Abstract
Emerging water pollutants such as pharmaceutical contaminants are suspected to induce adverse effects to human health. These molecules became worrisome due to their increasingly high concentrations in surface waters. Despite this alarming situation, available data about actual concentrations in the environment is rather scarce, as it is not commonly monitored or regulated. This is aggravated even further by the absence of portable and reliable methods for their determination in the field. A promising way to tackle these issues is the use of enzyme-based and miniaturized biosensors for their electrochemical detection. Here, we present an overview of the latest developments in amperometric microfluidic biosensors that include, modeling and multiphysics simulation, design, manufacture, testing, and operation methods. Different types of biosensors are described, highlighting those based on oxidases/peroxidases and the integration with microfluidic platforms. Finally, issues regarding the stability of the biosensors and the enzyme molecules are discussed, as well as the most relevant approaches to address these obstacles.
1. Introduction
Many places around the world currently face water pollution due to the increasing volumes of waste discharged to superficial waters. This is in turn, the result of intensified industrial, agricultural and household activities. Consequently, the world has witnessed a growing scarcity of potable water and a decrease of aquatic biodiversity. A number of treatment processes have been devised to tackle these issues, which are mainly based on a proper determination of physicochemical parameters, the presence of microorganisms, and the possible uses of the recovered waste [1,2,3,4]. A complete characterization of polluted water resources is still elusive mainly due to the absence of tools for in situ, rapid, and sensitive monitoring of field samples. Pharmaceutical derivatives are some of the most difficult contaminants to detect and remove from water. One of the main pharmaceutical products found in wastewater are antibiotics, which are generally originated from both human and animal therapeutic and nutraceutical preparations [5].
In recent decades, various methods have been reported for the detection of pharmaceutical waste in wastewater including optical methods, immunoassay tests, molecular spectroscopy-based techniques and devices that rely on optical fiber technology [6]. In general, these methods incorporate antibodies, antigens, and enzymes as active components for the detection of specific antibiotics. This is mainly done by tracking changes in the intensity of the reflected light and/or the fluorescence of conjugated markers that become active upon triggering specific recognition mechanisms [6]. Electrochemical strategies for the detection of pharmaceutical residues have been reported to be sensitive, selective, robust, efficient and rapid alternatives in wastewater. In general, these strategies rely on the measurement of changes in ions or electrons generated by enzymes or antibodies immobilized on electrodes [7,8,9]. Over the past few years, there has been increasing interest in simultaneously detecting several water contaminants. This has been intended to reduce analysis times and production costs of biosensors.
This chapter is dedicated to describe both electrochemical biosensors based on enzymes such as laccase and oxidase/peroxidase, and amperometric microfluidic biosensors. Additionally, we provide a set of recommendations for the design and testing of biosensor prototypes.
2. Enzymatic Biosensors Applied to Microfluidic Systems
Sensing platforms are generally composed of a recognition element and a transducing device as seen in Figure 1 [10,11,12]. In the particular case of biosensors, the recognition element is a biomolecule with stimuli-responsive capabilities. The transducing device converts, therefore, a measurable biological response or physicochemical change into an electrical signal [11,12]. Among the most common recognition molecules are: enzymes, antibodies, phages, or single-stranded DNA [10,11,13]. The recognition element in affinity biosensors binds permanently or semi-permanently with the analyte of interest, while for catalytic biosensors, the element maintains a non-permanent interaction in the form of a chemical reaction or electron transfer [10].
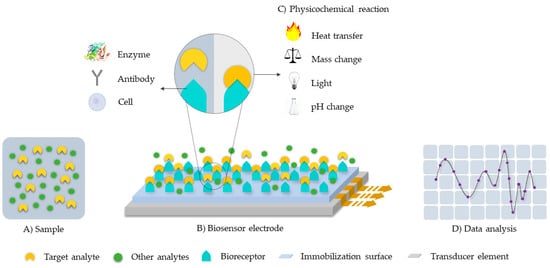
Figure 1.
Schematic of a typical biosensor system architecture. The target analyte is detected by the bioreceptor and translated into a signal for analysis with the aid of a transducer. (A) Sample. (B) Biosensor electrode: composed by the bioreceptor, the immobilization surface and the transducer element. (C) Physicochemical reaction. (D) Data analysis.
Enzyme-based sensors are highly specific catalytic biosensors where the recognition elements are extremely selective enzyme molecules immobilized on transducing surfaces known as electrodes [10,14,15]. Under the presence of a suitable substrate, the enzymes catalyze electrochemical reactions involving electroactive products or measurable electric changes on the transducer and the sample [15,16]. The concentration or redox potential of an analyte in a sample can be determined via measurements of potential, charge, or current. The devices to conduct these measurements apply potentiometric, impedimetric and amperometric techniques, respectively [14,16,17].
The amperometric methods are the most suitable to perform the transduction of the enzymatic response into a quantifiable signal [14]. In this case, the number of species involved in the redox process is related to the electrons generated when a fixed potential is applied between two electrodes, which can be ultimately detected as current change. Additionally, amperometric methods offer the possibility for real time detection, as well as ease for mass production [10,17,18]. The amperometric electrochemical biosensors can conduct measurements with the aid of two or three electrodes. In the two-electrode configuration, one functions as the working electrode while the other works as the reference. The sample is usually measured with the working electrode where the enzyme molecules are immobilized. Generated currents are evaluated by comparing the two electrodes. In the case of the three-electrode configuration, current flows between the working and a third electrode called counter while voltage is applied between the reference and the working electrodes (Figure 2) [19]. The surfaces of the electrodes are generally made of carbon and noble metals [17].
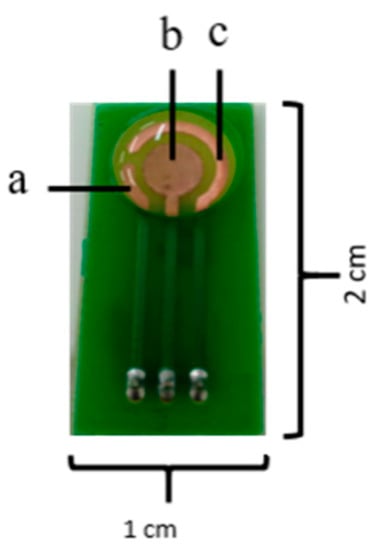
Figure 2.
Three electrode sensor configuration. (a) reference, (b) working and (c) counter electrodes.
2.1. Immobilization Methods
An essential procedure for the manufacture of electrodes in an electrochemical-enzymatic biosensor is the immobilization of enzyme molecules [16,20]. This can be achieved via physical adsorption, covalent binding, entrapment, encapsulation or cross-linking (Figure 3) [16,18]. Immobilization not only helps to prolong the shelf life of enzymes over time but also reduces the enzymatic time response. The process of immobilization of laccase molecules on electrode surfaces plays a key role in the performance of the biosensors. This is because during immobilization, the enzyme molecules are prone to conformational changes that might ultimately lead to partial or total loss of activity [18,21,22,23,24,25]. To enable these methods, activation of functional groups on the support surface might be required prior to immobilization to assure high immobilization yields [21,22].
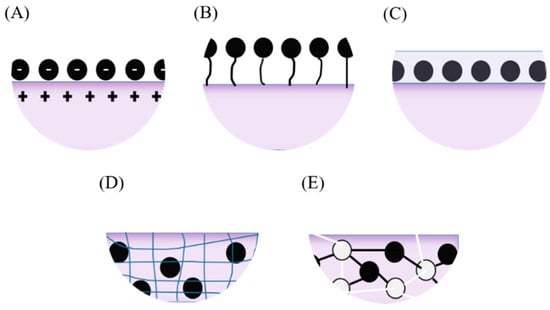
Figure 3.
Enzyme immobilization onto electrodes. (A) Physically adsorbed through electrostatic interactions, (B) covalently bound to the surface, (C) entrapped within a film, (D) encapsulated within a porous surface and (E) cross-linked within the surface.
