Detection of Breast Cancer Cells Using Acoustics Aptasensor Specific to HER2 Receptors
Abstract
1. Introduction
2. Materials and Methods
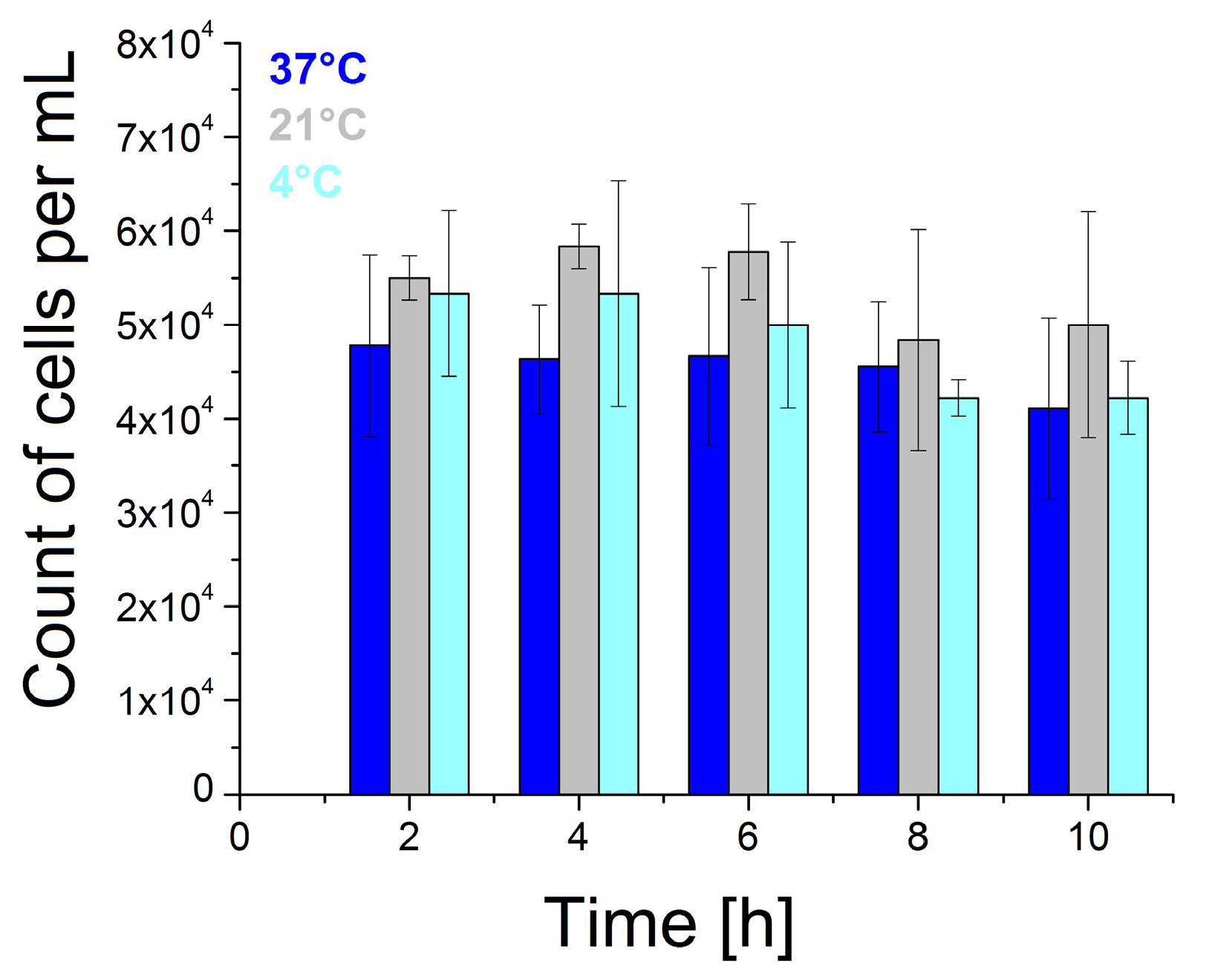
2.1. Preparation of the Cells and Their Viability
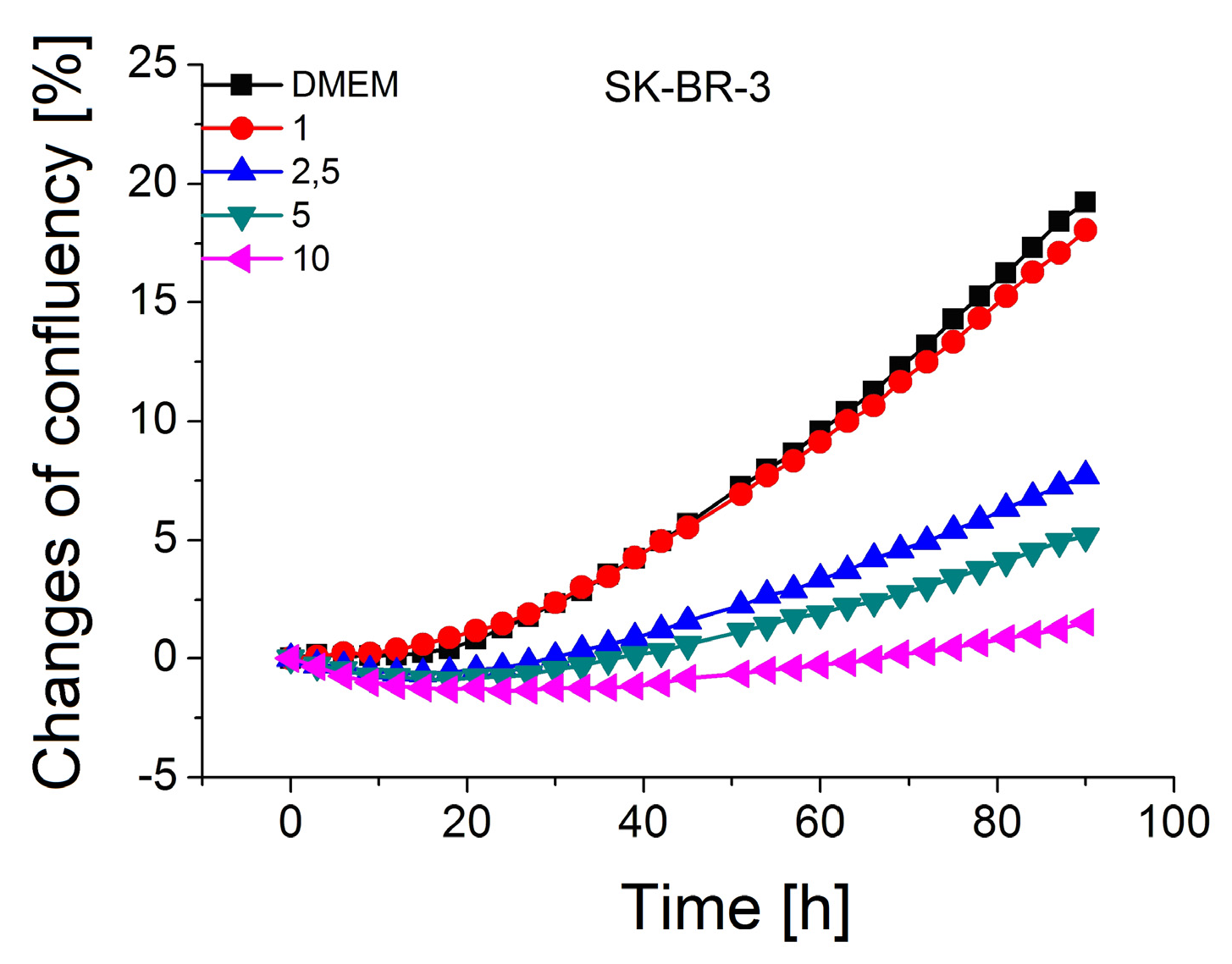
2.2. Cell Proliferation Study at Presence of Gold Nanoparticles
2.3. Thickness Shear Mode Method
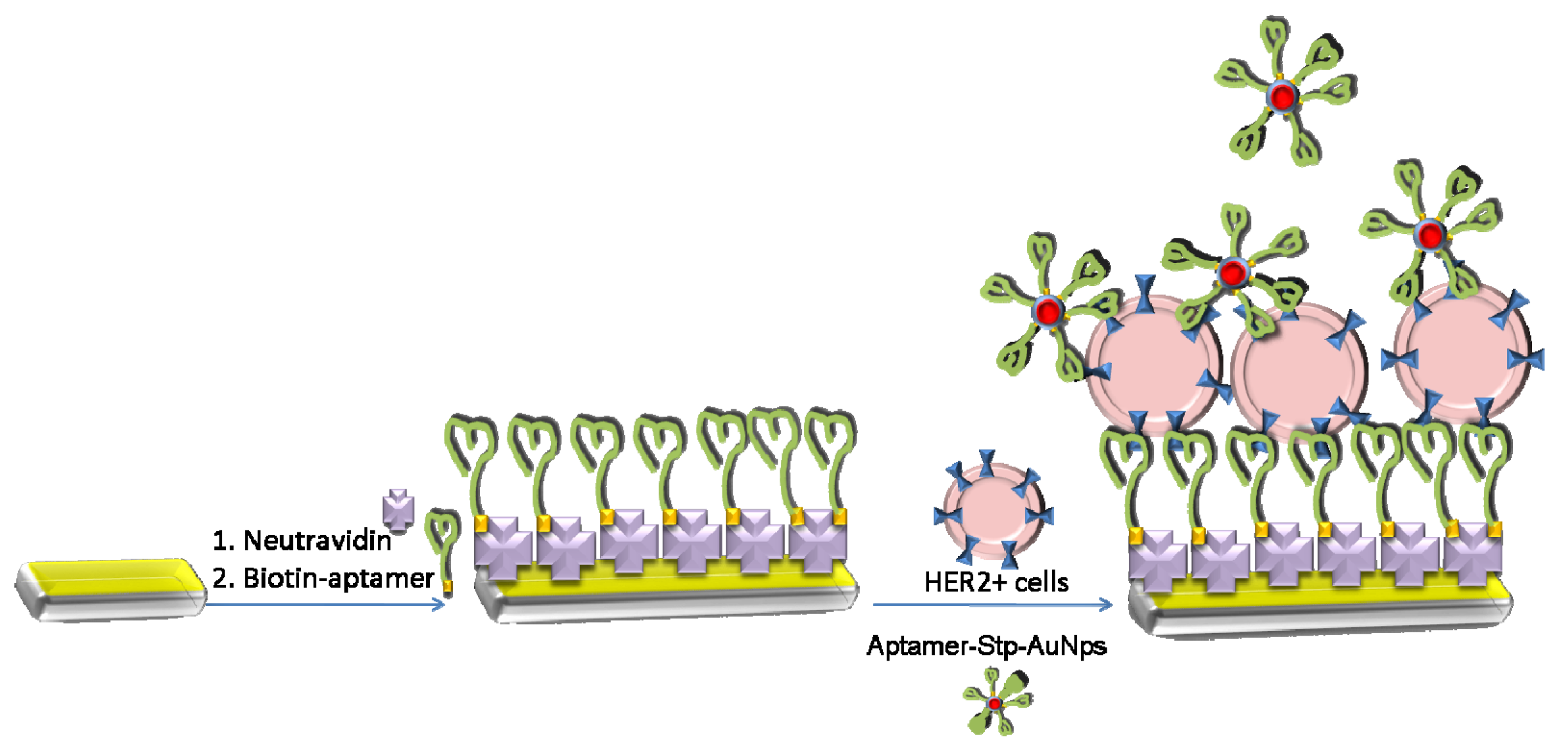
2.4. Preparation of the Aptasensor
2.5. Modification of Gold Nanoparticles by DNA Aptamers
3. Results and Discussion
3.1. The Study of Cell Viability
3.2. The Study of the Toxicity of Gold Nanoparticles on the Breast Cancer Cells
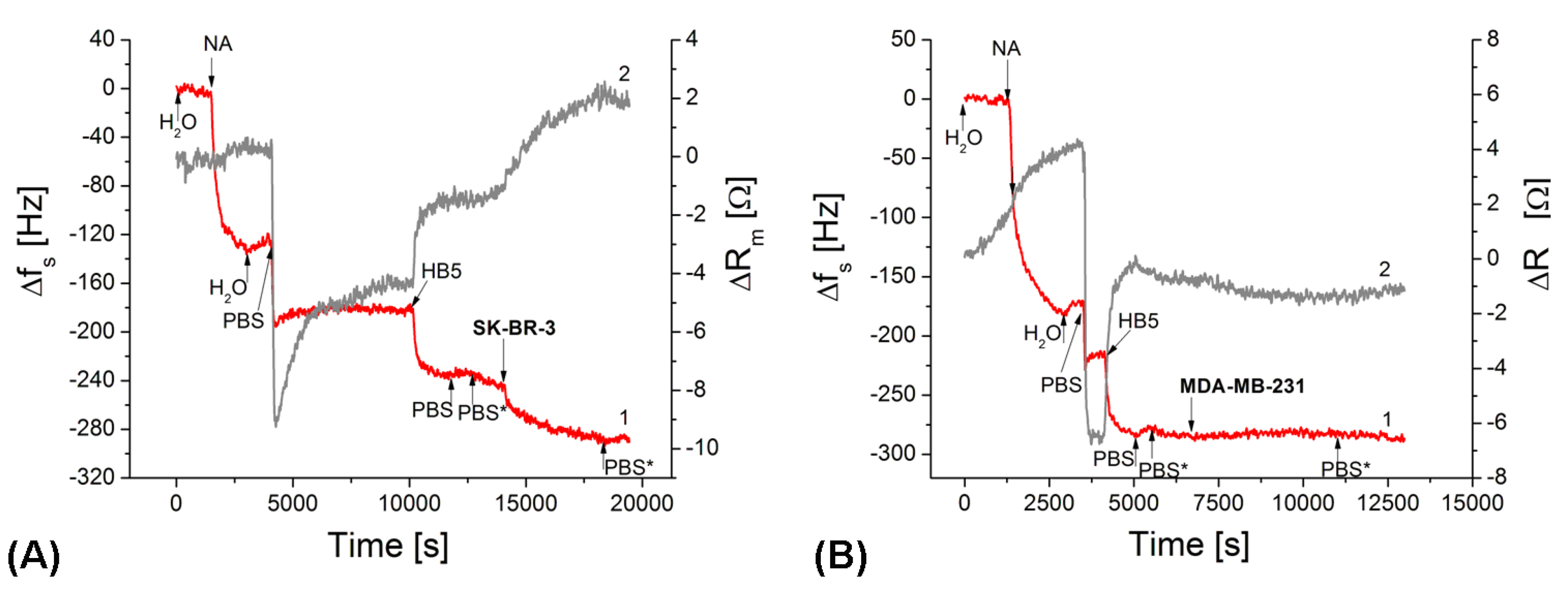
3.3. Detection of Breast Cancer Cells by TSM Method
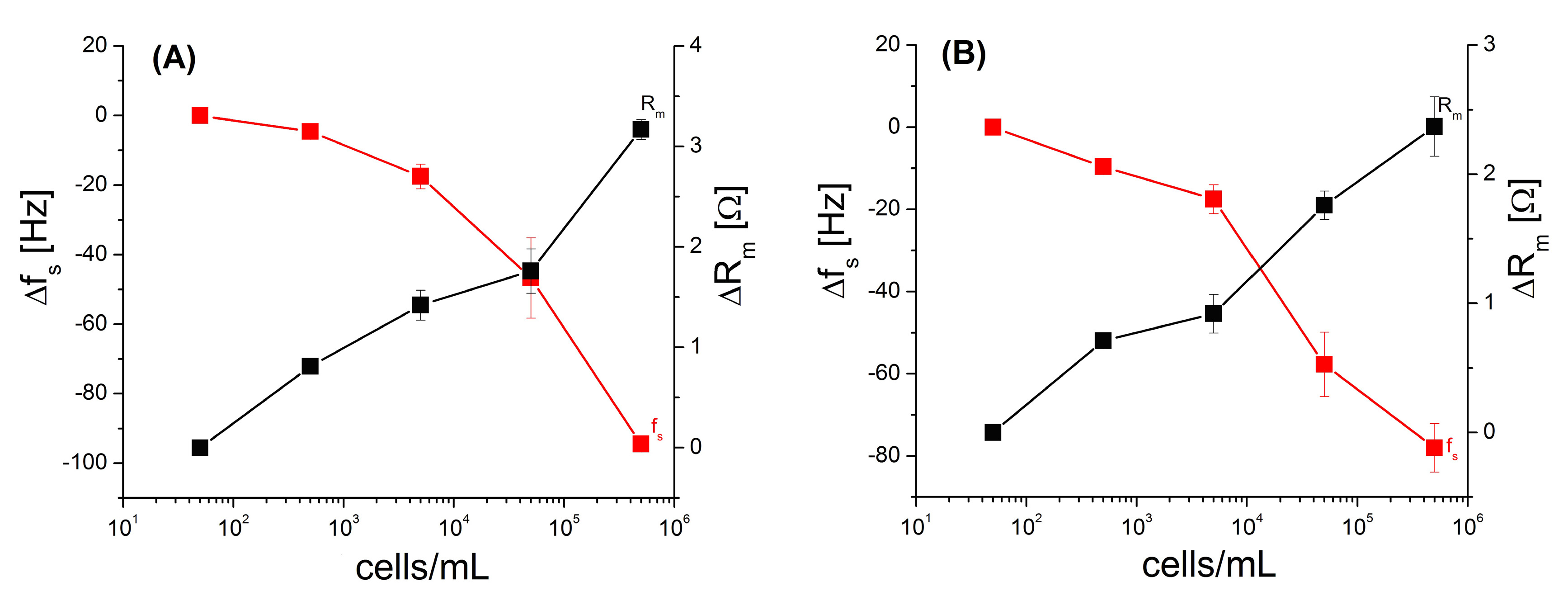
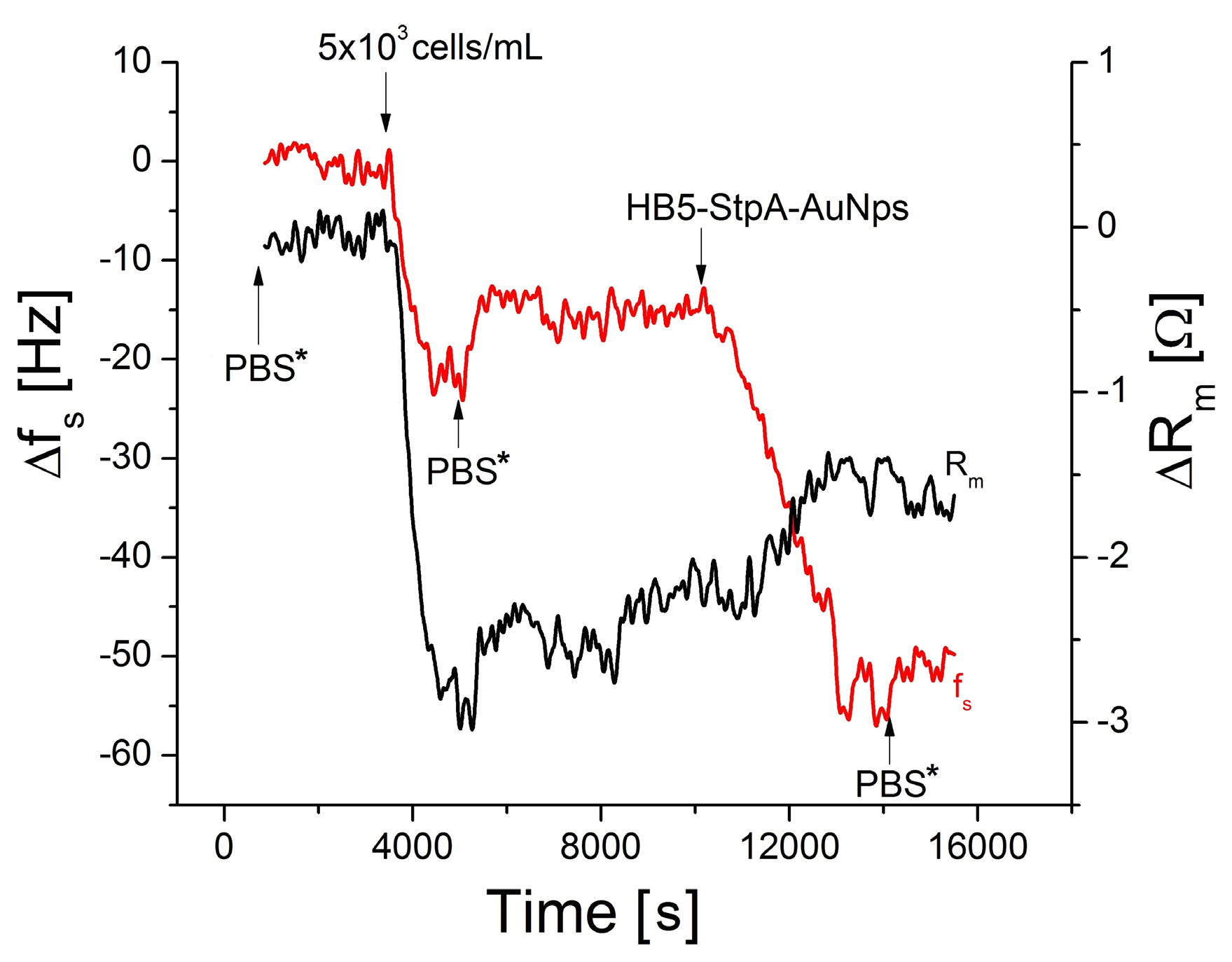
3.4. Amplification of Detection of Breast Cancer Cells by Gold Nanoparticles
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Yarden, Y. The EGFR family and its ligands in human cancer: Signalling mechanisms and therapeutic opportunities. Eur. J. Cancer 2001, 37, 3–8. [Google Scholar] [CrossRef]
- Chun, L.; Kim, S.-E.; Cho, M.; Choe, W.-S.; Nam, J.; Lee, D.W.; Lee, Y. Electrochemical detection of HER2 using single stranded DNA aptamer modified gold nanoparticles electrode. Sens. Actuators B 2013, 186, 446–450. [Google Scholar] [CrossRef]
- Wang, K.; Yao, H.; Meng, Y.; Wang, Y.; Yan, X.; Huang, R. Specific aptamer-conjugated mesoporous silica–carbon nanoparticles for HER2-targeted chemo-photothermal combined therapy. Acta Biomater. 2015, 16, 196–205. [Google Scholar] [CrossRef]
