; Morinaka, B.I.; Daolio, C.; Godejohann, M.; Miao, V.P.W.; Piel, J.; Andresson, O.S. Metagenomic Natural Product Discovery in Lichen Provides Evidence for a Family of Biosynthetic Pathways in Diverse Symbioses. Proc. Natl. Acad. Sci. USA 2013, 110, E3129–E3137. [Google Scholar] [CrossRef]
- Shishido, T.K.; Wahlsten, M.; Laine, P.; Rikkinen, J.; Lundell, T.; Auvinen, P. Microbial Communities of Cladonia lichens and Their Biosynthetic Gene Clusters Potentially Encoding Natural Products. Microorganisms 2021, 9, 1347. [Google Scholar] [CrossRef] [PubMed]
- Bertrand, R.L.; Abdel-Hameed, M.; Sorensen, J.L. Lichen Biosynthetic Gene Clusters. Part I. Genome Sequencing Reveals a Rich Biosynthetic Potential. J. Nat. Prod. 2018, 81, 723–731. [Google Scholar] [CrossRef] [PubMed]
- Calcott, M.J.; Ackerley, D.F.; Knight, A.; Keyzers, R.A.; Owen, J.G. Secondary Metabolism in the Lichen Symbiosis. Chem. Soc. Rev. 2018, 47, 1730–1760. [Google Scholar] [CrossRef]
- Spribille, T.; Tuovinen, V.; Resl, P.; Vanderpool, D.; Wolinski, H.; Aime, M.C.; Schneider, K.; Stabentheiner, E.; Toome-Heller, M.; Thor, G.; et al. Basidiomycete Yeasts in the Cortex of Ascomycete Macrolichens. Science 2016, 353, 488–492. [Google Scholar] [CrossRef] [PubMed]
- Grimm, M.; Grube, M.; Schiefelbein, U.; Zühlke, D.; Bernhardt, J.; Riedel, K. The Lichens’ Microbiota, Still a Mystery? Front. Microbiol. 2021, 12, 623839. [Google Scholar] [CrossRef] [PubMed]
- Sayed, A.M.; Hassan, M.H.A.; Alhadrami, H.A.; Hassan, H.M.; Goodfellow, M.; Rateb, M.E. Extreme Environments: Microbiology Leading to Specialized Metabolites. J. Appl. Microbiol. 2020, 128, 630–657. [Google Scholar] [CrossRef] [PubMed]
- Hei, Y.; Zhang, H.; Tan, N.; Zhou, Y.; Wei, X.; Hu, C.; Liu, Y.; Wang, L.; Qi, J.; Gao, J.-M. Antimicrobial Activity and Biosynthetic Potential of Cultivable Actinomycetes Associated with Lichen Symbiosis from Qinghai-Tibet Plateau. Microbiol. Res. 2021, 244, 126652. [Google Scholar] [CrossRef]
- Parrot, D.; Legrave, N.; Delmail, D.; Grube, M.; Suzuki, M.; Tomasi, S. Review—Lichen-Associated Bacteria as a Hot Spot of Chemodiversity: Focus on Uncialamycin, a Promising Compound for Future Medicinal Applications. Planta Med. 2016, 82, 1143–1152. [Google Scholar] [CrossRef] [PubMed]
- Golakoti, T.; Yoshida, W.Y.; Chaganty, S.; Moore, R.E. Isolation and Structure Determination of Nostocyclopeptides A1 and A2 from the Terrestrial Cyanobacterium Nostoc sp. ATCC53789. J. Nat. Prod. 2001, 64, 54–59. [Google Scholar] [CrossRef]
- Kaasalainen, U.; Fewer, D.P.; Jokela, J.; Wahlsten, M.; Sivonen, K.; Rikkinen, J. Cyanobacteria Produce a High Variety of Hepatotoxic Peptides in Lichen Symbiosis. Proc. Natl. Acad. Sci. USA 2012, 109, 5886–5891. [Google Scholar] [CrossRef] [PubMed]
- Fernández-Moriano, C.; Gómez-Serranillos, M.P.; Crespo, A. Antioxidant Potential of Lichen Species and Their Secondary Metabolites. A Systematic Review. Pharm. Biol. 2016, 54, 1–17. [Google Scholar] [CrossRef]
- Vaez, M.; Javad Davarpanah, S. New Insights into the Biological Activity of Lichens: Bioavailable Secondary Metabolites of Umbilicaria decussata as Potential Anticoagulants. Chem. Biodivers. 2021, 18. [Google Scholar] [CrossRef]
- Ingelfinger, R.; Henke, M.; Roser, L.; Ulshöfer, T.; Calchera, A.; Singh, G.; Parnham, M.J.; Geisslinger, G.; Fürst, R.; Schmitt, I.; et al. Unraveling the Pharmacological Potential of Lichen Extracts in the Context of Cancer and Inflammation With a Broad Screening Approach. Front. Pharmacol. 2020, 11, 1322. [Google Scholar] [CrossRef]
- Zhang, W.; Xu, L.; Yang, L.; Huang, Y.; Li, S.; Shen, Y. Phomopsidone A, a Novel Depsidone Metabolite from the Mangrove Endophytic Fungus Phomopsis sp. A123. Fitoterapia 2014, 96, 146–151. [Google Scholar] [CrossRef]
- Araújo, A.A.S.; de Melo, M.G.D.; Rabelo, T.K.; Nunes, P.S.; Santos, S.L.; Serafini, M.R.; Santos, M.R.V.; Quintans-Júnior, L.J.; Gelain, D.P. Review of the Biological Properties and Toxicity of Usnic Acid. Nat. Prod. Res. 2015, 29, 2167–2180. [Google Scholar] [CrossRef]
- Habrant, D.; Poigny, S.; Ségur-Derai, M.; Brunel, Y.; Heurtaux, B.; Le Gall, T.; Strehle, A.; Saladin, R.; Meunier, S.; Mioskowski, C.; et al. Evaluation of Antioxidant Properties of Monoaromatic Derivatives of Pulvinic Acids. J. Med. Chem. 2009, 52, 2454–2464. [Google Scholar] [CrossRef] [PubMed]
- Abdel-Hameed, M.; Bertrand, R.L.; Piercey-Normore, M.D.; Sorensen, J.L. Putative Identification of the Usnic Acid Biosynthetic Gene Cluster by de Novo Whole-Genome Sequencing of a Lichen-Forming Fungus. Fungal Biol. 2016, 120, 306–316. [Google Scholar] [CrossRef] [PubMed]
- Kim, W.; Liu, R.; Woo, S.; Kang, K.B.; Park, H.; Yu, Y.H.; Ha, H.-H.; Oh, S.-Y.; Yang, J.H.; Kim, H.; et al. Linking a Gene Cluster to Atranorin, a Major Cortical Substance of Lichens, through Genetic Dereplication and Heterologous Expression. mBio 2021, 12. [Google Scholar] [CrossRef] [PubMed]
- Singh, G.; Armaleo, D.; Dal Grande, F.; Schmitt, I. Depside and Depsidone Synthesis in Lichenized Fungi Comes into Focus through a Genome-Wide Comparison of the Olivetoric Acid and Physodic Acid Chemotypes of Pseudevernia Furfuracea. Biomolecules 2021, 11, 1445. [Google Scholar] [CrossRef]
- Armaleo, D.; Sun, X.; Culberson, C. Insights from the First Putative Biosynthetic Gene Cluster for a Lichen Depside and Depsidone. Mycologia 2011, 103, 741–754. [Google Scholar] [CrossRef]