When immobilizing laccase molecules via adsorption, surface attachment is achieved by the interplay of Vander Waals forces, hydrogen bridges, and hydrophobic, hydrophilic or ionic interactions [21,22,24]. In the case of covalent immobilization, the enzyme molecules react with the activated functional groups on the surface to form strong covalent bonds [22,25]. This immobilization approach generally leads to high conformational stability for the immobilized molecules [21]. Unfortunately, a risk of permanently damaging the active site of the enzyme molecules exists [25]. Moreover, under certain conditions, this approach has been reported to produce self-assembling, which might lead to detrimental intramolecular bonding. Some of the crosslinkers responsible for this include homo- and hetero-bifunctional molecules such as (3-Aminopropyl)triethoxysilane (APTES) and Glutaraldehyde [22,25]. In the case of entrapment, laccase molecules are trapped into a polymeric matrix to try to maintain their structural stability [22,25]. Encapsulation works by confining the enzyme molecules into semipermeable polymeric microspheres [22,25]. Finally, in the electrochemical immobilization, the enzyme molecules are oxidized by applying an electric potential difference to induce electron losses that are ultimately compensated by forming bonds with the support surface [18] as seen in Table 1.

Table 1.
Laccase-based biosensors for pharmaceutical analytes as classified by the employed transduction and immobilization methods.
The interaction between a target molecule in the sample and a specific enzyme is fairly well approximated by the Michaelis-Menten equation [14,16,20] (Equation (1)):
Rates of reaction () are experimentally obtained at different substrate concentrations ( as described by the kinetics equation where is the reaction rate at enzymatic saturation concentration and is the enzyme-substrate affinity. A good enzymatic biosensor seeks to be sensitive enough to show high values in response to low substrate concentrations.
After immobilization, the amount of conjugated enzyme can be estimated by cyclic voltammetry analysis (CV), electrochemical impedance spectroscopy (EIS), Fourier transform infrared (FTIR) spectroscopy, and atomic force microscopy (AFM) [18,26]. For the case of CV and EIS, immobilization is demonstrated if a characteristic electron transfer from laccase catalytic turnover is observed [18]. FTIR spectroscopy confirms covalent binding of enzymes and crosslinkers by collecting information on the surface functional groups. Topography features via AFM allow to collect information on the surface disposition of the enzyme molecules and possible clustering processes during immobilization [26]. At the same time, it is important to evaluate the long-term stability of the enzyme-surface attachment to determine whether the biosensor can be reused and possibly be incorporated into a cyclic operation.
The correct operation of the biosensor is evaluated by characterizing the response signal in the presence of commercially available oxidizing enzyme substrates. A concentration curve is built for each substrate to determine the limit of detection, the signal stability upon concentration changes, the sensitivity range of the device, and the repeatability. In the case of Laccase, oxidizing standards are 2,2′-azino-bis(3-ethylbenzothiazoline-6-sulphonic acid) (ABTS) and syringaldazine [20,27,28,29]. Selectivity can be probed by calculating the selectivity coefficient, which can be recovered from concentration curve analysis of possible interference analytes at a constant concentration of substrate [30,31].
Operational stability of the biosensor can be estimated by conducting measurements with the device under extreme conditions of pH and temperature [20,30] Storage stability assures the robustness of the sensor response in the long-term. Optimal storage conditions strongly depend on the type of immobilization approach and the variability of the environmental conditions [22,31,32].
The enzymatic component of the biosensor is perhaps the most sensitive to large swings in operational conditions. To tackle this issue some of the strategies include chemical modifications of pendant groups on the enzyme surface, conjugation with stabilizing polymer molecules, and activation via increased hydrostatic pressure [21].
2.2. Device Prototyping and Testing
New strategies and methods for the development of electrochemical biosensors rely on microfabrication techniques and integrated electronics. The portability and ultra-low volume requirements of microfluidic analytical devices reduce sample manipulation, transfer, and storage; that ultimately, help prevent contamination [18,33]. Understanding momentum transport phenomena for fluids inside the microfluidic systems is the key to define proper manufacturing processes and protocols. In the section below, we discussed the equations that govern these phenomena.
2.2.1. Microfluidics Fundamentals
The conservation of mass over time within a system is described by the continuity equation (Equation (2)):
where is the density of the fluid and , and are the components of the velocity in x, y and z, respectively. A conventional simplification for Equation (2) is to consider that the system is at a steady state, which means the system is invariant on time [34,35]. Additionally, if the fluid is considered incompressible ( = constant), then the expression can be simplified to Equation (3).
Conservation of momentum transfer led to the Navier-Stokes equations (Equation (4)), which can be solved along with (Equation (3)) to find the velocity profile of the fluid flowing within a microfluidic system.
where V is the velocity vector, P is the pressure, g is the gravitational field and μ is the viscosity of the fluid [34,35,36]. Due to the small scale of channels in microfluidic systems, gravitational forces are negligible. Furthermore, in microfluidics viscous forces dominate over convective forces. This can be seen with the aid of the Reynolds number (Re), which is defined according to the relationship in Equation (5).
where is the hydraulic diameter, is the density of the fluid, is the dynamic viscosity, and is the mean velocity. In the case of microfluidics, the Re numbers are typically below 1 (Re < 1) [37]. This flow regime is known as a laminar and under this condition; the convective terms of Equation (4) are negligible. Finally, if the steady state is assumed, Equation (5) can be simplified to:
If for instance, we considered a microfluidic system with squared channels, it is conventional to find that the width of the channels (W) is considerably smaller than the length of the channels (L). Likewise, the height of the channel (H) is shorter than W (Figure 4). Under these geometry considerations, the problem can be considered unidimensional with molecular momentum transfer only in the y direction (Equation (7)):
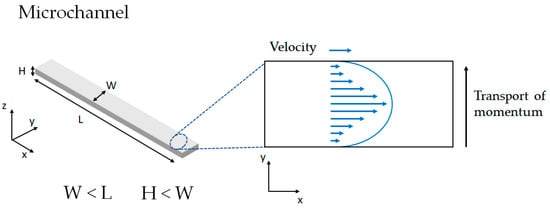
Figure 4.
Example of momentum transport and velocity profile generated in a microfluidic channel.
By solving Equation (7) it is possible to determine the required inlet and outlet pressures on the channel to generate a specific velocity profile in the fluid [34,37].
2.2.2. Prototypes Circuit Approximation
An easy and quick way to estimate the correlation between the pressure gradient, flow rate and resistance for a fluid within a microfluidic system is by using an electrical equivalent approach. To solve the equations, one can consider an analogy between the parameters in fluid mechanics and those of the electrical systems. Accordingly, the flow rate (Q) within the microsystem is due to a change in pressure (P) between inlets and outlets, which correspond to current flow (I) and the voltage potential (V) in circuits. In the two cases, an extra parameter must be considered, the resistance (R), which is understood as a restraining imposed by a system or material to the flow of fluids or electrons [34,35]. Figure 5 shows an example of different configurations of microsystems and the corresponding circuit analogy.
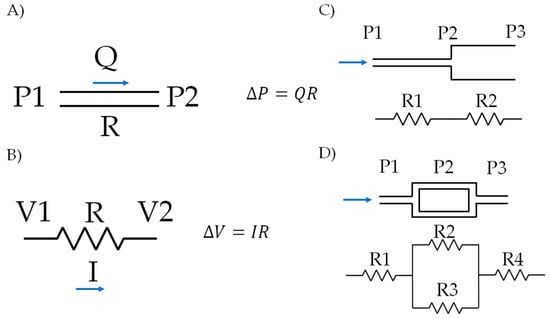
Figure 5.
Circuit approach for modeling fluid flow in microsystems: (A) pressure and flow ratio in a microchannel, (B) relationship for voltages and currents in a circuit, (C,D) examples of different microsystems configurations and their corresponding circuit analog.
If we consider a microchannel with a circular cross-sectional area, the fluid flow equations are solved with the aid of the Hagen-Poiseuille relationship (Equation (8)):
where A is the cross-sectional area perpendicular to fluid flow, R is the radius of the channel and L is the length of the channel. Manufacturing of microfluidics usually leads to channels with geometries far from circular and for that reason, Equation (8) no longer applies. A modified version of Equation (8) includes the hydraulic radio () (Equation (10)), which is defined according to the expression in Equation (9):
where is the perimeter of the cross-sectional area. This approach allows to consider the fluid flow in microfluidic channels with any cross-sectional area. With this new parameter, we can now rewrite Equation (8) and subsequently define the hydraulic resistor (Equations (11) and (12)):
The mass balance for the system is also achieved if we consider that for the electric circuit analogy, the sum of the currents in each node must equal zero. This leads to Equation (13) for the system:
This simple yet powerful approach is always used by us to estimate changes of pressure and flow within a microsystem without necessarily knowing the details of velocity distribution inside it [35].
