Detection of Circulating Tumor Cells Using Membrane-Based SERS Platform: A New Diagnostic Approach for ‘Liquid Biopsy’
Abstract
:1. Introduction
2. Experimental Section
2.1. Leucocyte Isolation
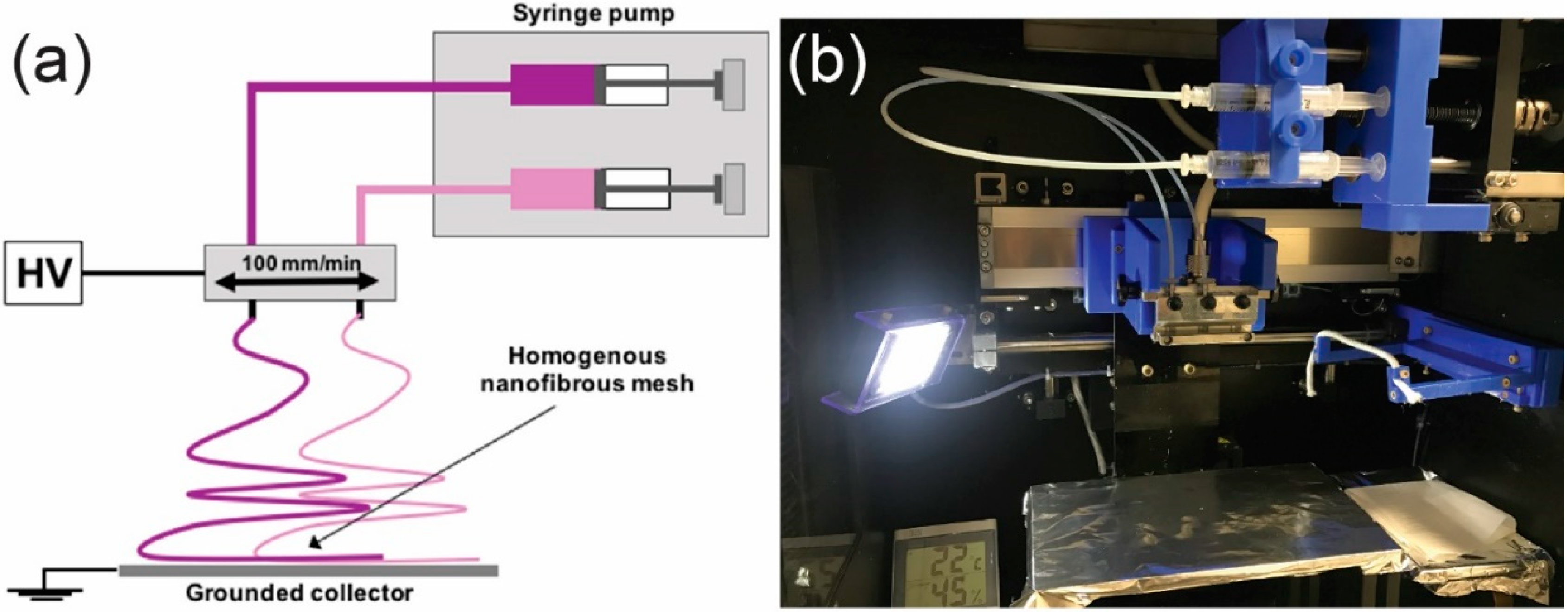
2.2. Fabrication of Membrane-Based SERS Platforms (MBSP) via Electrospinning
2.3. Sputtering of the Thin Layer of SERS Active Metal
3. Instrumentation
3.1. SERS Measurements
3.2. SEM Measurements
3.3. Chemometrics—Principal Component Analysis
4. Results and Discussion
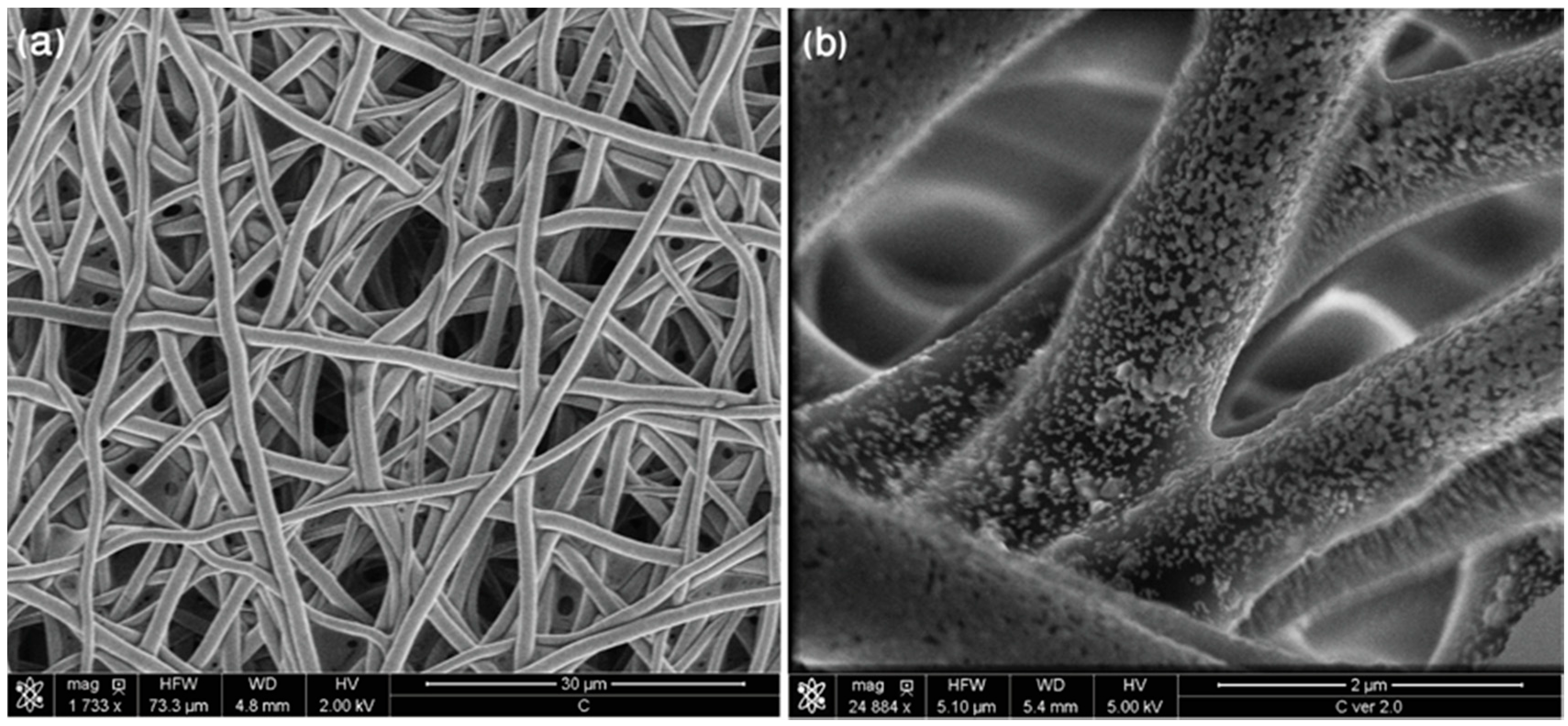
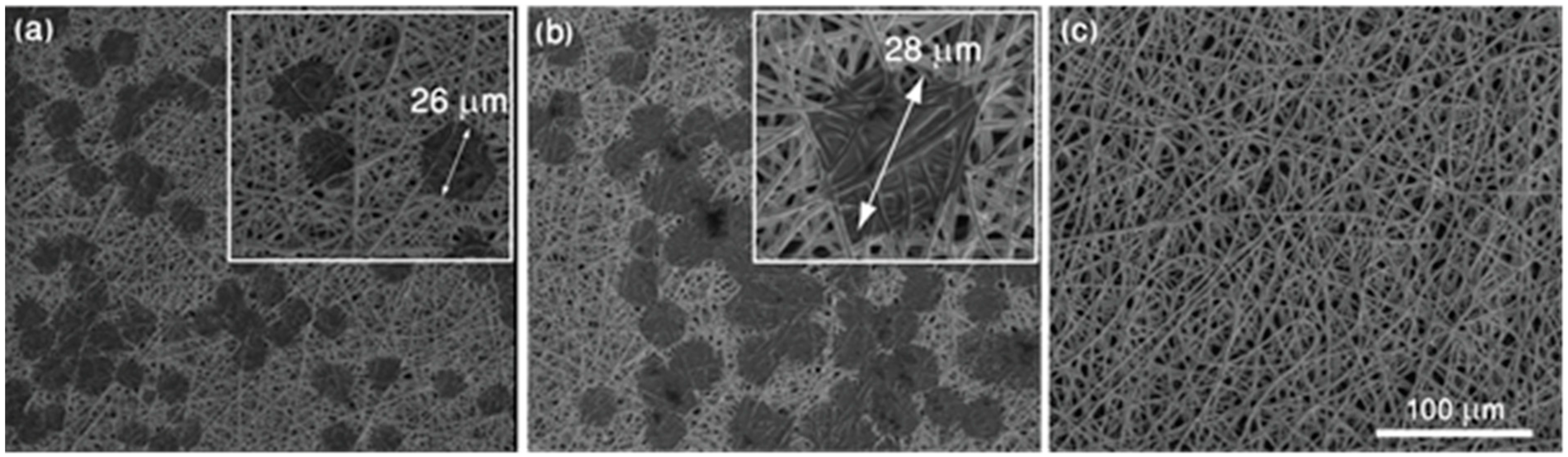
4.1. Preparation of the SERS Platform
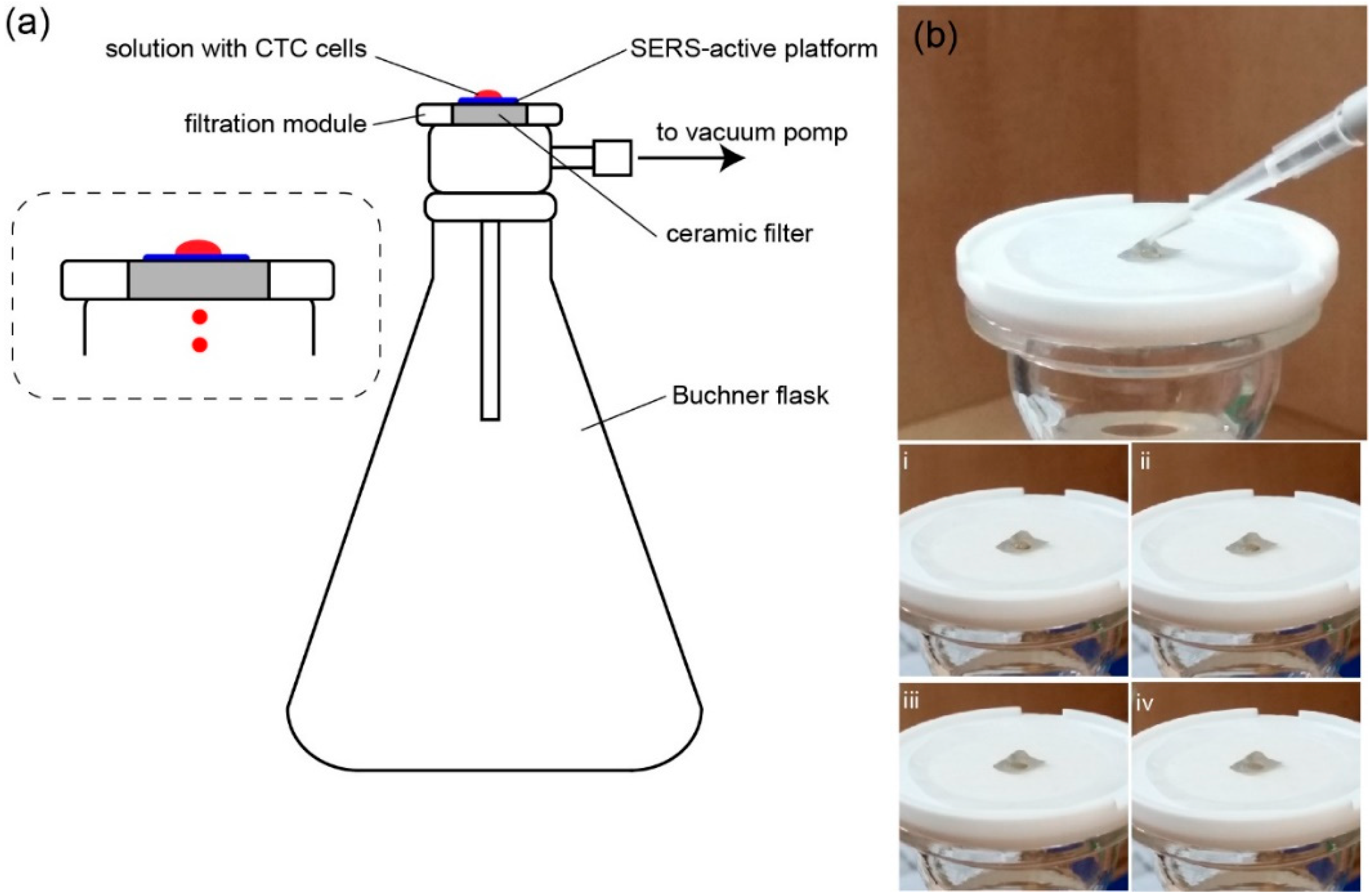
4.2. SERS Investigations of Circulating Tumor Cells
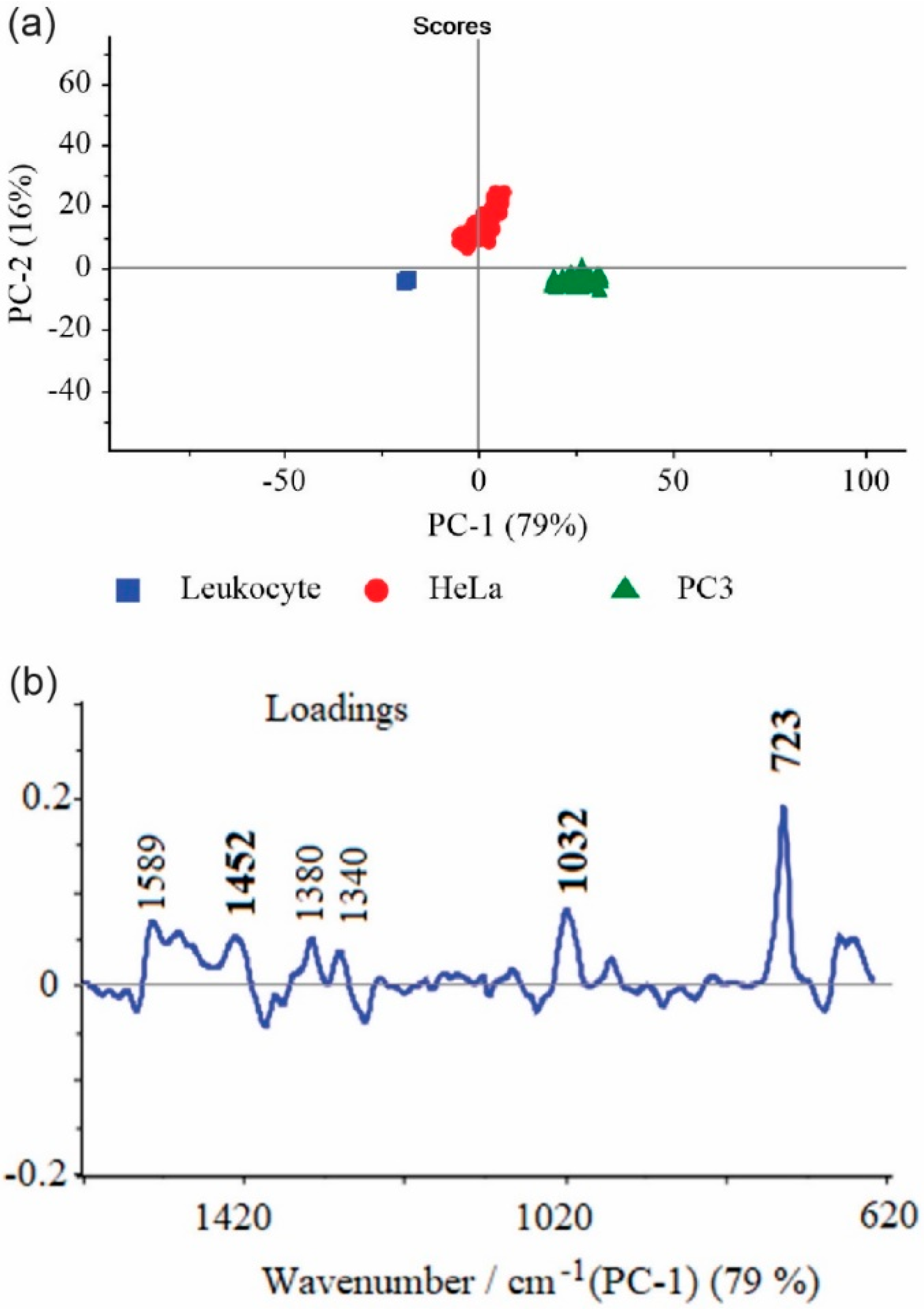
4.3. Principal Component Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Ghossein, R.A.; Bhattacharya, S.; Rosai, J. Molecular Detection of Micrometastases and Circulating Tumor Cells in Solid Tumors. Clin. Cancer Res. 1999, 5, 1950–1960. [Google Scholar] [PubMed]
- Swaby, R.F.; Cristofanilli, M. Circulating tumor cells in breast cancer: A tool whose time has come of age. BMC Med. 2011, 9, 43. [Google Scholar] [CrossRef] [PubMed]
- Danila, D.C.; Fleisher, M.; Scher, H.I. Circulating Tumor Cells as Biomarkers in Prostate Cancer. Clin. Cancer Res. 2011, 17, 3903–3912. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, F.; Yoneda, K.; Kondo, N.; Hashimoto, M.; Takuwa, T.; Matsumoto, S.; Okumura, Y.; Rahman, S.; Tsubota, N.; Tsujimura, T.; et al. Circulating Tumor Cell as a Diagnostic Marker in Primary Lung Cancer. Clin. Cancer Res. 2009, 15, 6980–6986. [Google Scholar] [CrossRef] [PubMed]
- Alix-Panabieres, C.; Pantel, K. Challenges in circulating tumour cell research. Nat. Rev. Cancer 2014, 14, 623–631. [Google Scholar] [CrossRef] [PubMed]
- Nagrath, S.; Sequist, L.V.; Maheswaran, S.; Bell, D.W.; Irimia, D.; Ulkus, L.; Smith, M.R.; Kwak, E.L.; Digumarthy, S.; Muzikansky, A.; et al. Isolation of rare circulating tumour cells in cancer patients by microchip technology. Nature 2007, 450, 1235–1239. [Google Scholar] [CrossRef] [PubMed]
