Thioredoxin Reductase Activity Predicts Gold Nanoparticle Radiosensitization Effect
Abstract
1. Introduction
2. Material and Methods
2.1. GNP Synthesis
2.2. Cell Culture
2.3. GNP Uptake
2.4. X-ray Irradiation
2.5. Clonogenic Assay
2.6. TrxR Activity Assay
2.7. Patient Survival Analysis
2.8. Statistical Analysis
3. Results
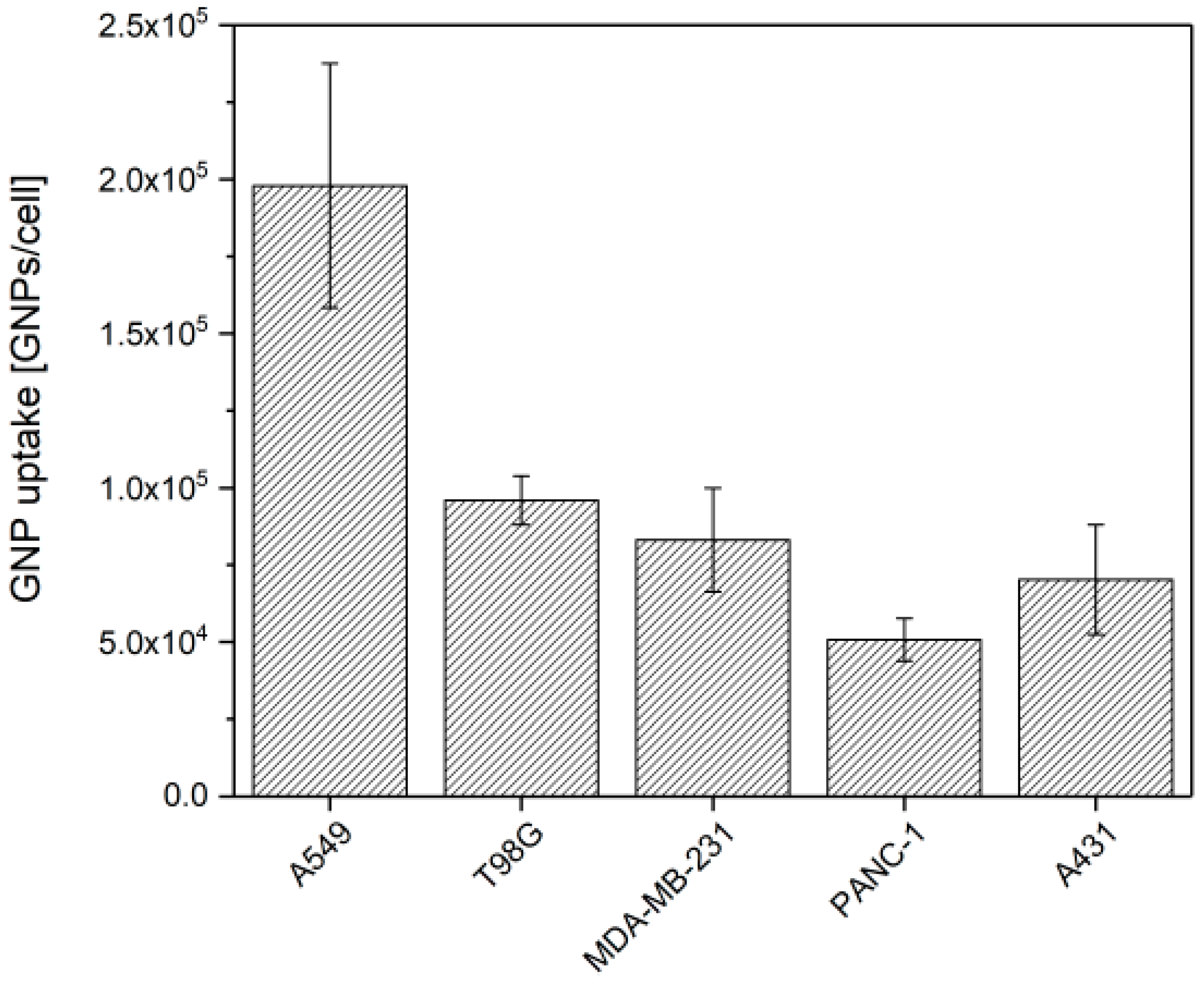
3.1. GNP Uptake
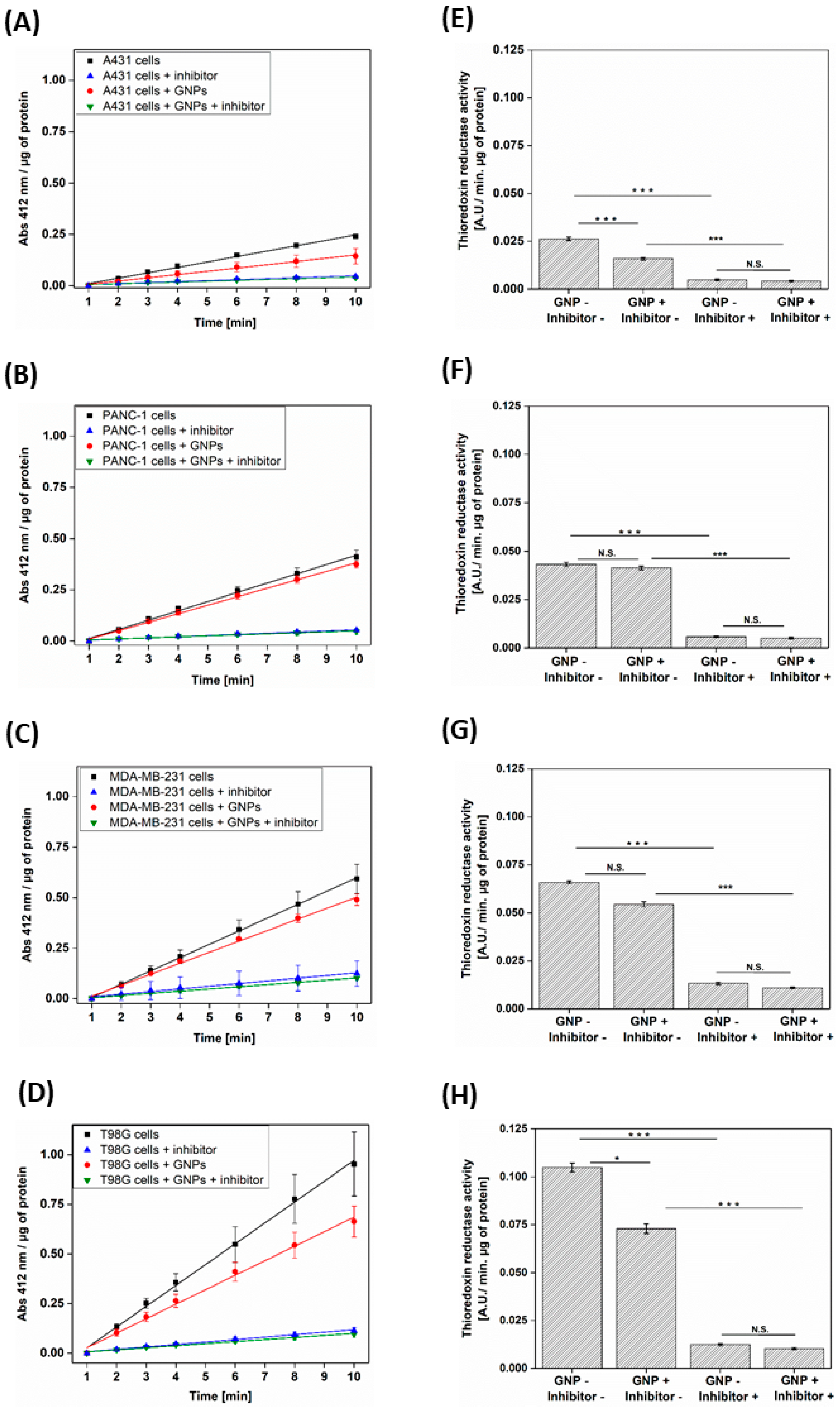
3.2. GNPs Decrease the TrxR Activity in Different Cell Lines
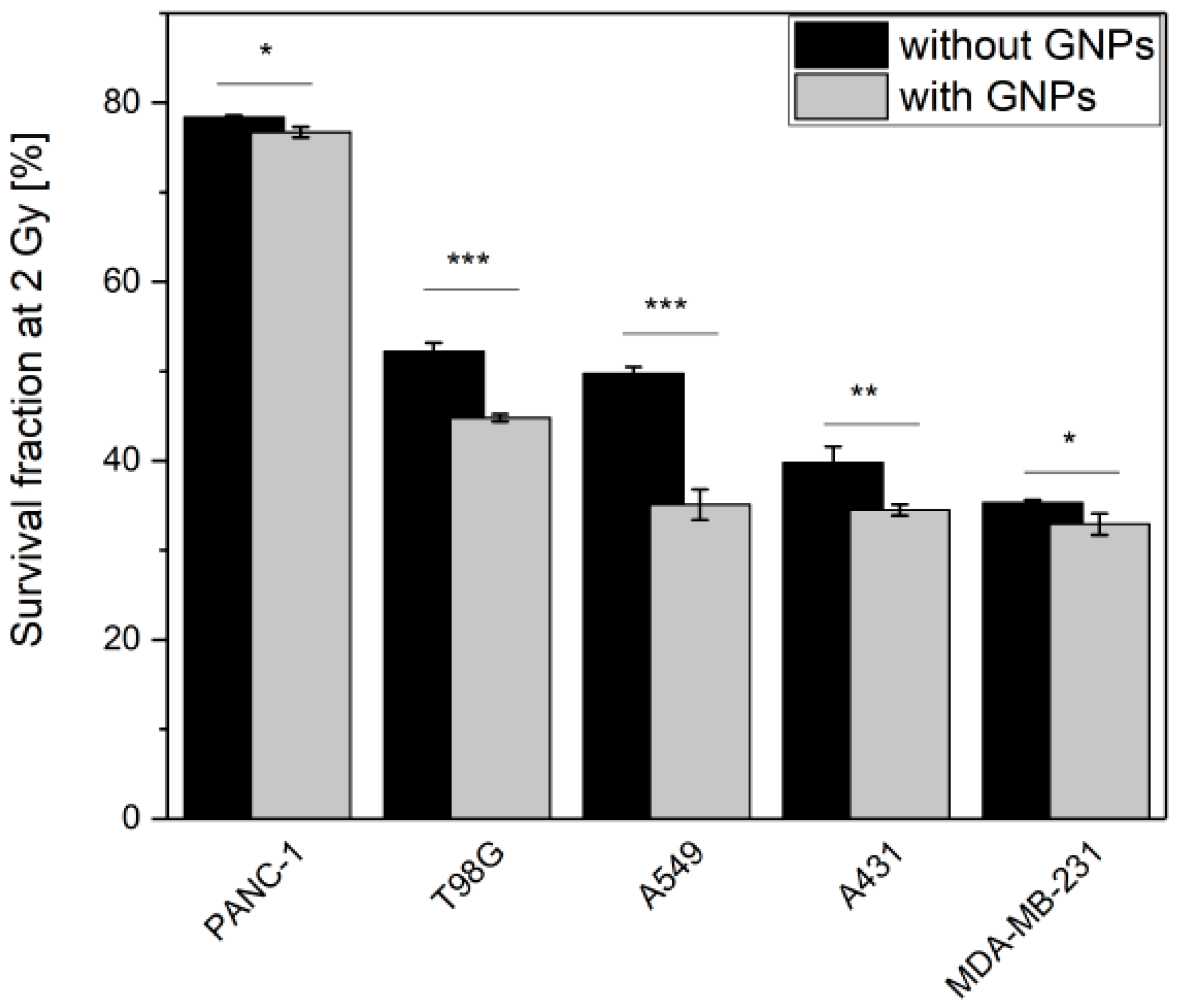
3.3. GNPs Enhance the Cell Death upon Irradiation
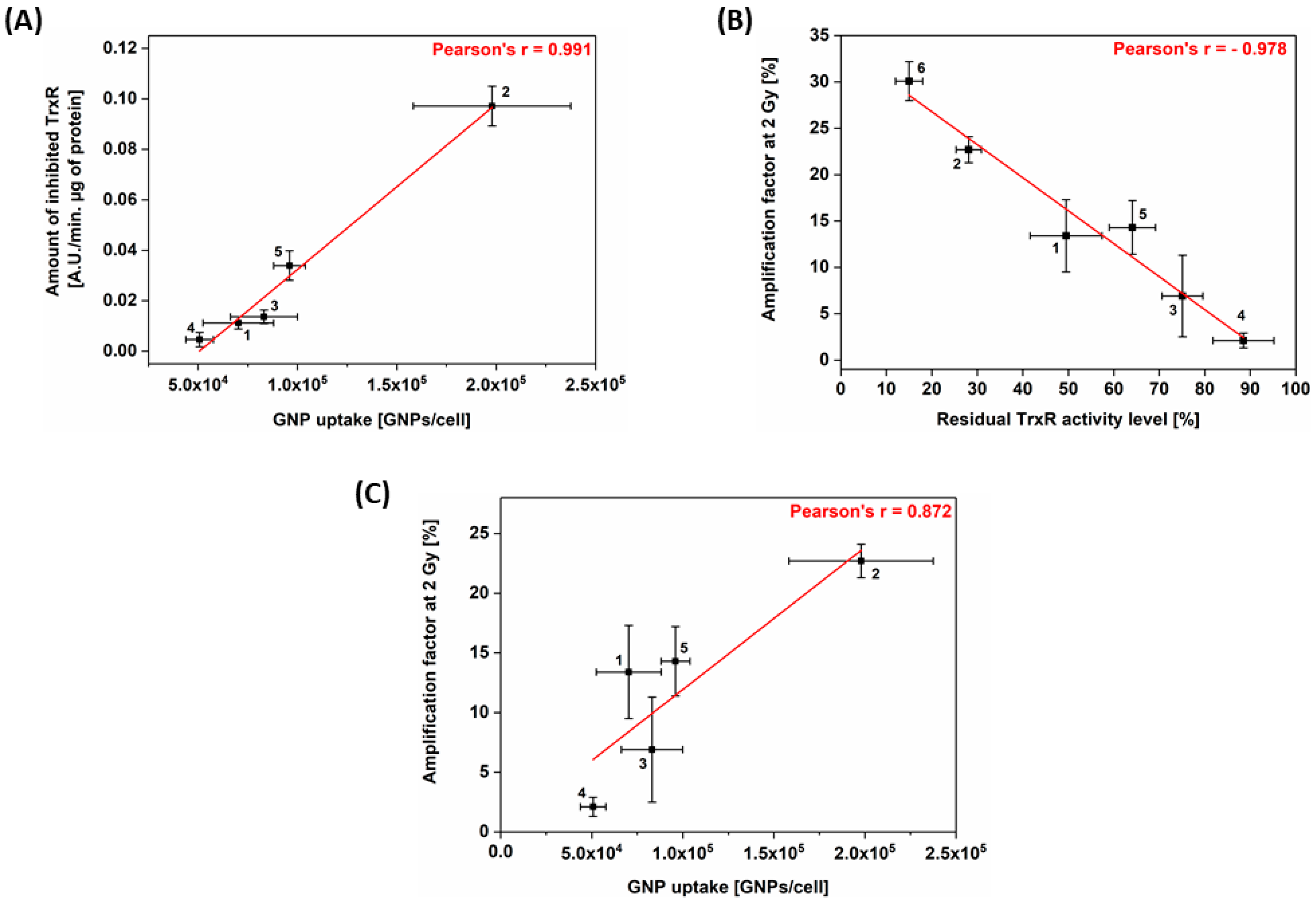
3.4. Correlation between Cell Response to Radiation, TrxR Activity and GNP Uptake
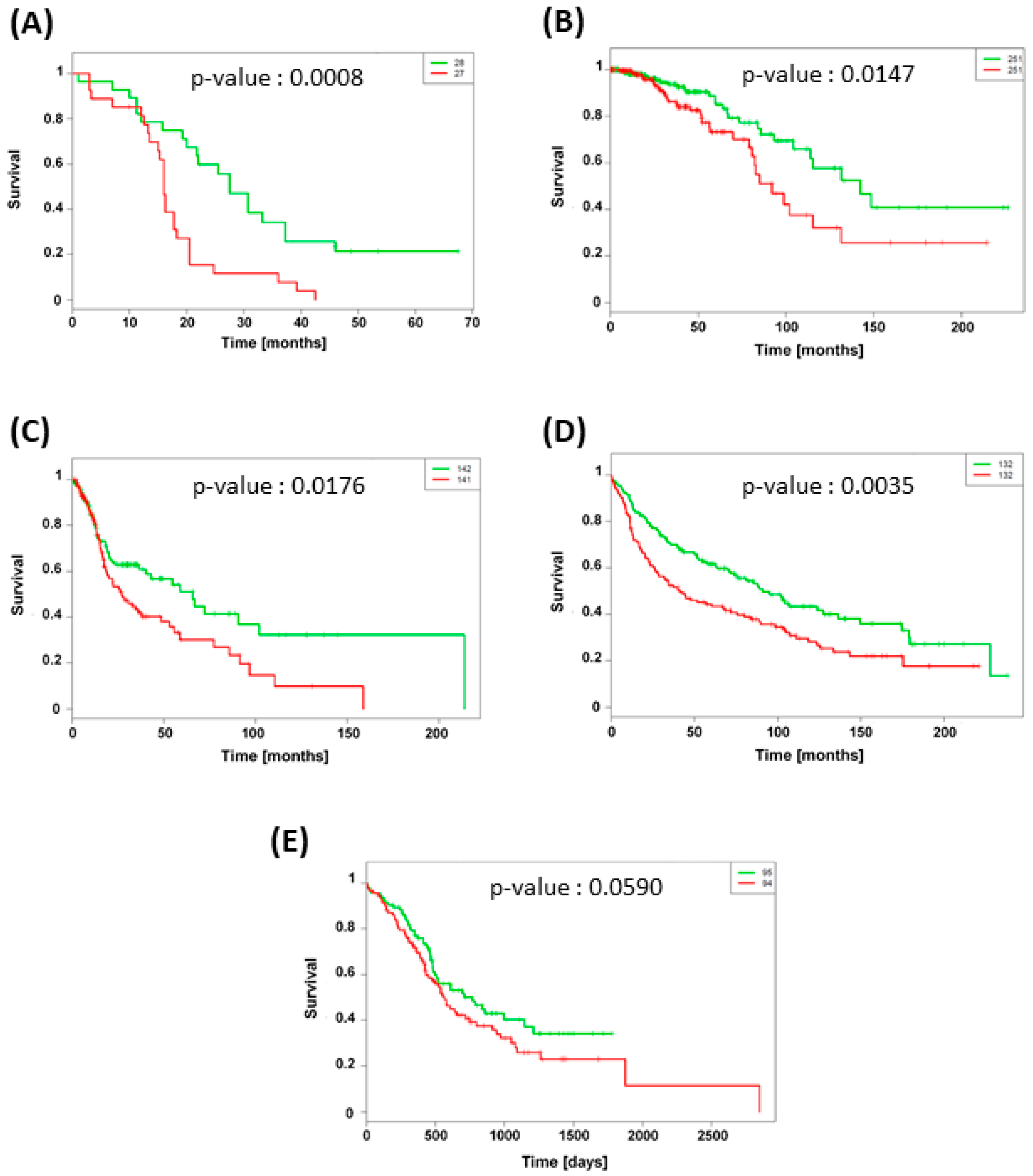
3.5. TXNRD1 Is an Unfavorable Prognostic Factor
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Delaney, G.; Jacob, S.; Featherstone, C.; Barton, M. The role of radiotherapy in cancer treatment: Estimating optimal utilization from a review of evidence-based clinical guidelines. Cancer 2005, 104, 1129–1137. [Google Scholar] [CrossRef] [PubMed]
- Laprise-Pelletier, M.; Simao, T.; Fortin, M.A. Gold Nanoparticles in Radiotherapy and Recent Progress in Nanobrachytherapy. Adv. Healthc. Mater. 2018, 7, e1701460. [Google Scholar] [CrossRef] [PubMed]
- Genard, G.; Wera, A.C.; Huart, C.; le Calve, B.; Penninckx, S.; Fattaccioli, A.; Tabarrant, T.; Demazy, C.; Ninane, N.; Heuskin, A.C.; et al. Proton irradiation orchestrates macrophage reprogramming through NFkappaB signaling. Cell Death Dis. 2018, 9, 728. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, K.; Usami, N.; Porcel, E.; Lacombe, S.; le Sech, C. Enhancement of radiation effect by heavy elements. Mutat. Res. 2010, 704, 123–131. [Google Scholar] [CrossRef] [PubMed]
- Currell, F.; Villagomez-Bernabe, B. Physical and Chemical Processes for Gold Nanoparticles and Ionising Radiation in Medical Contexts. In Gold Nanoparticles for Physics, Chemistry and Biology; World Scientific: Singapore, 2017; pp. 509–536. [Google Scholar]
- Li, S.; Penninckx, S.; Karmani, L.; Heuskin, A.C.; Watillon, K.; Marega, R.; Zola, J.; Corvaglia, V.; Genard, G.; Gallez, B.; et al. LET-dependent radiosensitization effects of gold nanoparticles for proton irradiation. Nanotechnology 2016, 27, 455101. [Google Scholar] [CrossRef] [PubMed]