2.2.3. Mixing and Separation
Some of the most common applications of microfluidic systems include separation and mixing processes, which are useful to concentrate or remove some components of the analyzed mixtures. Depending on the application, it is possible to separate micrometer (e.g., cells) to nanometer size (e.g., proteins) particles from complex mixtures. Separation of particles with significant differences in sizes or densities, generally rely on devices capable of generating sufficient inertial forces [38,39]. This can be achieved by spiral geometries because in this case the drag and lift forces acting on the particles force them to separate from each other within the microchannel. The separation efficiency strongly depends on properties of the particles such as mass, size, and density. Another efficient way to separate particles or molecules is by means of aqueous two-phase systems (ATPS) [39,40]. These systems take advantage of the electrostatic potential at the interface between two immiscible liquids and the tendency of charged molecules to preferentially partition into one of the fluids in contact [40,41]. The net superficial charge can be also exploited for separation by making use of electrophoretic mechanisms. This is also the case when the molecules of interest exhibit magnetism, as they can be separated via magnetophoresis [42,43].
A major challenge when working with microfluidic systems is the difficulty to generate efficient mixing patterns mainly due to the low Reynolds numbers (Re < 1). To tackle this issue, it is necessary to generate turbulence through special geometries such as Zigzag channels, 3-D serpentine structures, and twisted channels [44]. Micromixing can be achieved either through active mixing or passive mixing [45,46]. In the case of active mixing, integration of components is accomplished via an external energy source such as ultrasound, acoustic vibrations, small impellers, or electrokinetic instability. In contrast, passive micromixing requires a device to intimately put in contact the components through disturbances in the mixing patterns. These devices include chaotic flow configuration, flow recirculation configuration, colliding jet, split and recombine flow configurations.
2.2.4. Simulations
An easy and economical way to make and test prototypes is by Computational Fluid Dynamics (CFD) simulations. There are different software packages to implement CFD simulations, one of which is Comsol Multiphysics®. This software solves partial differential equations (PDE) associated with physical phenomena via the finite element method (FEM) [47]. In this approach, PDE equations are solved by putting the functions of interests in the integral form, where the unknowns are discretized as summations over the functions defined on the finite elements. Depending on the topology of the geometry, i.e., 2D or 3D, the elements are assembled into polygonal forms, thereby creating a mesh over the computational domain. The FEM is adaptable to complex geometries and can handle discontinuous gradients of a variable. Nevertheless, like any other discrete numerical method for solving continuous PDE, it introduces some numerical error. This can be minimized by varying the meshing approach or by refining the mesh on those spots where the flux of gradients is the greatest [47].
Figure 6 shows an example of the simulated fluid flow in a microfluidic channel after implementation in the CFD module of Comsol Multiphysics®. The Navier-Stokes equations were solved to find the velocity distribution (see above). Maximum velocity values are shown for each section of the microchannel in Figure 6C. Also, it is possible to recover data on pressure changes within the system, shear stress, and shear rate.
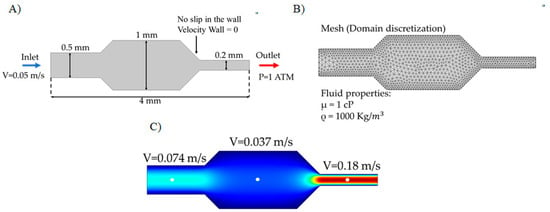
Figure 6.
Simulation of microfluidic system in Comsol Multiphysics ® (A) Simulation domain with the corresponding boundary conditions (inlet speed, outlet pressure and non-slip in the walls), (B) discretization of the domain by a meshing process and the list of physical properties of the fluid, (C) velocity profile in the channel as recovered from Computational Fluid Dynamics (CFD) simulations.
This simulation tool has been used in different studies to optimize the response of different types of biosensors [48,49,50]. This optimization can be from the operation values of the biosensor (voltages, resistance values, flow rates, etc.) or changes in the geometry.
2.2.5. Fabrication of Prototypes
Chemical etching is based on eroding a surface to generate microchannels with depths ranging from 5–15 µm [51,52]. Glass is the preferred substrate to manufacture microsystems by this technique [53]. Chemical etching on glass starts by covering the surface with a sacrificial layer known as photoresist by means of a spin coating machine. Depending on the properties of the photoresist and the speed of the spin coating process, different sacrificial layer thicknesses can be generated. Next, the photoresist covered glass sheet is exposed to a mask with a layout of the microsystem. Subsequently, the two surfaces (photoresist and glass) are immersed in hydrofluoric acid. In areas with absence of photoresist, the glass surface is eroded, thereby generating the microchannel. Finally, the microchannel is sealed with a layer of PDMS [54]. Figure 7A shows a schematic of the etching process. While this technique offers a high resolution, the involved costs are relatively high due to the multistage processing required.
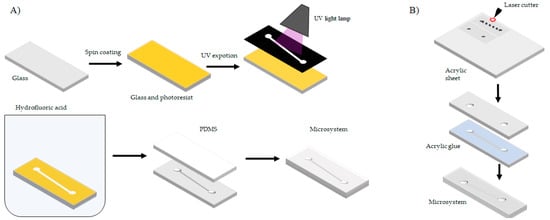
Figure 7.
(A) Manufacturing process by photolithographic techniques and (B) manufacturing process by laser cutting.
Physical etching is based on making laser cuts and laser eroding of surfaces to achieve channels with depths in the range of 1–0.3 mm. The preferred material to apply this technique is Poly(methylmethacrylate) (PMMA) [55,56]. To obtain a PMMA sheet with the desired depths, surface erosion and cuts are conducted with the aid of a programmed laser beam. Then the cut pieces are glued together with an acrylic adhesive and maintained under constant pressure for several hours. Figure 7B shows a schematic of the laser cutting process. This technique is inexpensive, simpler and faster when compared with wet etching, soft lithography and thermoforming [57]. Laser cutting precision and fidelity is limited when working below 0.5 mm.
3. Oxidase/Peroxidase Based-Biosensors
A family of enzymes with a well-recognized ability for the detection of substances including pharmaceuticals are oxidases and peroxidases [58,59]. They have been reported to work either independently or in combination. Through oxidation-reduction reactions (REDOX), these enzymes catalyze the biotransformation of numerous compounds (Figure 8) [60,61,62].
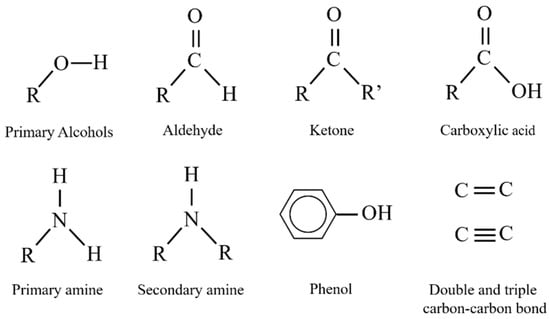
Figure 8.
Functional groups catalyzed in redox reactions by oxidase and peroxidase enzymes.
Oxidase/Peroxidase based-biosensors have been successfully used to detect molecules such as glucose, alcohol, putrescine, oxygen, hydrogen peroxide and other small metabolites. This has been useful for applications in the food industry, medical diagnosis and in vitro assays of pharmaceutical trials [21,59,63,64,65]. Even though these enzymatic biosensors exhibit high sensitivity and selectivity toward the detection of small molecules, their use is limited due to issues regarding low stability of the enzyme molecules during the fabrication, special storage conditions, and complicated implementation protocols [63]. One avenue to overcome these obstacles is to immobilize the enzyme molecules onto the device surfaces. This approach has been reported to enhance operational and storage stability, sensitivity, selectivity, response time and reproducibility [66].
Physical adsorption was the immobilization approach used by Somasekhar et al. to develop an amperometric biosensor for alcohol detection [30]. In their contribution, a protein matrix of alcohol oxidase (AOx) with entrapped ferrocene was prepared via a combination of microwaves and physical adsorption on a sol-gel chitosan system [30]. Covalent binding has also been used in the fabrication of glucose biosensors to co-immobilize glucose oxidase (GOx) and HRP into polyacrylic acid deposited on a transmission electron microscope (TEM) grid with the aid of EDC and NHS as crosslinkers [67]. The same approach was also implemented in a biosensor for putrescine detection. In this case, putrescine oxidase/peroxidase was co-immobilized via glutaraldehyde crosslinking on a Ketjen Black mesoporous electrode [68].
