Tracing Antibiotic Resistance Genes along the Irrigation Water Chain to Chive: Does Tap or Surface Water Make a Difference?
Abstract
:1. Introduction
2. Results
2.1. Comparing Reservoir- to Tap Water-Irrigated Chive
2.2. Comparing Sprinkler Water to Corresponding Chives
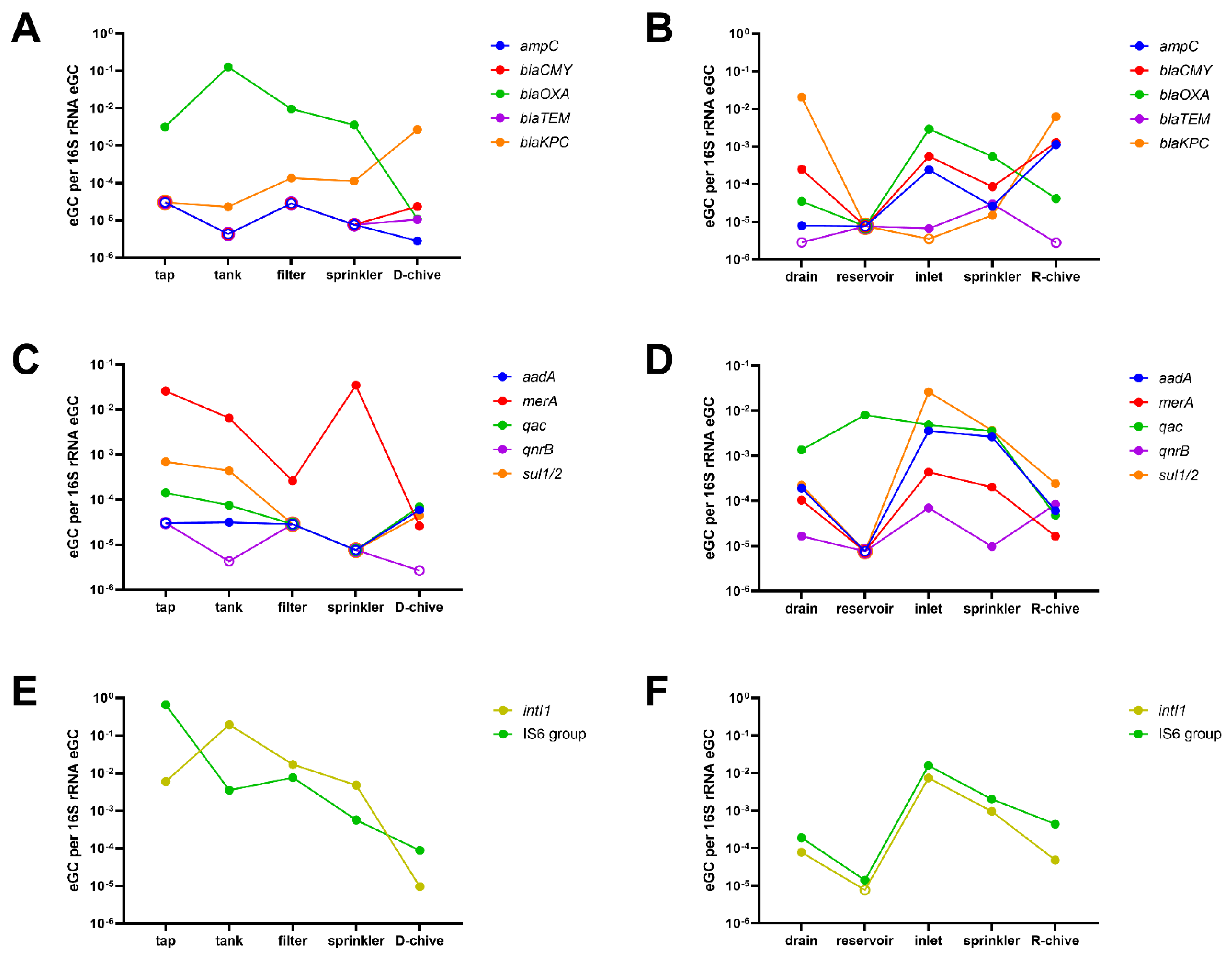
2.3. Relative ARG and MGE Abundance along the Irrigation Chains to Chive
3. Discussion
4. Materials and Methods
4.1. Experimental Field Trial Setup and Sampling
4.2. DNA Extraction and qPCR
4.3. Data Analysis
4.4. Statistics
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Centers for Disease Control and Prevention. Antibiotic Resistance Threats in the United States; U.S. Department of Health and Human Services, CDC: Atlanta, GA, USA, 2019. [Google Scholar]
- Mindlin, S.; Soina, V.; Petrova, M.; Gorlenko, Z.M. Isolation of antibiotic resistance bacterial strains from Eastern Siberia permafrost sediments. Russ. J. Genet. 2008, 44, 27–34. [Google Scholar] [CrossRef]
- Magiorakos, A.P.; Srinivasan, A.; Carey, R.B.; Carmeli, Y.; Falagas, M.E.; Giske, C.G.; Harbarth, S.; Hindler, J.F.; Kahlmeter, G.; Olsson-Liljequist, B. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: An international expert proposal for interim standard definitions for acquired resistance. Clin. Microbiol. Infect. 2012, 18, 268–281. [Google Scholar] [CrossRef] [Green Version]
- Guenther, S.; Ewers, C.; Wieler, L.H. Extended-spectrum beta-lactamases producing E. coli in wildlife, yet another form of environmental pollution? Front. Microbiol. 2011, 2, 246. [Google Scholar] [CrossRef] [Green Version]
- Neu, H. Penicillin-binding proteins and beta-lactamases: Their effects on the use of cephalosporins and other new beta-lactams. Curr. Clin. Top. Infect. Dis. 1987, 8, 37–61. [Google Scholar]
- Thomson, K.S.; Hayden, M.E.; Sanders, C.C.; Bradford, P.A. Detection of extended spectrum β-lactamases of Enterobacteriaceae in routine disk diffusion susceptibility tests. In Proceedings of the 92nd Annual Meeting of the American Society for Microbiology, New Orleans, LA, USA; 1992. [Google Scholar]
- Thanner, S.; Drissner, D.; Walsh, F. Antimicrobial resistance in agriculture. MBio 2016, 7, e02227-15. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vogt, D.; Overesch, G.; Endimiani, A.; Collaud, A.; Thomann, A.; Perreten, V. Occurrence and genetic characteristics of third-generation cephalosporin-resistant Escherichia coli in Swiss retail meat. Microb. Drug Resist. 2014, 20, 485–494. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pesavento, G.; Calonico, C.; Ducci, B.; Magnanini, A.; Lo Nostro, A. Prevalence and antibiotic resistance of Enterococcus spp. isolated from retail cheese, ready-to-eat salads, ham, and raw meat. Food Microbiol. 2014, 41, 1–7. [Google Scholar] [CrossRef]