- Wu, X.; Shaikh, A.B.; Yu, Y.; Li, Y.; Ni, S.; Lu, A.; Zhang, G. Potential diagnostic and therapeutic applications of oligonucleotide aptamers in breast cancer. Int. J. Mol. Sci. 2017, 18, 1851. [Google Scholar] [CrossRef]
- Elmore, J.G.; Barton, M.B.; Moceri, V.M.; Polk, S.; Arena, P.J.; Fletcher, S.W. Ten-year risk of false positive screening mammograms and clinical breast examinations. N. Engl. J. Med. 1998, 338, 1089–1096. [Google Scholar] [CrossRef]
- Hianik, T.; Wang, J. Electrochemical aptasensors–recent achievements and perspectives. Electroanalysis 2009, 21, 1223–1235. [Google Scholar] [CrossRef]
- Ellington, A.D.; Szostak, J.W. In vitro selection of RNA molecules that bind specific ligands. Nature 1990, 346, 818–822. [Google Scholar] [CrossRef]
- Tuerk, C.; Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 1990, 249, 505–510. [Google Scholar] [CrossRef]
- Mercier, M.C.; Dontenwill, M.; Choulier, L. Selection of nucleic acid aptamers targeting tumor cell-surface protein biomarkers. Cancers 2017, 69, 69. [Google Scholar] [CrossRef]
- Šnejdárková, M.; Svobodová, L.; Polohová, V.; Hianik, T. The study of surface properties of an IgE-sensitive aptasensor using an acoustic method. Anal. Bioanal. Chem. 2008, 390, 1087–1091. [Google Scholar] [CrossRef]