In addition, the electrical responses of Oxidase/Peroxidase amperometric biosensors to a specific substrate can be measured either by direct or indirect methods. Hervás et al. implemented an indirect method to measure glucose by correlating it with the amount of H2O2 released by the GOx molecules immobilized into polymethacrylic acid microparticles. The released H2O2 was detected upon oxidation on an electrode by measuring the produced current changes [69]. In contrast, a direct method is illustrated by direct electron transfer in an amperometric alcohol biosensor coupled with a putrescine detector. The system consists of an electroactive surface for facile electron exchange made of multi-walled carbon nanotubes. In this case, electrons transfer between the modified glassy carbon electrode and the HRP for H2O2 reduction [30]. Putrescine oxidation by putrescine oxidase was coupled to the reduction of H2O2 by putrescine peroxidase while electrons needed for the reaction are provided by the electrode [68]. The PANI film within the Halal verification biosensor provides not only electrical responses but colorimetric changes to the presence of ethanol in beverages. This is accomplished by a redox reaction with H2O2 that promotes a color change from green to blue [31]. Detection of glucose has also been achieved qualitatively by changes in the optical appearance of 4-cyano-40-pentylbiphenyl. This is attributed to a molecular orientation change from planar to homeotropic when increasing amounts of glucose are present [67].
The most commonly found pharmaceutical residues are antibiotics, analgesics, anti-inflammatories, steroid hormones, antihypertensives, and antidepressants, which originate from both human and veterinary therapeutics [70,71]. Oxidase/Peroxidase biosensors have become increasingly popular for detecting pharmaceutical compounds due to the sensitivity of these enzymes to such molecules [72]. Analgesics such as paracetamol have been detected by the HPR-mediated oxidation of the compound, as reported by Narang et al. [73] HPR immobilized on core-Shell ZrO@ Fe3O4 nanoparticles and embedded on a chitosan hybrid film were electrodeposited on Au electrodes for detection of paracetamol via cyclic voltammetry. Other enzyme-based biosensors have been reported for the detection of antibiotics such as penicillins and tetracyclines by means of peroxidases, laccases, and tyrosinases as the recognition elements [5,73].
4. Laccase Based-Biosensors
Laccase (EC 1.10.3.2, p-benzenediol: oxygen oxidoreductase) is an oxidoreductase enzyme, capable of oxidizing phenolic compounds into phenoxyl radicals, with the aid of the 4 copper electrons in its structure and the presence of molecular oxygen [27,74]. These enzymes are produced principally by fungi but they have been reported to be present in plants and some bacteria as well [75,76]. Due to their potent catalysis capabilities, laccases are widely used in the synthesis of organic compounds [28] and bioremediation of wastewater effluents. They have been indeed tested in the remediation of effluents from pharmaceutical manufacturing, production of textiles, paints, and logging [22,23].
This case analyzes biosensors for the detection and/or quantification of phenol compounds where the bioreceptor is the laccase enzyme. The chemical or physical reaction, called output response, needs to be converted to a measurable electronic signal by a transducer element (Figure 1) [77,78]. To achieve a selective detection, the bioreceptor needs to have a great affinity for the desired analyte, namely the phenol compounds [22].
According to the transduction method, biosensors can be categorized into electrochemical, optical and thermal [22,24].
In general, electrochemical biosensors measure the electric potential difference or current due to electrons produced by the oxidation of analytes upon catalysis. As a result, molecular oxygen present in the medium is reduced to produce water molecules (See Figure 9A) [20,22,24]. Depending on the detection system used in the electrode, this type of biosensors can also be classified by the electrical magnitude at which the potential difference is measured [22,24]. The first type is termed voltammetric sensors in which the current flow passing through an electrode is measured as a response to an applied electric potential. In this case, the analyte-enzyme reaction is performed at a specific potential range where current changes are observable [22,24]. A special case of voltammetric sensors are the amperometric sensors where the potential is kept at a constant value [18,22,24]. The second type is the conductometric sensors, which analyze changes in the conductance of the sample medium during redox reactions. The third type is the potentiometric sensors, which incorporate two non-polarizable electrodes to measure the potential difference, the two electrodes are called working and reference [22,24].
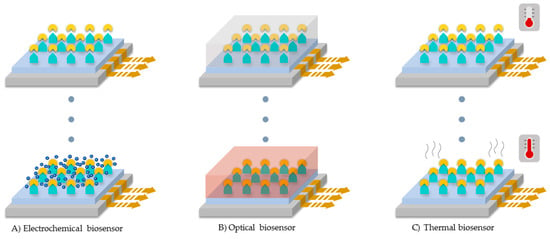
Figure 9.
Laccase-based biosensors categories according to the transduction method used. (A) Electrochemical biosensor, (B) optical biosensor, and (C) thermal biosensor.
Optical biosensors measure changes in spectroscopic properties of enzymatic catalysis products such as fluorescence and/or absorption. Some of the spectroscopic techniques include UV-VIS, FTIR, Raman and surface plasmon resonance, as seen in Figure 9B [22,25]. Finally, thermal biosensors measure enthalpy changes as a result of reactions conducted on the electrode surface (See Figure 9C) [22].
Table 1 shows the latest developments in biosensors for measuring analytes common to the pharmaceutical industry. Some of these analytes are feedstocks for the manufacture of antibiotics, analgesics, steroids and antioxidants [22,24,75]. For example, adrenaline, dopamine, norepinephrine and L-DOPA are often incorporated as active compounds into process for the manufacture of neurotransmitters [79].
The vast majority of experimental trials for enzyme-based electrochemical biosensors have been conducted in artificial wastewaters where the contents of the substances to be measured are fully known. Real wastewater matrices contain, however, a large variety of compounds with unknown concentrations including alkylphenols, cresols, aniline 2,4-dichlorophenol, some catechols, and phenolic resins [103]. The origin of many of these compounds is attributed to industrial, agricultural and domestic waste [104]. The presence of these compounds is largely responsible for the reduced output electrical signals of this type of biosensors when dealing with real samples. Additionally, Zilly et al. reported disruption of laccases’ active sites due to the presence of NaCl and Na2SO4, which are thought to promote conformational changes and even the blockage of the electron transfer machinery [105,106]. Enzyme degradation has also been reported by the presence of nitrite, thiosulfate, and cyanide [25]. Finally, it is crucial to take into account that in situ monitoring of field samples might require analyses at extreme temperatures and pH values, which are generally responsible for detrimental conformational changes.
5. Concluding Remarks and Future Perspectives
In a broader perspective, when compared to the conventional methods, laccase-based biosensors have a tremendous potential for the detection of phenolic compounds in wastewater thanks to a fast and in situ sample analysis, cheaper cost per analysis and the possibility for “real time” measurements [22]. Despite these advantages, the implementation of laccase-based biosensors has been limited due to issues regarding enzyme stability, which are generally exacerbated by sudden changes in pH, temperature, and ionic force [21,25]. As a result, an area of increasingly growing interest is the development of novel immobilization strategies that rely on a deeper understanding of surface-protein and protein-protein interactions.
An additional challenge in the application of laccase-based biosensors is their limited ability to exclusively detect one of the compounds present in a mixture, and especially if they are all substrates of laccase. An avenue to overcome this issue is to incorporate surface modifications capable of preferentially immobilizing the enzyme molecules in a conformation/orientation that favors the interaction with only the molecules of interest. This is important for industrial applications where, by identifying each analyte separately, a focused intervention can be performed in an in-line and real-time manner. This approach has proven useful to improve the regio- and enantio-selectivity of various enzymes, as a direct consequence of increasing the rigidity of the active site. Active sites with suppressed flexibility show a significant reduction in their ability toward catalyzing different types of compounds thereby providing higher selectivities [107,108]. While increasing selectivity may be attractive for some applications, one might be cautious when altering the active site 3D conformation as the enzyme turnover may decrease considerably. This could potentially lead to a decline in the detection limits and sensitivity of the biosensing platforms.