- Geser, N.; Stephan, R.; Hächler, H. Occurrence and characteristics of extended-spectrum β-lactamase (ESBL) producing Enterobacteriaceae in food producing animals, minced meat and raw milk. BMC Vet. Res. 2012, 8, 21. [Google Scholar] [CrossRef] [Green Version]
- Blau, K.; Bettermann, A.; Jechalke, S.; Fornefeld, E.; Vanrobaeys, Y.; Stalder, T.; Top, E.M.; Smalla, K. The Transferable Resistome of Produce. MBio 2018, 9, e01300-18. [Google Scholar] [CrossRef] [Green Version]
- Al-Kharousi, Z.S.; Guizani, N.; Al-Sadi, A.M.; Al-Bulushi, I.M. Antibiotic resistance of Enterobacteriaceae isolated from fresh fruits and vegetables and characterization of their AmpC β-lactamases. J. Food Prot. 2019, 82, 1857–1863. [Google Scholar] [CrossRef] [PubMed]
- Drissner, D.; Zuercher, U. Microbial safety of fresh fruits and vegetables. In Encyclopedia of Food Safety; Motarjemi, Y., Todd, E., Moy, G., Eds.; Elsevier: Oxford, UK, 2014; Volume 3. [Google Scholar]
- Gekenidis, M.-T.; Rigotti, S.; Hummerjohann, J.; Walsh, F.; Drissner, D. Long-Term Persistence of blaCTX-M-15 in Soil and Lettuce after Introducing Extended-Spectrum β-Lactamase (ESBL)-Producing Escherichia coli via Manure or Water. Microorganisms 2020, 8, 1646. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.-H.; Qiao, M.; Su, J.-Q.; Chen, Z.; Zhou, X.; Zhu, Y.-G. High throughput profiling of antibiotic resistance genes in urban park soils with reclaimed water irrigation. Environ. Sci. Technol. 2014, 48, 9079–9085. [Google Scholar] [CrossRef] [PubMed]
- Martinez, J.L. General principles of antibiotic resistance in bacteria. Drug Discov. Today Technol. 2014, 11, 33–39. [Google Scholar] [CrossRef]
- De la Cruz, F. Horizontal Gene Transfer: Methods and Protocols; Springer: Berlin, Germany, 2020. [Google Scholar] [CrossRef] [Green Version]
- Ensink, J.H.J.; Mahmood, T.; Dalsgaard, A. Wastewater-irrigated vegetables: Market handling versus irrigation water quality. Trop. Med. Int. Health 2007, 12, 2–7. [Google Scholar] [CrossRef]
- Maimon, A.; Tal, A.; Friedler, E.; Gross, A. Safe on-site reuse of greywater for irrigation-a critical review of current guidelines. Environ. Sci. Technol. 2010, 44, 3213–3220. [Google Scholar] [CrossRef]