- Bertrand, R.L.; Sorensen, J.L. A Comprehensive Catalogue of Polyketide Synthase Gene Clusters in Lichenizing Fungi. J. Ind. Microbiol. Biotechnol. 2018, 45, 1067–1081. [Google Scholar] [CrossRef]
- Nguyen, D.M.T.; Do, L.M.T.; Nguyen, V.T.; Chavasiri, W.; Mortier, J.; Nguyen, P.P.K. Phenolic Compounds from the Lichen Lobaria orientalis. J. Nat. Prod. 2017, 80, 261–268. [Google Scholar] [CrossRef]
- Oksanen, I.; Jokela, J.; Fewer, D.P.; Wahlsten, M.; Rikkinen, J.; Sivonen, K. Discovery of Rare and Highly Toxic Microcystins from Lichen-Associated Cyanobacterium Nostoc sp. Strain IO-102-I. Appl. Environ. Microbiol. 2004, 70, 5756–5763. [Google Scholar] [CrossRef]
- Garg, N.; Zeng, Y.; Edlund, A.; Melnik, A.V.; Sanchez, L.M.; Mohimani, H.; Gurevich, A.; Miao, V.; Schiffler, S.; Lim, Y.W.; et al. Spatial Molecular Architecture of the Microbial Community of a Peltigera Lichen. mSystems 2016, 1. [Google Scholar] [CrossRef]
- Krespach, M.K.C.; García-Altares, M.; Flak, M.; Schoeler, H.; Scherlach, K.; Netzker, T.; Schmalzl, A.; Mattern, D.J.; Schroeckh, V.; Komor, A.; et al. Lichen-like Association of Chlamydomonas Reinhardtii and Aspergillus Nidulans Protects Algal Cells from Bacteria. ISME J. 2020, 14, 2794–2805. [Google Scholar] [CrossRef]
- Noh, H.-J.; Park, Y.; Hong, S.G.; Lee, Y.M. Diversity and Physiological Characteristics of Antarctic Lichens-Associated Bacteria. Microorganisms 2021, 9, 607. [Google Scholar] [CrossRef]
- Noël, A.; Garnier, A.; Clément, M.; Rouaud, I.; Sauvager, A.; Bousarghin, L.; Vásquez-Ocmín, P.; Maciuk, A.; Tomasi, S. Lichen-Associated Bacteria Transform Antibacterial Usnic Acid to Products of Lower Antibiotic Activity. Phytochemistry 2021, 181, 112535. [Google Scholar] [CrossRef]
- Silby, M.W.; Winstanley, C.; Godfrey, S.A.C.; Levy, S.B.; Jackson, R.W. Pseudomonas Genomes: Diverse and Adaptable. FEMS Microbiol. Rev. 2011, 35, 652–680. [Google Scholar] [CrossRef]
- Passarelli-Araujo, H.; Franco, G.R.; Venancio, T.M. Network Analysis of Ten Thousand Genomes Shed Light on Pseudomonas Diversity and Classification. Microbiol. Res. 2022, 254, 126919. [Google Scholar] [CrossRef]
- Gross, H.; Loper, J.E. Genomics of Secondary Metabolite Production by Pseudomonas spp. Nat. Prod. Rep. 2009, 26, 1408. [Google Scholar] [CrossRef]
- Nunes da Silva, M.; Carvalho, S.M.P.; Rodrigues, A.M.; Gómez-Cadenas, A.; António, C.; Vasconcelos, M.W. Defence-related Pathways, Phytohormones and Primary Metabolism Are Key Players in Kiwifruit Plant Tolerance to Pseudomonas syringae Pv. actinidiae. Plant. Cell Environ. 2021, 45, 528–541. [Google Scholar] [CrossRef]
- Tumewu, S.A.; Matsui, H.; Yamamoto, M.; Noutoshi, Y.; Toyoda, K.; Ichinose, Y. Identification of Chemoreceptor Proteins for Amino Acids Involved in Host Plant Infection in Pseudomonas syringae Pv. Tabaci 6605. Microbiol. Res. 2021, 253, 126869. [Google Scholar] [CrossRef]
- Krishna, P.S.; Woodcock, S.D.; Pfeilmeier, S.; Bornemann, S.; Zipfel, C.; Malone, J.G. Pseudomonas syringae Addresses Distinct Environmental Challenges during Plant Infection through the Coordinated Deployment of Polysaccharides. J. Exp. Bot. 2021, erab550. [Google Scholar] [CrossRef] [PubMed]
- Zboralski, A.; Filion, M. Genetic Factors Involved in Rhizosphere Colonization by Phytobeneficial Pseudomonas spp. Comput. Struct. Biotechnol. J. 2020, 18, 3539–3554. [Google Scholar] [CrossRef] [PubMed]
- Vick, S.H.W.; Fabian, B.K.; Dawson, C.J.; Foster, C.; Asher, A.; Hassan, K.A.; Midgley, D.J.; Paulsen, I.T.; Tetu, S.G. Delving into Defence: Identifying the Pseudomonas Protegens Pf-5 Gene Suite Involved in Defence against Secreted Products of Fungal, Oomycete and Bacterial Rhizosphere Competitors. Microb. Genom. 2021, 7. [Google Scholar] [CrossRef]
- Yang, R.; Li, S.; Li, Y.; Yan, Y.; Fang, Y.; Zou, L.; Chen, G. Bactericidal Effect of Pseudomonas Oryziphila sp. Nov., a Novel Pseudomonas Species Against Xanthomonas oryzae Reduces Disease Severity of Bacterial Leaf Streak of Rice. Front. Microbiol. 2021, 12, 759536. [Google Scholar] [CrossRef]
- Pacheco-Moreno, A.; Stefanato, F.L.; Ford, J.J.; Trippel, C.; Uszkoreit, S.; Ferrafiat, L.; Grenga, L.; Dickens, R.; Kelly, N.; Kingdon, A.D.; et al. Pan-Genome Analysis Identifies Intersecting Roles for Pseudomonas Specialized Metabolites in Potato Pathogen Inhibition. eLife 2021, 10, e71900. [Google Scholar] [CrossRef]
- Gomila, M.; Peña, A.; Mulet, M.; Lalucat, J.; García Valdés, E. Phylogenomics and Systematics in Pseudomonas. Front. Microbiol. 2015, 6. [Google Scholar] [CrossRef] [PubMed]
- Vior, N.M.; Lacret, R.; Chandra, G.; Dorai-Raj, S.; Trick, M.; Truman, A.W. Discovery and Biosynthesis of the Antibiotic Bicyclomycin in Distantly Related Bacterial Classes. Appl. Environ. Microbiol. 2018, 84. [Google Scholar] [CrossRef] [PubMed]
- Bown, L.; Li, Y.; Berrué, F.; Verhoeven, J.T.P.; Dufour, S.C.; Bignell, D.R.D. Coronafacoyl Phytotoxin Biosynthesis and Evolution in the Common Scab Pathogen Streptomyces Scabiei. Appl. Environ. Microbiol. 2017, 83. [Google Scholar] [CrossRef]
- Kim, S.Y.; Ju, K.-S.; Metcalf, W.W.; Evans, B.S.; Kuzuyama, T.; van der Donk, W.A. Different Biosynthetic Pathways to Fosfomycin in Pseudomonas Syringae and Streptomyces Species. Antimicrob. Agents Chemother. 2012, 56, 4175–4183. [Google Scholar] [CrossRef]