- Andergassen, U.; Kolbl, A.C.; Mahner, S.; Jeschke, U. Real-time RT-PCR systems for CTC detection from blood samples of breast cancer and gynaecological tumour patients (Review). Oncol. Rep. 2016, 35, 1905–1915. [Google Scholar] [CrossRef] [PubMed]
- Stott, S.L.; Hsu, C.H.; Tsukrov, D.I.; Yu, M.; Miyamoto, D.T.; Waltman, B.A.; Rothenberg, S.M.; Shah, A.M.; Smas, M.E.; Korir, G.K.; et al. Isolation of circulating tumor cells using a microvortex-generating herringbone-chip. Proc. Natl. Acad. Sci. USA 2010, 107, 18392–18397. [Google Scholar] [CrossRef] [PubMed]
- Gleghorn, J.P.; Pratt, E.D.; Denning, D.; Liu, H.; Bander, N.H.; Tagawa, S.T.; Nanus, D.M.; Giannakakou, P.A.; Kirby, B.J. Capture of circulating tumor cells from whole blood of prostate cancer patients using geometrically enhanced differential immunocapture (GEDI) and a prostate-specific antibody. Lab Chip 2010, 10, 27–29. [Google Scholar] [CrossRef] [PubMed]
- Ozkumur, E.; Shah, A.M.; Ciciliano, J.C.; Emmink, B.L.; Miyamoto, D.T.; Brachtel, E.; Yu, M.; Chen, P.-i.; Morgan, B.; Trautwein, J.; et al. Inertial Focusing for Tumor Antigen–Dependent and –Independent Sorting of Rare Circulating Tumor Cells. Sci. Transl. Med. 2013, 5, ra47–ra179. [Google Scholar] [CrossRef] [PubMed]
- Krebs, M.G.; Hou, J.-M.; Ward, T.H.; Blackhall, F.H.; Dive, C. Circulating tumour cells: Their utility in cancer management and predicting outcomes. Ther. Adv. Med. Oncol. 2010, 2, 351–365. [Google Scholar] [CrossRef] [PubMed]
- Yu, M.; Stott, S.; Toner, M.; Maheswaran, S.; Haber, D.A. Circulating tumor cells: Approaches to isolation and characterization. J. Cell Biol. 2011, 192, 373–382. [Google Scholar] [CrossRef] [PubMed]
- Pantel, K.; Alix-Panabieres, C. Real-time liquid biopsy in cancer patients: Fact or fiction? Cancer Res. 2013, 73, 6384–6388. [Google Scholar] [CrossRef] [PubMed]
- Gorges, T.M.; Pantel, K. Circulating tumor cells as therapy-related biomarkers in cancer patients. Cancer Immunol. Immunother. CII 2013, 62, 931–939. [Google Scholar] [CrossRef] [PubMed]
- Jin, C.; McFaul, S.M.; Duffy, S.P.; Deng, X.; Tavassoli, P.; Black, P.C.; Ma, H. Technologies for label-free separation of circulating tumor cells: From historical foundations to recent developments. Lab Chip 2014, 14, 32–44. [Google Scholar] [CrossRef] [PubMed]