- Hainfeld, J.F.; Slatkin, D.N.; Smilowitz, H.M. The use of gold nanoparticles to enhance radiotherapy in mice. Phys. Med. Biol. 2004, 49, N309–N315. [Google Scholar] [CrossRef] [PubMed]
- Xu, R.; Ma, J.; Sun, X.; Chen, Z.; Jiang, X.; Guo, Z.; Huang, L.; Li, Y.; Wang, M.; Wang, C.; et al. Ag nanoparticles sensitize IR-induced killing of cancer cells. Cell Res. 2009, 19, 1031–1034. [Google Scholar] [CrossRef]
- Liu, P.; Jin, H.; Guo, Z.; Ma, J.; Zhao, J.; Li, D.; Wu, H.; Gu, N. Silver nanoparticles outperform gold nanoparticles in radiosensitizing U251 cells in vitro and in an intracranial mouse model of glioma. Int. J. Nanomed. 2016, 11, 5003–5014. [Google Scholar] [CrossRef]
- Miladi, I.; Aloy, M.T.; Armandy, E.; Mowat, P.; Kryza, D.; Magne, N.; Tillement, O.; Lux, F.; Billotey, C.; Janier, M.; et al. Combining ultrasmall gadolinium-based nanoparticles with photon irradiation overcomes radioresistance of head and neck squamous cell carcinoma. Nanomed. Nanotechnol. Biol. Med. 2015, 11, 247–257. [Google Scholar] [CrossRef]
- Porcel, E.; Tillement, O.; Lux, F.; Mowat, P.; Usami, N.; Kobayashi, K.; Furusawa, Y.; le Sech, C.; Li, S.; Lacombe, S. Gadolinium-based nanoparticles to improve the hadrontherapy performances. Nanomed. Nanotechnol. Biol. Med. 2014, 10, 1601–1608. [Google Scholar] [CrossRef]
- Detappe, A.; Kunjachan, S.; Rottmann, J.; Robar, J.; Tsiamas, P.; Korideck, H.; Tillement, O.; Berbeco, R. AGuIX nanoparticles as a promising platform for image-guided radiation therapy. Cancer Nanotechnol. 2015, 6, 4. [Google Scholar] [CrossRef] [PubMed]
- McGinnity, T.L.; Dominguez, O.; Curtis, T.E.; Nallathamby, P.D.; Hoffman, A.J.; Roeder, R.K. Hafnia (HfO2) nanoparticles as an X-ray contrast agent and mid-infrared biosensor. Nanoscale 2016, 8, 13627–13637. [Google Scholar] [CrossRef] [PubMed]
- Pottier, A.; Borghi, E.; Levy, L. New use of metals as nanosized radioenhancers. Anticancer Res. 2014, 34, 443–453. [Google Scholar] [PubMed]
- Li, S.; Porcel, E.; Remita, H.; Marco, S.; Réfrégiers, M.; Dutertre, M.; Confalonieri, F.; Lacombe, S. Platinum nanoparticles: An exquisite tool to overcome radioresistance. Cancer Nanotechnol. 2017, 8, 4. [Google Scholar] [CrossRef]