- Gijs, M.; Penner, G.; Blackler, G.B.; Impens, N.R.E.N.; Baatout, S.; Luxen, A.; Aerts, M. Improved aptamers for the diagnosis and potential treatment of HER2-positive cancer. Pharmaceuticals 2016, 9, 29. [Google Scholar] [CrossRef]
- Li, X.; Zhang, W.; Liu, L.; Zhu, Z.; Ouyang, G.; An, Y.; Zhao, C.; James, C. In vitro selection of DNA aptamers for metastasis breast cancer cell recognition and tissue imaging. Anal. Chem. 2014, 86, 6596–6603. [Google Scholar] [CrossRef]
- Liu, Z.; Duan, J.-H.; Song, Y.-M.; Ma, J.; Wang, F.-D.; Lu, X.; Yang, X.-D. Novel HER2 aptamer selectively delivers cytotoxic drug to HER2-positive breast cancer cells in vitro. J. Transl. Med. 2012, 10, 148. [Google Scholar] [CrossRef]
- Shahdordizadeh, M.; Yazdian-Robati, R.; Ramezani, M.; Abnous, K.; Taghdisi, S.M. Aptamer application in targeted delivery systems for diagnosis and treatment of breast cancer. J. Mat. Chem. B 2016, 4, 7766–7778. [Google Scholar] [CrossRef]
- Medley, C.D.; Smith, J.E.; Tang, Z.; Wu, Y.; Bamrungsap, S.; Tan, W. Gold nanoparticle-based colorimetric assay for the direct detection of cancerous cells. Anal. Chem. 2008, 80, 1067–1072. [Google Scholar] [CrossRef]
- Zhou, T.; Marx, K.A.; Dewilde, A.H.; McIntosh, D.; Braunhut, S.J. Dynamic cell adhesion and viscoelastic signatures distinguish normal from malignant human mammary cells using quartz crystal microbalance. Anal. Biochem. 2012, 421, 164–171. [Google Scholar] [CrossRef]
- Skládal, P. Piezoelectric biosensors. Trends in Anal. Chem. 2016, 79, 127–133. [Google Scholar] [CrossRef]