- Yoon, H.J.; Kozminsky, M.; Nagrath, S. Emerging role of nanomaterials in circulating tumor cell isolation and analysis. ACS Nano 2014, 8, 1995–2017. [Google Scholar] [CrossRef] [PubMed]
- Whitesides, G.M. The origins and the future of microfluidics. Nature 2006, 442, 368. [Google Scholar] [CrossRef] [PubMed]
- Cristofanilli, M.; Budd, G.T.; Ellis, M.J.; Stopeck, A.; Matera, J.; Miller, M.C.; Reuben, J.M.; Doyle, G.V.; Allard, W.J.; Terstappen, L.W.M.M.; et al. Circulating Tumor Cells, Disease Progression, and Survival in Metastatic Breast Cancer. N. Engl. J. Med. 2004, 351, 781–791. [Google Scholar] [CrossRef] [PubMed]
- Olmos, D.; Arkenau, H.T.; Ang, J.E.; Ledaki, I.; Attard, G.; Carden, C.P.; Reid, A.H.; A’Hern, R.; Fong, P.C.; Oomen, N.B.; et al. Circulating tumour cell (CTC) counts as intermediate end points in castration-resistant prostate cancer (CRPC): A single-centre experience. Ann. Oncol. 2009, 20, 27–33. [Google Scholar] [CrossRef] [PubMed]
- Cohen, S.J.; Punt, C.J.; Iannotti, N.; Saidman, B.H.; Sabbath, K.D.; Gabrail, N.Y.; Picus, J.; Morse, M.A.; Mitchell, E.; Miller, M.C.; et al. Prognostic significance of circulating tumor cells in patients with metastatic colorectal cancer. Ann. Oncol. 2009, 20, 1223–1229. [Google Scholar] [CrossRef] [PubMed]
- Allard, W.J.; Matera, J.; Miller, M.C.; Repollet, M.; Connelly, M.C.; Rao, C.; Tibbe, A.G.; Uhr, J.W.; Terstappen, L.W. Tumor cells circulate in the peripheral blood of all major carcinomas but not in healthy subjects or patients with nonmalignant diseases. Clin. Cancer Res. 2004, 10, 6897–6904. [Google Scholar] [CrossRef] [PubMed]
- Neugebauer, U.; Bocklitz, T.; Clement, J.H.; Krafft, C.; Popp, J. Towards detection and identification of circulating tumour cells using Raman spectroscopy. Analyst 2010, 135, 3178–3182. [Google Scholar] [CrossRef] [PubMed]
- Campion, A.; Kambhampati, P. Surface-enhanced Raman scattering. Chem. Soc. Rev. 1998, 27, 241–250. [Google Scholar] [CrossRef]
- Ru, E.C.L.; Etchegoin, P.G. Single-Molecule Surface-Enhanced Raman Spectroscopy. Annu. Rev. Phys. Chem. 2012, 63, 65–87. [Google Scholar] [PubMed]