- Uyttendaele, M.; Jaykus, L.A.; Amoah, P.; Chiodini, A.; Cunliffe, D.; Jacxsens, L.; Holvoet, K.; Korsten, L.; Lau, M.; McClure, P. Microbial hazards in irrigation water: Standards, norms, and testing to manage use of water in fresh produce primary production. Compr. Rev. Food Sci. Food Saf. 2015, 14, 336–356. [Google Scholar] [CrossRef]
- Araújo, S.; Silva, I.A.T.; Tacão, M.; Patinha, C.; Alves, A.; Henriques, I. Characterization of antibiotic resistant and pathogenic Escherichia coli in irrigation water and vegetables in household farms. Int. J. Food Microbiol. 2017, 257, 192–200. [Google Scholar] [CrossRef]
- Zhu, Y.G.; Johnson, T.A.; Su, J.Q.; Qiao, M.; Guo, G.X.; Stedtfeld, R.D.; Hashsham, S.A.; Tiedje, J.M. Diverse and abundant antibiotic resistance genes in Chinese swine farms. Proc. Natl. Acad. Sci. USA 2013, 110, 3435–3440. [Google Scholar] [CrossRef] [Green Version]
- Panesso, D.; Abadía-Patiño, L.; Vanegas, N.; Reynolds, P.E.; Courvalin, P.; Arias, C.A. Transcriptional analysis of the vanC cluster from Enterococcus gallinarum strains with constitutive and inducible vancomycin resistance. Antimicrob. Agents Chemother. 2005, 49, 1060–1066. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alegbeleye, O.O.; Singleton, I.; Sant’Ana, A.S. Sources and contamination routes of microbial pathogens to fresh produce during field cultivation: A review. Food Microbiol. 2018, 73, 177–208. [Google Scholar] [CrossRef] [PubMed]
- Machado-Moreira, B.; Richards, K.; Brennan, F.; Abram, F.; Burgess, C.M. Microbial contamination of fresh produce: What, where, and how? Compr. Rev. Food Sci. Food Saf. 2019, 18, 1727–1750. [Google Scholar] [CrossRef] [Green Version]
- Gekenidis, M.T.; Schoner, U.; von Ah, U.; Schmelcher, M.; Walsh, F.; Drissner, D. Tracing back multidrug-resistant bacteria in fresh herb production: From chive to source through the irrigation water chain. FEMS Microbiol. Ecol. 2018, 94, fiy149. [Google Scholar] [CrossRef]
- Cuzon, G.; Naas, T.; Nordmann, P. KPC carbapenemases: What is at stake in clinical microbiology? Pathol. Biol. 2010, 58, 39–45. [Google Scholar] [CrossRef] [PubMed]
- Holvoet, K.; Sampers, I.; Callens, B.; Dewulf, J.; Uyttendaele, M. Moderate prevalence of antimicrobial resistance in Escherichia coli isolates from lettuce, irrigation water, and soil. Appl. Environ. Microbiol. 2013, 79, 6677–6683. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Njage, P.M.; Buys, E.M. Pathogenic and commensal Escherichia coli from irrigation water show potential in transmission of extended spectrum and AmpC beta-lactamases determinants to isolates from lettuce. Microb. Biotechnol. 2014, 8, 462–473. [Google Scholar] [CrossRef]