The limited access to potable water and considering that industrialization has accelerated in emerging economies around the world increasing the amount of discharged residues, represents a difficult challenge for modern science and engineering. There is a growing concern with a class of wastewater residues known as Pharmaceuticals and Personal Care Products (PPCPs) as a major threat to human health. In this chapter, we reviewed the latest developments in electrochemical methods for the detection of pharmaceutical residues, and particularly those associated with enzyme-based biosensors. Also, we detailed amperometric systems coupled to microfluidic platforms for sample processing. Most of the progress in the field of enzymatic biosensors is mainly due to the advancement of technologies to enable miniaturization of prototypes, ease of handling and portability, and the reduction of cost for field implementation. Despite the encouraging results, important drawbacks are yet to be addressed including low sensitivity and selectivity in multicomponent mixtures, low resilience to changes in pH and temperature, limited immobilization yields for the bioactive components, and largely constrained long-term and operational stability. These are commonly found samples in real field wastewater samples where complex mixtures usually contain more than one analyte of interest and at very low concentrations. These drawbacks can be overcome by implementing low-cost microfluidic devices capable of selectively separating and increasing the concentration of the analytes of interest prior to sensing.
Future developments include multienzyme portable devices capable of detecting several compounds at the parts per billion level in real-time. This approach is, however, challenging due to issues regarding proper surface orientation and high conformational stability for the immobilized enzyme molecules, limited long-term and operational stability, processing and deconvoluting complex data signals, providing reproducible and reproducible manufacturing protocols, and most critically—shorter response times. Research in the biosensors area should be compelled to manufacture faster, smaller and more efficient devices through the integration of electronics and biological systems. To meet this goal, biosensors may include the use of new integrated circuits that optimize miniaturization and profitability, with the implementation of wireless and Internet of Things (IoT) technology for remote control and data handling.
Author Contributions
Conceptualization, J.F.O. and J.C.C.; methodology, A.L.C., S.L.F., M.J.N., O.P.F. and P.R.P.; software, S.L.F., A.L.C. and M.J.N.; validation S.L.F., A.L.C. and M.J.N.; formal analysis, A.L.C., S.L.F., M.J.N., O.P.F., P.R.P., J.F.O. and J.C.C.; investigation, A.L.C., S.L.F., M.J.N., O.P.F. and P.R.P.; resources, J.F.O.; data curation, M.J.N. and A.L.C.; writing—original draft preparation, A.L.C., S.L.F., M.J.N., O.P.F., P.R.P., J.F.O. and J.C.C.; writing—review and editing, A.L.C., S.L.F., M.J.N., O.P.F., P.R.P., J.F.O. and J.C.C.; visualization, M.J.N. and A.L.C.; supervision, J.F.O. and J.C.C.; project administration, J.F.O. and J.C.C.; funding acquisition, J.F.O.
Funding
This research was supported by Colciencias Grant “Formación de capital humano de alto nivel para el departamento del Cesar” to O.P.F.
Acknowledgments
We like to thank the Departments of Biomedical Engineering and Electrical and Electronics Engineering for access to simulation, CAD software and manufacture facilities.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Ejeian, F.; Etedali, P.; Mansouri-Tehrani, H.A.; Soozanipour, A.; Low, Z.X.; Asadnia, M.; Taheri-Kafrani, A.; Razmjou, A. Biosensors for wastewater monitoring: A review. Biosens. Bioelectron. 2018, 118, 66–79. [Google Scholar] [CrossRef] [PubMed]
- Morais, R.L.; Garcia, L.F.; Moreno, E.K.; Thomaz, D.V.; Rodrigues, M.F.; Vaz, B.G.; de Souza Gi, E. Electrochemical remediation of industrial pharmaceutical wastewater containing hormones in a pilot scale treatment system. Eclética Quím. J. 2019, 44, 40–52. [Google Scholar] [CrossRef]
- Coleman, C.; Huang, C.-H.; Lee, W.-N.; Nowack, K.; Carter, J.; Fu, J. Removal of pharmaceuticals and personal care products by two-stage biofiltration for drinking water treatment. Sci. Total Environ. 2019, 664, 240–248. [Google Scholar]
- Paucar, N.E.; Kim, I.; Tanaka, H.; Sato, C. Ozone treatment process for the removal of pharmaceuticals and personal care products in wastewater. Ozone Sci. Eng. 2019, 41, 3–16. [Google Scholar] [CrossRef]
- Gaudin, V. Advances in biosensor development for the screening of antibiotic residues in food products of animal origin—A comprehensive review. Biosens. Bioelectron. 2017, 90, 363–377. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Zhao, W.-M. Optical methods of antibiotic residues detections: A comprehensive review. Sens. Actuators B Chem. 2018, 269, 238–256. [Google Scholar] [CrossRef]
- Wang, S.; Xu, L.-P.; Zhang, X. Ultrasensitive Electrochemical Biosensor Based on Noble Metal Nanomaterials. Sci. Adv. Mater. 2015, 7, 2084–2102. [Google Scholar] [CrossRef]
- Lan, L.; Yao, Y.; Ping, J.; Ying, Y. Recent advances in nanomaterial-based biosensors for antibiotics detection. Biosens. Bioelectron. 2017, 91, 504–514. [Google Scholar] [CrossRef] [PubMed]
- Nikoleli, G.P.; Nikolelis, D.P.; Siontorou, C.G.; Karapetis, S.; Nikolelis, M.T. Application of biosensors based on lipid membranes for the rapid detection of toxins. Biosensors 2018, 8, 61. [Google Scholar] [CrossRef] [PubMed]
- Rackus, D.G.; Shamsi, M.H.; Wheeler, A.R. Electrochemistry, biosensors and microfluidics: A convergence of fields. Chem. Soc. Rev. 2015, 44, 5320–5340. [Google Scholar] [CrossRef] [PubMed]
- Bahadır, E.B.; Sezgintürk, M.K. Applications of commercial biosensors in clinical, food, environmental, and biothreat/biowarfare analyses. Anal. Biochem. 2015, 478, 107–120. [Google Scholar] [CrossRef] [PubMed]
- Huet, A.-C.; Delahaut, P.; Fodey, T.; Haughey, S.A.; Elliott, C.; Weigel, S. Advances in biosensor-based analysis for antimicrobial residues in foods. TrAC Trends Anal. Chem. 2010, 29, 1281–1294. [Google Scholar] [CrossRef]
- Pullano, S.A.; Critello, C.D.; Mahbub, I.; Tasneem, N.T.; Shamsir, S.; Islam, S.K.; Greco, M.; Fiorillo, A.S. EGFET-based sensors for bioanalytical applications: A review. Sensors 2018, 18, 4042. [Google Scholar] [CrossRef] [PubMed]
- Monteiro, T.; Almeida, M.G. Electrochemical Enzyme Biosensors Revisited: Old Solutions for New Problems. Crit. Rev. Anal. Chem. 2018, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Zhu, C.; Yang, G.; Li, H.; Du, D.; Lin, Y. Electrochemical Sensors and Biosensors Based on Nanomaterials and Nanostructures. Anal. Chem. 2015, 87, 230–249. [Google Scholar] [CrossRef] [PubMed]
- Kurbanoglu, S.; Ozkan, S.A.; Merkoçi, A. Nanomaterials-based enzyme electrochemical biosensors operating through inhibition for biosensing applications. Biosens. Bioelectron. 2017, 89, 886–898. [Google Scholar] [CrossRef] [PubMed]