- Simon, M.A.; Ongpipattanakul, C.; Nair, S.K.; van der Donk, W.A. Biosynthesis of Fosfomycin in Pseudomonads Reveals an Unexpected Enzymatic Activity in the Metallohydrolase Superfamily. Proc. Natl. Acad. Sci. USA 2021, 118, e2019863118. [Google Scholar] [CrossRef]
- Chen, X.; Xu, M.; Lü, J.; Xu, J.; Wang, Y.; Lin, S.; Deng, Z.; Tao, M. Biosynthesis of Tropolones in Streptomyces spp.: Interweaving Biosynthesis and Degradation of Phenylacetic Acid and Hydroxylations on the Tropone Ring. Appl. Environ. Microbiol. 2018, 84. [Google Scholar] [CrossRef]
- Muzio, F.M.; Agaras, B.C.; Masi, M.; Tuzi, A.; Evidente, A.; Valverde, C. 7-hydroxytropolone Is the Main Metabolite Responsible for the Fungal Antagonism of Pseudomonas donghuensis Strain SVBP6. Environ. Microbiol. 2020, 22, 2550–2563. [Google Scholar] [CrossRef]
- Moffat, A.D.; Elliston, A.; Patron, N.J.; Truman, A.W.; Carrasco Lopez, J.A. A Biofoundry Workflow for the Identification of Genetic Determinants of Microbial Growth Inhibition. Synth. Biol. 2021, 6, ysab004. [Google Scholar] [CrossRef]
- Quinn, R.J.; Barraza, I.; García-Diéguez, L.; Pajon, C.; Krausfeldt, L.E.; Ibrahim, K.; Enzinna, L.A.; Thorn, M.E.; Eldakar, O.T.; Craddock, T.J.A.; et al. Periodically Disturbing the Spatial Structure of Biofilms Can Affect the Production of an Essential Virulence Factor in Pseudomonas aeruginosa. mSystems 2021, 6, e00961-21. [Google Scholar] [CrossRef]
- Kang, D.; Revtovich, A.V.; Deyanov, A.E.; Kirienko, N.V. Pyoverdine Inhibitors and Gallium Nitrate Synergistically Affect Pseudomonas aeruginosa. mSphere 2021, 6. [Google Scholar] [CrossRef]
- Meyer, J.-M. Pyoverdines: Pigments, Siderophores and Potential Taxonomic Markers of Fluorescent Pseudomonas Species. Arch. Microbiol. 2000, 174, 135–142. [Google Scholar] [CrossRef]
- Ringel, M.T.; Brüser, T. The Biosynthesis of Pyoverdines. Microb. Cell 2018, 5, 424–437. [Google Scholar] [CrossRef]
- Götze, S.; Stallforth, P. Structure, Properties, and Biological Functions of Nonribosomal Lipopeptides from Pseudomonads. Nat. Prod. Rep. 2020, 37, 29–54. [Google Scholar] [CrossRef]
- Berti, A.D.; Greve, N.J.; Christensen, Q.H.; Thomas, M.G. Identification of a Biosynthetic Gene Cluster and the Six Associated Lipopeptides Involved in Swarming Motility of Pseudomonas syringae Pv. Tomato DC3000. J. Bacteriol. 2007, 189, 6312–6323. [Google Scholar] [CrossRef]
- Pauwelyn, E.; Huang, C.-J.; Ongena, M.; Leclère, V.; Jacques, P.; Bleyaert, P.; Budzikiewicz, H.; Schäfer, M.; Höfte, M. New Linear Lipopeptides Produced by Pseudomonas cichorii SF1-54 Are Involved in Virulence, Swarming Motility, and Biofilm Formation. MPMI 2013, 26, 585–598. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Berry, C.L.; Brassinga, A.K.C.; Donald, L.J.; Fernando, W.G.D.; Loewen, P.C.; de Kievit, T.R. Chemical and Biological Characterization of Sclerosin, an Antifungal Lipopeptide. Can. J. Microbiol. 2012, 58, 1027–1034. [Google Scholar] [CrossRef][Green Version]
- Bassarello, C.; Lazzaroni, S.; Bifulco, G.; Lo Cantore, P.; Iacobellis, N.S.; Riccio, R.; Gomez-Paloma, L.; Evidente, A. Tolaasins A−E, Five New Lipodepsipeptides Produced by Pseudomonas tolaasii. J. Nat. Prod. 2004, 67, 811–816. [Google Scholar] [CrossRef]
- Zhang, S.; Mukherji, R.; Chowdhury, S.; Reimer, L.; Stallforth, P. Lipopeptide-Mediated Bacterial Interaction Enables Cooperative Predator Defense. Proc. Natl. Acad. Sci. USA 2021, 118, e2013759118. [Google Scholar] [CrossRef]
- Girard, L.; Höfte, M.; Mot, R.D. Lipopeptide Families at the Interface between Pathogenic and Beneficial Pseudomonas -Plant Interactions. Crit. Rev. Microbiol. 2020, 46, 397–419. [Google Scholar] [CrossRef] [PubMed]
- Geudens, N.; Martins, J.C. Cyclic Lipodepsipeptides From Pseudomonas spp.—Biological Swiss-Army Knives. Front. Microbiol. 2018, 9, 1867. [Google Scholar] [CrossRef]
- Nguyen, D.D.; Melnik, A.V.; Koyama, N.; Lu, X.; Schorn, M.; Fang, J.; Aguinaldo, K.; Lincecum, T.L.; Ghequire, M.G.K.; Carrion, V.J.; et al. Indexing the Pseudomonas Specialized Metabolome Enabled the Discovery of Poaeamide B and the Bananamides. Nat. Microbiol. 2017, 2, 16197. [Google Scholar] [CrossRef] [PubMed]
- Zheng, W.; Wang, X.; Chen, Y.; Dong, Y.; Zhou, D.; Liu, R.; Zhou, H.; Bian, X.; Wang, H.; Tu, Q.; et al. Recombineering Facilitates the Discovery of Natural Product Biosynthetic Pathways in Pseudomonas parafulva. Biotechnol. J. 2021, 16, 2000575. [Google Scholar] [CrossRef]
- Götze, S.; Herbst-Irmer, R.; Klapper, M.; Görls, H.; Schneider, K.R.A.; Barnett, R.; Burks, T.; Neu, U.; Stallforth, P. Structure, Biosynthesis, and Biological Activity of the Cyclic Lipopeptide Anikasin. ACS Chem. Biol. 2017, 12, 2498–2502. [Google Scholar] [CrossRef] [PubMed]
- Kinscherf, T.G.; Willis, D.K. The Biosynthetic Gene Cluster for the β-Lactam Antibiotic Tabtoxin in Pseudomonas syringae. J. Antibiot. 2005, 58, 817–821. [Google Scholar] [CrossRef] [PubMed]
- Ikeda, Y.; Idemoto, H.; Hirayama, F.; Yamamoto, K.; Iwao, K.; Asao, T.; Munakata, T. Safracins, New Antitumor Antibiotics I. Producing Organism, Fermentation and Isolation. J. Antibiot. 1983, 36, 1279–1283. [Google Scholar] [CrossRef] [PubMed]
- Velasco, A.; Acebo, P.; Gomez, A.; Schleissner, C.; Rodríguez, P.; Aparicio, T.; Conde, S.; Muñoz, R.; De La Calle, F.; Garcia, J.L.; et al. Molecular Characterization of the Safracin Biosynthetic Pathway from Pseudomonas fluorescens A2-2: Designing New Cytotoxic Compounds: Safracin Biosynthetic Pathway. Mol. Microbiol. 2005, 56, 144–154. [Google Scholar] [CrossRef] [PubMed]