- Hu, Q.; Zhang, X.-X.; Jia, S.; Huang, K.; Tang, J.; Shi, P.; Ye, L.; Ren, H. Metagenomic insights into ultraviolet disinfection effects on antibiotic resistome in biologically treated wastewater. Water Res. 2016, 101, 309–317. [Google Scholar] [CrossRef]
- Fatta-Kassinos, D.; Dionysiou, D.D.; Kümmerer, K. Advanced Treatment Technologies for Urban Wastewater Reuse; Springer: Berlin, Germany, 2015. [Google Scholar] [CrossRef]
- Wallmann, L.; Krampe, J.; Lahnsteiner, J.; Radu, E.; van Rensburg, P.; Slipko, K.; Wögerbauer, M.; Kreuzinger, N. Fate and persistence of antibiotic-resistant bacteria and genes through a multi-barrier treatment facility for direct potable reuse. J. Water Reuse Desal. 2021. [Google Scholar] [CrossRef]
- Di Cesare, A.; Eckert, E.M.; Rogora, M.; Corno, G. Rainfall increases the abundance of antibiotic resistance genes within a riverine microbial community. Environ. Pollut. 2017, 226, 473–478. [Google Scholar] [CrossRef]
- Zhang, S.; Pang, S.; Wang, P.; Wang, C.; Han, N.; Liu, B.; Han, B.; Li, Y.; Anim-Larbi, K. Antibiotic concentration and antibiotic-resistant bacteria in two shallow urban lakes after stormwater event. Environ. Sci. Pollut. Res. 2016, 23, 9984–9992. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Han, M.; Li, E.; Liu, X.; Wei, H.; Yang, C.; Lu, S.; Ning, K. Distribution of antibiotic resistance genes in an agriculturally disturbed lake in China: Their links with microbial communities, antibiotics, and water quality. J. Hazard. Mater. 2020, 393, 122426. [Google Scholar] [CrossRef] [PubMed]
- Di Cesare, A.; Eckert, E.M.; Teruggi, A.; Fontaneto, D.; Bertoni, R.; Callieri, C.; Corno, G. Constitutive presence of antibiotic resistance genes within the bacterial community of a large subalpine lake. Mol. Ecol. 2015, 24, 3888–3900. [Google Scholar] [CrossRef] [PubMed]
- Andronache, C. Estimated variability of below-cloud aerosol removal by rainfall for observed aerosol size distributions. Atmos. Chem. Phys. 2003, 3, 131–143. [Google Scholar] [CrossRef] [Green Version]
- Bio Suisse Standards for the Production, Processing and Marketing of ‘bud’ Products; Association of Swiss Organic Agriculture Organisations: Basel, Switzerland, 2015.