- Maiti, K.K.; Samanta, A.; Vendrell, M.; Soh, K.S.; Olivo, M.; Chang, Y.T. Multiplex cancer cell detection by SERS nanotags with cyanine and triphenylmethine Raman reporters. Chem. Commun. 2011, 47, 3514–3516. [Google Scholar] [CrossRef] [PubMed]
- Nima, Z.A.; Mahmood, M.; Xu, Y.; Mustafa, T.; Watanabe, F.; Nedosekin, D.A.; Juratli, M.A.; Fahmi, T.; Galanzha, E.I.; Nolan, J.P.; et al. Circulating tumor cell identification by functionalized silver-gold nanorods with multicolor, super-enhanced SERS and photothermal resonances. Sci. Rep. 2014, 4, 4752. [Google Scholar] [CrossRef] [PubMed]
- Hoonejani, M.R.; Pallaoro, A.; Braun, G.B.; Moskovits, M.; Meinhart, C.D. Quantitative multiplexed simulated-cell identification by SERS in microfluidic devices. Nanoscale 2015, 7, 16834–16840. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Zhang, R.; Gao, M.; Zhang, X. Novel Nitrocellulose Membrane Substrate for Efficient Analysis of Circulating Tumor Cells Coupled with Surface-Enhanced Raman Scattering Imaging. ACS Appl. Mater. Interfaces 2014, 6, 370–376. [Google Scholar] [CrossRef] [PubMed]
- Jun, B.H.; Noh, M.S.; Kim, J.; Kim, G.; Kang, H.; Kim, M.S.; Seo, Y.T.; Baek, J.; Kim, J.H.; Park, J.; et al. Multifunctional Silver-Embedded Magnetic Nanoparticles as SERS Nanoprobes and Their Applications. Small 2010, 6, 119–125. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Qian, X.M.; Beitler, J.J.; Chen, Z.G.; Khuri, F.R.; Lewis, M.M.; Shin, H.J.C.; Nie, S.M.; Shin, D.M. Detection of Circulating Tumor Cells in Human Peripheral Blood Using Surface-Enhanced Raman Scattering Nanoparticles. Cancer Res. 2011, 71, 1526–1532. [Google Scholar] [CrossRef] [PubMed]
- Chang, Z.M.; Wang, Z.; Shao, D.; Yue, J.; Xing, H.; Li, L.; Ge, M.; Li, M.; Yan, H.; Hu, H.; et al. Shape Engineering Boosts Magnetic Mesoporous Silica Nanoparticle-Based Isolation and Detection of Circulating Tumor Cells. ACS Appl. Mater. Interfaces 2018, 10, 10656–10663. [Google Scholar] [CrossRef] [PubMed]
- Shi, W.; Paproski, R.J.; Moore, R.B.; Zemp, R. Detection of Circulating Tumor Cells Using Targeted Surface-Enhanced Raman Scattering Nanoparticles and Magnetic Enrichment; SPIE: Bellingham, WA, USA, 2014; p. 8. [Google Scholar]