- Wells, J.S.; Trejo, W.H.; Principe, P.A.; Sykes, R.B. Obafluorin, a Novel.BETA.-Lactone Produced by Pseudomonas fluorescens. Taxonomy, Fermentation and Biological Properties. J. Antibiot. 1984, 37, 802–803. [Google Scholar] [CrossRef] [PubMed]
- Scott, T.A.; Heine, D.; Qin, Z.; Wilkinson, B. An L-Threonine Transaldolase Is Required for L-Threo-β-Hydroxy-α-Amino Acid Assembly during Obafluorin Biosynthesis. Nat. Commun. 2017, 8, 15935. [Google Scholar] [CrossRef]
- Schaffer, J.E.; Reck, M.R.; Prasad, N.K.; Wencewicz, T.A. β-Lactone Formation during Product Release from a Nonribosomal Peptide Synthetase. Nat. Chem. Biol. 2017, 13, 737–744. [Google Scholar] [CrossRef] [PubMed]
- Patteson, J.B.; Lescallette, A.R.; Li, B. Discovery and Biosynthesis of Azabicyclene, a Conserved Nonribosomal Peptide in Pseudomonas aeruginosa. Org. Lett. 2019, 21, 4955–4959. [Google Scholar] [CrossRef]
- El-Sayed, A.K.; Hothersall, J.; Cooper, S.M.; Stephens, E.; Simpson, T.J.; Thomas, C.M. Characterization of the Mupirocin Biosynthesis Gene Cluster from Pseudomonas fluorescens NCIMB 10586. Chem. Biol. 2003, 10, 419–430. [Google Scholar] [CrossRef]
- Gurney, R.; Thomas, C.M. Mupirocin: Biosynthesis, Special Features and Applications of an Antibiotic from a Gram-Negative Bacterium. Appl. Microbiol. Biotechnol. 2011, 90, 11–21. [Google Scholar] [CrossRef]
- Matthijs, S.; Vander Wauven, C.; Cornu, B.; Ye, L.; Cornelis, P.; Thomas, C.M.; Ongena, M. Antimicrobial Properties of Pseudomonas Strains Producing the Antibiotic Mupirocin. Res. Microbiol. 2014, 165, 695–704. [Google Scholar] [CrossRef] [PubMed]
- Zhao, M.-M.; Lyu, N.; Wang, D.; Wu, X.-G.; Zhao, Y.-Z.; Zhang, L.-Q.; Zhou, H.-Y. PhlG Mediates the Conversion of DAPG to MAPG in Pseudomonas fluorescens 2P24. Sci. Rep. 2020, 10, 4296. [Google Scholar] [CrossRef] [PubMed]
- Meyer, S.L.F.; Halbrendt, J.M.; Carta, L.K.; Skantar, A.M.; Liu, T.; Abdelnabby, H.M.E.; Vinyard, B.T. Toxicity of 2,4-Diacetylphloroglucinol (DAPG) to Plant-Parasitic and Bacterial-Feeding Nematodes. J. Nematol. 2009, 41, 274–280. [Google Scholar] [PubMed]
- Haas, D.; Keel, C. Regulation of antibiotic production in root-colonizing Pseudomonas spp. and relevance for the biological control of plant disease. Annu. Rev. Phytopathol. 2003, 41, 117–153. [Google Scholar] [CrossRef]
- Bignell, D.R.D.; Cheng, Z.; Bown, L. The Coronafacoyl Phytotoxins: Structure, Biosynthesis, Regulation and Biological Activities. Antonie Leeuwenhoek 2018, 111, 649–666. [Google Scholar] [CrossRef] [PubMed]
- Ramel, C.; Tobler, M.; Meyer, M.; Bigler, L.; Ebert, M.-O.; Schellenberg, B.; Dudler, R. Biosynthesis of the Proteasome Inhibitor Syringolin A: The Ureido Group Joining Two Amino Acids Originates from Bicarbonate. BMC BioChem. 2009, 10, 26. [Google Scholar] [CrossRef]
- Schellenberg, B.; Ramel, C.; Dudler, R. Pseudomonas syringae Virulence Factor Syringolin A Counteracts Stomatal Immunity by Proteasome Inhibition. MPMI 2010, 23, 1287–1293. [Google Scholar] [CrossRef]
- Blankenfeldt, W.; Parsons, J.F. The Structural Biology of Phenazine Biosynthesis. Curr. Opin. Struct. Biol. 2014, 29, 26–33. [Google Scholar] [CrossRef]
- Pierson, L.S.; Pierson, E.A. Metabolism and Function of Phenazines in Bacteria: Impacts on the Behavior of Bacteria in the Environment and Biotechnological Processes. Appl. Microbiol. Biotechnol. 2010, 86, 1659–1670. [Google Scholar] [CrossRef] [PubMed]
- Schmitz, S.; Rosenbaum, M.A. Controlling the Production of Pseudomonas Phenazines by Modulating the Genetic Repertoire. ACS Chem. Biol. 2020, 15, 3244–3252. [Google Scholar] [CrossRef] [PubMed]
- Castric, P.A. Hydrogen Cyanide, a Secondary Metabolite of Pseudomonas aeruginosa. Can. J. Microbiol. 1975, 21, 613–618. [Google Scholar] [CrossRef]
- Hotter, V.; Zopf, D.; Kim, H.J.; Silge, A.; Schmitt, M.; Aiyar, P.; Fleck, J.; Matthäus, C.; Hniopek, J.; Yan, Q.; et al. A Polyyne Toxin Produced by an Antagonistic Bacterium Blinds and Lyses a Chlamydomonad Alga. Proc. Natl. Acad. Sci. USA 2021, 118, e2107695118. [Google Scholar] [CrossRef]
- Mullins, A.J.; Webster, G.; Kim, H.J.; Zhao, J.; Petrova, Y.D.; Ramming, C.E.; Jenner, M.; Murray, J.A.H.; Connor, T.R.; Hertweck, C.; et al. Discovery of the Pseudomonas Polyyne Protegencin by a Phylogeny-Guided Study of Polyyne Biosynthetic Gene Cluster Diversity. mBio 2021, 12, e00715-21. [Google Scholar] [CrossRef]
- Mevers, E.; Saurí, J.; Helfrich, E.J.N.; Henke, M.; Barns, K.J.; Bugni, T.S.; Andes, D.; Currie, C.R.; Clardy, J. Pyonitrins A–D: Chimeric Natural Products Produced by Pseudomonas protegens. J. Am. Chem. Soc. 2019, 141, 17098–17101. [Google Scholar] [CrossRef]
- Klapper, M.; Schlabach, K.; Paschold, A.; Zhang, S.; Chowdhury, S.; Menzel, K.; Rosenbaum, M.A.; Stallforth, P. Biosynthesis of Pseudomonas -Derived Butenolides. Angew. Chem. Int. Ed. 2020, 59, 5607–5610. [Google Scholar] [CrossRef] [PubMed]
- Ting, C.P.; Funk, M.A.; Halaby, S.L.; Zhang, Z.; Gonen, T.; van der Donk, W.A. Use of a Scaffold Peptide in the Biosynthesis of Amino Acid–Derived Natural Products. Science 2019, 365, 280–284. [Google Scholar] [CrossRef] [PubMed]
- Loeschcke, A.; Thies, S. Pseudomonas Putida—A Versatile Host for the Production of Natural Products. Appl. Microbiol. Biotechnol. 2015, 99, 6197–6214. [Google Scholar] [CrossRef] [PubMed]