- Porcel, E.; Liehn, S.; Remita, H.; Usami, N.; Kobayashi, K.; Furusawa, Y.; le Sech, C.; Lacombe, S. Platinum nanoparticles: A promising material for future cancer therapy? Nanotechnology 2010, 21, 085103. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.-D.; Luo, Z.; Chen, J.; Song, S.; Yuan, X.; Shen, X.; Wang, H.; Sun, Y.; Gao, K.; Zhang, L. Ultrasmall glutathione-protected gold nanoclusters as next generation radiotherapy sensitizers with high tumor uptake and high renal clearance. Sci. Rep. 2015, 5, 8669. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Liu, X.; Jin, X.; He, P.; Zheng, X.; Dai, Z.; Ye, F.; Zhao, T.; Chen, W.; Li, Q. The dependence of radiation enhancement effect on the concentration of gold nanoparticles exposed to low- and high-LET radiations. Phys. Med. 2015, 31, 210–218. [Google Scholar] [CrossRef] [PubMed]
- Hossain, M.; Luo, Y.; Sun, Z.; Wang, C.; Zhang, M.; Fu, H.; Qiao, Y.; Su, M. X-ray enabled detection and eradication of circulating tumor cells with nanoparticles. Biosens. Bioelectron. 2012, 38, 348–354. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Ma, M.; Chen, S.; Dai, J.; Chen, F.; Wang, Z. Construction of multifunctional organic–inorganic hybrid Bi2S3–PLGA capsules for highly efficient ultrasound-guided radiosensitization of brachytherapy. RSC Adv. 2014, 4, 26861–26865. [Google Scholar] [CrossRef]
- Zhang, X.D.; Wu, D.; Shen, X.; Chen, J.; Sun, Y.M.; Liu, P.X.; Liang, X.J. Size-dependent radiosensitization of PEG-coated gold nanoparticles for cancer radiation therapy. Biomaterials 2012, 33, 6408–6419. [Google Scholar] [CrossRef] [PubMed]
- Villanueva, A.; Canete, M.; Roca, A.G.; Calero, M.; Veintemillas-Verdaguer, S.; Serna, C.J.; Mdel, P.M.; Miranda, R. The influence of surface functionalization on the enhanced internalization of magnetic nanoparticles in cancer cells. Nanotechnology 2009, 20, 115103. [Google Scholar] [CrossRef] [PubMed]
- Ma, N.; Wu, F.-G.; Zhang, X.; Jiang, Y.-W.; Jia, H.-R.; Wang, H.-Y.; Li, Y.-H.; Liu, P.; Gu, N.; Chen, Z. Shape-Dependent Radiosensitization Effect of Gold Nanostructures in Cancer Radiotherapy: Comparison of Gold Nanoparticles, Nanospikes, and Nanorods. ACS Appl. Mater. Interfaces 2017, 9, 13037–13048. [Google Scholar] [CrossRef] [PubMed]