- Elmlund, L.; Kack, C.; Aastrup, T.; Nicholls, I.A. Study of the interaction of trastuzumab and SKOV3 epithelial cancer cells using a quartz crystal microbalance sensor. Sensors 2015, 15, 5884–5894. [Google Scholar] [CrossRef]
- Poturnayová, A.; Buríková, M.; Bizík, J.; Hianik, T. DNA aptamers in detection of leukemia cells by thickness shear mode acoustics method. ChemPhysChem 2019, 20, 545–554. [Google Scholar] [CrossRef]
- Shangguan, D.; Li, Y.; Tang, Z.; Cao, Z.C.; Chen, H.W.; Mallikaratchy, P.; Sefah, K.; Yang, C.J.; Tan, W. Aptamers evolved from live cells as effective molecular probes for cancer study. Proc. Natl. Acad. Sci. USA 2006, 103, 11838–11843. [Google Scholar] [CrossRef]
- Huang, Y.-F.; Shangguan, D.; Liu, H.; Phillips, J.A.; Zhang, X.; Chen, Y.; Tan, W. Molecular assembly of an aptamer-drug conjugate for targeted drug delivery to tumor cells. ChemBioChem 2009, 10, 862–868. [Google Scholar] [CrossRef]
- Avelar-Freitas, B.A.; Almeida, V.G.; Pinto, M.C.X.; Mourao, F.A.G.; Massensini, A.R.; Martins-Filho, O.A.; Rocha-Vieira, E.; Brito-Melo, G.E.A. Trypan blue exclusion assay by flow cytometry. Braz. J. Med. Biol. Res. 2014, 47, 307–3015. [Google Scholar] [CrossRef]
- Mascotti, K.; Mccullough, J.; Burger, S.R. HPC viability measurement: Trypan blue versus acridine orange and propidium iodide. Transfusion 2000, 40, 693–696. [Google Scholar] [CrossRef]
- Tassew, N.; Thompson, M. Kinetic characterization of TAR RNA-Tat peptide and neomycin interactions by acoustic wave biosensor. Biophys. Chem. 2003, 106, 241–252. [Google Scholar] [CrossRef]
