Transmission of Similar Mcr-1 Carrying Plasmids among Different Escherichia coli Lineages Isolated from Livestock and the Farmer
Abstract
1. Introduction
2. Results and Discussion
2.1. Antimicrobial Susceptibility Testing
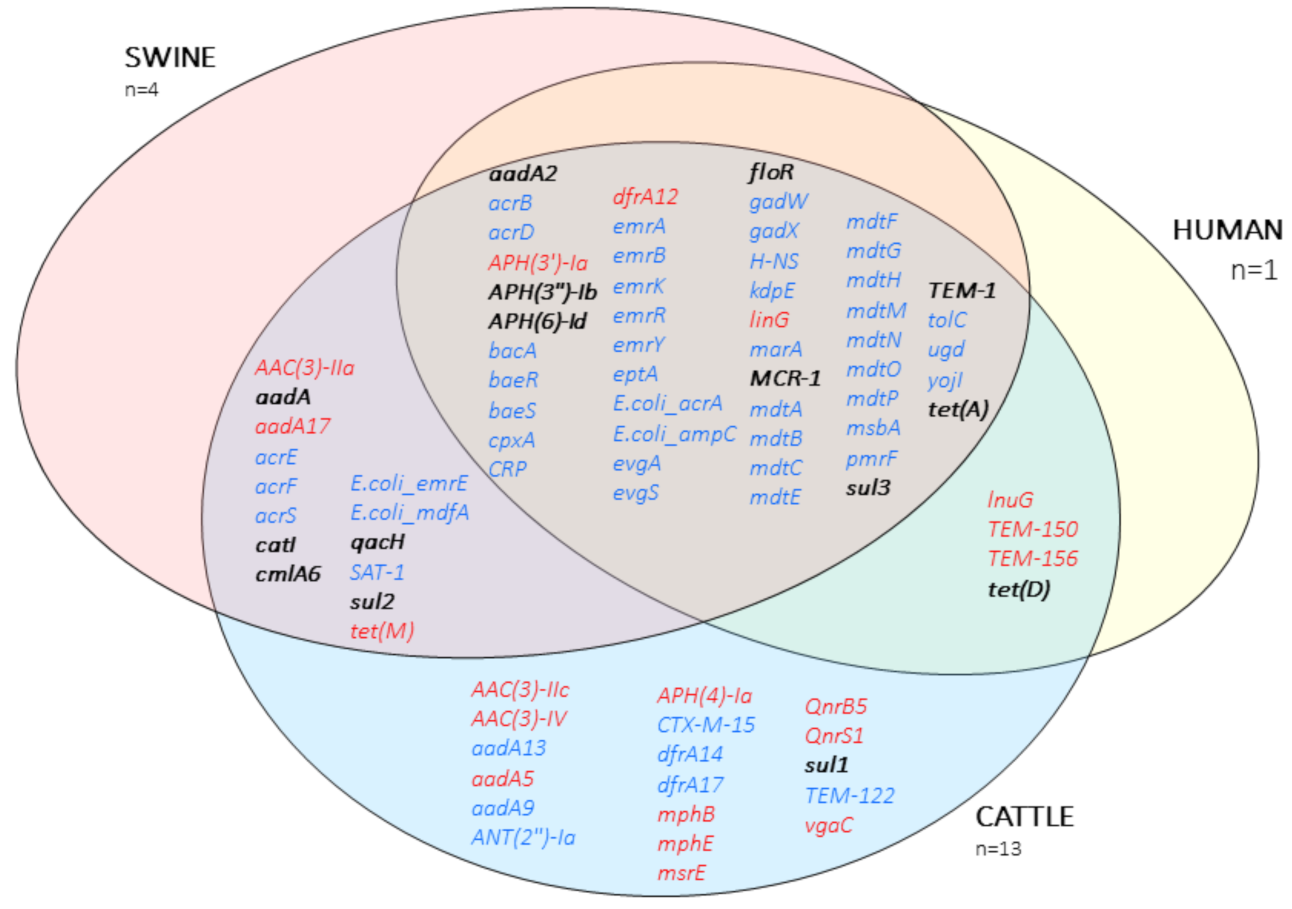
2.2. Antibiotic Resistance Genes
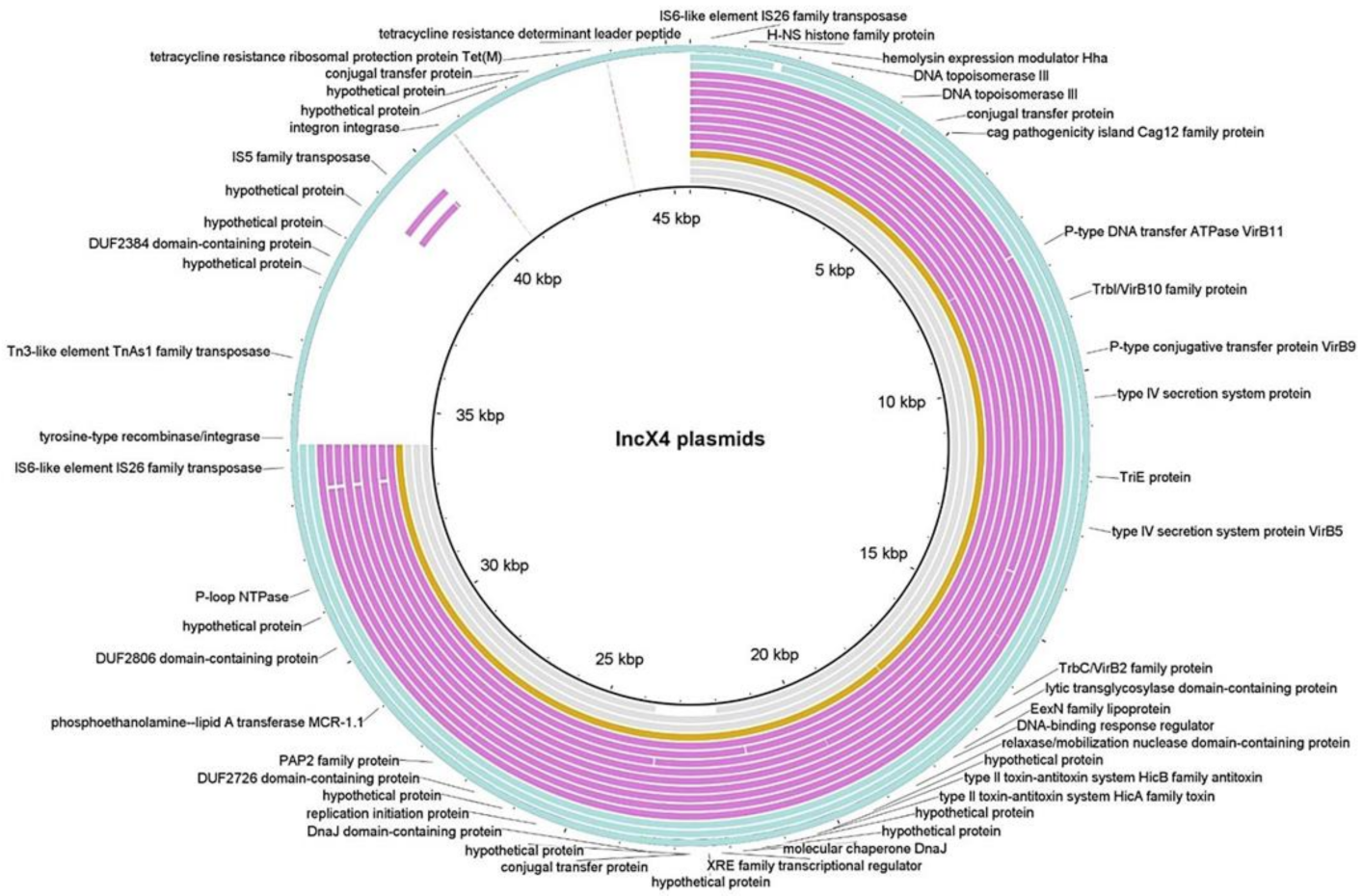
2.3. Genetic Context of mcr-1
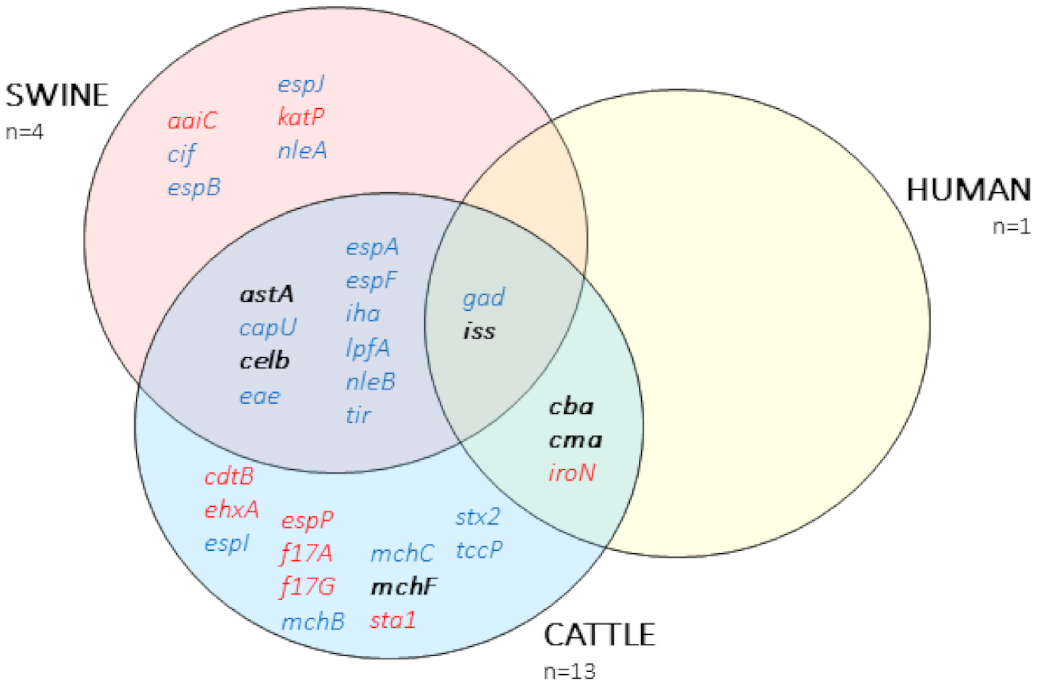
2.4. Virulence Factors
3. Discussion
4. Materials and Methods
4.1. Study Design
4.2. Antimicrobial Susceptibility Testing
4.3. Whole Genome Sequencing and Data Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Koyama, Y.; Kurosasa, A.; Tsuchiya, A.; Takakuta, K. A new antibiotic “colistin” produced by spore-forming soil bacteria. J. Antibiot. 1950, 3, 457–458. [Google Scholar]