- Penninckx, S.; Heuskin, A.C.; Michiels, C.; Lucas, S. The role of thioredoxin reductase in gold nanoparticle radiosensitization effects. Nanomedicine (Lond. Engl.) 2018. [Google Scholar] [CrossRef] [PubMed]
- Aguirre-Gamboa, R.; Gomez-Rueda, H.; Martinez-Ledesma, E.; Martinez-Torteya, A.; Chacolla-Huaringa, R.; Rodriguez-Barrientos, A.; Tamez-Pena, J.G.; Trevino, V. SurvExpress: An online biomarker validation tool and database for cancer gene expression data using survival analysis. PLoS ONE 2013, 8, e74250. [Google Scholar] [CrossRef] [PubMed]
- Joo, K.M.; Kim, J.; Jin, J.; Kim, M.; Seol, H.J.; Muradov, J.; Yang, H.; Choi, Y.L.; Park, W.Y.; Kong, D.S.; et al. Patient-specific orthotopic glioblastoma xenograft models recapitulate the histopathology and biology of human glioblastomas in situ. Cell Rep. 2013, 3, 260–273. [Google Scholar] [CrossRef] [PubMed]
- Rousseaux, S.; Debernardi, A.; Jacquiau, B.; Vitte, A.L.; Vesin, A.; Nagy-Mignotte, H.; Moro-Sibilot, D.; Brichon, P.Y.; Lantuejoul, S.; Hainaut, P.; et al. Ectopic activation of germline and placental genes identifies aggressive metastasis-prone lung cancers. Sci. Transl. Med. 2013, 5, 186ra166. [Google Scholar] [CrossRef] [PubMed]
- La Thangue, N.B.; Kerr, D.J. Predictive biomarkers: A paradigm shift towards personalized cancer medicine. Nat. Rev. Clin. Oncol. 2011, 8, 587–596. [Google Scholar] [CrossRef]
- Leone, A.; Roca, M.S.; Ciardiello, C.; Costantini, S.; Budillon, A. Oxidative Stress Gene Expression Profile Correlates with Cancer Patient Poor Prognosis: Identification of Crucial Pathways Might Select Novel Therapeutic Approaches. Oxid. Med. Cell. Longev. 2017, 2017, 2597581. [Google Scholar] [CrossRef]
- Arner, E.S.; Holmgren, A. The thioredoxin system in cancer, Seminars in cancer biology. 2006, 16, 420–426. [Google Scholar]
- Dunn, L.L.; Buckle, A.M.; Cooke, J.P.; Ng, M.K. The emerging role of the thioredoxin system in angiogenesis. Arterioscler. Thromb. Vasc. Biol. 2010, 30, 2089–2098. [Google Scholar] [CrossRef]
- Yoo, M.H.; Xu, X.M.; Carlson, B.A.; Patterson, A.D.; Gladyshev, V.N.; Hatfield, D.L. Targeting thioredoxin reductase 1 reduction in cancer cells inhibits self-sufficient growth and DNA replication. PLoS ONE 2007, 2, e1112. [Google Scholar] [CrossRef] [PubMed]