- Cai, W.J.; Chen, L.; Chen, B.; Gao, Z.; Lee, S.H.; Chen, J.; Pierrot, D.; Sullivan, K.; Wang, Y.; Hu, X.; et al. Decrease in the CO2 uptake capacity in an ice-free arctic ocean basin. Science 2010, 329, 556–559. [Google Scholar] [CrossRef]
- Han, Q.; Chen, R.; Wang, F.; Chen, S.; Sun, X.; Guan, X.; Yang, Y.; Peng, B.; Pan, X.; Li, J.; et al. Pre-exposure to 50 Hz-electromagnetic fields enhanced the antiproliferative efficacy of 5-fluorouracil in breast cancer MCF-7 cells. PLoS ONE 2018, 13, e0192888. [Google Scholar] [CrossRef]
- Lewinski, N.; Colvin, V.; Drezek, R. Cytotoxicity of nanoparticles. Small 2008, 4, 26–49. [Google Scholar] [CrossRef]
- Alkilany, A.M.; Murphy, C.J. Toxicity and cellular uptake of gold nanoparticles: What we have learned so far? J. Nanoparticle Res. 2010, 12, 2313–2333. [Google Scholar] [CrossRef]
- Smolkova, B.; Miklikova, S.; Kajabova, V.H.; Babelova, A.; Yamani, N.E.; Zduriencikova, M.; Fridrichova, I.; Zmetakova, I.; Krivulcik, T.; Kalinkova, L.; et al. Global and gene specific DNA methylation in breast cancer cells was not affected during epithelial-to-mesenchymal transition in vitro. Neoplasma 2016, 63, 901–910. [Google Scholar] [CrossRef]
- Giljohann, D.A.; Seferos, D.S.; Daniel, W.L.; Massich, M.D.; Patel, P.C.; Mirkin, C.A. Gold nanoparticles for biology and medicine. Angew. Chem. Int. Ed. Engl. 2010, 49, 3280–3294. [Google Scholar] [CrossRef]
