Micronuclei Detection by Flow Cytometry as a High-Throughput Approach for the Genotoxicity Testing of Nanomaterials
Abstract
1. Introduction
2. Material and Methods
2.1. Selected Nanomaterials
2.2. Cell Culture
2.3. Cell Viability
2.4. The FCMN Assay
2.5. Statistical Analysis
3. Results and Discussion
3.1. Nanoparticles Characterization
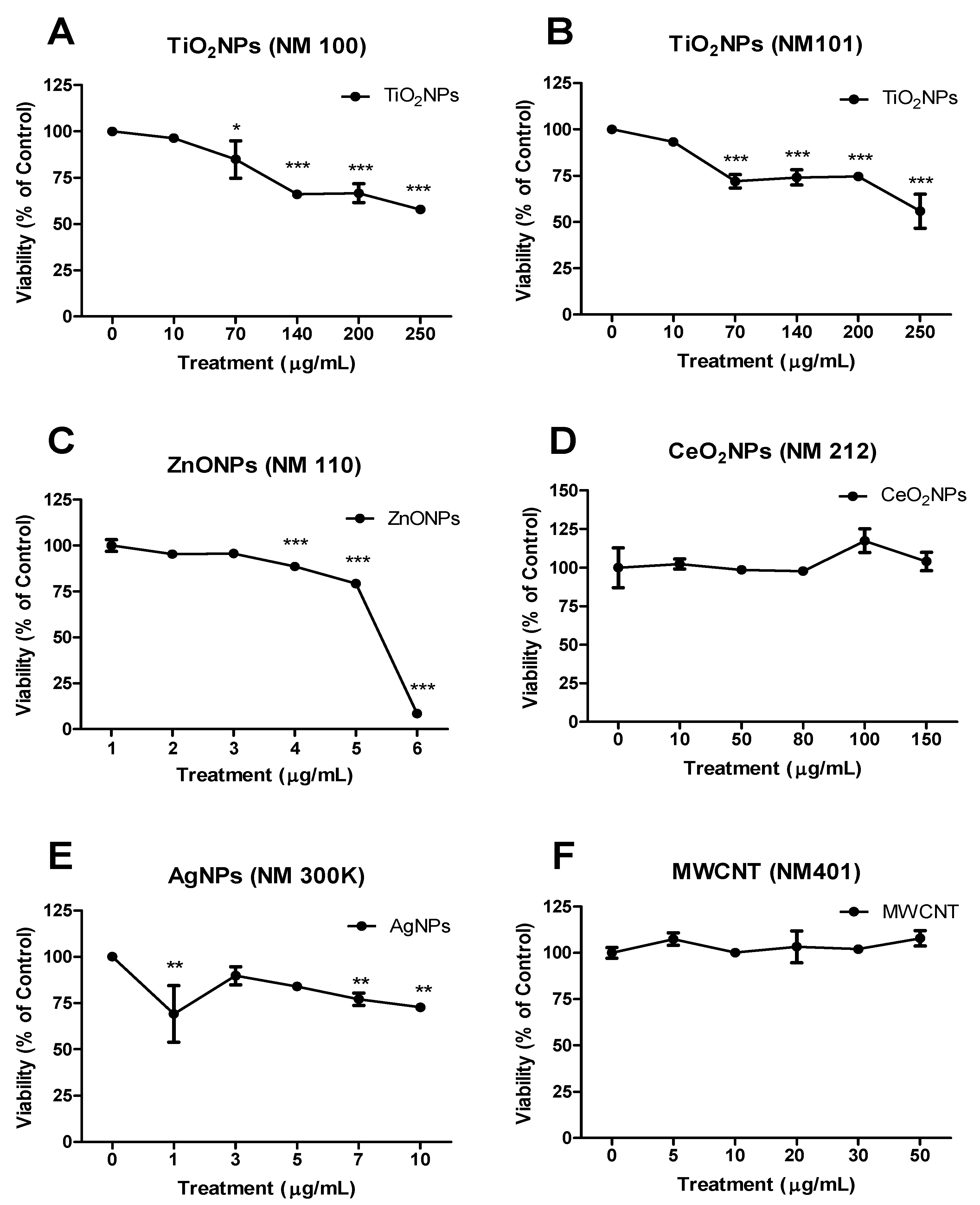
3.2. Cell Viability
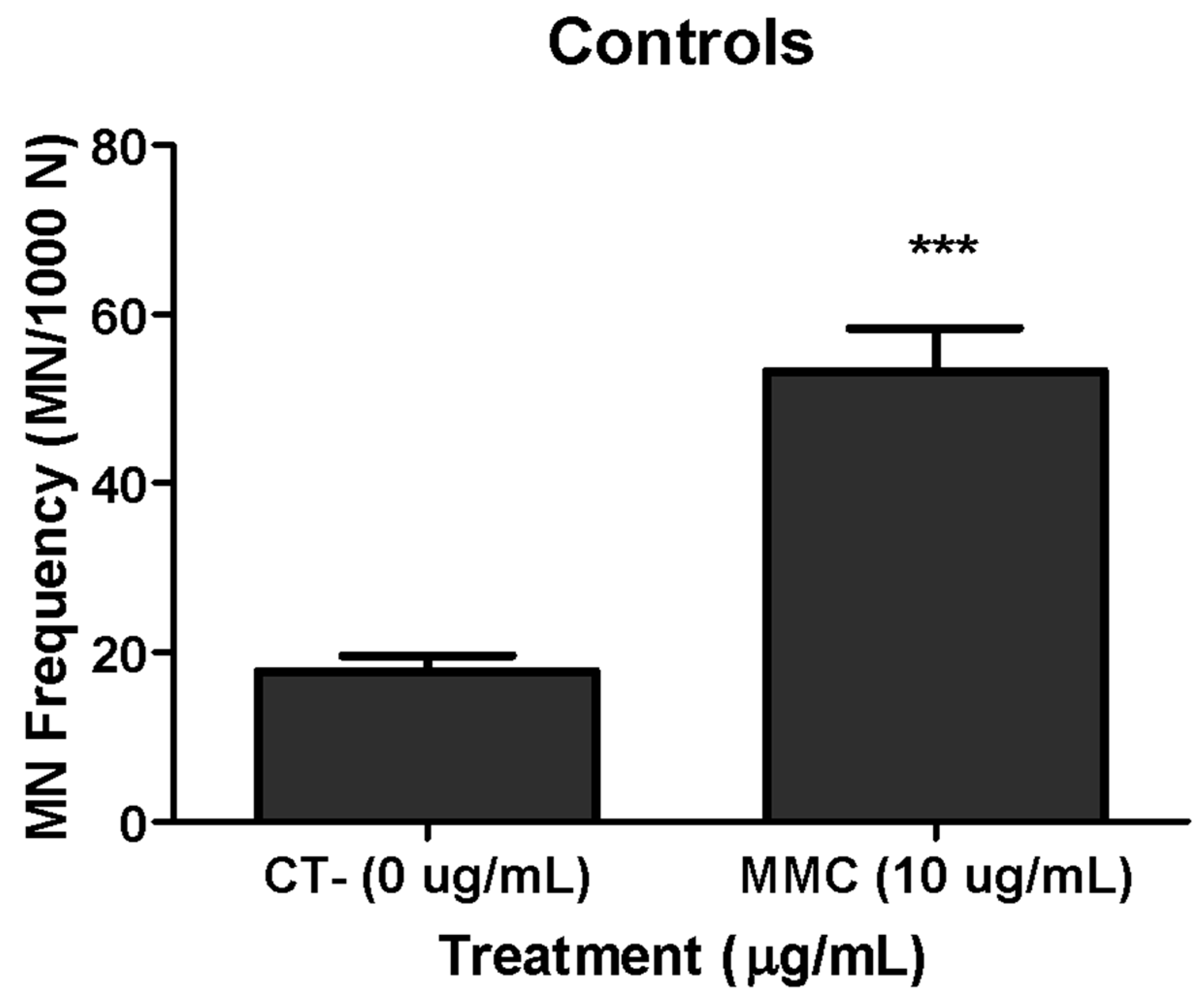
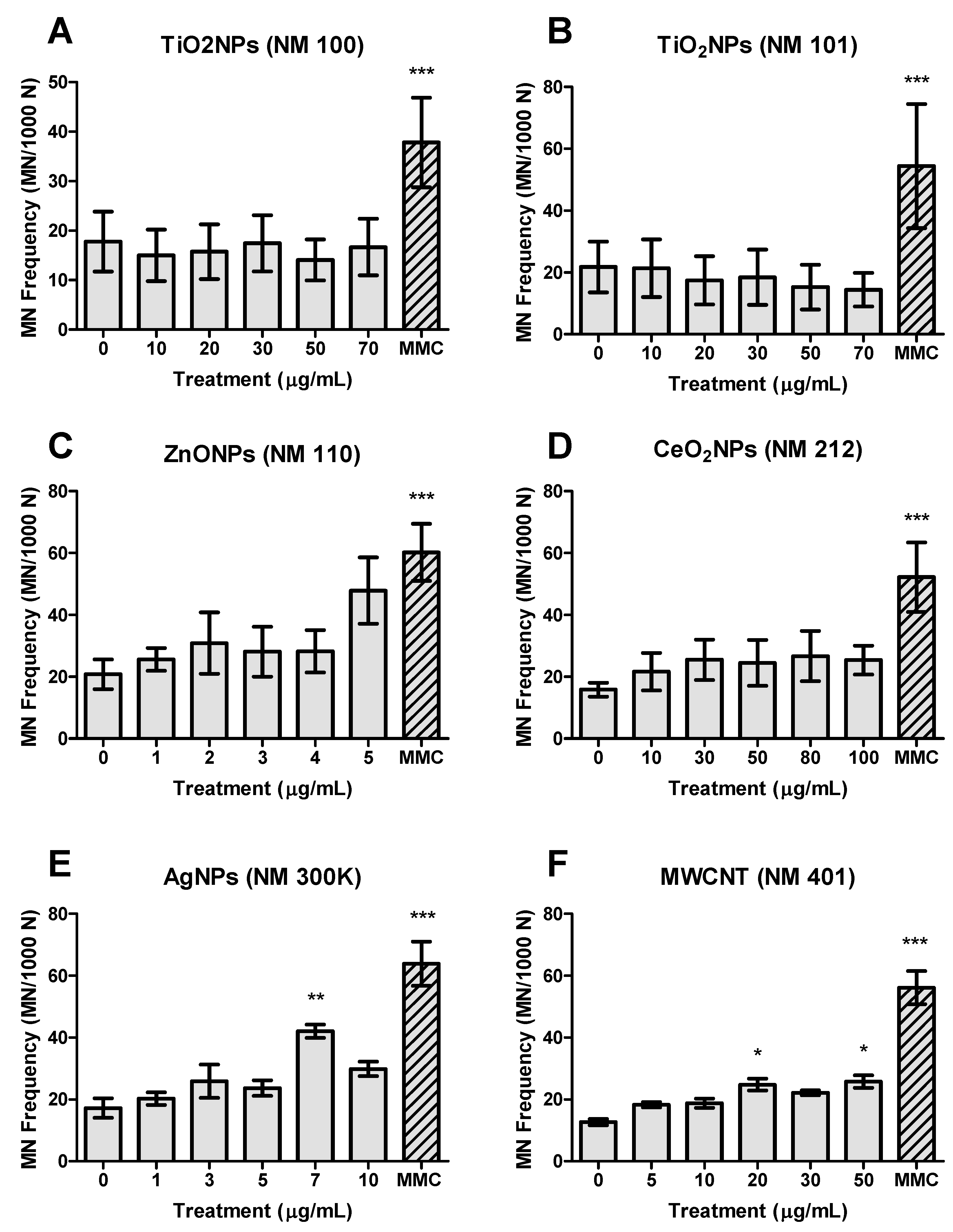
3.3. Genotoxicity
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Hansen, S.F.; Heggelund, L.R.; Revilla Besora, P.; Mackevica, A.; Boldrin, A.; Baun, A. Nanoproducts -what is available to European consumers? Environ. Sci. Nano 2016, 3, 169–180. [Google Scholar] [CrossRef]
- Vance, M.E.; Kuiken, T.; Vejerano, E.P.; McGinnis, S.P.; Hochella, M.F., Jr.; Rejeski, D.; Hull, M.S. Nanotechnology in the real world: Redeveloping the nanomaterial consumer products inventory. Beilstein J. Nanotechnol. 2015, 6, 1769–1780. [Google Scholar] [CrossRef] [PubMed]
- Zhu, M.; Nie, G.; Meng, H.; Xia, T.; Nel, A.; Zhao, Y. Physicochemical properties determine nanomaterial cellular uptake, transport, and fate. ACC Chem. Res. 2013, 46, 622–631. [Google Scholar] [CrossRef] [PubMed]
- Gebel, T.; Foth, H.; Damm, G.; Freyberger, A.; Kramer, P.J.; Lilienblum, W.; Röhl, C.; Schupp, T.; Weiss, C.; Wollin, K.M.; et al. Manufactured nanomaterials: Categorization and approaches to hazard assessment. Arch. Toxicol. 2014, 88, 2191–2211. [Google Scholar] [CrossRef]