- Yoo, M.H.; Xu, X.M.; Carlson, B.A.; Gladyshev, V.N.; Hatfield, D.L. Thioredoxin reductase 1 deficiency reverses tumor phenotype and tumorigenicity of lung carcinoma cells. J. Biol. Chem. 2006, 281, 13005–13008. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.J.; Miyoshi, Y.; Taguchi, T.; Tamaki, Y.; Nakamura, H.; Yodoi, J.; Kato, K.; Noguchi, S. High thioredoxin expression is associated with resistance to docetaxel in primary breast cancer. Clin. Cancer Res. 2005, 11, 8425–8430. [Google Scholar] [CrossRef] [PubMed]
- Iwao-Koizumi, K.; Matoba, R.; Ueno, N.; Kim, S.J.; Ando, A.; Miyoshi, Y.; Maeda, E.; Noguchi, S.; Kato, K. Prediction of docetaxel response in human breast cancer by gene expression profiling. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2005, 23, 422–431. [Google Scholar] [CrossRef] [PubMed]
- Fiskus, W.; Saba, N.; Shen, M.; Ghias, M.; Liu, J.; Gupta, S.D.; Chauhan, L.; Rao, R.; Gunewardena, S.; Schorno, K.; et al. Auranofin induces lethal oxidative and endoplasmic reticulum stress and exerts potent preclinical activity against chronic lymphocytic leukemia. Cancer Res. 2014, 74, 2520–2532. [Google Scholar] [CrossRef] [PubMed]
- Zou, P.; Chen, M.; Ji, J.; Chen, W.; Chen, X.; Ying, S.; Zhang, J.; Zhang, Z.; Liu, Z.; Yang, S.; et al. Auranofin induces apoptosis by ROS-mediated ER stress and mitochondrial dysfunction and displayed synergistic lethality with piperlongumine in gastric cancer. Oncotarget 2015, 6, 36505–36521. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Bouzakoura, S.; de Mey, S.; Jiang, H.; Law, K.; Dufait, I.; Corbet, C.; Verovski, V.; Gevaert, T.; Feron, O.; et al. Auranofin radiosensitizes tumor cells through targeting thioredoxin reductase and resulting overproduction of reactive oxygen species. Oncotarget 2017, 8, 35728–35742. [Google Scholar] [CrossRef]
- Fan, C.; Zheng, W.; Fu, X.; Li, X.; Wong, Y.S.; Chen, T. Enhancement of auranofin-induced lung cancer cell apoptosis by selenocystine, a natural inhibitor of TrxR1 in vitro and in vivo. Cell Death Dis. 2014, 5, e1191. [Google Scholar] [CrossRef] [PubMed]
- Sirolimus and Auranofin in Treating Patients with Advanced or Recurrent Non-Small Cell Lung Cancer or Small Cell Lung Cancer. Available online: Clinicaltrials.org (accessed on 19 February 2019).