- Krafft, C.; Beleites, C.; Schie, I.W.; Clement, J.H.; Popp, J. Raman-based Identification of Circulating Tumor Cells for Cancer Diagnosis. In Biomedical Vibrational Spectroscopy 2016: Advances in Research and Industry; Mahadevan-Jansen, A., Petrich, W., Eds.; Spie-Int Soc Optical Engineering: Bellingham, WA, USA, 2016; Volume 9704. [Google Scholar]
- Wang, H.H.; Liu, C.Y.; Wu, S.B.; Liu, N.W.; Peng, C.Y.; Chan, T.H.; Hsu, C.F.; Wang, J.K.; Wang, Y.L. Highly Raman-Enhancing Substrates Based on Silver Nanoparticle Arrays with Tunable Sub-10 nm Gaps. Adv. Mater. 2006, 18, 491–495. [Google Scholar] [CrossRef]
- Dick, L.A.; McFarland, A.D.; Haynes, C.L.; Van Duyne, R.P. Metal Film over Nanosphere (MFON) Electrodes for Surface-Enhanced Raman Spectroscopy (SERS): Improvements in Surface Nanostructure Stability and Suppression of Irreversible Loss. J. Phys. Chem. B 2002, 106, 853–860. [Google Scholar] [CrossRef]
- Fromm, D.P.; Sundaramurthy, A.; Kinkhabwala, A.; Schuck, P.J.; Kino, G.S.; Moerner, W.E. Exploring the chemical enhancement for surface-enhanced Raman scattering with Au bowtie nanoantennas. J. Chem. Phys. 2006, 124, 61101. [Google Scholar] [CrossRef] [PubMed]
- Alvarez-Puebla, R.; Cui, B.; Bravo-Vasquez, J.-P.; Veres, T.; Fenniri, H. Nanoimprinted SERS-Active Substrates with Tunable Surface Plasmon Resonances. J. Phys. Chem. C 2007, 111, 6720–6723. [Google Scholar] [CrossRef]
- Semin, D.J.; Rowlen, K.L. Influence of vapor deposition parameters on SERS active Ag film morphology and optical properties. Anal. Chem. 1994, 66, 4324–4331. [Google Scholar] [CrossRef]
- Faulds, K.; Smith, W.E.; Graham, D. Evaluation of Surface-Enhanced Resonance Raman Scattering for Quantitative DNA Analysis. Anal. Chem. 2004, 76, 412–417. [Google Scholar] [CrossRef] [PubMed]
- Szymborski, T.; Witkowska, E.; Adamkiewicz, W.; Waluk, J.; Kamińska, A. Electrospun polymer mat as a SERS platform for the immobilization and detection of bacteria from fluids. Analyst 2014, 139, 5061–5064. [Google Scholar] [CrossRef] [PubMed]
- Witkowska, E.; Szymborski, T.; Kamińska, A.; Waluk, J. Polymer mat prepared via Forcespinning™ as a SERS platform for immobilization and detection of bacteria from blood plasma. Mater. Sci. Eng. C 2017, 71, 345–350. [Google Scholar] [CrossRef] [PubMed]