- Estrada-de los Santos, P.; Vinuesa, P.; Martínez-Aguilar, L.; Hirsch, A.M.; Caballero-Mellado, J. Phylogenetic Analysis of Burkholderia Species by Multilocus Sequence Analysis. Curr. Microbiol. 2013, 67, 51–60. [Google Scholar] [CrossRef] [PubMed]
- Kunakom, S.; Eustáquio, A.S. Burkholderia as a Source of Natural Products. J. Nat. Prod. 2019, 82, 2018–2037. [Google Scholar] [CrossRef]
- Holden, M.T.G.; Titball, R.W.; Peacock, S.J.; Cerdeno-Tarraga, A.M.; Atkins, T.; Crossman, L.C.; Pitt, T.; Churcher, C.; Mungall, K.; Bentley, S.D.; et al. Genomic Plasticity of the Causative Agent of Melioidosis, Burkholderia pseudomallei. Proc. Natl. Acad. Sci. USA 2004, 101, 14240–14245. [Google Scholar] [CrossRef] [PubMed]
- Jenner, M.; Jian, X.; Dashti, Y.; Masschelein, J.; Hobson, C.; Roberts, D.M.; Jones, C.; Harris, S.; Parkhill, J.; Raja, H.A.; et al. An Unusual Burkholderia gladioli Double Chain-Initiating Nonribosomal Peptide Synthetase Assembles ‘Fungal’ Icosalide Antibiotics. Chem. Sci. 2019, 10, 5489–5494. [Google Scholar] [CrossRef] [PubMed]
- Partida-Martinez, L.P.; Hertweck, C. A Gene Cluster Encoding Rhizoxin Biosynthesis in “Burkholderia rhizoxina”, the Bacterial Endosymbiont of the FungusRhizopus Microsporus. ChemBioChem 2007, 8, 41–45. [Google Scholar] [CrossRef]
- Scherlach, K.; Partida-Martinez, L.P.; Dahse, H.-M.; Hertweck, C. Antimitotic Rhizoxin Derivatives from a Cultured Bacterial Endosymbiont of the Rice Pathogenic Fungus Rhizopus microsporus. J. Am. Chem. Soc. 2006, 128, 11529–11536. [Google Scholar] [CrossRef] [PubMed]
- Partida-Martinez, L.P.; Hertweck, C. Pathogenic Fungus Harbours Endosymbiotic Bacteria for Toxin Production. Nature 2005, 437, 884–888. [Google Scholar] [CrossRef] [PubMed]
- Kusebauch, B.; Busch, B.; Scherlach, K.; Roth, M.; Hertweck, C. Polyketide-Chain Branching by an Enzymatic Michael Addition. Angew. Chem. Int. Ed. 2009, 48, 5001–5004. [Google Scholar] [CrossRef] [PubMed]
- Scherlach, K.; Busch, B.; Lackner, G.; Paszkowski, U.; Hertweck, C. Symbiotic Cooperation in the Biosynthesis of a Phytotoxin. Angew. Chem. Int. Ed. 2012, 51, 9615–9618. [Google Scholar] [CrossRef] [PubMed]
- Mahenthiralingam, E.; Song, L.; Sass, A.; White, J.; Wilmot, C.; Marchbank, A.; Boaisha, O.; Paine, J.; Knight, D.; Challis, G.L. Enacyloxins Are Products of an Unusual Hybrid Modular Polyketide Synthase Encoded by a Cryptic Burkholderia ambifaria Genomic Island. Chem. Biol. 2011, 18, 665–677. [Google Scholar] [CrossRef] [PubMed]
- Nakou, I.T.; Jenner, M.; Dashti, Y.; Romero-Canelón, I.; Masschelein, J.; Mahenthiralingam, E.; Challis, G.L. Genomics-Driven Discovery of a Novel Glutarimide Antibiotic from Burkholderia gladioli Reveals an Unusual Polyketide Synthase Chain Release Mechanism. Angew. Chem. Int. Ed. 2020, 59, 23145–23153. [Google Scholar] [CrossRef]
- Parker, W.L.; Rathnum, M.L.; Seiner, V.; Trejo, W.H.; Principe, P.A.; Sykes, R.B. Cepacin A and Cepacin B, Two New Antibiotics Produced by Pseudomonas cepacia. J. Antibiot. 1984, 37, 431–440. [Google Scholar] [CrossRef]
- Mullins, A.J.; Murray, J.A.H.; Bull, M.J.; Jenner, M.; Jones, C.; Webster, G.; Green, A.E.; Neill, D.R.; Connor, T.R.; Parkhill, J.; et al. Genome Mining Identifies Cepacin as a Plant-Protective Metabolite of the Biopesticidal Bacterium Burkholderia ambifaria. Nat. Microbiol. 2019, 4, 996–1005. [Google Scholar] [CrossRef]
- Masschelein, J.; Jenner, M.; Challis, G.L. Antibiotics from Gram-Negative Bacteria: A Comprehensive Overview and Selected Biosynthetic Highlights. Nat. Prod. Rep. 2017, 34, 712–783. [Google Scholar] [CrossRef]
- Foxfire, A.; Buhrow, A.R.; Orugunty, R.S.; Smith, L. Drug Discovery through the Isolation of Natural Products from Burkholderia. Expert Opin. Drug Discov. 2021, 16, 807–822. [Google Scholar] [CrossRef]
- Fuerst, J.A.; Sagulenko, E. Beyond the Bacterium: Planctomycetes Challenge Our Concepts of Microbial Structure and Function. Nat. Rev. Microbiol. 2011, 9, 403–413. [Google Scholar] [CrossRef] [PubMed]
- Boedeker, C.; Schüler, M.; Reintjes, G.; Jeske, O.; van Teeseling, M.C.F.; Jogler, M.; Rast, P.; Borchert, D.; Devos, D.P.; Kucklick, M.; et al. Determining the Bacterial Cell Biology of Planctomycetes. Nat. Commun. 2017, 8, 14853. [Google Scholar] [CrossRef] [PubMed]
- Mahajan, M.; Seeger, C.; Yee, B.; Andersson, S.G.E. Evolutionary Remodeling of the Cell Envelope in Bacteria of the Planctomycetes Phylum. Genome Biol. Evol. 2020, 12, 1528–1548. [Google Scholar] [CrossRef] [PubMed]
- Jeske, O.; Schüler, M.; Schumann, P.; Schneider, A.; Boedeker, C.; Jogler, M.; Bollschweiler, D.; Rohde, M.; Mayer, C.; Engelhardt, H.; et al. Planctomycetes Do Possess a Peptidoglycan Cell Wall. Nat. Commun. 2015, 6, 7116. [Google Scholar] [CrossRef] [PubMed]
- Van Teeseling, M.C.F.; Mesman, R.J.; Kuru, E.; Espaillat, A.; Cava, F.; Brun, Y.V.; VanNieuwenhze, M.S.; Kartal, B.; van Niftrik, L. Anammox Planctomycetes Have a Peptidoglycan Cell Wall. Nat. Commun. 2015, 6, 6878. [Google Scholar] [CrossRef]
- Jetten, M.S.M.; Niftrik, L.; van Strous, M.; Kartal, B.; Keltjens, J.T.; Op den Camp, H.J.M. Biochemistry and Molecular Biology of Anammox Bacteria. Crit. Rev. Biochem. Mol. Biol. 2009, 44, 65–84. [Google Scholar] [CrossRef] [PubMed]
- Wiegand, S.; Jogler, M.; Boedeker, C.; Pinto, D.; Vollmers, J.; Rivas-Marín, E.; Kohn, T.; Peeters, S.H.; Heuer, A.; Rast, P.; et al. Cultivation and Functional Characterization of 79 Planctomycetes Uncovers Their Unique Biology. Nat. Microbiol. 2020, 5, 126–140. [Google Scholar] [CrossRef] [PubMed]