- Phase I and II Study of Auranofin in Chronic Lymphocytic Leukemia (CLL). Available online: Clinicaltrials.org (accessed on 19 February 2019).
- Auranofin in Treating Patients with Recurrent Epithelial Ovarian, Primary Peritoneal, or Fallopian Tube Cancer. Available online: Clinicaltrials.org (accessed on 19 February 2019).
- Butterworth, K.T.; Coulter, J.A.; Jain, S.; Forker, J.; McMahon, S.J.; Schettino, G.; Prise, K.M.; Currell, F.J.; Hirst, D.G. Evaluation of cytotoxicity and radiation enhancement using 1.9 nm gold particles: Potential application for cancer therapy. Nanotechnology 2010, 21, 295101. [Google Scholar] [CrossRef] [PubMed]
- Polf, J.C.; Bronk, L.F.; Driessen, W.H.; Arap, W.; Pasqualini, R.; Gillin, M. Enhanced relative biological effectiveness of proton radiotherapy in tumor cells with internalized gold nanoparticles. Appl. Phys. Lett. 2011, 98, 193702. [Google Scholar] [CrossRef] [PubMed]
- Hespeels, F.; Heuskin, A.C.; Scifoni, E.; Kraemer, M.; Lucas, S. Backscattered electron emission after proton impact on carbon and gold films: Experiments and simulations. Nucl. Instrum. Methods Phys. Res. Sect. B Beam Interact. Mater. Atmos 2017, 401, 8–17. [Google Scholar] [CrossRef]
- Misawa, M.; Takahashi, J. Generation of reactive oxygen species induced by gold nanoparticles under x-ray and UV Irradiations. Nanomed. Nanotechnol. Biol. Med. 2011, 7, 604–614. [Google Scholar] [CrossRef] [PubMed]
- Sicard-Roselli, C.; Brun, E.; Gilles, M.; Baldacchino, G.; Kelsey, C.; McQuaid, H.; Polin, C.; Wardlow, N.; Currell, F. A new mechanism for hydroxyl radical production in irradiated nanoparticle solutions. Small 2014, 10, 3338–3346. [Google Scholar] [CrossRef] [PubMed]
- Sotiropoulos, M.; Henthorn, N.T.; Warmenhoven, J.W.; Mackay, R.I.; Kirkby, K.J.; Merchant, M.J. Modelling direct DNA damage for gold nanoparticle enhanced proton therapy. Nanoscale 2017, 9, 18413–18422. [Google Scholar] [CrossRef] [PubMed]