- Schmittgen, T.D.; Livak, K.J. Analyzing real-time PCR data by the comparative CT method. Nat. Prot. 2008, 3, 1101. [Google Scholar] [CrossRef] [PubMed]
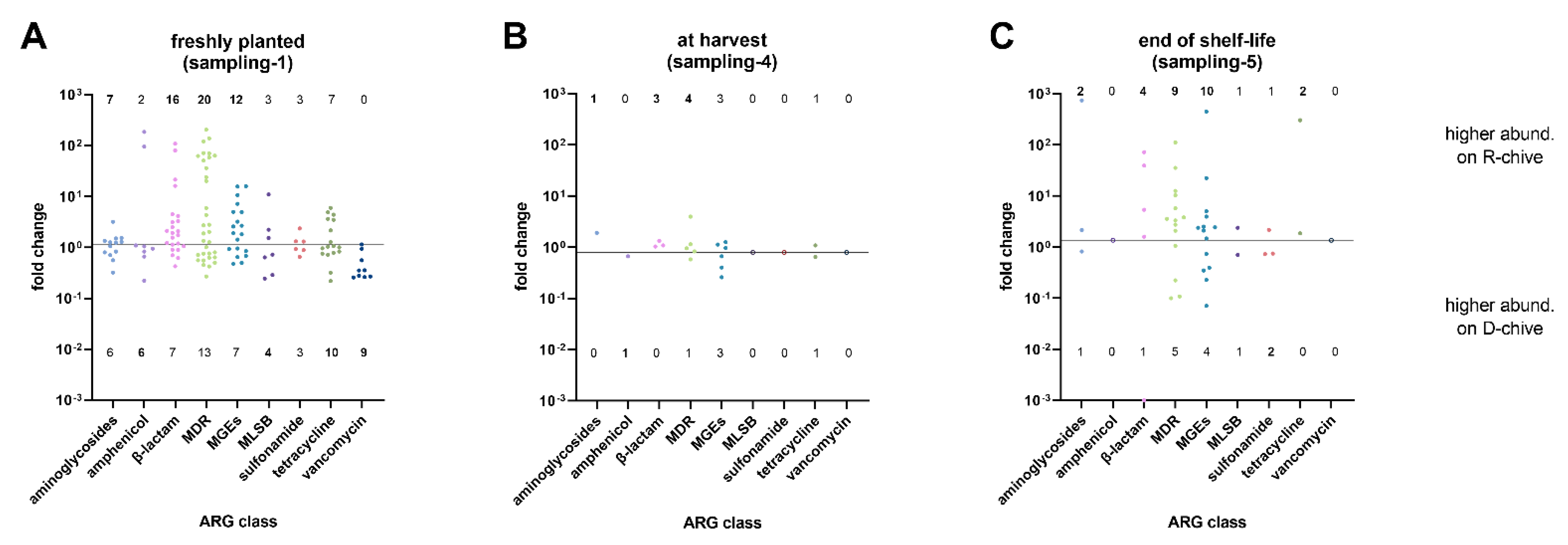

| Sampling-1 | Sampling-4 | Sampling-5 | ||||
|---|---|---|---|---|---|---|
| R-chive | D-chive | R-chive | D-chive | R-chive | D-chive | |
| aminoglycosides | 7 | 6 | 1 | 0 | 2 | 1 |
| amphenicol | 2 | 6 | 0 | 1 | 0 | 0 |
| β-lactam | 16 | 7 | 3 | 0 | 4 | 1 |
| MDR | 20 | 13 | 4 | 1 | 9 | 5 |
| MGE | 12 | 7 | 3 | 3 | 10 | 4 |
| MLSB | 3 | 4 | 0 | 0 | 1 | 1 |
| sulfonamide | 3 | 3 | 0 | 0 | 1 | 2 |
| tetracycline | 7 | 10 | 1 | 1 | 2 | 0 |
| vancomycin | 0 | 9 | 0 | 0 | 0 | 0 |
| p-value(Fisher’s Exact Test) | 0.01302 (*) | 0.4605 (ns) | 0.8739 (ns) | |||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gekenidis, M.-T.; Walsh, F.; Drissner, D. Tracing Antibiotic Resistance Genes along the Irrigation Water Chain to Chive: Does Tap or Surface Water Make a Difference? Antibiotics 2021, 10, 1100. https://doi.org/10.3390/antibiotics10091100
Gekenidis M-T, Walsh F, Drissner D. Tracing Antibiotic Resistance Genes along the Irrigation Water Chain to Chive: Does Tap or Surface Water Make a Difference? Antibiotics. 2021; 10(9):1100. https://doi.org/10.3390/antibiotics10091100
Chicago/Turabian StyleGekenidis, Maria-Theresia, Fiona Walsh, and David Drissner. 2021. "Tracing Antibiotic Resistance Genes along the Irrigation Water Chain to Chive: Does Tap or Surface Water Make a Difference?" Antibiotics 10, no. 9: 1100. https://doi.org/10.3390/antibiotics10091100
APA StyleGekenidis, M.-T., Walsh, F., & Drissner, D. (2021). Tracing Antibiotic Resistance Genes along the Irrigation Water Chain to Chive: Does Tap or Surface Water Make a Difference? Antibiotics, 10(9), 1100. https://doi.org/10.3390/antibiotics10091100