- Vollmers, J.; Frentrup, M.; Rast, P.; Jogler, C.; Kaster, A.-K. Untangling Genomes of Novel Planctomycetal and Verrucomicrobial Species from Monterey Bay Kelp Forest Metagenomes by Refined Binning. Front. Microbiol. 2017, 8. [Google Scholar] [CrossRef]
- Kallscheuer, N.; Jogler, C. The Bacterial Phylum Planctomycetes as Novel Source for Bioactive Small Molecules. Biotechnol. Adv. 2021, 53, 107818. [Google Scholar] [CrossRef] [PubMed]
- Panter, F.; Garcia, R.; Thewes, A.; Zaburannyi, N.; Bunk, B.; Overmann, J.; Gutierrez, M.V.; Krug, D.; Müller, R. Production of a Dibrominated Aromatic Secondary Metabolite by a Planctomycete Implies Complex Interaction with a Macroalgal Host. ACS Chem. Biol. 2019, 14, 2713–2719. [Google Scholar] [CrossRef]
- Sandargo, B.; Jeske, O.; Boedeker, C.; Wiegand, S.; Wennrich, J.-P.; Kallscheuer, N.; Jogler, M.; Rohde, M.; Jogler, C.; Surup, F. Stieleriacines, N-Acyl Dehydrotyrosines From the Marine Planctomycete Stieleria neptunia sp. nov. Front. Microbiol. 2020, 11, 1408. [Google Scholar] [CrossRef] [PubMed]
- Kallscheuer, N.; Jeske, O.; Sandargo, B.; Boedeker, C.; Wiegand, S.; Bartling, P.; Jogler, M.; Rohde, M.; Petersen, J.; Medema, M.H.; et al. The Planctomycete Stieleria maiorica Mal15T Employs Stieleriacines to Alter the Species Composition in Marine Biofilms. Commun. Biol. 2020, 3, 303. [Google Scholar] [CrossRef] [PubMed]
- Bäckhed, F.; Ley, R.E.; Sonnenburg, J.L.; Peterson, D.A.; Gordon, J.I. Host-Bacterial Mutualism in the Human Intestine. Science 2005, 307, 1915–1920. [Google Scholar] [CrossRef] [PubMed]
- Whitman, W.B.; Coleman, D.C.; Wiebe, W.J. Prokaryotes: The Unseen Majority. Proc. Natl. Acad. Sci. USA 1998, 95, 6578–6583. [Google Scholar] [CrossRef] [PubMed]
- Hugenholtz, P.; Goebel, B.M.; Pace, N.R. Impact of Culture-Independent Studies on the Emerging Phylogenetic View of Bacterial Diversity. J. Bacteriol. 1998, 180, 4765–4774. [Google Scholar] [CrossRef]
- Donia, M.S.; Cimermancic, P.; Schulze, C.J.; Wieland Brown, L.C.; Martin, J.; Mitreva, M.; Clardy, J.; Linington, R.G.; Fischbach, M.A. A Systematic Analysis of Biosynthetic Gene Clusters in the Human Microbiome Reveals a Common Family of Antibiotics. Cell 2014, 158, 1402–1414. [Google Scholar] [CrossRef]
- Dabard, J.; Bridonneau, C.; Phillipe, C.; Anglade, P.; Molle, D.; Nardi, M.; Ladiré, M.; Girardin, H.; Marcille, F.; Gomez, A.; et al. Ruminococcin A, a New Lantibiotic Produced by aRuminococcus gnavus Strain Isolated from Human Feces. Appl. Environ. Microbiol. 2001, 67, 4111–4118. [Google Scholar] [CrossRef]
- Hatziioanou, D.; Gherghisan-Filip, C.; Saalbach, G.; Horn, N.; Wegmann, U.; Duncan, S.H.; Flint, H.J.; Mayer, M.J.; Narbad, A. Discovery of a Novel Lantibiotic Nisin O from Blautia obeum A2-162, Isolated from the Human Gastrointestinal Tract. Microbiology 2017, 163, 1292–1305. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Gutierrez, E.; O’Connor, P.M.; Saalbach, G.; Walsh, C.J.; Hegarty, J.W.; Guinane, C.M.; Mayer, M.J.; Narbad, A.; Cotter, P.D. First Evidence of Production of the Lantibiotic Nisin P. Sci. Rep. 2020, 10, 3738. [Google Scholar] [CrossRef]
- Salomón, R.A.; Farías, R.N. Microcin 25, a Novel Antimicrobial Peptide Produced by Escherichia coli. J. Bacteriol. 1992, 174, 7428–7435. [Google Scholar] [CrossRef] [PubMed]
- Zha, L.; Jiang, Y.; Henke, M.T.; Wilson, M.R.; Wang, J.X.; Kelleher, N.L.; Balskus, E.P. Colibactin Assembly Line Enzymes Use S-Adenosylmethionine to Build a Cyclopropane Ring. Nat. Chem. Biol. 2017, 13, 1063–1065. [Google Scholar] [CrossRef] [PubMed]
- Nougayrède, J.-P.; Homburg, S.; Taieb, F.; Boury, M.; Brzuszkiewicz, E.; Gottschalk, G.; Buchrieser, C.; Hacker, J.; Dobrindt, U.; Oswald, E. Escherichia coli Induces DNA Double-Strand Breaks in Eukaryotic Cells. Science 2006, 313, 848–851. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.; Stornetta, A.; Villalta, P.W.; Wilson, M.R.; Boudreau, P.D.; Zha, L.; Balbo, S.; Balskus, E.P. Reactivity of an Unusual Amidase May Explain Colibactin’s DNA Cross-Linking Activity. J. Am. Chem. Soc. 2019, 141, 11489–11496. [Google Scholar] [CrossRef] [PubMed]
- Wilson, M.R.; Jiang, Y.; Villalta, P.W.; Stornetta, A.; Boudreau, P.D.; Carrá, A.; Brennan, C.A.; Chun, E.; Ngo, L.; Samson, L.D.; et al. The Human Gut Bacterial Genotoxin Colibactin Alkylates DNA. Science 2019, 363, eaar7785. [Google Scholar] [CrossRef] [PubMed]
- Unterhauser, K.; Pöltl, L.; Schneditz, G.; Kienesberger, S.; Glabonjat, R.A.; Kitsera, M.; Pletz, J.; Josa-Prado, F.; Dornisch, E.; Lembacher-Fadum, C.; et al. Klebsiella oxytoca Enterotoxins Tilimycin and Tilivalline Have Distinct Host DNA-Damaging and Microtubule-Stabilizing Activities. Proc. Natl. Acad. Sci. USA 2019, 116, 3774–3783. [Google Scholar] [CrossRef] [PubMed]
- Dornisch, E.; Pletz, J.; Glabonjat, R.A.; Martin, F.; Lembacher-Fadum, C.; Neger, M.; Högenauer, C.; Francesconi, K.; Kroutil, W.; Zangger, K.; et al. Biosynthesis of the Enterotoxic Pyrrolobenzodiazepine Natural Product Tilivalline. Angew. Chem. Int. Ed. 2017, 56, 14753–14757. [Google Scholar] [CrossRef] [PubMed]
- Sardelli, L.; Perottoni, S.; Tunesi, M.; Boeri, L.; Fusco, F.; Petrini, P.; Albani, D.; Giordano, C. Technological Tools and Strategies for Culturing Human Gut Microbiota in Engineered in Vitro Models. Biotechnol. Bioeng. 2021, 118, 2886–2905. [Google Scholar] [CrossRef]