- Catry, B.; Cavaleri, M.; Baptiste, K.; Grave, K.; Grein, K.; Holm, A.; Jukes, H.; Liebana, E.; Navas, A.L.; Mackay, D.; et al. Use of colistin-containing products within the European Union and European Economic Area (EU/EEA): Development of resistance in animals and possible impact on human and animal health. Int. J. Antimicrob. Agents 2015, 46, 297–306. [Google Scholar] [CrossRef] [PubMed]
- Moreno, M.A. Survey of quantitative antimicrobial consumption per production stage in farrow-to-finish pig farms in Spain. Vet. Rec. Open 2014, 1, e000002. [Google Scholar] [CrossRef] [PubMed]
- Van Rennings, L.; von Münchhausen, C.; Ottilie, H.; Hartmann, M.; Merle, R.; Honscha, W.; Käsbohrer, A.; Kreienbrock, L. Cross-Sectional Study on Antibiotic Usage in Pigs in Germany. PLoS ONE 2015, 10, e0119114. [Google Scholar] [CrossRef]
- Sjölund, M.; Backhans, A.; Greko, C.; Emanuelson, U.; Lindberg, A. Antimicrobial usage in 60 Swedish farrow-to-finish pig herds. Prev. Vet. Med. 2015, 121, 257–264. [Google Scholar] [CrossRef] [PubMed]
- Callens, B.; Persoons, D.; Maes, D.; Laanen, M.; Postma, M.; Boyen, F.; Haesebrouck, F.; Butaye, P.; Catry, B.; Dewulf, J. Prophylactic and metaphylactic antimicrobial use in Belgian fattening pig herds. Prev. Vet. Med. 2012, 106, 53–62. [Google Scholar] [CrossRef] [PubMed]
- European Medicines Agency. European Medicines Agency Sales of Veterinary Antimicrobial Agents in 29 European Countries in 2014; European Medicines Agency: London, UK, 2014. [Google Scholar]
- Informe JIACRA España. Primer Análisis Integrado del Consumo de Antibióticos y su Relación con la Aparición de Resistencia. 2018. Available online: https://resistenciaantibioticos.es/es/system/files/field/files/informe_jiacra-espana.pdf?file=1&type=node&id=410&force=0 (accessed on 17 March 2021).