- Vila, L.; Rubio, L.; Annangi, B.; García-Rodríguez, A.; Marcos, R.; Hernández, A. Frozen dispersions of nanomaterials are a useful operational procedure in nanotoxicology. Nanotoxicology 2017, 11, 31–40. [Google Scholar] [CrossRef]
- Nel, A.; Xia, T.; Meng, H.; Wang, X.; Lin, S.; Ji, Z.; Zhang, H. Nanomaterial toxicity testing in the 21st century: Use of a predictive toxicological approach and high-throughput screening. ACC Chem. Res. 2013, 46, 607–621. [Google Scholar] [CrossRef]
- Nelson, B.C.; Wright, C.W.; Ibuki, Y.; Moreno-Villanueva, M.; Karlsson, H.L.; Hendriks, G.; Sims, C.M.; Singh, N.; Doak, S.H. Emerging metrology for high-throughput nanomaterial genotoxicology. Mutagenesis 2017, 32, 215–232. [Google Scholar] [CrossRef]
- Collins, A.R.; Annangi, B.; Rubio, L.; Marcos, R.; Dorn, M.; Merker, C.; Estrela-Lopis, I.; Cimpan, M.R.; Ibrahim, M.; Cimpan, E.; et al. High throughput toxicity screening and intracellular detection of nanomaterials. Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 2017, 9, e1413. [Google Scholar] [CrossRef]
- Taurozzi, J.S.; Hackley, V.A.; Wiesner, M.R. Ultrasonic dispersion of nanoparticles for environmental, health and safety assessment -issues and recommendations. Nanotoxicology 2011, 5, 711–729. [Google Scholar] [CrossRef]