- Wang, L.; Ravichandran, V.; Yin, Y.; Yin, J.; Zhang, Y. Natural Products from Mammalian Gut Microbiota. Trends Biotechnol. 2019, 37, 492–504. [Google Scholar] [CrossRef]
- D’Onofrio, A.; Crawford, J.M.; Stewart, E.J.; Witt, K.; Gavrish, E.; Epstein, S.; Clardy, J.; Lewis, K. Siderophores from Neighboring Organisms Promote the Growth of Uncultured Bacteria. Chem. Biol. 2010, 17, 254–264. [Google Scholar] [CrossRef] [PubMed]
- Kaeberlein, T.; Lewis, K.; Epstein, S.S. Isolating “Uncultivable” Microorganisms in Pure Culture in a Simulated Natural Environment. Science 2002, 296, 1127–1129. [Google Scholar] [CrossRef]
- Ling, L.L.; Schneider, T.; Peoples, A.J.; Spoering, A.L.; Engels, I.; Conlon, B.P.; Mueller, A.; Schäberle, T.F.; Hughes, D.E.; Epstein, S.; et al. A New Antibiotic Kills Pathogens without Detectable Resistance. Nature 2015, 517, 455–459. [Google Scholar] [CrossRef] [PubMed]
- Wirtz, D.A.; Ludwig, K.C.; Arts, M.; Marx, C.E.; Krannich, S.; Barac, P.; Kehraus, S.; Josten, M.; Henrichfreise, B.; Müller, A.; et al. Biosynthesis and Mechanism of Action of the Cell Wall Targeting Antibiotic Hypeptin. Angew. Chem. Int. Ed. 2021, 60, 13579–13586. [Google Scholar] [CrossRef] [PubMed]
- Shoji, J.; Hinoo, H.; Hattori, T.; Hirooka, K.; Kimura, Y.; Yoshida, T. Isolation and Characterization of Hypeptin from Pseudomonas sp. J. Antibiot. 1989, 42, 1460–1464. [Google Scholar] [CrossRef] [PubMed]
- Merino, N.; Aronson, H.S.; Bojanova, D.P.; Feyhl-Buska, J.; Wong, M.L.; Zhang, S.; Giovannelli, D. Corrigendum: Living at the Extremes: Extremophiles and the Limits of Life in a Planetary Context. Front. Microbiol. 2019, 10, 1785. [Google Scholar] [CrossRef] [PubMed]
- Gabani, P.; Singh, O.V. Radiation-Resistant Extremophiles and Their Potential in Biotechnology and Therapeutics. Appl. Microbiol. Biotechnol. 2013, 97, 993–1004. [Google Scholar] [CrossRef] [PubMed]
- Wilson, Z.E.; Brimble, M.A. Molecules Derived from the Extremes of Life: A Decade Later. Nat. Prod. Rep. 2021, 38, 24–82. [Google Scholar] [CrossRef] [PubMed]
- Wilson, Z.E.; Brimble, M.A. Molecules Derived from the Extremes of Life. Nat. Prod. Rep. 2009, 26, 44–71. [Google Scholar] [CrossRef] [PubMed]
- Tripathi, V.C.; Satish, S.; Horam, S.; Raj, S.; Lal, A.; Arockiaraj, J.; Pasupuleti, M.; Dikshit, D.K. Natural Products from Polar Organisms: Structural Diversity, Bioactivities and Potential Pharmaceutical Applications. Polar Sci. 2018, 18, 147–166. [Google Scholar] [CrossRef]
- Hui, M.L.-Y.; Tan, L.T.-H.; Letchumanan, V.; He, Y.-W.; Fang, C.-M.; Chan, K.-G.; Law, J.W.-F.; Lee, L.-H. The Extremophilic Actinobacteria: From Microbes to Medicine. Antibiotics 2021, 10, 682. [Google Scholar] [CrossRef]
- Proteau, P.J.; Gerwick, W.H.; Garcia-Pichel, F.; Castenholz, R. The Structure of Scytonemin, an Ultraviolet Sunscreen Pigment from the Sheaths of Cyanobacteria. Experientia 1993, 49, 825–829. [Google Scholar] [CrossRef] [PubMed]
- Sorrels, C.M.; Proteau, P.J.; Gerwick, W.H. Organization, Evolution, and Expression Analysis of the Biosynthetic Gene Cluster for Scytonemin, a Cyanobacterial UV-Absorbing Pigment. Appl. Environ. Microbiol. 2009, 75, 4861–4869. [Google Scholar] [CrossRef]
- Baltz, R.H. Gifted Microbes for Genome Mining and Natural Product Discovery. J. Ind. Microbiol. Biotechnol. 2017, 44, 573–588. [Google Scholar] [CrossRef] [PubMed]
- Cowan, D.; Ramond, J.-B.; Makhalanyane, T.; De Maayer, P. Metagenomics of Extreme Environments. Curr. Opin. Microbiol. 2015, 25, 97–102. [Google Scholar] [CrossRef]
- Andrianasolo, E.H.; Haramaty, L.; Rosario-Passapera, R.; Bidle, K.; White, E.; Vetriani, C.; Falkowski, P.; Lutz, R. Ammonificins A and B, Hydroxyethylamine Chroman Derivatives from a Cultured Marine Hydrothermal Vent Bacterium, Thermovibrio ammonificans. J. Nat. Prod. 2009, 72, 1216–1219. [Google Scholar] [CrossRef] [PubMed]
- Andrianasolo, E.H.; Haramaty, L.; Rosario-Passapera, R.; Vetriani, C.; Falkowski, P.; White, E.; Lutz, R. Ammonificins C and D, Hydroxyethylamine Chromene Derivatives from a Cultured Marine Hydrothermal Vent Bacterium, Thermovibrio ammonificans. Mar. Drugs 2012, 10, 2300–2311. [Google Scholar] [CrossRef]
- Akiyama, H.; Oku, N.; Kasai, H.; Shizuri, Y.; Matsumoto, S.; Igarashi, Y. Metabolites from Thermophilic Bacteria I: N-Propionylanthranilic Acid, a Co-Metabolite of the Bacillamide Class Antibiotics and Tryptophan Metabolites with Herbicidal Activity from Laceyella sacchari. J. Antibiot. 2014, 67, 795–798. [Google Scholar] [CrossRef]
- Della Sala, G.; Mangoni, A.; Costantino, V.; Teta, R. Identification of the Biosynthetic Gene Cluster of Thermoactinoamides and Discovery of New Congeners by Integrated Genome Mining and MS-Based Molecular Networking. Front. Chem. 2020, 8, 397. [Google Scholar] [CrossRef]
- Teta, R.; Marteinsson, V.T.; Longeon, A.; Klonowski, A.M.; Groben, R.; Bourguet-Kondracki, M.-L.; Costantino, V.; Mangoni, A. Thermoactinoamide A, an Antibiotic Lipophilic Cyclopeptide from the Icelandic Thermophilic Bacterium Thermoactinomyces vulgaris. J. Nat. Prod. 2017, 80, 2530–2535. [Google Scholar] [CrossRef] [PubMed]
- Park, J.-S.; Kagaya, N.; Hashimoto, J.; Izumikawa, M.; Yabe, S.; Shin-ya, K.; Nishiyama, M.; Kuzuyama, T. Identification and Biosynthesis of New Acyloins from the Thermophilic Bacterium Thermosporothrix hazakensis SK20-1 T. ChemBioChem 2014, 15, 527–532. [Google Scholar] [CrossRef] [PubMed]