- Rocchitta, G.; Spanu, A.; Babudieri, S.; Latte, G.; Madeddu, G.; Galleri, G.; Nuvoli, S.; Bagella, P.; Demartis, M.; Fiore, V.; et al. Enzyme Biosensors for Biomedical Applications: Strategies for Safeguarding Analytical Performances in Biological Fluids. Sensors 2016, 16, 780. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez-Rivera, J.C.; Osma, J.F. Fabrication of an Amperometric Flow-Injection Microfluidic Biosensor Based on Laccase for In Situ Determination of Phenolic Compounds. BioMed Res. Int. 2015, 2015, 845261. [Google Scholar] [CrossRef] [PubMed]
- Pohanka, M.; Skládal, P. Electrochemical Biosensors—Principles and Applications—A Review. J. Appl. Biomed. 2008, 6, 57–64. [Google Scholar]
- García-Morales, R.; García-García, A.; Orona-Navar, C.; Osma, J.F.; Nigam, K.D.P.; Ornelas-Soto, N. Biotransformation of emerging pollutants in groundwater by laccase from P. sanguineus CS43 immobilized onto titania nanoparticles. J. Environ. Chem. Eng. 2018, 6, 710–717. [Google Scholar] [CrossRef]
- Reyes-De-Corcuera, J.I.; Olstad, H.E.; García-Torres, R. Stability and Stabilization of Enzyme Biosensors: The Key to Successful Application and Commercialization. Annu. Rev. Food Sci. Technol. 2018, 9, 293–322. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Delgado, M.M.; Alemán-Nava, G.S.; Rodríguez-Delgado, J.M.; Dieck-Assad, G.; Martínez-Chapa, S.O.; Barceló, D.; Parra, R. Laccase-based biosensors for detection of phenolic compounds. TrAC Trends Anal. Chem. 2015, 74, 21–45. [Google Scholar] [CrossRef]
- Yashas, S.R.; Shivakumara, B.R.; Udayashankara, T.H.; Krishna, B.M. Laccase biosensor: Green technique for quantification of phenols in wastewater (a review). Orient. J. Chem. 2018, 34, 631–637. [Google Scholar]
- Gul, I.; Ahmad, M.S.; Naqvi, S.S.; Hussain, A.; Wali, R.; Farooqi, A.A.; Ahmed, I. Polyphenol oxidase (PPO) based biosensors for detection of phenolic compounds: A Review. J. Appl. Biol. Biotechnol. 2017, 5, 72–85. [Google Scholar]
- Rodríguez-Delgado, M.; Ornelas-Soto, N. Laccases: A blue enzyme for greener alternative technologies in the detection and treatment of emerging pollutants. In Green Technologies and Environmental Sustainability; Springer International Publishing: Cham, Switzerland, 2017; pp. 45–65. [Google Scholar]
- Nazari, M.; Kashanian, S.; Rafipour, R. Laccase immobilization on the electrode surface to design a biosensor for the detection of phenolic compound such as catechol. Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 2015, 145, 130–138. [Google Scholar] [CrossRef] [PubMed]
- Morozova, O.V.; Shumakovich, G.P.; Gorbacheva, M.A.; Shleev, S.V.; Yaropolov, A.I. ‘Blue’ laccases. Biochemistry 2007, 72, 1136–1150. [Google Scholar] [CrossRef] [PubMed]
- Riva, S. Laccases: Blue enzymes for green chemistry. Trends Biotechnol. 2006, 24, 219–226. [Google Scholar] [CrossRef] [PubMed]
- Bai, X.; Gu, H.; Chen, W.; Shi, H.; Yang, B.; Huang, X.; Zhang, Q. Immobilized laccase on activated poly(vinyl alcohol) microspheres for enzyme thermistor application. Appl. Biochem. Biotechnol. 2014, 173, 1097–1107. [Google Scholar] [CrossRef] [PubMed]
- Chinnadayyala, S.R.; Kakoti, A.; Santhosh, M.; Goswami, P. A novel amperometric alcohol biosensor developed in a 3rd generation bioelectrode platform using peroxidase coupled ferrocene activated alcohol oxidase as biorecognition system. Biosens. Bioelectron. 2014, 55, 120–126. [Google Scholar] [CrossRef] [PubMed]
- Kuswandi, B.; Irmawati, T.; Hidayat, M.A.; Jayus; Ahmad, M. A simple visual ethanol biosensor based on alcohol oxidase immobilized onto polyaniline film for halal verification of fermented beverage samples. Sensors 2014, 14, 2135–2149. [Google Scholar] [CrossRef] [PubMed]
- Mazlan, S.Z.; Lee, Y.H.; Hanifah, S.A. A new Laccase based biosensor for tartrazine. Sensors 2017, 17, 2859. [Google Scholar] [CrossRef] [PubMed]
- Reverté, L.; Prieto-Simón, B.; Campàs, M. New advances in electrochemical biosensors for the detection of toxins: Nanomaterials, magnetic beads and microfluidics systems. A review. Anal. Chim. Acta 2016, 908, 8–21. [Google Scholar] [CrossRef] [PubMed]
- Thakur, S.; Rao, S.N. Theoretical Microfluidics; Oxford University Press: Oxford, UK, 2014. [Google Scholar]
- Kirby, B.J. Micro- and Nanoscale Fluid Mechanics: Transport in Microfluidic Devices: Species and Charge Transport; Cambridge University Press: Cambridge, MA, USA, 2010. [Google Scholar]
- Fox, A.T.; McDonald, W.R.; Pritchard, P.J. Fox and McDonald’s Introduction to Fluid Mechanics, 8th ed.; Wiley: Hoboken, NJ, USA, 1999. [Google Scholar]
- Prakash, S.; Yeom, J. Nanofluidics and Microfluidics: Systems and Applications; Elsevier: Amsterdam, The Netherlands, 2014. [Google Scholar]
- Martel, J.M.; Toner, M. Inertial focusing dynamics in spiral microchannels. Phys. Fluids 2012, 24, 371–396. [Google Scholar] [CrossRef] [PubMed]
- Salafi, T.; Zeming, K.K.; Zhang, Y. Advancements in microfluidics for nanoparticle separation. Lab Chip 2017, 17, 11–33. [Google Scholar] [CrossRef] [PubMed]
- SooHoo, J.R.; Walker, G.M. Microfluidic aqueous two phase system for leukocyte concentration from whole blood. Biomed. Microdevices 2009, 11, 323–329. [Google Scholar] [CrossRef] [PubMed]
- Tsukamoto, M.; Taira, S.; Yamamura, S.; Morita, Y.; Nagatani, N.; Takamura, Y.; Tamiya, E. Cell separation by an aqueous two-phase system in a microfluidic device. Analyst 2009, 134, 1994–1998. [Google Scholar] [CrossRef] [PubMed]
- Yung, C.W.; Fiering, J.; Mueller, A.J.; Ingber, D.E. Micromagnetic-microfluidic blood cleansing device. Lab Chip 2009, 9, 1171–1177. [Google Scholar] [CrossRef] [PubMed]
- Wu, D.; Qin, J.; Lin, B. Electrophoretic separations on microfluidic chips. J. Chromatogr. A 2008, 1184, 542–559. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.Y.; Chang, C.L.; Wang, Y.N.; Fu, L.M. Microfluidic mixing: A review. Int. J. Mol. Sci. 2011, 12, 3263–3287. [Google Scholar] [CrossRef] [PubMed]
- Hessel, V.; Löwe, H.; Schönfeld, F. Micromixers—A review on passive and active mixing principles. Chem. Eng. Sci. 2005, 60, 2479–2501. [Google Scholar] [CrossRef]
- Meijer, H.E.H.; Singh, M.K.; Kang, T.G.; den Toonder, J.M.J.; Anderson, P.D. Passive and active mixing in microfluidic devices. Macromol. Symp. 2009, 279, 201–209. [Google Scholar] [CrossRef]
- Dickinson, E.J.F.; Ekström, H.; Fontes, E. COMSOL Multiphysics®: Finite element software for electrochemical analysis. A mini-review. Electrochem. Commun. 2014, 40, 71–74. [Google Scholar] [CrossRef]
- Kaffash, A.; Rostami, K.; Zare, H.R. Modeling of an electrochemical nanobiosensor in COMSOL Multiphysics to determine phenol in the presence of horseradish peroxidase enzyme. Enzym. Microb. Technol. 2019, 121, 23–28. [Google Scholar] [CrossRef] [PubMed]
- Hossen, M.N.; Ferdous, M.; Khalek, M.A.; Chakma, S.; Paul, B.K.; Ahmed, K. Design and analysis of biosensor based on surface plasmon resonance. Sens. Bio-Sensing Res. 2018, 21, 1–6. [Google Scholar] [CrossRef]