- Cohen, J.M.; Beltran-Huarac, J.; Pyrgiotakis, G.; Demokritou, P. Effective delivery of sonication energy to fast settling and agglomerating nanomaterial suspensions for cellular studies: Implications for stability, particle kinetics, dosimetry and toxicity. NanoImpact 2018, 10, 81–86. [Google Scholar] [CrossRef]
- Kaur, I.; Ellis, L.J.; Romer, I.; Tantra, R.; Carriere, M.; Allard, S.; Mayne-L’Hermite, M.; Minelli, C.; Unger, W.; Potthoff, A.; et al. Dispersion of nanomaterials in aqueous media: Towards protocol optimization. J. Vis. Exp. 2017, 130, 56074. [Google Scholar] [CrossRef] [PubMed]
- Gliga, A.R.; Skoglund, S.; Wallinder, I.O.; Fadeel, B.; Karlsson, H.L. Size-dependent cytotoxicity of silver nanoparticles in human lung cells: The role of cellular uptake, agglomeration and Ag release. Part. Fibre Toxicol. 2014, 11, 11. [Google Scholar] [CrossRef] [PubMed]
- Singh, N.; Manshian, B.; Jenkins, G.J.; Griffiths, S.M.; Williams, P.M.; Maffeis, T.G.; Wright, C.J.; Doak, S.H. NanoGenotoxicology: The DNA damaging potential of engineered nanomaterials. Biomaterials 2009, 30, 3891–3914. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, L.; Cundari, E.; Leyns, L.; Kirsch-Volders, M. Towards a new paradigm in nano-genotoxicology: Facing complexity of nanomaterials’ cellular interactions and effects. Basic Clin. Pharmacol. Toxicol. 2017, 121 (Suppl. S3), 23–29. [Google Scholar] [CrossRef] [PubMed]