- Kijeńska, E.; Swieszkowski, W. 2–General requirements of electrospun materials for tissue engineering: Setups and strategy for successful electrospinning in laboratory and industry. In Electrospun Materials for Tissue Engineering and Biomedical Applications; Uyar, T., Kny, E., Eds.; Woodhead Publishing: Sawston, UK, 2017; pp. 43–56. [Google Scholar]
- Zhang, F.; Chen, P.; Zhang, L.; Mao, S.C.; Lin, L.; Tang, Y.G.; Cui, J.C.; Qi, X.D.; Yang, J.H.; Ma, Y.F. Enhancement of Raman scattering by field superposition of rough submicrometer silver particles. Appl. Phys. Lett. 2012, 100, 4. [Google Scholar] [CrossRef]
- Zheng, S.; Lin, H.K.; Lu, B.; Williams, A.; Datar, R.; Cote, R.J.; Tai, Y.C. 3D microfilter device for viable circulating tumor cell (CTC) enrichment from blood. Biomed. Microdevices 2011, 13, 203–213. [Google Scholar] [CrossRef] [PubMed]
- Cheng, W.T.; Liu, M.T.; Liu, H.N.; Lin, S.Y. Micro-Raman spectroscopy used to identify and grade human skin pilomatrixoma. Microsc. Res. Tech. 2005, 68, 75–79. [Google Scholar] [CrossRef] [PubMed]
- Neugebauer, U.; Clement, J.H.; Bocklitz, T.; Krafft, C.; Popp, J. Identification and differentiation of single cells from peripheral blood by Raman spectroscopic imaging. J. Biophotonics 2010, 3, 579–587. [Google Scholar] [CrossRef] [PubMed]
- Stone, N.; Kendall, C.; Smith, J.; Crow, P.; Barr, H. Raman spectroscopy for identification of epithelial cancers. Faraday Discuss. 2004, 126, 141–157. [Google Scholar] [CrossRef] [PubMed]
- Notingher, I.; Green, C.; Dyer, C.; Perkins, E.; Hopkins, N.; Lindsay, C.; Hench, L.L. Discrimination between ricin and sulphur mustard toxicity in vitro using Raman spectroscopy. J. R. Soc. Interface 2004, 1, 79–90. [Google Scholar] [CrossRef] [PubMed]
- Stone, N.; Kendall, C.; Shepherd, N.; Crow, P.; Barr, H. Near-infrared Raman spectroscopy for the classification of epithelial pre-cancers and cancers. J. Raman Spectrosc. 2002, 33, 564–573. [Google Scholar] [CrossRef]
- Fung, M.F.K.; Senterman, M.K.; Mikhael, N.Z.; Lacelle, S.; Wong, P.T.T. Pressure-tuning Fourier transform infrared spectroscopic study of carcinogenesis in human endometrium. Biospectroscopy 1996, 2, 155–165. [Google Scholar] [CrossRef]