- Wu, G.; Meyyappan, M.; Lai, K.W.C. Simulation of graphene field-effect transistor biosensors for bacterial detection. Sensors 2018, 18, 1715. [Google Scholar] [CrossRef] [PubMed]
- Berkowski, K.L.; Plunkett, K.N.; Yu, Q.; Moore, J.S. Introduction to Photolithography: Preparation of Microscale Polymer Silhouettes. J. Chem. Educ. 2005, 82, 1365. [Google Scholar] [CrossRef]
- Sun, S.; Leggett, G.J. Generation of Nanostructures by Scanning Near-Field Photolithography of Self-Assembled Monolayers and Wet Chemical Etching. Nano Lett. 2002, 2, 1223–1227. [Google Scholar] [CrossRef]
- Brower, K.; White, A.K.; Fordyce, P.M. Multi-step Variable Height Photolithography for Valved Multilayer Microfluidic Devices. J. Vis. Exp. 2017, 55276. [Google Scholar] [CrossRef] [PubMed]
- Szilasi, S.Z.; Cserháti, C. Selective etching of PDMS: Etching technique for application as a positive tone resist. Appl. Surf. Sci. 2018, 457, 662–669. [Google Scholar] [CrossRef]
- Huang, Y.; Liu, S.; Yang, W.; Yu, C. Surface roughness analysis and improvement of PMMA-based microfluidic chip chambers by CO2laser cutting. Appl. Surf. Sci. 2010, 256, 1675–1678. [Google Scholar] [CrossRef]
- Carbaugh, D.J.; Wright, J.T.; Rahman, F. Negative tone photolithography with photo-sensitised polymethyl methacrylate (PMMA). Microelectron. Eng. 2017, 171, 53–59. [Google Scholar] [CrossRef]
- Trantidou, T.; Friddin, M.S.; Gan, K.B.; Han, L.; Bolognesi, G.; Brooks, N.J.; Ces, O. Mask-free laser lithography for rapid and low-cost microfluidic device fabrication. Anal. Chem. 2018, 90, 13915–13921. [Google Scholar] [CrossRef] [PubMed]
- Stasyuk, N.; Gayda, G.; Zakalskiy, A.; Zakalska, O.; Serkiz, R.; Gonchar, M. Amperometric biosensors based on oxidases and PtRu nanoparticles as artificial peroxidase. Food Chem. 2019, 285, 213–220. [Google Scholar] [CrossRef] [PubMed]
- Soylemez, S.; Kaya, H.Z.; Udum, Y.A.; Toppare, L. A multipurpose conjugated polymer: Electrochromic device and biosensor construction for glucose detection. Org. Electron. Phys. Mater. Appl. 2019, 65, 327–333. [Google Scholar] [CrossRef]
- Monti, D.; Ottolina, G.; Carrea, G.; Riva, S. Redox Reactions Catalyzed by Isolated Enzymes. Chem. Rev. 2011, 111, 4111–4140. [Google Scholar] [CrossRef] [PubMed]
- Colmati, F.; Sgobbi, L.F.; Teixeira, G.F.; Vilela, R.S.; Martins, T.D.; Figueiredo, G.O. Electrochemical Biosensors Containing Pure Enzymes or Crude Extracts as Enzyme Sources for Pesticides and Phenolic Compounds with Pharmacological Property Detection and Quantification. In Environmental Biosensors; IntechOpen: Goiânia, Brazil, 2019. [Google Scholar]
- Gayda, G.Z.; Demkiv, O.M.; Stasyuk, N.Y.; Serkiz, R.Y.; Lootsik, M.D.; Errachid, A.; Gonchar, M.V.; Nisnevitch, M. Metallic Nanoparticles Obtained via ‘Green’ Synthesis as a Platform for Biosensor Construction. Appl. Sci. 2019, 9, 720. [Google Scholar] [CrossRef]
- Xiao, F.; Wang, L.; Duan, H. Nanomaterial based electrochemical sensors for in vitro detection of small molecule metabolites. Biotechnol. Adv. 2016, 34, 234–249. [Google Scholar] [CrossRef] [PubMed]
- Pitschmann, V.; Urban, M.; Dědič, J.; Matějovský, L.; Dymák, M.; Lobotka, M. Modified Biosensor for Cholinesterase Inhibitors with Guinea Green B as the Color Indicator. Biosensors 2018, 8, 81. [Google Scholar] [CrossRef] [PubMed]
- Antunes, R.; Ferraz, D.; Garcia, L.; Thomaz, D.; Luque, R.; Lobón, G.; Gil, E.; Lopes, F. Development of a Polyphenol Oxidase Biosensor from Jenipapo Fruit Extract (Genipa americana L.) and Determination of Phenolic Compounds in Textile Industrial Effluents. Biosensors 2018, 8, 47. [Google Scholar] [CrossRef] [PubMed]
- Sassolas, A.; Blum, L.J.; Leca-Bouvier, B.D. Immobilization strategies to develop enzymatic biosensors. Biotechnol. Adv. 2012, 30, 489–511. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.; Park, S. Glucose biosensor based on GOx/HRP bienzyme at liquid–crystal/aqueous interface. J. Colloid Interface Sci. 2015, 457, 281–288. [Google Scholar] [CrossRef] [PubMed]
- Xia, H.Q.; Kitazumi, Y.; Shirai, O.; Ohta, H.; Kurihara, S.; Kano, K. Putrescine oxidase/peroxidase-co-immobilized and mediator-less mesoporous microelectrode for diffusion-controlled steady-state amperometric detection of putrescine. J. Electroanal. Chem. 2017, 804, 128–132. [Google Scholar] [CrossRef]
- Pérez, J.P.H.; López-Ruiz, B.; López-Cabarcos, E. Synthesis and characterization of microparticles based on poly-methacrylic acid with glucose oxidase for biosensor applications. Talanta 2016, 149, 310–318. [Google Scholar] [CrossRef] [PubMed]
- Bottoni, P.; Caroli, S. Detection and quantification of residues and metabolites of medicinal products in environmental compartments, food commodities and workplaces. A review. J. Pharm. Biomed. Anal. 2015, 106, 3–24. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, M.J.; Hameed, B.H. Removal of emerging pharmaceutical contaminants by adsorption in a fixed-bed column: A review. Ecotoxicol. Environ. Saf. 2018, 149, 257–266. [Google Scholar] [CrossRef] [PubMed]
- Rebollar-Pérez, G.; Campos-Terán, J.; Ornelas-Soto, N.; Méndez-Albores, A.; Torres, E. Biosensors based on oxidative enzymes for detection of environmental pollutants. Biocatalysis 2016, 1, 118–129. [Google Scholar] [CrossRef]
- Narang, J.; Malhotra, N.; Singh, S.; Singh, G.; Pundir, C.S. Monitoring analgesic drug using sensing method based on nanocomposite. RSC Adv. 2015, 5, 2396–2404. [Google Scholar] [CrossRef]
- Jaouani, A.; Guillén, F.; Penninckx, M.J.; Martínez, A.T.; Martínez, M.J. Role of Pycnoporus coccineus laccase in the degradation of aromatic compounds in olive oil mill wastewater. Enzym. Microb. Technol. 2005, 36, 478–486. [Google Scholar] [CrossRef]
- Sheikhi, F.; Ardakani, M.R.; Enayatizamir, N.; Rodriguez-Couto, S. The Determination of Assay for Laccase of Bacillus subtilis WPI with Two Classes of Chemical Compounds as Substrates. Indian J. Microbiol. 2012, 52, 701–707. [Google Scholar] [CrossRef] [PubMed]
- Viswanath, B.; Rajesh, B.; Janardhan, A.; Kumar, A.P.; Narasimha, G. Fungal laccases and their applications in bioremediation. Enzym. Res. 2014, 2014, 163242. [Google Scholar] [CrossRef] [PubMed]
- Bhalla, N.; Jolly, P.; Formisano, N.; Estrela, P. Introduction to biosensors. Essays Biochem. 2016, 60, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Leonardo, S.; Toldrà, A. Trends and Prospects on Electrochemical Biosensors for the Detection of Marine Toxins. Compr. Anal. Chem. 2017, 78, 303–341. [Google Scholar]
- Brondani, D.; Scheeren, C.W.; Dupont, J.; Vieira, I.C. Biosensor based on platinum nanoparticles dispersed in ionic liquid and laccase for determination of adrenaline. Sens. Actuators B Chem. 2009, 140, 252–259. [Google Scholar] [CrossRef]
- Baluta, S.; Lesiak, A.; Graphene, J.C. Quantum Dots-based Electrochemical Biosensor for Catecholamine Neurotransmitters Detection. Electroanalysis 2018, 30, 1773–1782. [Google Scholar] [CrossRef]