- Park, J.-S.; Yabe, S.; Shin-ya, K.; Nishiyama, M.; Kuzuyama, T. New 2-(1′H-Indole-3′-Carbonyl)-Thiazoles Derived from the Thermophilic Bacterium Thermosporothrix hazakensis SK20-1T. J. Antibiot. 2015, 68, 60–62. [Google Scholar] [CrossRef] [PubMed]
- Igarashi, Y.; Yamamoto, K.; Ueno, C.; Yamada, N.; Saito, K.; Takahashi, K.; Enomoto, M.; Kuwahara, S.; Oikawa, T.; Tashiro, E.; et al. Ktedonoketone and 2’-Oxosattabacin, Benzenoid Metabolites from a Thermophilic Bacterium Thermosporothrix hazakensis in the Phylum Chloroflexi. J. Antibiot. 2019, 72, 653–660. [Google Scholar] [CrossRef] [PubMed]
- Yabe, S.; Sakai, Y.; Abe, K.; Yokota, A. Diversity of Ktedonobacteria with Actinomycetes-Like Morphology in Terrestrial Environments. Microbes Environ. 2017, 32, 61–70. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.; Saitou, A.; Wang, C.-M.; Toyoda, A.; Minakuchi, Y.; Sekiguchi, Y.; Ueda, K.; Takano, H.; Sakai, Y.; Abe, K.; et al. Genome Features and Secondary Metabolites Biosynthetic Potential of the Class Ktedonobacteria. Front. Microbiol. 2019, 10, 893. [Google Scholar] [CrossRef] [PubMed]
- Xu, B.; Aitken, E.J.; Baker, B.P.; Turner, C.A.; Harvey, J.E.; Stott, M.B.; Power, J.F.; Harris, P.W.R.; Keyzers, R.A.; Brimble, M.A. Genome Mining, Isolation, Chemical Synthesis and Biological Evaluation of a Novel Lanthipeptide, Tikitericin, from the Extremophilic Microorganism Thermogemmatispora Strain T81. Chem. Sci. 2018, 9, 7311–7317. [Google Scholar] [CrossRef]
- Tedesco, P.; Maida, I.; Palma Esposito, F.; Tortorella, E.; Subko, K.; Ezeofor, C.; Zhang, Y.; Tabudravu, J.; Jaspars, M.; Fani, R.; et al. Antimicrobial Activity of Monoramnholipids Produced by Bacterial Strains Isolated from the Ross Sea (Antarctica). Mar. Drugs 2016, 14, 83. [Google Scholar] [CrossRef]
- Zhang, H.L.; Hua, H.M.; Pei, Y.H.; Yao, X.S. Three New Cytotoxic Cyclic Acylpeptides from Marine Bacillus sp. Chem. Pharm. Bull. 2004, 52, 1029–1030. [Google Scholar] [CrossRef]
- Du, Y.E.; Bae, E.S.; Lim, Y.; Cho, J.-C.; Nam, S.-J.; Shin, J.; Lee, S.K.; Nam, S.-I.; Oh, D.-C. Svalbamides A and B, Pyrrolidinone-Bearing Lipodipeptides from Arctic Paenibacillus sp. Mar. Drugs 2021, 19, 229. [Google Scholar] [CrossRef]
- Homann, V.V.; Sandy, M.; Tincu, J.A.; Templeton, A.S.; Tebo, B.M.; Butler, A. Loihichelins A−F, a Suite of Amphiphilic Siderophores Produced by the Marine Bacterium Halomonas LOB-5. J. Nat. Prod. 2009, 72, 884–888. [Google Scholar] [CrossRef]
- Son, S.; Ko, S.-K.; Jang, M.; Kim, J.; Kim, G.; Lee, J.; Jeon, E.; Futamura, Y.; Ryoo, I.-J.; Lee, J.-S.; et al. New Cyclic Lipopeptides of the Iturin Class Produced by Saltern-Derived Bacillus sp. KCB14S006. Mar. Drugs 2016, 14, 72. [Google Scholar] [CrossRef]
- Li, W.; Tang, X.-X.; Yan, X.; Wu, Z.; Yi, Z.-W.; Fang, M.-J.; Su, X.; Qiu, Y.-K. A New Macrolactin Antibiotic from Deep Sea-Derived Bacteria Bacillus subtilis B5. Nat. Prod. Res. 2016, 30, 2777–2782. [Google Scholar] [CrossRef]
- Yan, X.; Zhou, Y.-X.; Tang, X.-X.; Liu, X.-X.; Yi, Z.-W.; Fang, M.-J.; Wu, Z.; Jiang, F.-Q.; Qiu, Y.-K. Macrolactins from Marine-Derived Bacillus subtilis B5 Bacteria as Inhibitors of Inducible Nitric Oxide and Cytokines Expression. Mar. Drugs 2016, 14, 195. [Google Scholar] [CrossRef] [PubMed]
- Xie, C.-L.; Xia, J.-M.; Su, R.-Q.; Li, J.; Liu, Y.; Yang, X.-W.; Yang, Q. Bacilsubteramide A, a New Indole Alkaloid, from the Deep-Sea-Derived Bacillus subterraneus 11593. Nat. Prod. Res. 2018, 32, 2553–2557. [Google Scholar] [CrossRef] [PubMed]
- Park, H.B.; Kwon, H.C.; Lee, C.-H.; Yang, H.O. Glionitrin A, an Antibiotic−Antitumor Metabolite Derived from Competitive Interaction between Abandoned Mine Microbes. J. Nat. Prod. 2009, 72, 248–252. [Google Scholar] [CrossRef] [PubMed]
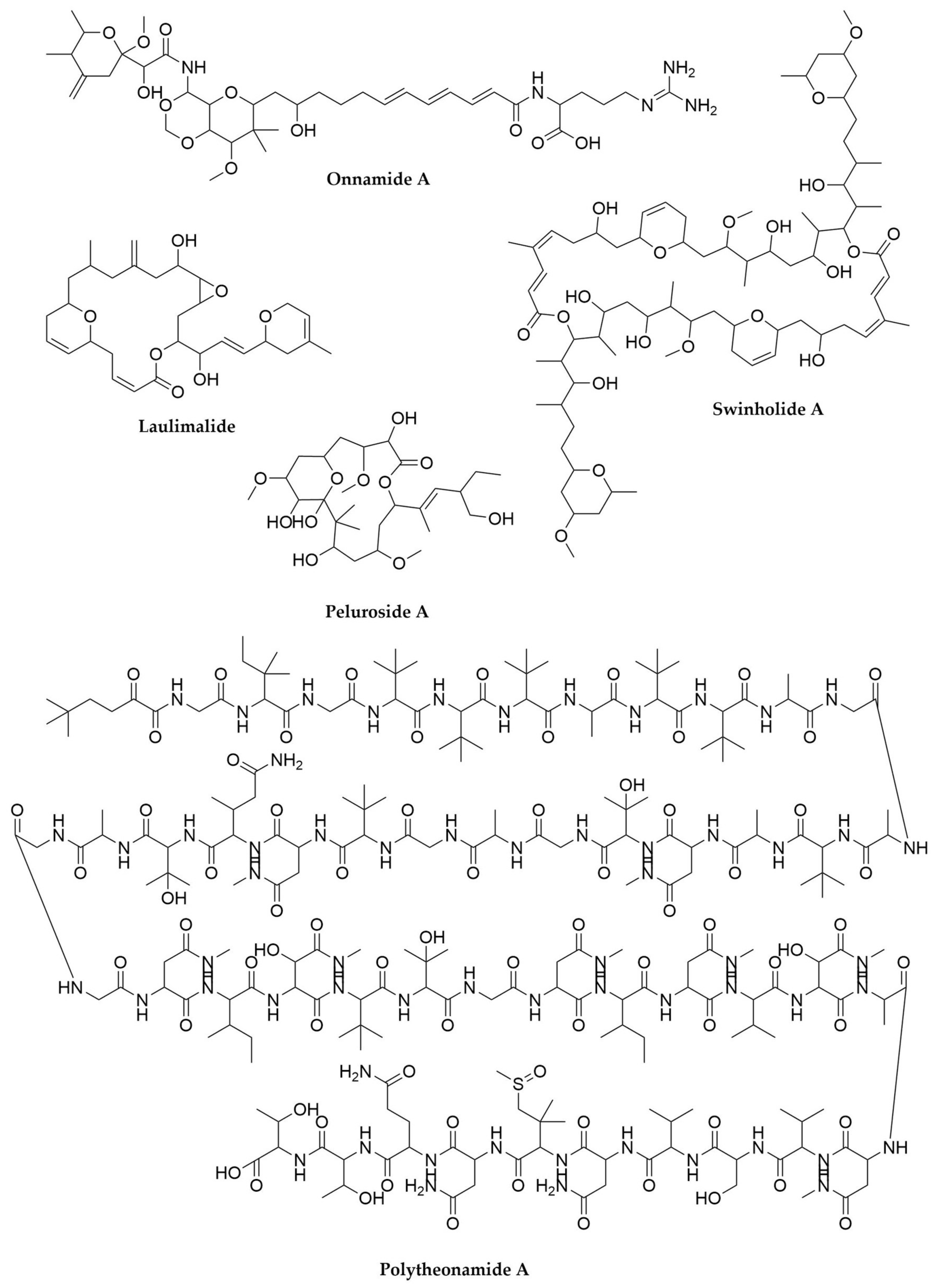
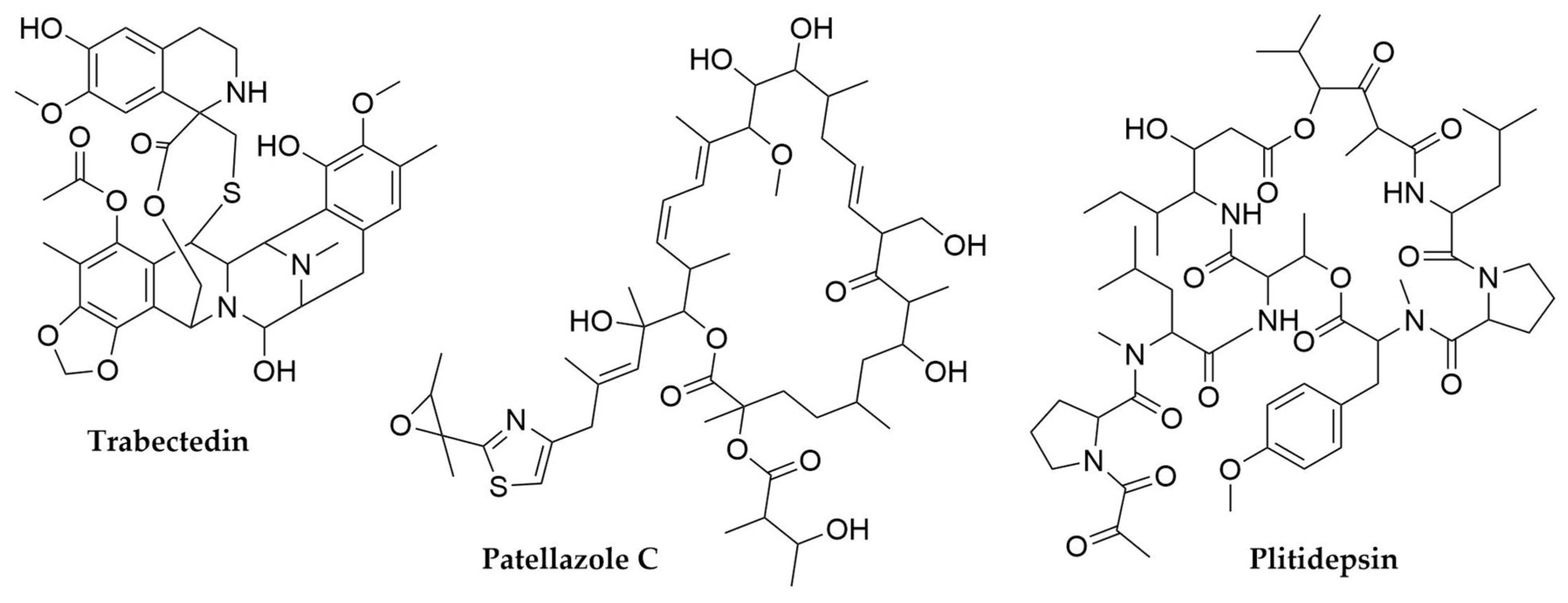
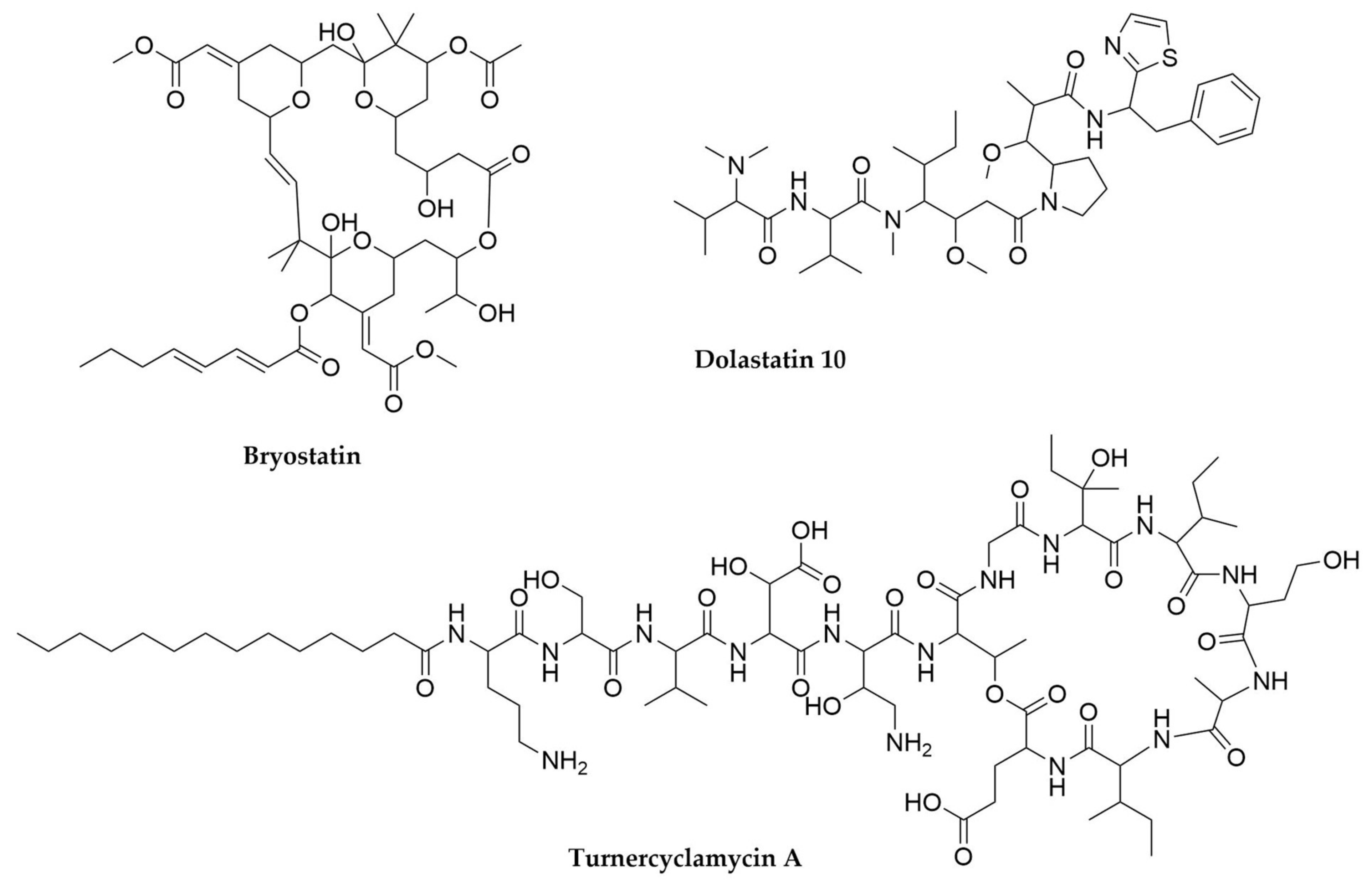

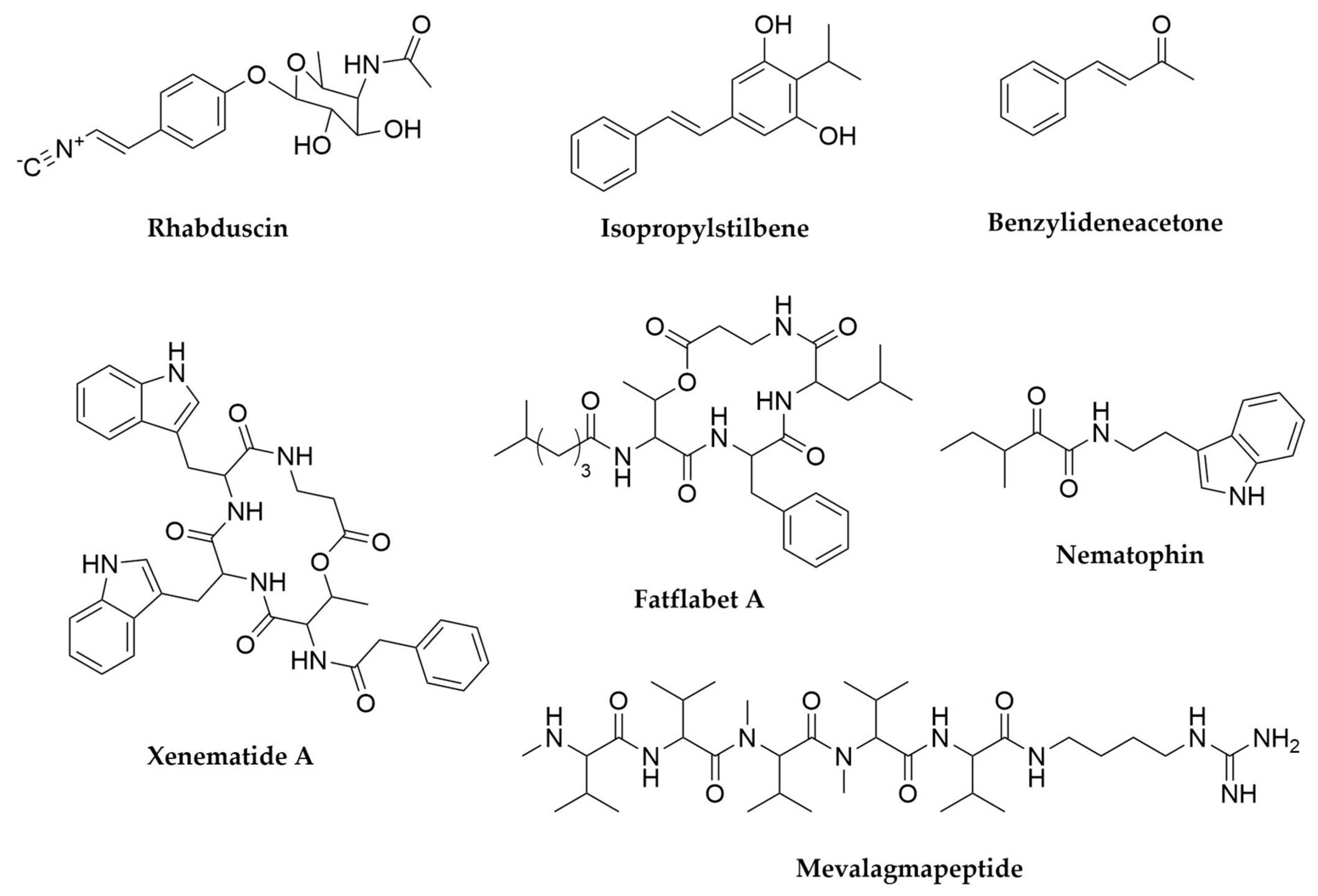
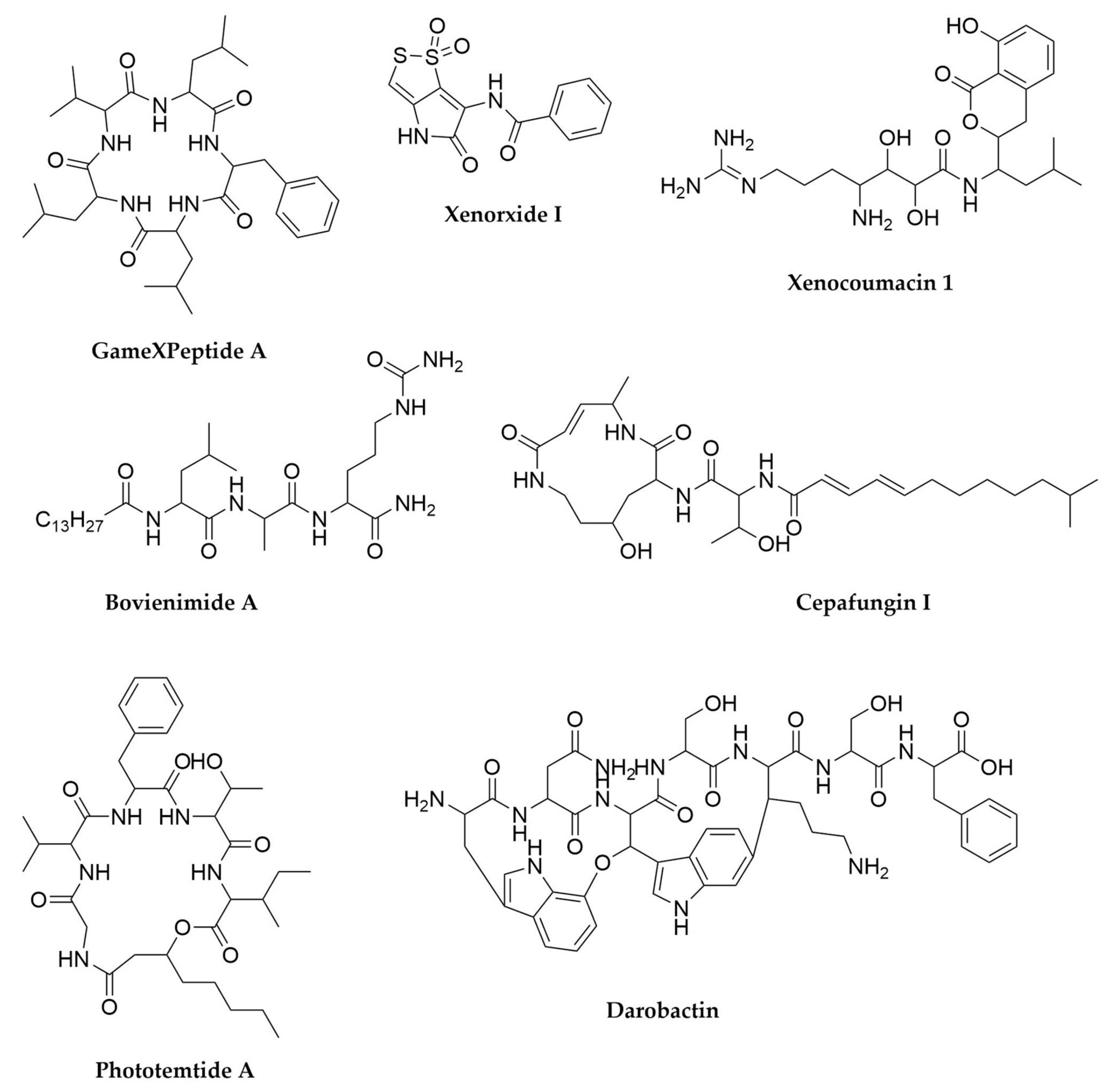
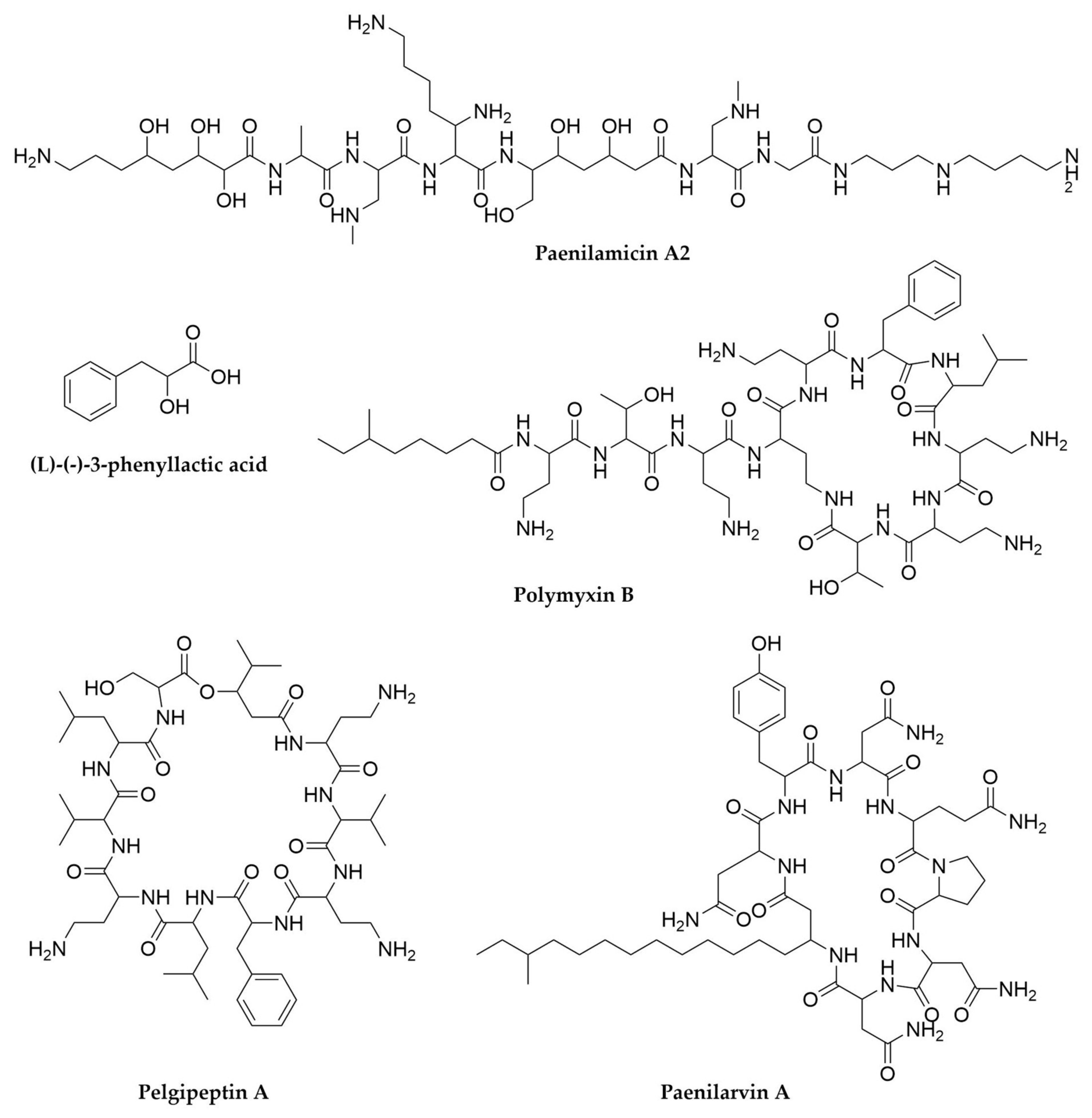
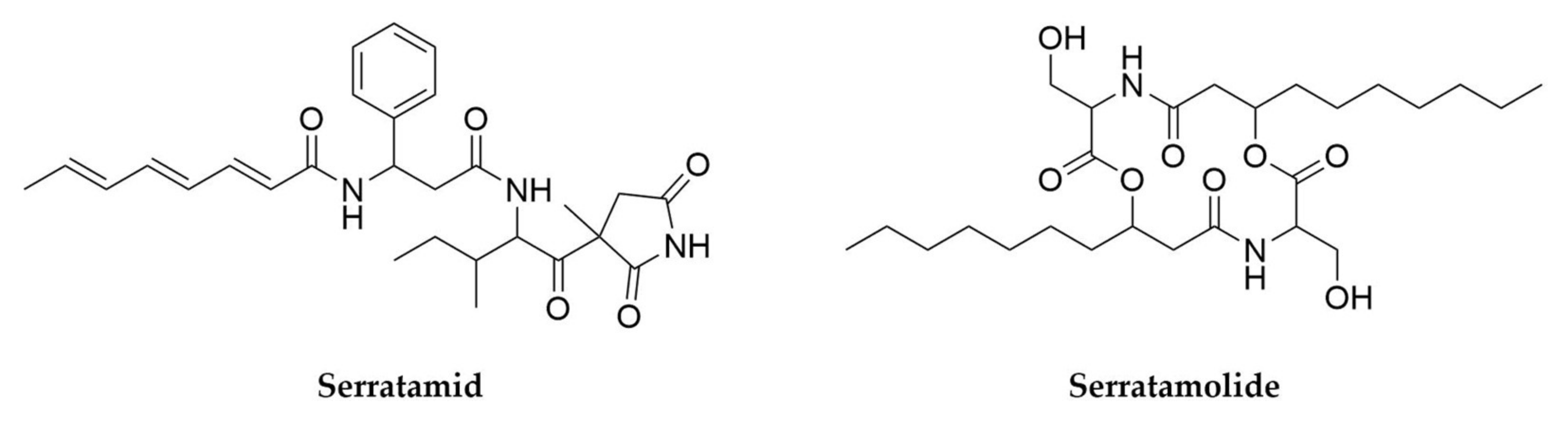
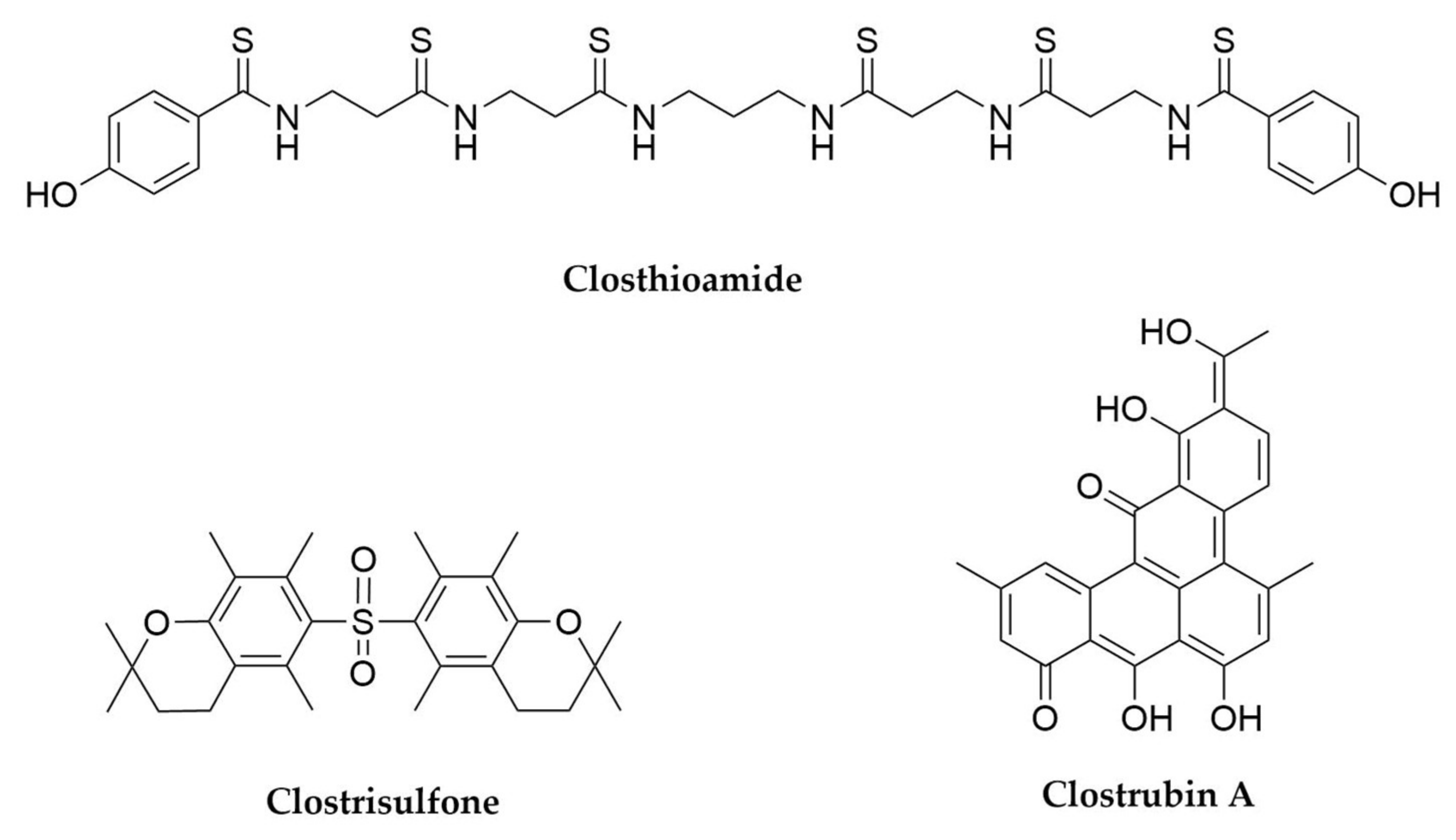
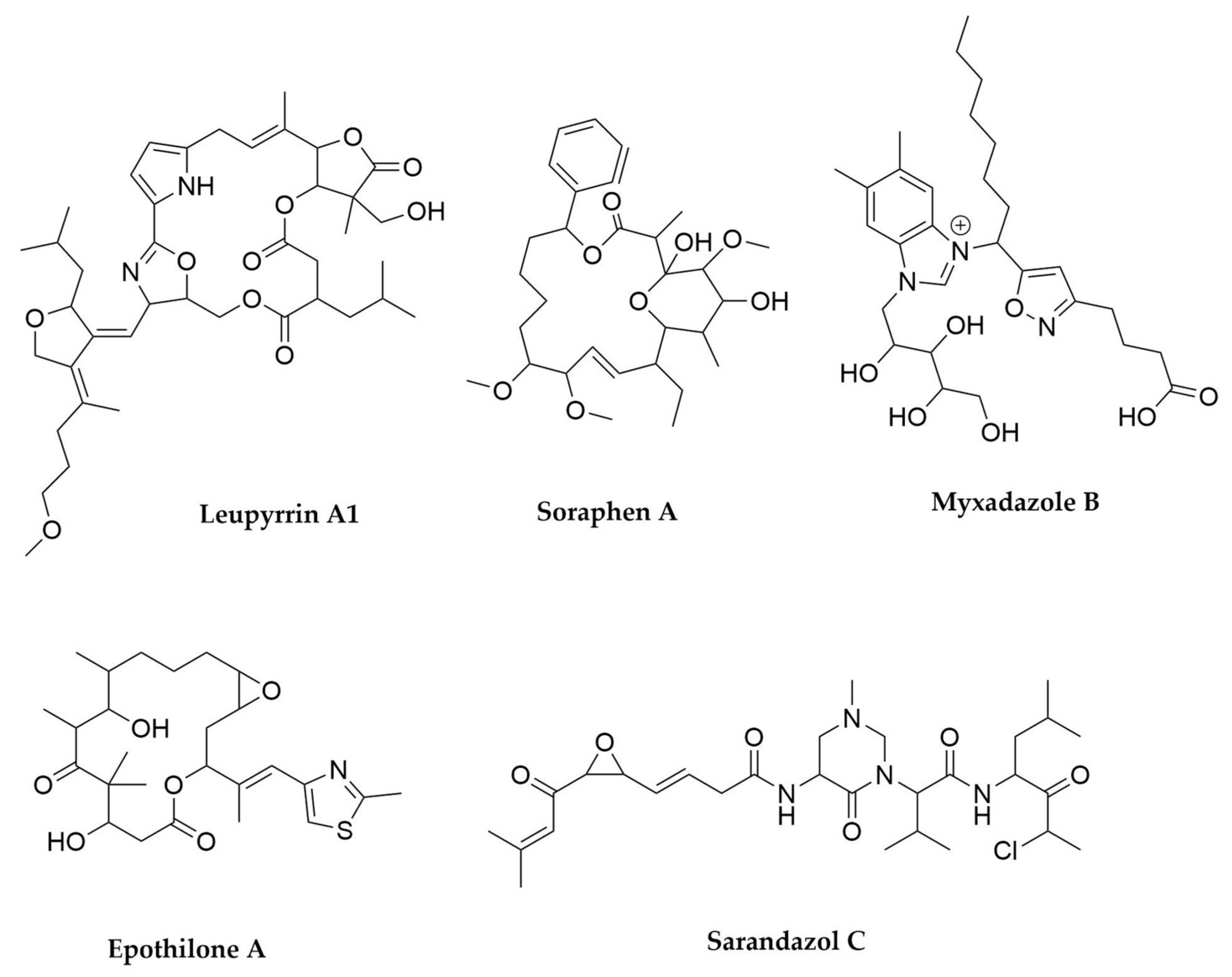
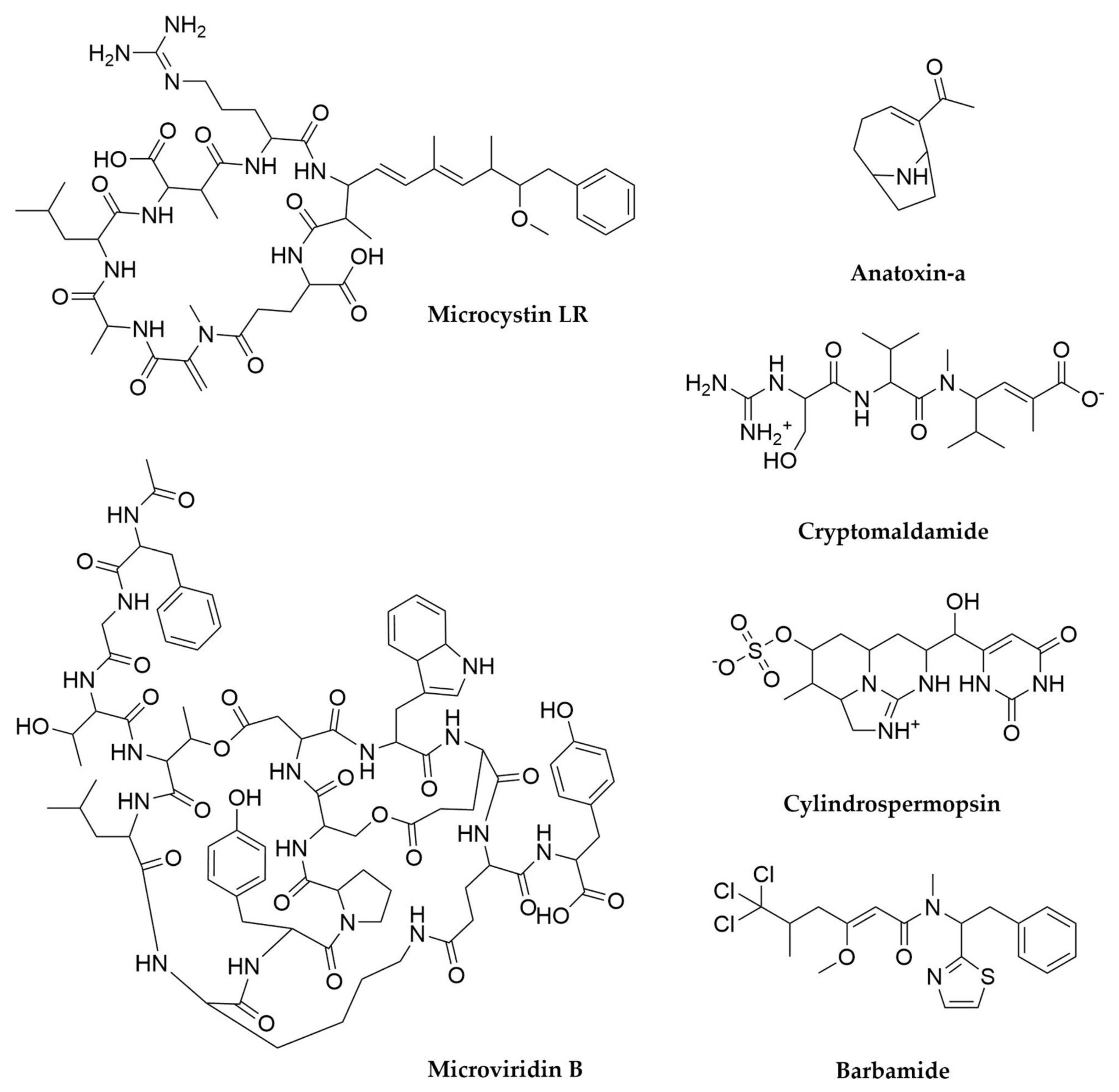
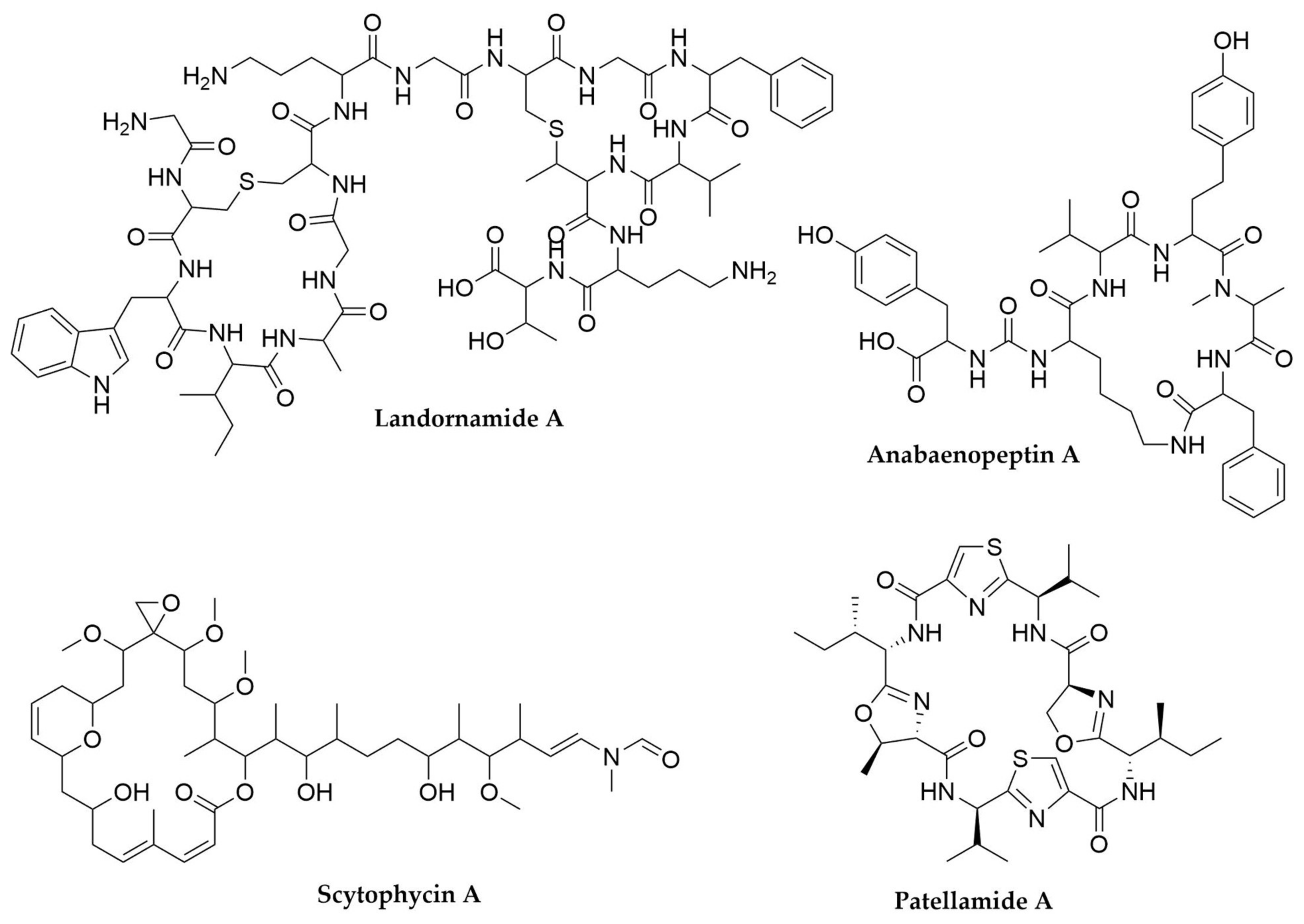
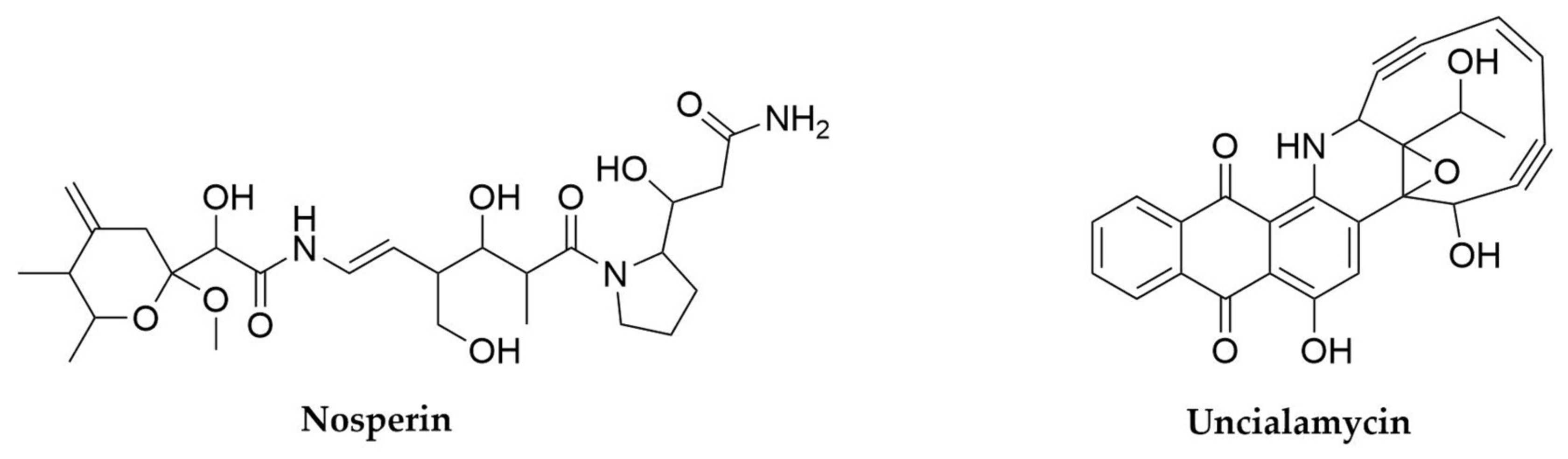
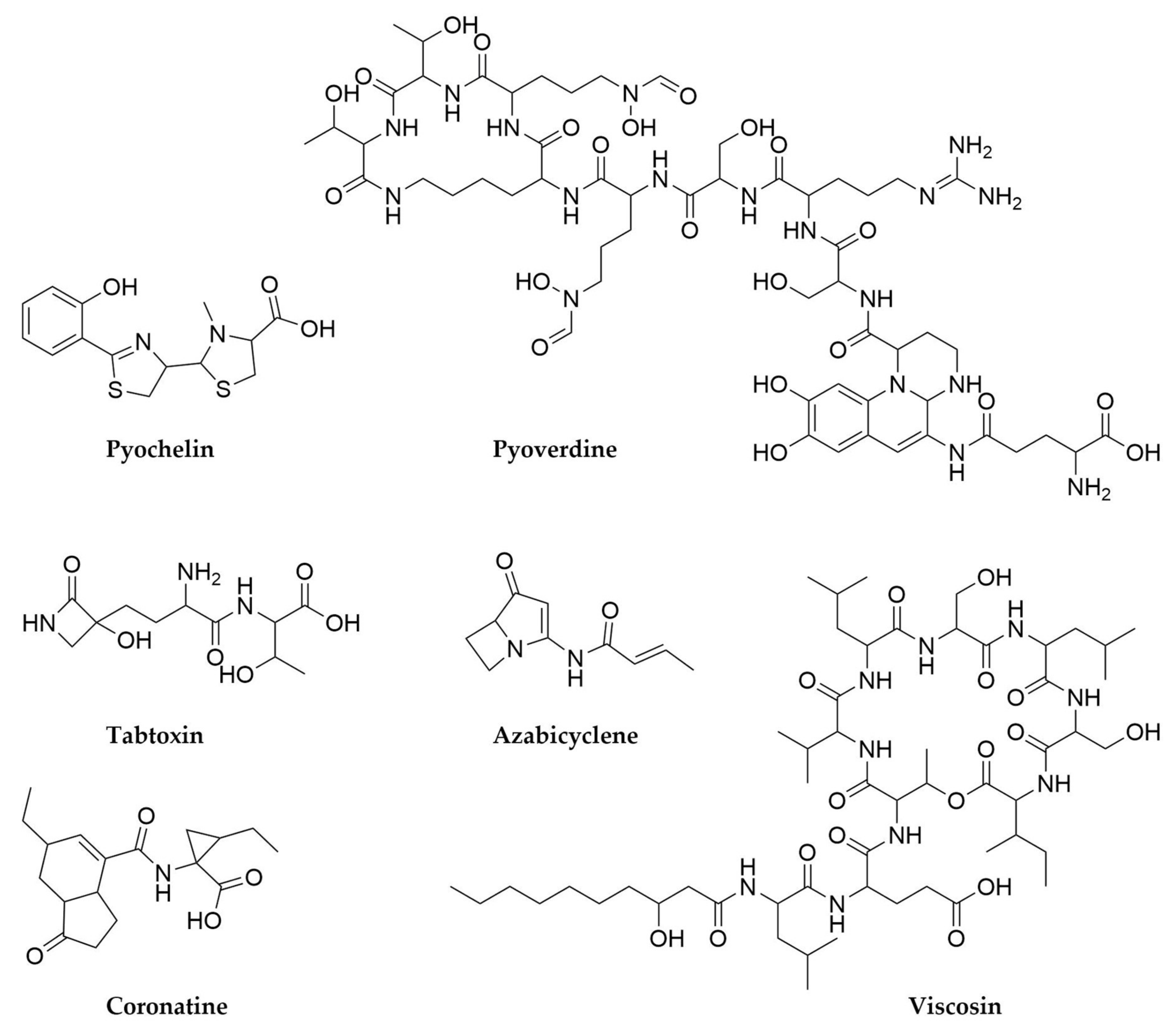


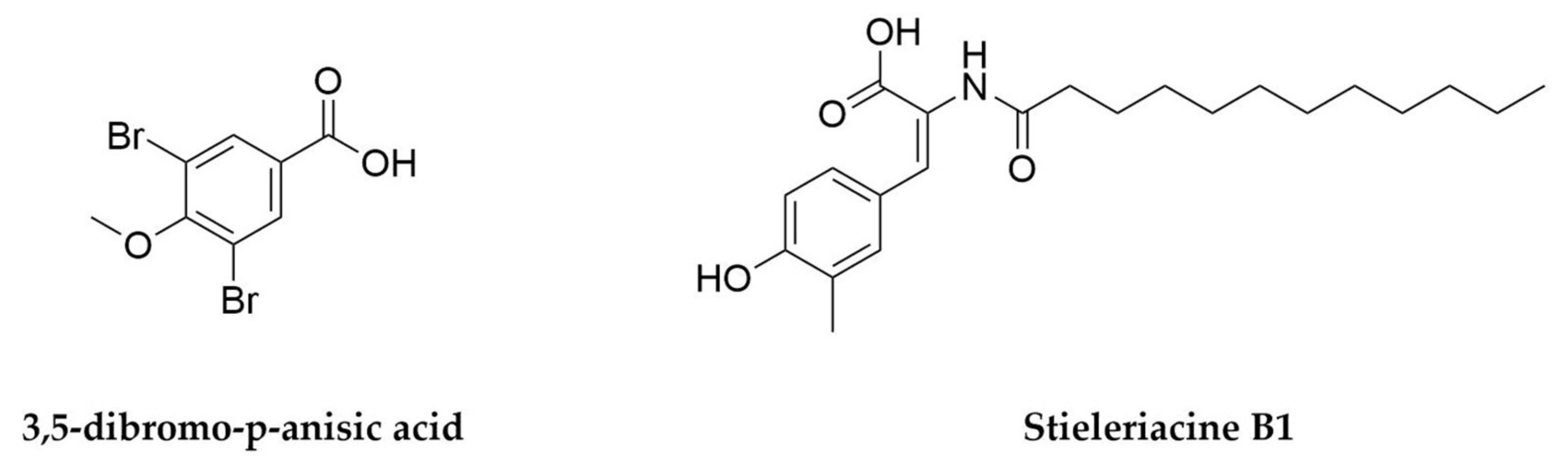
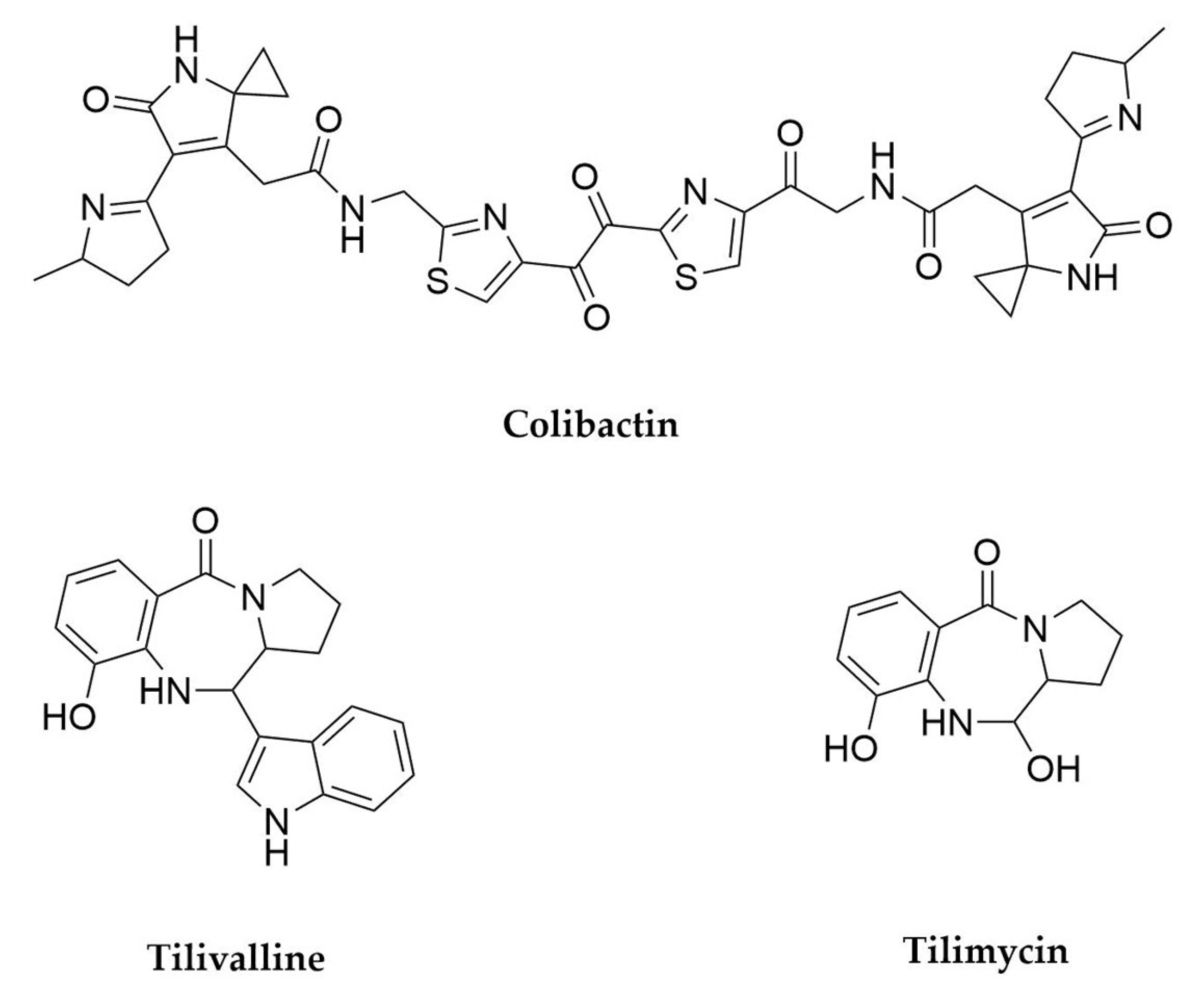

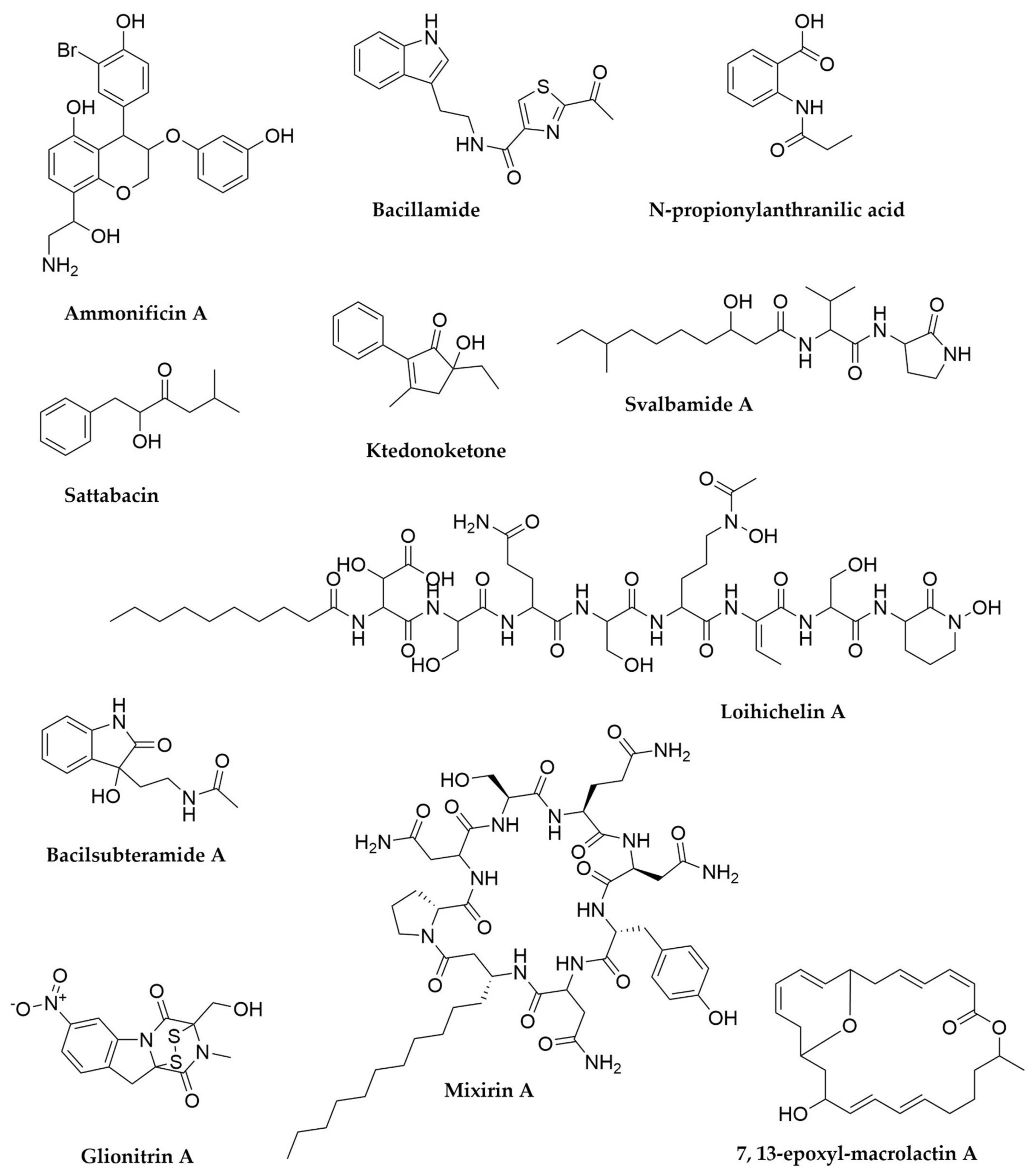
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Santos-Aberturas, J.; Vior, N.M. Beyond Soil-Dwelling Actinobacteria: Fantastic Antibiotics and Where to Find Them. Antibiotics 2022, 11, 195. https://doi.org/10.3390/antibiotics11020195
Santos-Aberturas J, Vior NM. Beyond Soil-Dwelling Actinobacteria: Fantastic Antibiotics and Where to Find Them. Antibiotics. 2022; 11(2):195. https://doi.org/10.3390/antibiotics11020195
Chicago/Turabian StyleSantos-Aberturas, Javier, and Natalia M. Vior. 2022. "Beyond Soil-Dwelling Actinobacteria: Fantastic Antibiotics and Where to Find Them" Antibiotics 11, no. 2: 195. https://doi.org/10.3390/antibiotics11020195
APA StyleSantos-Aberturas, J., & Vior, N. M. (2022). Beyond Soil-Dwelling Actinobacteria: Fantastic Antibiotics and Where to Find Them. Antibiotics, 11(2), 195. https://doi.org/10.3390/antibiotics11020195