- Ellis, J.S.; Thompson, M. Acoustic coupling atmultiple interfaces and the liquid phase response of the thickness shear-mode acoustic wave sensor. Chem. Commun. 2004, 11, 1310–1311. [Google Scholar] [CrossRef]
- Hianik, T. Detection of protein-aptamer interactions by means of electrochemical indicators and transverse shear mode method. In Aptamer in Bioanalysis; Mascini, M., Ed.; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2009; Volume 313, pp. 101–128. ISBN 9780470148303. [Google Scholar]
- Hayashi, N.; Nakamura, S.; Tokuda, Y.; Shimoda, Y.; Yagata, H.; Yoshida, A.; Ota, H.; Hortobagyi, G.N.; Cristofanilli, M.; Ueno, N.T. Prognostic value of HER2-positive circulating tumor cells in patients with metastatic breast cancer. Int. J. Clin. Oncol. 2012, 17, 96–104. [Google Scholar] [CrossRef]
- Glassford, A.P.M. Response of a quartz crystal microbalance to a liquid deposit. J. Vac. Sci. Technol. 1978, 15, 1836–1843. [Google Scholar] [CrossRef]
- Zhu, Y.; Chandra, P.; Shim, Y.-B. Ultrasensitive and selective electrochemical diagnosis of breast cancer based on a hydrazine−Au nanoparticle−aptamer bioconjugate. Anal. Chem. 2013, 85, 1058–1064. [Google Scholar] [CrossRef]
- Chen, Q.; Tang, W.; Wang, D.; Wu, X.; Li, N.; Liu, F. Amplified QCM-D biosensor for protein based on aptamer-functionalized gold nanoparticles. Biosens. Bioelectr. 2010, 26, 575–579. [Google Scholar] [CrossRef]
- Sauerbrey, G. Verwendung von schwingquarzen zur wägung dünner schichten und zur mikrowägung. Z. Phys. 1959, 155, 206–222. [Google Scholar] [CrossRef]