- Liu, Y.-Y.; Wang, Y.; Walsh, T.R.; Yi, L.-X.; Zhang, R.; Spencer, J.; Doi, Y.; Tian, G.; Dong, B.; Huang, X.; et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: A microbiological and molecular biological study. Lancet Infect. Dis. 2016, 16, 161–168. [Google Scholar] [CrossRef]
- Xavier, B.B.; Lammens, C.; Ruhal, R.; Kumar-Singh, S.; Butaye, P.; Goossens, H.; Malhotra-Kumar, S. Identification of a novel plasmid-mediated colistin-resistance gene, mcr-2, in Escherichia coli, Belgium, June 2016. Euro Surveill. 2016, 21, 30280. [Google Scholar] [CrossRef]
- Yin, W.; Li, H.; Shen, Y.; Liu, Z.; Wang, S.; Shen, Z.; Zhang, R.; Walsh, T.R.; Shen, J.; Wang, Y. Novel Plasmid-Mediated Colistin Resistance Gene mcr-3 in Escherichia coli. MBio 2017, 8. [Google Scholar] [CrossRef]
- Borowiak, M.; Fischer, J.; Hammerl, J.A.; Hendriksen, R.S.; Szabo, I.; Malorny, B. Identification of a novel transposon-associated phosphoethanolamine transferase gene, mcr-5, conferring colistin resistance in d-tartrate fermenting Salmonella enterica subsp. enterica serovar Paratyphi B. J. Antimicrob. Chemother. 2017, 72, 3317–3324. [Google Scholar] [CrossRef]
- Carattoli, A.; Villa, L.; Feudi, C.; Curcio, L.; Orsini, S.; Luppi, A.; Pezzotti, G.; Magistrali, C.F. Novel plasmid-mediated colistin resistance mcr-4 gene in Salmonella and Escherichia coli, Italy 2013, Spain and Belgium, 2015 to 2016. Euro Surveill. 2017, 22, 30589. [Google Scholar] [CrossRef]
- Wang, R.; van Dorp, L.; Shaw, L.P.; Bradley, P.; Wang, Q.; Wang, X.; Jin, L.; Zhang, Q.; Liu, Y.; Rieux, A.; et al. The global distribution and spread of the mobilized colistin resistance gene mcr-1. Nat. Commun. 2018, 9, 1179. [Google Scholar] [CrossRef]
- Hernández, M.; Iglesias, M.R.; Rodríguez-Lázaro, D.; Gallardo, A.; Quijada, N.; Miguela-Villoldo, P.; Campos, M.J.; Píriz, S.; López-Orozco, G.; de Frutos, C.; et al. Co-occurrence of colistin-resistance genes mcr-1 and mcr-3 among multidrug-resistant Escherichia coli isolated from cattle, Spain, September 2015. Euro Surveill. 2017, 22, 30586. [Google Scholar] [CrossRef] [PubMed]
- Partridge, S.R.; Kwong, S.M.; Firth, N.; Jensen, S.O. Mobile genetic elements associated with antimicrobial resistance. Clin. Microbiol. Rev. 2018, 31. [Google Scholar] [CrossRef]
- Bai, F.; Li, X.; Niu, B.; Zhang, Z.; Malakar, P.K.; Liu, H.; Pan, Y.; Zhao, Y. A mcr-1-Carrying Conjugative IncX4 Plasmid in Colistin-Resistant Escherichia coli ST278 Strain Isolated from Dairy Cow Feces in Shanghai, China. Front. Microbiol. 2018, 9, 1–9. [Google Scholar] [CrossRef]
- Zurfluh, K.; Nüesch-Inderbinen, M.; Klumpp, J.; Poirel, L.; Nordmann, P.; Stephan, R. Key features of mcr-1-bearing plasmids from Escherichia coli isolated from humans and food. Antimicrob. Resist. Infect. Control 2017, 6, 1–6. [Google Scholar] [CrossRef]