- Bonassi, S.; Znaor, A.; Ceppi, M.; Lando, C.; Chang, W.P.; Holland, N.; Kirsch-Volders, M.; Zeiger, E.; Ban, S.; Barale, R.; et al. An increased micronucleus frequency in peripheral blood lymphocytes predicts the risk of cancer in humans. Carcinogenesis 2007, 28, 625–631. [Google Scholar] [CrossRef] [PubMed]
- Fenech, M. Cytokinesis-block micronucleus cytome assay. Nat. Protoc. 2007, 2, 1084–1104. [Google Scholar] [CrossRef] [PubMed]
- Nusse, M.; Kramer, J. Flow cytometric analysis of micronuclei found in cells after irradiation. Cytometry 1984, 5, 20–25. [Google Scholar] [CrossRef]
- Avlasevich, S.; Bryce, S.; De Boeck, M.; Elhajouji, A.; Van Goethem, F.; Lynch, A.; Nicolette, J.; Shi, J.; Dertinger, S. Flow cytometric analysis of micronuclei in mammalian cell cultures: Past, present and future. Mutagenesis 2011, 26, 147–152. [Google Scholar] [CrossRef]
- Di Bucchianico, S.; Cappellini, F.; Le Bihanic, F.; Zhang, Y.; Dreij, K.; Karlsson, H.L. Genotoxicity of TiO2 nanoparticles assessed by mini-gel comet assay and micronucleus scoring with flow cytometry. Mutagenesis 2017, 32, 127–137. [Google Scholar] [CrossRef]
- Li, Y.; Doak, S.H.; Yan, J.; Chen, D.H.; Zhou, M.; Mittelstaedt, R.A.; Chen, Y.; Li, C.; Chen, T. Factors affecting the in vitro micronucleus assay for evaluation of nanomaterials. Mutagenesis 2017, 32, 151–159. [Google Scholar] [CrossRef]