- Heuskin, A.C.; Gallez, B.; Feron, O.; Martinive, P.; Michiels, C.; Lucas, S. Metallic nanoparticles irradiated by low-energy protons for radiation therapy: Are there significant physical effects to enhance the dose delivery? Med. Phys. 2017, 44, 4299–4312. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Li, X.; Wang, Y.; Liu, Z.; Fu, L.; Hu, L. Enhancement of radiation effect and increase of apoptosis in lung cancer cells by thio-glucose-bound gold nanoparticles at megavoltage radiation energies. J. Nanopart. Res. 2013, 15, 1642. [Google Scholar] [CrossRef]
- Saberi, A.; Shahbazi-Gahrouei, D.; Abbasian, M.; Fesharaki, M.; Baharlouei, A.; Arab-Bafrani, Z. Gold nanoparticles in combination with megavoltage radiation energy increased radiosensitization and apoptosis in colon cancer HT-29 cells. Int. J. Radiat. Biol. 2017, 93, 315–323. [Google Scholar] [CrossRef]
- Liu, C.J.; Wang, C.H.; Chien, C.C.; Yang, T.Y.; Chen, S.T.; Leng, W.H.; Lee, C.F.; Lee, K.H.; Hwu, Y.; Lee, Y.C.; et al. Enhanced x-ray irradiation-induced cancer cell damage by gold nanoparticles treated by a new synthesis method of polyethylene glycol modification. Nanotechnology 2008, 19, 295104. [Google Scholar] [CrossRef]
- Liu, Y.; Zhang, P.; Li, F.; Jin, X.; Li, J.; Chen, W.; Li, Q. Metal-based NanoEnhancers for Future Radiotherapy: Radiosensitizing and Synergistic Effects on Tumor Cells. Theranostics 2018, 8, 1824–1849. [Google Scholar] [CrossRef] [PubMed]
- Butterworth, K.T.; McMahon, S.J.; Taggart, L.E.; Prise, K.M. Radiosensitization by gold nanoparticles: Effective at megavoltage energies and potential role of oxidative stress. Transl. Cancer Res. 2013, 2, 269–279. [Google Scholar]
- Jeynes, J.C.; Merchant, M.J.; Spindler, A.; Wera, A.C.; Kirkby, K.J. Investigation of gold nanoparticle radiosensitization mechanisms using a free radical scavenger and protons of different energies. Phys. Med. Biol. 2014, 59, 6431–6443. [Google Scholar] [CrossRef] [PubMed]
- Schlathölter, T.; Eustache, P.; Porcel, E.; Salado, D.; Stefancikova, L.; Tillement, O.; Lux, F.; Mowat, P.; Biegun, A.K.; van Goethem, M.J.; et al. Improving proton therapy by metal-containing nanoparticles: Nanoscale insights. Int. J. Nanomed. 2016, 11, 1549–1556. [Google Scholar] [CrossRef] [PubMed]
- Taggart, L.E.; McMahon, S.J.; Butterworth, K.T.; Currell, F.J.; Schettino, G.; Prise, K.M. Protein disulphide isomerase as a target for nanoparticle-mediated sensitisation of cancer cells to radiation. Nanotechnology 2016, 27, 215101. [Google Scholar] [CrossRef] [PubMed]