- Wu, R.; Yi, L.X.; Yu, L.F.; Wang, J.; Liu, Y.; Chen, X.; Lv, L.; Yang, J.; Liu, J.H. Fitness advantage of mcr-1-bearing IncI2 and IncX4 plasmids in vitro. Front. Microbiol. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Xie, M.; Zhang, J.; Yang, Z.; Liu, L.; Liu, X.; Zheng, Z.; Chan, E.W.C.; Chen, S. Genetic characterization of mcr-1-bearing plasmids to depict molecular mechanisms underlying dissemination of the colistin resistance determinant. J. Antimicrob. Chemother. 2017, 72, 393–401. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Fang, L.X.; Wu, Z.; Deng, H.; Yang, R.S.; Li, X.P.; Li, S.M.; Liao, X.P.; Feng, Y.; Liu, Y.H. Genetic Analysis of the IncX4 Plasmids: Implications for a Unique Pattern in the mcr-1 Acquisition. Sci. Rep. 2017, 7, 424. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Ke, B.; Zhao, X.; Guo, Y.; Wang, W.; Wang, X.; Zhu, H. Antimicrobial resistance profile of mcr-1 positive clinical isolates of Escherichia coli in China from 2013 to 2016. Front. Microbiol. 2018, 9, 1–10. [Google Scholar] [CrossRef]
- Shen, Y.; Wu, Z.; Wang, Y.; Zhang, R.; Zhou, H.W.; Wang, S.; Lei, L.; Li, M.; Cai, J.; Tyrrell, J.; et al. Heterogeneous and flexible transmission of mcr-1 in hospital-associated Escherichia coli. MBio 2018, 9. [Google Scholar] [CrossRef]
- Dunn, S.J.; Connor, C.; McNally, A. The evolution and transmission of multi-drug resistant Escherichia coli and Klebsiella pneumoniae: The complexity of clones and plasmids. Curr. Opin. Microbiol. 2019, 51, 51–56. [Google Scholar] [CrossRef]
- San Millan, A. Evolution of Plasmid-Mediated Antibiotic Resistance in the Clinical Context. Trends Microbiol. 2018, 26, 978–985. [Google Scholar] [CrossRef]
- Ludden, C.; Raven, K.E.; Jamrozy, D.; Gouliouris, T.; Blane, B.; Coll, F.; de Goffau, M.; Naydenova, P.; Horner, C.; Hernandez-Garcia, J.; et al. One Health Genomic Surveillance of Escherichia coli Demonstrates Distinct Lineages and Mobile Genetic Elements in Isolates from Humans versus Livestock. MBio 2019, 10. [Google Scholar] [CrossRef] [PubMed]
- Saliu, E.-M.; Vahjen, W.; Zentek, J. Types and prevalence of extended–spectrum beta–lactamase producing Enterobacteriaceae in poultry. Anim. Heal. Res. Rev. 2017, 18, 46–57. [Google Scholar] [CrossRef] [PubMed]
- McDaniel, C.J.; Cardwell, D.M.; Moeller, R.B., Jr.; Gray, G.C. Humans and Cattle: A Review of Bovine Zoonoses. Vector Borne Zoonotic Dis. 2014, 14, 1. [Google Scholar] [CrossRef]
- Tomley, F.M.; Shirley, M.W. Livestock infectious diseases and zoonoses. Philos. Trans. R. Soc. Lond. B. Biol. Sci. 2009, 364, 2637–2642. [Google Scholar] [CrossRef] [PubMed]
- Fan, R.; Shao, K.; Yang, X.; Bai, X.; Fu, S.; Sun, H.; Xu, Y.; Wang, H.; Li, Q.; Hu, B.; et al. High prevalence of non-O157 Shiga toxin-producing Escherichia coli in beef cattle detected by combining four selective agars. BMC Microbiol. 2019, 19, 213. [Google Scholar] [CrossRef]
- Thomas, N.A.; Navarro-Garcia, F.; Jorge Blanco, M.; Robins-Browne, R.M.; Holt, K.E.; Ingle, D.J.; Hocking, D.M.; Yang, J.; Tauschek, M. Are Escherichia coli Pathotypes Still Relevant in the Era of Whole-Genome Sequencing? Front. Cell. Infect. Microbiol. 2016, 1, 141. [Google Scholar] [CrossRef]
- Hussein, H.S.; Bollinger, L.M. Prevalence of Shiga toxin-producing Escherichia coli in beef cattle. J. Food Prot. 2005, 68, 2224–2241. [Google Scholar] [CrossRef]
- Padola, N.L.; Etcheverría, A.I. Shiga toxin-producing Escherichia coli in human, cattle, and foods. Strategies for detection and control. Front. Cell. Infect. Microbiol. 2014, 4, 89. [Google Scholar] [CrossRef]
- Yasui, H.; Kurosawa, Y. Measurement of recombination frequencies between two homologous DNA segments embedded in a YAC vector. Gene 1993, 129, 135–139. [Google Scholar] [CrossRef]