- Lodish, H.; Berk, A.; Zipursky, S.L.; Matsudaira, P.; Baltimore, D.; Darnell, J.E. Molecular Cell Biology, 4th ed.; W. H. Freeman: New York, NY, USA, 2000; Volume 1184, ISBN 9780716737063. [Google Scholar]
- Fredriksson, C.; Kihlman, S.; Rodahl, M.; Kasemo, B. The piezoelectric quartz crystal mass and dissipation sensor: A means of studying cell adhesion. Langmuir 1998, 14, 248–251. [Google Scholar] [CrossRef]
- Pan, Y.L.; Guo, M.; Nie, Z.; Huang, Y.; Pan, C.; Zeng, K.; Zhang, Y.; Yao, S. Selective collection and detection of leukemia cells on a magnet-quartz crystal microbalance system using aptamer-conjugated magnetic beads. Biosens. Bioelectr. 2010, 25, 1609–1614. [Google Scholar] [CrossRef]
- Shan, W.; Pan, Y.; Fang, H.; Guo, M.; Nie, Z.; Huang, Y.; Yao, S. An aptamer-based quartz crystal microbalance biosensor for sensitive and selective detection of leukemia cells using silver-enhanced gold nanoparticle label. Talanta 2014, 126, 130–135. [Google Scholar] [CrossRef]
- Zhang, S.; Bai, H.; Luo, J.; Yang, P.; Cai, J. A recyclable chitosan-based QCM biosensor for sensitive and selective detection of breast cancer cells in real time. Analyst 2014, 139, 6259–6265. [Google Scholar] [CrossRef]

| Target Protein | Aptamer | Sequence 5′-3′ | Kd, nM | Reference |
|---|---|---|---|---|
| Epitope peptide HER2 | HB5 | AACCGCCCAAATCCCTAAGAGTCTGCACT TGTCATTTTGTATATGTATTTGGTTTTTGGC TCTCACAGACACACTACACACGCACA | 18.9 | [13] |