- Huang, Z.; McWilliams, A.; Lui, H.; McLean, D.I.; Lam, S.; Zeng, H. Near-infrared Raman spectroscopy for optical diagnosis of lung cancer. Int. J. Cancer 2003, 107, 1047–1052. [Google Scholar] [CrossRef] [PubMed]
- Utzinger, U.; Heintzelman, D.L.; Mahadevan-Jansen, A.; Malpica, A.; Follen, M.; Richards-Kortum, R. Near-Infrared Raman Spectroscopy for in Vivo Detection of Cervical Precancers. Appl. Spectrosc. 2001, 55, 955–959. [Google Scholar] [CrossRef]
- Krafft, C.; Knetschke, T.; Siegner, A.; Funk, R.H.W.; Salzer, R. Mapping of single cells by near infrared Raman microspectroscopy. Vib. Spectrosc. 2003, 32, 75–83. [Google Scholar] [CrossRef]
- Taleb, A.; Diamond, J.; McGarvey, J.J.; Beattie, J.R.; Toland, C.; Hamilton, P.W. Raman Microscopy for the Chemometric Analysis of Tumor Cells. J. Phys. Chem. B 2006, 110, 19625–19631. [Google Scholar] [CrossRef] [PubMed]
- Nijssen, A.; Bakker Schut, T.C.; Heule, F.; Caspers, P.J.; Hayes, D.P.; Neumann, M.H.; Puppels, G.J. Discriminating basal cell carcinoma from its surrounding tissue by Raman spectroscopy. J. Investig. Dermatol. 2002, 119, 64–69. [Google Scholar] [PubMed]
- Notingher, I.; Verrier, S.; Haque, S.; Polak, J.M.; Hench, L.L. Spectroscopic study of human lung epithelial cells (A549) in culture: Living cells versus dead cells. Biopolymers 2003, 72, 230–240. [Google Scholar] [CrossRef] [PubMed]
- Uzunbajakava, N.; Lenferink, A.; Kraan, Y.; Willekens, B.; Vrensen, G.; Greve, J.; Otto, C. Nonresonant Raman imaging of protein distribution in single human cells. Biopolymers 2003, 72, 1–9. [Google Scholar] [CrossRef] [PubMed]
- El-Said, W.A.; Kim, T.H.; Kim, H.; Choi, J.W. Analysis of intracellular state based on controlled 3D nanostructures mediated surface enhanced Raman scattering. PLoS ONE 2011, 6, e15836. [Google Scholar] [CrossRef] [PubMed]

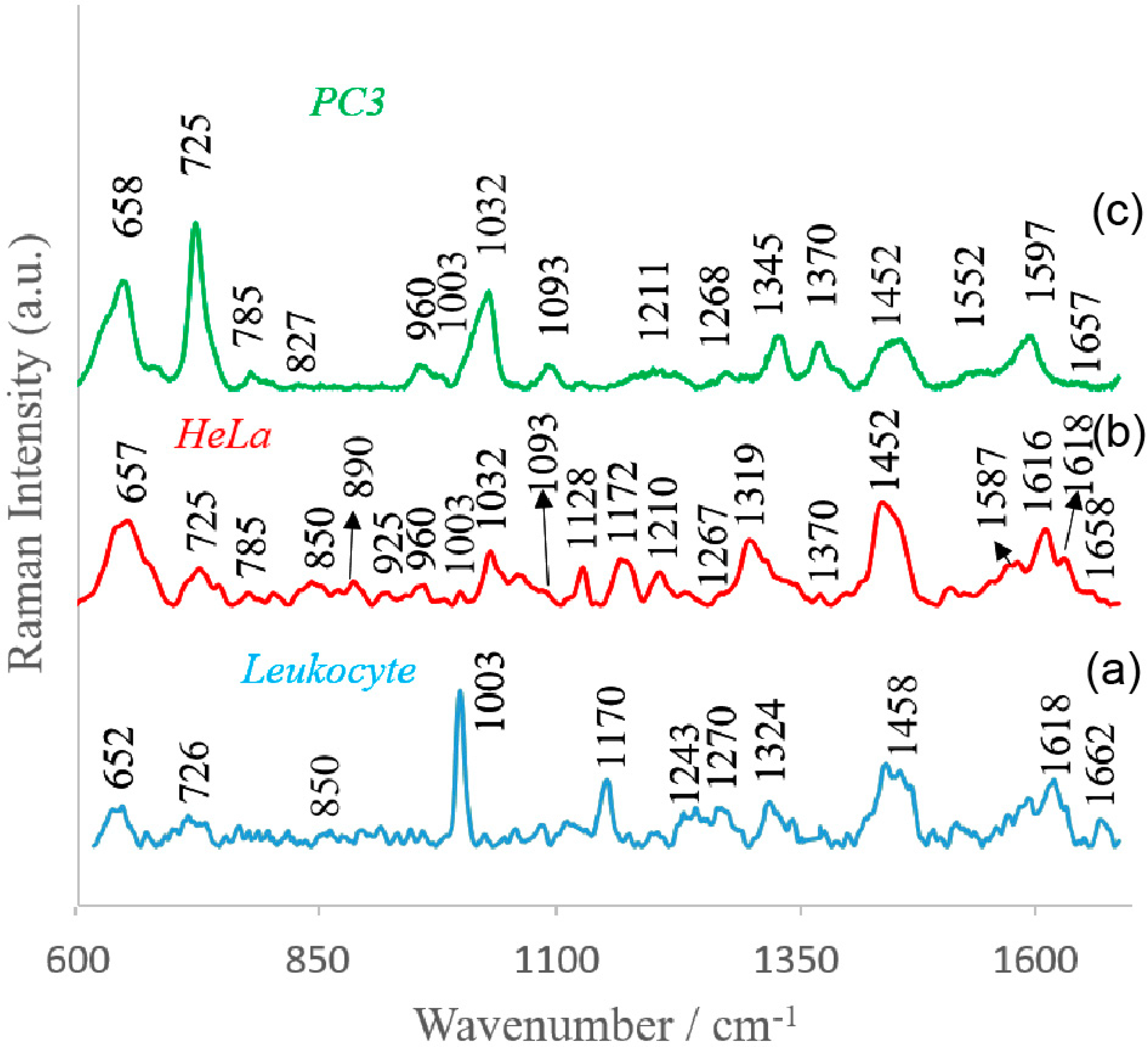

| Observed SERS Band (cm−1) | Protein | Lipids | Nucleic Acid |
|---|---|---|---|
| 652–658 | Tyr (C–C twist) | ||
| 725–730 | Trp | C–N head group choline (H3C)3N+ | A |
| 785 | PO2 symm | ||
| 827 | RNA backbone | ||
| 850 | Tyr, Pro | ||