- Komoda, Y.; Enomoto, M.; Tominaga, A. Large Inversion in Escherichia coli K-12 1485IN Between Inversely Oriented IS3 Elements Near lac and cdd. Genetics 1991, 129, 639–645. [Google Scholar] [CrossRef]
- Cantón, R.; Coque, T.M. The CTX-M β-lactamase pandemic. Curr. Opin. Microbiol. 2006, 9, 466–475. [Google Scholar] [CrossRef]
- Chong, Y.; Shimoda, S.; Shimono, N. Current epidemiology, genetic evolution and clinical impact of extended-spectrum β-lactamase-producing Escherichia coli and Klebsiella pneumoniae. Infect. Genet. Evol. 2018, 61, 185–188. [Google Scholar] [CrossRef] [PubMed]
- Bevan, E.R.; Jones, A.M.; Hawkey, P.M. Global epidemiology of CTX-M b-lactamases: Temporal and geographical shifts in genotype. J. Antimicrob. Chemother. 2017, 72, 2145–2155. [Google Scholar] [CrossRef] [PubMed]
- Boyd, D.A.; Tyler, S.; Christianson, S.; Mcgeer, A.; Muller, M.P.; Willey, B.M.; Bryce, E.; Gardam, M.; Nordmann, P.; Mulvey, M.R.; et al. Complete Nucleotide Sequence of a 92-Kilobase Plasmid Harboring the CTX-M-15 Extended-Spectrum Beta-Lactamase Involved in an Outbreak in Long-Term-Care Facilities in Toronto, Canada. Antimicrob. Agents Chemother. 2004, 48, 3758–3764. [Google Scholar] [CrossRef]
- Yasir, M.; Farman, M.; Shah, M.W.; Jiman-Fatani, A.A.; Othman, N.A.; Almasaudi, S.B.; Alawi, M.; Shakil, S.; Al-Abdullah, N.; Ismaeel, N.A.; et al. Genomic and antimicrobial resistance genes diversity in multidrug-resistant CTX-M-positive isolates of Escherichia coli at a health care facility in Jeddah. J. Infect. Public Health 2020, 13, 94–100. [Google Scholar] [CrossRef] [PubMed]
- Mehrad, B.; Clark, N.M.; Zhanel, G.G.; Lynch, J.P. Antimicrobial resistance in hospital-acquired gram-negative bacterial infections. Chest 2015, 147, 1413–1421. [Google Scholar] [CrossRef] [PubMed]
- Heininger, A.; Binder, M.; Schmidt, S.; Unertl, K.; Botzenhart, K.; Döring, G. PCR and blood culture for detection of Escherichia coli bacteremia in rats. J. Clin. Microbiol. 1999, 37, 2479–2482. [Google Scholar] [CrossRef] [PubMed]
- Rebelo, A.R.; Bortolaia, V.; Kjeldgaard, J.S.; Pedersen, S.K.; Leekitcharoenphon, P.; Hansen, I.M.; Guerra, B.; Malorny, B.; Borowiak, M.; Hammerl, J.A.; et al. Multiplex PCR for detection of plasmid-mediated colistin resistance determinants, mcr-1, mcr-2, mcr-3, mcr-4 and mcr-5 for surveillance purposes. Euro Surveill. 2018, 23. [Google Scholar] [CrossRef]
- Schwarz, S.; Silley, P.; Simjee, S.; Woodford, N.; van Duijkeren, E.; Johnson, A.P.; Gaastra, W. Assessing the antimicrobial susceptibility of bacteria obtained from animals. Vet. Microbiol. 2010, 141, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Porechop. Available online: https://github.com/rrwick/Porechop (accessed on 17 March 2021).
- Kolmogorov, M.; Yuan, J.; Lin, Y.; Pevzner, P.A. Assembly of long, error-prone reads using repeat graphs. Nat. Biotechnol. 2019, 37, 540–546. [Google Scholar] [CrossRef] [PubMed]
- Li, H. Minimap2: Pairwise alignment for nucleotide sequences. Bioinformatics 2018, 34, 3094–3100. [Google Scholar] [CrossRef] [PubMed]
- Vaser, R.; Sovic, I.; Nagarajan, N.; Sikic, M. Fast and accurate de novo genome assembly from long uncorrected reads. Genome Res. 2017, 27. [Google Scholar] [CrossRef]
- Abricate. Available online: https://github.com/tseemann/abricate (accessed on 17 March 2021).