- Sahu, S.C.; Njoroge, J.; Bryce, S.M.; Yourick, J.J.; Sprando, R.L. Comparative genotoxicity of nanosilver in human liver HepG2 and colon Caco2 cells evaluated by a flow cytometric in vitro micronucleus assay. J. Appl. Toxicol. 2014, 34, 1226–1234. [Google Scholar] [CrossRef] [PubMed]
- Sahu, S.C.; Njoroge, J.; Bryce, S.M.; Zheng, J.; Ihrie, J. Flow cytometric evaluation of the contribution of ionic silver to genotoxic potential of nanosilver in human liver HepG2 and colon Caco2 cells. J. Appl. Toxicol. 2016, 36, 521–531. [Google Scholar] [CrossRef] [PubMed]
- Butler, K.S.; Peeler, D.J.; Casey, B.J.; Dair, B.J.; Elespuru, R.K. Silver nanoparticles: Correlating nanoparticle size and cellular uptake with genotoxicity. Mutagenesis 2015, 30, 577–591. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, L.; Lukamowicz-Rajska, M.; Thomassen, L.C.; Kirschhock, C.E.; Leyns, L.; Lison, D.; Martens, J.A.; Elhajouji, A.; Kirsch-Volders, M. Co-assessment of cell cycle and micronucleus frequencies demonstrates the influence of serum on the in vitro genotoxic response to amorphous monodisperse silica nanoparticles of varying sizes. Nanotoxicology 2014, 8, 876–884. [Google Scholar] [CrossRef] [PubMed]
- NanoReg. Available online: http://www.rivm.nl/en/About_RIVM/Mission_and_strategy/Inter national_Affairs/International_Projects/Completed/NANoREG/NANoREG_Results_Repository_sub_page_Final_Report (accessed on 22 January 2019).