| 890 | Structural protein modes of tumors | ||
| 925 | C–C str alpha-helix, Pro, Val | ||
| 960 | CH3 def | CH3 def | |
| 1003 | Phe | ||
| 1030–1032 | CH2CH3 bending modes of lipids | ||
| 1093 | C–N stretch | CC str chain, C–O str | PO2 symm |
| 1128 | C–N str bk | porphyrin | |
| 1170–1172 | Tyr C–H inplane | T | |
| 1210 | C–C6H5 str in phenylalanine tyrosine | ||
| 1243 | Amide III (beta sheet) | ||
| 1267–1270 | Amide III ( random coil) | =CH def | |
| 1319 | CH3 def, collagen | CH3CH2 twist | G |
| 1324 | purine bases of DNA | ||
| 1345 | A, G | ||
| 1370 | sphingoglycolipids | ||
| 1452 | structural protein modes of tumors | ||
| 1458 | A, G | ||
| 1552 | A, G | ||
| 1597–1600 | Phe, Tyr | ||
| 1616–1618 | C≡C str of Tyr and Trp | ||
| 1657–1665 | Amide I | C=C str |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kamińska, A.; Szymborski, T.; Witkowska, E.; Kijeńska-Gawrońska, E.; Świeszkowski, W.; Niciński, K.; Trzcińska-Danielewicz, J.; Girstun, A. Detection of Circulating Tumor Cells Using Membrane-Based SERS Platform: A New Diagnostic Approach for ‘Liquid Biopsy’. Nanomaterials 2019, 9, 366. https://doi.org/10.3390/nano9030366
Kamińska A, Szymborski T, Witkowska E, Kijeńska-Gawrońska E, Świeszkowski W, Niciński K, Trzcińska-Danielewicz J, Girstun A. Detection of Circulating Tumor Cells Using Membrane-Based SERS Platform: A New Diagnostic Approach for ‘Liquid Biopsy’. Nanomaterials. 2019; 9(3):366. https://doi.org/10.3390/nano9030366
Chicago/Turabian StyleKamińska, Agnieszka, Tomasz Szymborski, Evelin Witkowska, Ewa Kijeńska-Gawrońska, Wojciech Świeszkowski, Krzysztof Niciński, Joanna Trzcińska-Danielewicz, and Agnieszka Girstun. 2019. "Detection of Circulating Tumor Cells Using Membrane-Based SERS Platform: A New Diagnostic Approach for ‘Liquid Biopsy’" Nanomaterials 9, no. 3: 366. https://doi.org/10.3390/nano9030366
APA StyleKamińska, A., Szymborski, T., Witkowska, E., Kijeńska-Gawrońska, E., Świeszkowski, W., Niciński, K., Trzcińska-Danielewicz, J., & Girstun, A. (2019). Detection of Circulating Tumor Cells Using Membrane-Based SERS Platform: A New Diagnostic Approach for ‘Liquid Biopsy’. Nanomaterials, 9(3), 366. https://doi.org/10.3390/nano9030366