- Carattoli, A.; Zankari, E.; García-Fernández, A.; Voldby Larsen, M.; Lund, O.; Villa, L.; Møller Aarestrup, F.; Hasman, H. In Silico Detection and Typing of Plasmids using PlasmidFinder and Plasmid Multilocus Sequence Typing. Antimicrob. Agents Chemother. 2014, 58, 3895–3903. [Google Scholar] [CrossRef]
- Jia, B.; Raphenya, A.R.; Alcock, B.; Waglechner, N.; Guo, P.; Tsang, K.K.; Lago, B.A.; Dave, B.M.; Pereira, S.; Sharma, A.N.; et al. CARD 2017: Expansion and model-centric curation of the comprehensive antibiotic resistance database. Nucleic Acids Res. 2017, 45, D566–D573. [Google Scholar] [CrossRef]
- Chen, L.; Zheng, D.; Liu, B.; Yang, J.; Jin, Q. VFDB 2016: Hierarchical and refined dataset for big data analysis—10 years on. Nucleic Acids Res. 2016, 44, D694–D697. [Google Scholar] [CrossRef]
- Siguier, P.; Perochon, J.; Lestrade, L.; Mahillon, J.; Chandler, M. ISfinder: The reference centre for bacterial insertion sequences. Nucleic Acids Res. 2006, 34, D32–D36. [Google Scholar] [CrossRef] [PubMed]
- Arndt, D.; Grant, J.R.; Marcu, A.; Sajed, T.; Pon, A.; Liang, Y.; Wishart, D.S. PHASTER: A better, faster version of the PHAST phage search tool. Nucleic Acids Res. 2016, 44. [Google Scholar] [CrossRef]
- Li, X.; Xie, Y.; Liu, M.; Tai, C.; Sun, J.; Deng, Z.; Ou, H.-Y. oriTfinder: A web-based tool for the identification of origin of transfers in DNA sequences of bacterial mobile genetic elements. Nucleic Acids Res. 2018, 46, 229–234. [Google Scholar] [CrossRef]
- Tatusova, T.; DiCuccio, M.; Badretdin, A.; Chetvernin, V.; Nawrocki, E.P.; Zaslavsky, L.; Lomsadze, A.; Pruitt, K.D.; Borodovsky, M.; Ostell, J. NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res. 2016, 44, 6614–6624. [Google Scholar] [CrossRef] [PubMed]
- Seemann, T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics 2014, 30, 2068–2069. [Google Scholar] [CrossRef] [PubMed]
- Seppey, M.; Manni, M.; Zdobnov, E.M. BUSCO: Assessing Genome Assembly and Annotation Completeness. Methods Mol. Biol. 2019, 1962, 227–245. [Google Scholar]
- Joensen, K.G.; Tetzschner, A.M.M.; Iguchi, A.; Aarestrup, F.M.; Scheutz, F. Rapid and Easy In Silico Serotyping of Escherichia coli Isolates by Use of Whole-Genome Sequencing Data. J. Clin. Microbiol. 2015, 53, 2410–2426. [Google Scholar] [CrossRef] [PubMed]
- Larsen, M.V.; Cosentino, S.; Rasmussen, S.; Friis, C.; Hasman, H.; Marvig, R.L.; Jelsbak, L.; Sicheritz-Ponten, T.; Ussery, D.W.; Aarestrup, F.M.; et al. Multilocus Sequence Typing of Total-Genome-Sequenced Bacteria. J. Clin. Microbiol. 2012, 50, 1355–1361. [Google Scholar] [CrossRef]
- Kaas, R.S.; Leekitcharoenphon, P.; Aarestrup, F.M.; Lund, O. Solving the Problem of Comparing Whole Bacterial Genomes across Different Sequencing Platforms. PLoS ONE 2014, 9, e104984. [Google Scholar] [CrossRef] [PubMed]
- Beghain, J.; Bridier-Nahmias, A.; Le Nagard, H.; Denamur, E.; Clermont, O. ClermonTyping: An easy-to-use and accurate in silico method for Escherichia genus strain phylotyping. Microb. Genom. 2018, 4. [Google Scholar] [CrossRef]
- Wick, R.R.; Schultz, M.B.; Zobel, J.; Holt, K.E. Bandage: Interactive visualization of de novo genome assemblies. Bioinformatics 2015, 31, 3350–3352. [Google Scholar] [CrossRef] [PubMed]
- Alikhan, N.F.; Petty, N.K.; Ben Zakour, N.L.; Beatson, S.A. BLAST Ring Image Generator (BRIG): Simple prokaryote genome comparisons. BMC Genomics 2011, 12, 402. [Google Scholar] [CrossRef] [PubMed]