- Vales, G.; Rubio, L.; Marcos, R. Long-term exposures to low doses of titanium dioxide nanoparticles induce cell transformation, but not genotoxic damage in BEAS-2B cells. Nanotoxicology 2015, 9, 568–578. [Google Scholar] [CrossRef]
- Bryce, S.M.; Bemis, J.C.; Avlasevich, S.L.; Dertinger, S.D. In vitro micronucleus assay scored by flow cytometry provides a comprehensive evaluation of cytogenetic damage and cytotoxicity. Mutat. Res. 2007, 630, 78–91. [Google Scholar] [CrossRef]
- Avlasevich, S.L.; Bryce, S.M.; Cairns, S.E.; Dertinger, S.D. In vitro micronucleus scoring by flow cytometry: Differential staining of micronuclei versus apoptotic and necrotic chromatin enhances assay reliability. Environ. Mol. Mutagen. 2006, 47, 56–66. [Google Scholar] [CrossRef]
- Johnston, H.; Pojana, G.; Zuin, S.; Jacobsen, N.R.; Møller, P.; Loft, S.; Semmler-Behnke, M.; McGuiness, C.; Balharry, D.; Marcomini, A.; et al. Engineered nanomaterial risk. Lessons learnt from completed nanotoxicology studies: Potential solutions to current and future challenges. Crit. Rev. Toxicol. 2013, 43, 1–20. [Google Scholar] [CrossRef]
- Nanogenotox. Available online: http://www.nanogenotox.eu/files/PDF/Deliverables/nanogeno tox%20deliverable%203_wp4_%20dispersion%20protocol.pdf (accessed on 22 January 2019).
- García-Rodríguez, A.; Rubio, L.; Vila, L.; Xamena, N.; Velázquez, A.; Marcos, R.; Hernández, A. The comet assay as a tool to detect the genotoxic potential of nanomaterials. Nanomaterials 2019, 9, 1385. [Google Scholar] [CrossRef]
- Riaz Ahmed, K.B.; Nagy, A.M.; Brown, R.P.; Zhang, Q.; Malghan, S.G.; Goering, P.L. Silver nanoparticles: Significance of physicochemical properties and assay interference on the interpretation of in vitro cytotoxicity studies. Toxicol. Vitr. 2017, 38, 179–192. [Google Scholar] [CrossRef]
- Martirosyan, A.; Schneider, Y.J. Engineered nanomaterials in food: Implications for food safety and consumer health. Int. J. Environ. Res. Public Health 2014, 11, 5720–5750. [Google Scholar] [CrossRef] [PubMed]
- Prasad, R.Y.; Wallace, K.; Daniel, K.M.; Tennant, A.H.; Zucker, R.M.; Strickland, J.; Dreher, K.; Kligerman, A.D.; Blackman, C.F.; Demarini, D.M. Effect of treatment media on the agglomeration of titanium dioxide nanoparticles: Impact on genotoxicity, cellular interaction, and cell cycle. ACS Nano 2013, 7, 1929–1942. [Google Scholar] [CrossRef] [PubMed]
- Prasad, R.Y.; Simmons, S.O.; Killius, M.G.; Zucker, R.M.; Kligerman, A.D.; Blackman, C.F.; Fry, R.C.; Demarini, D.M. Cellular interactions and biological responses to titanium dioxide nanoparticles in HepG2 and BEAS-2B cells: Role of cell culture media. Environ. Mol. Mutagen. 2014, 55, 336–342. [Google Scholar] [CrossRef] [PubMed]
- Andreoli, C.; Leter, G.; De Berardis, B.; Degan, P.; De Angelis, I.; Pacchierotti, F.; Crebelli, R.; Barone, F.; Zijno, A. Critical issues in genotoxicity assessment of TiO2 nanoparticles by human peripheral blood mononuclear cells. J. Appl. Toxicol. 2018, 38, 1471–1482. [Google Scholar] [CrossRef]