- Chiu, S.J.; Lee, M.Y.; Chou, W.G.; Lin, L.Y. Germanium oxide enhances the radiosensitivity of cells. Radiat. Res. 2003, 159, 391–400. [Google Scholar] [CrossRef]
- Mirjolet, C.; Papa, A.L.; Créhange, G.; Raguin, O.; Seignez, C.; Paul, C.; Truc, G.; Maingon, P.; Millot, N. The radiosensitization effect of titanate nanotubes as a new tool in radiation therapy for glioblastoma: A proof-of-concept. Radiother. Oncol. 2013, 108, 136–142. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Bouchy, S.; Penninckx, S.; Marega, R.; Fichera, O.; Gallez, B.; Feron, O.; Martinive, P.; Heuskin, A.C.; Michiels, C.; et al. Antibody-functionalized gold nanoparticles as tumor targeting radiosensitizers for proton therapy. Nanomedicine (Lond. Engl.) 2019. [Google Scholar] [CrossRef]
| Organ | Cancer Type | Number of Patients | Median Survival Time (Months) | Database | |
|---|---|---|---|---|---|
| Low TXNRD1 Expression | High TXNRD1 Expression | ||||
| Brain | Glioblastoma | 58 | 27 | 16 | GSE 42669 [26] |
| Breast | Invasive carcinoma | 502 | 142 | 92 | TCGA-BRCA |
| Head & Neck | Squamous cell carcinoma | 283 | 66 | 27 | TCGA-HNSC |
| Lung | Adenocarcinoma | 264 | 102 | 44 | GSE 30219 [27] |
| Pancreas | Ductal adenocarcinoma | 189 | 26 | 19 | PACA-AU |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Penninckx, S.; Heuskin, A.-C.; Michiels, C.; Lucas, S. Thioredoxin Reductase Activity Predicts Gold Nanoparticle Radiosensitization Effect. Nanomaterials 2019, 9, 295. https://doi.org/10.3390/nano9020295
Penninckx S, Heuskin A-C, Michiels C, Lucas S. Thioredoxin Reductase Activity Predicts Gold Nanoparticle Radiosensitization Effect. Nanomaterials. 2019; 9(2):295. https://doi.org/10.3390/nano9020295
Chicago/Turabian StylePenninckx, Sébastien, Anne-Catherine Heuskin, Carine Michiels, and Stéphane Lucas. 2019. "Thioredoxin Reductase Activity Predicts Gold Nanoparticle Radiosensitization Effect" Nanomaterials 9, no. 2: 295. https://doi.org/10.3390/nano9020295
APA StylePenninckx, S., Heuskin, A.-C., Michiels, C., & Lucas, S. (2019). Thioredoxin Reductase Activity Predicts Gold Nanoparticle Radiosensitization Effect. Nanomaterials, 9(2), 295. https://doi.org/10.3390/nano9020295