| Host | Isolate | Phyl. | MLST | mcr-1 | mcr-1 loc. | Pl. GC% | Pl. Size (bp) | mcr-1 Genomic Context |
|---|---|---|---|---|---|---|---|---|
| Human | Farmer | A | ST398 | yes | IncX4 | 41.9 | 33,270 | mcr-1-pap2 |
| Swine | P2_16 | B1 | ST5229 | Yes | Chr. | - | - | mcr-1-pap2 |
| P1_10 | B1 | ST20 | yes | IncX4 | 44.1 | 45,441 | mcr-1-pap2 | |
| P2_2 | A | ST10 | yes | IncX4 | 42.5 | 35,296 | mcr-1-pap2-ΔISApl1 | |
| P2_27 | A | ST10 | yes | IncX4 | 42.5 | 35,326 | mcr-1-pap2-ΔISApl1 | |
| Bovine | 15B_27 | A | ST6395 | yes | IncX4 | 42.2 | 34,706 | mcr-1-pap2-ΔISApl1 |
| 15B_17 | A | ST6395 | yes | IncX4 | 42.2 | 34,758 | mcr-1-pap2-ΔISApl1 | |
| 15B_4 | A | ST6395 | yes | IncX4 | 41.8 | 33,577 | mcr-1-pap2-ΔISApl1 | |
| 14_24 | B1 | ST224 | yes | IncX4 | 41.8 | 33,557 | mcr-1-pap2-ΔISApl1 | |
| V7_16 | A | ST206 | yes | IncX4 | 42.2 | 34,618 | mcr-1-pap2-ΔISApl1 | |
| 15A_16 | A | ST43 | yes | IncX4 | 41.8 | 33,242 | mcr-1-pap2 | |
| 14_20 | B1 | ST224 | yes | IncX4 | 41.8 | 33,283 | mcr-1-pap2 | |
| 14_4 | B1 | ST2539 | yes | IncX4 | 41.8 | 33,262 | mcr-1-pap2 | |
| V7_18 | B1 | ST224 | yes | IncX4 | 41.9 | 33,268 | mcr-1-pap2 | |
| 15B_22 15B_22 | A A | ST1638 ST1638 | yes yes | IncX4 IncHI2 | 41.9 45.4 | 33,268 234,156 | mcr-1-pap2 ISApl1-mcr-1-pap2 | |
| 15A_1 | A | ST1431 | yes | IncI2 | 42.5 | 61,766 | ISApl1-mcr-1-pap2 | |
| 15B_13 | B1 | ST101 | Yes | Chr. | - | - | ISApl1-mcr-1-pap2-ISApl1 | |
| 15A_11 | B1 | ST4981 | no | - | - | - | - |
| Isolate ID | Coverage (%) | Identity (%) |
|---|---|---|
| P1_10 | 74 | 99.99 |
| P2_2 | 94 | 99.87 |
| P2_27 | 94 | 100 |
| V7_16 | 95 | 99.97 |
| 15B_17 | 95 | 99.99 |
| 15B_27 | 95 | 100 |
| 15B_4 | 98 | 99.97 |
| 14_24 | 98 | 99.99 |
| 14_4 | 99 | 99.96 |
| V7_18 | 99 | 99.96 |
| 15A_16 | 99 | 99.97 |
| 14_20 | 99 | 99.97 |
| 15B_22 | 99 | 99.97 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Viñes, J.; Cuscó, A.; Napp, S.; Alvarez, J.; Saez-Llorente, J.L.; Rosàs-Rodoreda, M.; Francino, O.; Migura-Garcia, L. Transmission of Similar Mcr-1 Carrying Plasmids among Different Escherichia coli Lineages Isolated from Livestock and the Farmer. Antibiotics 2021, 10, 313. https://doi.org/10.3390/antibiotics10030313
Viñes J, Cuscó A, Napp S, Alvarez J, Saez-Llorente JL, Rosàs-Rodoreda M, Francino O, Migura-Garcia L. Transmission of Similar Mcr-1 Carrying Plasmids among Different Escherichia coli Lineages Isolated from Livestock and the Farmer. Antibiotics. 2021; 10(3):313. https://doi.org/10.3390/antibiotics10030313
Chicago/Turabian StyleViñes, Joaquim, Anna Cuscó, Sebastian Napp, Julio Alvarez, Jose Luis Saez-Llorente, Montserrat Rosàs-Rodoreda, Olga Francino, and Lourdes Migura-Garcia. 2021. "Transmission of Similar Mcr-1 Carrying Plasmids among Different Escherichia coli Lineages Isolated from Livestock and the Farmer" Antibiotics 10, no. 3: 313. https://doi.org/10.3390/antibiotics10030313
APA StyleViñes, J., Cuscó, A., Napp, S., Alvarez, J., Saez-Llorente, J. L., Rosàs-Rodoreda, M., Francino, O., & Migura-Garcia, L. (2021). Transmission of Similar Mcr-1 Carrying Plasmids among Different Escherichia coli Lineages Isolated from Livestock and the Farmer. Antibiotics, 10(3), 313. https://doi.org/10.3390/antibiotics10030313