- Santhiago, M.; Vieira, I.C. l-Cysteine determination in pharmaceutical formulations using a biosensor based on laccase from Aspergillus oryzae. Sens. Actuators B Chem. 2007, 128, 279–285. [Google Scholar] [CrossRef]
- Zhao, W.; Wang, K.; Wei, Y.; Ma, Y.; Liu, L.; Huang, X. Laccase biosensor based on phytic acid modification of nanostructured SiO2 surface for sensitive detection of dopamine. Langmuir 2014, 30, 11131–11137. [Google Scholar] [CrossRef] [PubMed]
- Leite, O.D.; Lupetti, K.O.; Fatibello-Filho, O.; Vieira, I.C.; Barbosa, A.D. Synergic effect studies of the bi-enzymatic system laccaseperoxidase in a voltammetric biosensor for catecholamines. Talanta 2003, 59, 889–896. [Google Scholar] [CrossRef]
- Das, P.; Barbora, L.; Das, M.; Goswami, P. Highly sensitive and stable laccase based amperometric biosensor developed on nano-composite matrix for detecting pyrocatechol in environmental samples. Sens. Actuators B Chem. 2014, 192, 737–744. [Google Scholar] [CrossRef]
- Rahman, M.A.; Noh, H.B.; Shim, Y.B. Direct electrochemistry of laccase immobilized on Au nanoparticles encapsulated-dendrimer bonded conducting polymer: Application for a catechin sensor. Anal. Chem. 2008, 80, 8020–8027. [Google Scholar] [CrossRef] [PubMed]
- Chen, T.; Xu, Y.; Wei, S.; Li, A.; Huang, L.; Liu, J. A signal amplification system constructed by bi-enzymes and bi-nanospheres for sensitive detection of norepinephrine and miRNA. Biosens. Bioelectron. 2019, 124–125, 224–232. [Google Scholar] [CrossRef] [PubMed]
- Portaccio, M.; Di Martino, S.; Maiuri, P.; Durante, D.; De Luca, P.; Lepore, M.; Bencivenga, U.; Rossi, S.; De Maio, A.; Mita, D.G. Biosensors for phenolic compounds: The catechol as a substrate model. J. Mol. Catal. B Enzym. 2006, 41, 97–102. [Google Scholar] [CrossRef]
- Kulys, J.; Vidziunaite, R. Amperometric biosensors based on recombinant laccases for phenols determination. Biosens. Bioelectron. 2002, 18, 319–325. [Google Scholar] [CrossRef]
- Roy, J.J.; Abraham, T.E.; Abhijith, K.S.; Kumar, P.V.S.; Thakur, M.S. Biosensor for the determination of phenols based on Cross-Linked Enzyme Crystals (CLEC) of laccase. Biosens. Bioelectron. 2005, 21, 206–211. [Google Scholar] [CrossRef] [PubMed]
- Odaci, D.; Timur, S.; Pazarlioğlu, N.; Kirgöz, Ü.A.; Telefoncu, A. Effects of mediators on the laccase biosensor response in paracetamol detection. Biotechnol. Appl. Biochem. 2006, 45, 23–28. [Google Scholar] [PubMed]
- Li, Y.; Zhang, L.; Li, M.; Pan, Z.; Li, D. A disposable biosensor based on immobilization of laccase with silica spheres on the MWCNTs-doped screen-printed electrode. Chem. Cent. J. 2012, 6, 103. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Li, D.; Li, G.; Luo, L.; Ullah, N.; Wei, Q.; Huang, F. Facile fabrication of gold nanoparticle on zein ultrafine fibers and their application for catechol biosensor. Appl. Surf. Sci. 2015, 328, 444–452. [Google Scholar] [CrossRef]
- Bauer, C.G.; Kühn, A.; Gajovic, N.; Skorobogatko, O.; Holt, P.J.; Bruce, N.C.; Makower, A.; Lowe, C.R.; Scheller, F.W. New enzyme sensors for morphine and codeine based on morphine dehydrogenase and laccase. Fresenius J. Anal. Chem. 1999, 364, 179–183. [Google Scholar] [CrossRef]
- Moraes, J.T.; Salamanca-Neto, C.A.; Švorc, Ĺ.; Schirmann, J.G.; Barbosa-Dekker, A.M.; Dekker, R.F.; Sartori, E.R. Laccase from Botryosphaeria rhodina MAMB-05 as a biological component in electrochemical biosensing devices. Anal. Methods 2019, 11, 717–720. [Google Scholar] [CrossRef]
- Liu, Y.; Qu, X.; Guo, H.; Chen, H.; Liu, B.; Dong, S. Facile preparation of amperometric laccase biosensor with multifunction based on the matrix of carbon nanotubes-chitosan composite. Biosens. Bioelectron. 2006, 21, 2195–2201. [Google Scholar] [CrossRef] [PubMed]
- Ferry, Y.; Leech, D. Amperometric Detection of Catecholamine Neurotransmitters Using Electrocatalytic Substrate Recycling at a Laccase Electrode. Electroanalysis 2005, 17, 113–119. [Google Scholar] [CrossRef]
- Sanz, J.; de Marcos, S.; Galbán, J. Autoindicating optical properties of laccase as the base of an optical biosensor film for phenol determination. Anal. Bioanal. Chem. 2012, 404, 351–359. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Fang, H.; Liu, C.; Gu, E.; Jiang, D. A novel fiber optic biosensor for the determination of adrenaline based on immobilized laccase catalysis. Anal. Lett. 2008, 41, 1430–1442. [Google Scholar] [CrossRef]
- Silva, L.I.B.; Ferreira, F.D.P.; Freitas, A.C.; Rocha-Santos, T.A.P.; Duarte, A.C. Optical fiber biosensor coupled to chromatographic separation for screening of dopamine, norepinephrine and epinephrine in human urine and plasma. Talanta 2009, 80, 853–857. [Google Scholar] [CrossRef] [PubMed]
- Abdullah, J.; Ahmad, M.; Lee, Y.H.; Karuppiah, N.; Sidek, H. An optical biosensor based on immobilization of lacease and MBTH in stacked films for the detection of catechol. Sensors 2007, 7, 2238–2250. [Google Scholar] [CrossRef] [PubMed]
- Karami, C.; Taher, M.A. A catechol biosensor based on immobilizing laccase to Fe3O4@Au core-shell nanoparticles. Int. J. Biol. Macromol. 2019, 129, 84–90. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, F.D.P.; Silva, L.I.B.; Freitas, A.C.; Rocha-Santos, T.A.P.; Duarte, A.C. High performance liquid chromatography coupled to an optical fiber detector coated with laccase for screening catecholamines in plasma and urine. J. Chromatogr. A 2009, 1216, 7049–7054. [Google Scholar] [CrossRef] [PubMed]
- Mohapatra, S.; Padhye, L.P.; Mukherji, S. Challenges in Detection of Antibiotics in Wastewater Matrix; Springer: Singapore, 2018; pp. 3–20. [Google Scholar]
- Anku, W.W.; Mamo, M.A.; Govender, P.P. Phenolic Compounds in Water: Sources, Reactivity, Toxicity and Treatment Methods. In Phenolic Compounds—Natural Sources, Importance and Applications; InTech: Johannesburg, South Africa, 2017. [Google Scholar]
- Zilly, A.; da Silva Coelho-Moreira, J.; Bracht, A.; De Souza, C.G.; Carvajal, A.E.; Koehnlein, E.A.; Peralta, R.M. Influence of NaCl and Na2SO4 on the kinetics and dye decolorization ability of crude laccase from Ganoderma lucidum. Int. Biodeterior. Biodegrad. 2011, 65, 340–344. [Google Scholar] [CrossRef]
- Margot, J.; Copin, P.J.; von Gunten, U.; Barry, D.A.; Holliger, C. Sulfamethoxazole and isoproturon degradation and detoxification by a laccase-mediator system: Influence of treatment conditions and mechanistic aspects. Biochem. Eng. J. 2015, 103, 47–59. [Google Scholar] [CrossRef]
- Mogharabi, M.; Rezaei, S.; Faramarzi, M.A. Trends in Peptide and Protein Sciences; Shahid Beheshti University of Medical Sciences, Laser Application in Medical Sciences Research Center: Tehran, Iran, 2017; Volume 1. [Google Scholar]
- Denard, C.A.; Ren, H.; Zhao, H. Improving and repurposing biocatalysts via directed evolution. Curr. Opin. Chem. Biol. 2015, 25, 55–64. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).