| HER2 protein | HeA2_3 | TCTAAAAGGATTCTTCCCAAGGGGATCCA ATTCAAACAGC | 6.2 | [11] |
| PTK7 protein | Sgc8c | ATCTAACTGCTGCGCCGCCGGGAAAATAC TGTACGGTTAGA | 1.0 | [20,21] |
| Cells/mL | HB5 Aptamer | Complex HB5-StpA-AuNPs | ||
|---|---|---|---|---|
| Δfs ± SD (Hz) | ΔRm ± SD (Ω) | Δfs ± SD (Hz) | ΔRm ± SD (Ω) | |
| 50 | 0 | 0 | 0 | 0 |
| 5 × 102 | 0 | 0 | −3.5 ± 3.25 | 0.4 ± 0.10 |
| 5 × 103 | −1 ± 4.2 | 0 ± 0.30 | −22.4 ± 3.54 | 0.9 ± 0.42 |
| 5 × 104 | −7.3 ± 3.54 | 1.17 ± 0.38 | −45.9 ± 4.50 | 1.8 ± 0.54 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Poturnayová, A.; Dzubinová, Ľ.; Buríková, M.; Bízik, J.; Hianik, T. Detection of Breast Cancer Cells Using Acoustics Aptasensor Specific to HER2 Receptors. Biosensors 2019, 9, 72. https://doi.org/10.3390/bios9020072
Poturnayová A, Dzubinová Ľ, Buríková M, Bízik J, Hianik T. Detection of Breast Cancer Cells Using Acoustics Aptasensor Specific to HER2 Receptors. Biosensors. 2019; 9(2):72. https://doi.org/10.3390/bios9020072
Chicago/Turabian StylePoturnayová, Alexandra, Ľudmila Dzubinová, Monika Buríková, Jozef Bízik, and Tibor Hianik. 2019. "Detection of Breast Cancer Cells Using Acoustics Aptasensor Specific to HER2 Receptors" Biosensors 9, no. 2: 72. https://doi.org/10.3390/bios9020072
APA StylePoturnayová, A., Dzubinová, Ľ., Buríková, M., Bízik, J., & Hianik, T. (2019). Detection of Breast Cancer Cells Using Acoustics Aptasensor Specific to HER2 Receptors. Biosensors, 9(2), 72. https://doi.org/10.3390/bios9020072