- Donner, E.M.; Myhre, A.; Brown, S.C.; Boatman, R.; Warheit, D.B. in vivo micronucleus studies with 6 titanium dioxide materials (3 pigment-grade & 3 nanoscale) in orally-exposed rats. Regul. Toxicol. Pharm. 2016, 74, 64–74. [Google Scholar]
- Vila, L.; García-Rodríguez, A.; Marcos, R.; Hernández, A. Titanium dioxide nanoparticles translocate through differentiated Caco-2 cell monolayers, without disrupting the barrier functionality or inducing genotoxic damage. J. Appl. Toxicol. 2018, 38, 1195–1205. [Google Scholar] [CrossRef]
- Kwon, J.Y.; Lee, S.Y.; Koedrith, P.; Lee, J.Y.; Kim, K.M.; Oh, J.M.; Yang, S.I.; Kim, M.K.; Lee, J.K.; Jeong, J.; et al. Lack of genotoxic potential of ZnO nanoparticles in in vitro and in vivo tests. Mutat. Res. Genet. Toxicol. Environ. Mutagen. 2014, 761, 1–9. [Google Scholar] [CrossRef]
- Li, C.H.; Shen, C.C.; Cheng, Y.W.; Huang, S.H.; Wu, C.C.; Kao, C.C.; Liao, J.W.; Kang, J.J. Organ biodistribution, clearance, and genotoxicity of orally administered zinc oxide nanoparticles in mice. Nanotoxicology 2012, 6, 746–756. [Google Scholar] [CrossRef]
- Zijno, A.; De Angelis, I.; De Berardis, B.; Andreoli, C.; Russo, M.T.; Pietraforte, D.; Scorza, G.; Degan, P.; Ponti, J.; Rossi, F.; et al. Different mechanisms are involved in oxidative DNA damage and genotoxicity induction by ZnO and TiO2 nanoparticles in human colon carcinoma cells. Toxicol. Vitr. 2015, 29, 1503–1512. [Google Scholar] [CrossRef]
- Senapati, V.A.; Kumar, A.; Gupta, G.S.; Pandey, A.K.; Dhawan, A. ZnO nanoparticles induced inflammatory response and genotoxicity in human blood cells: A mechanistic approach. Food Chem. Toxicol. 2015, 85, 61–70. [Google Scholar] [CrossRef]
- Annangi, B.; Rubio, L.; Alaraby, M.; Bach, J.; Marcos, R.; Hernández, A. Acute and long-term in vitro effects of zinc oxide nanoparticles. Arch. Toxicol. 2016, 90, 2201–2213. [Google Scholar] [CrossRef] [PubMed]
- Vandebriel, R.J.; De Jong, W.H. A review of mammalian toxicity of ZnO nanoparticles. Nanotechnol. Sci. Appl. 2012, 5, 61–71. [Google Scholar] [CrossRef] [PubMed]
- Pandurangan, M.; Kim, D.H. In vitro toxicity of zinc oxide nanoparticles: A review. J. Nanopart. Res. 2015, 17, 158. [Google Scholar] [CrossRef]
- Roszak, J.; Catalán, J.; Järventaus, H.; Lindberg, H.K.; Suhonen, S.; Vippola, M.; Stępnik, M.; Norppa, H. Effect of particle size and dispersion status on cytotoxicity and genotoxicity of zinc oxide in human bronchial epithelial cells. Mutat. Res. Genet. Toxicol. Environ. Mutagen. 2016, 805, 7–18. [Google Scholar] [CrossRef]
- Kinnula, V.L.; Yankaskas, J.R.; Chang, L.; Virtanen, I.; Linnala, A.; Kang, B.H.; Crapo, J.D. Primary and immortalized (BEAS 2B) human bronchial epithelial cells have significant antioxidative capacity in vitro. Am. J. Respir. Cell Mol. Biol. 1994, 11, 568–576. [Google Scholar] [CrossRef]
- Rubio, L.; Marcos, R.; Hernández, A. Nanoceria acts as antioxidant in tumoral and transformed cells. Chem. Biol. Interact. 2018, 291, 7–15. [Google Scholar] [CrossRef]
- Rubio, L.; Annangi, B.; Vila, L.; Hernández, A.; Marcos, R. Antioxidant and anti-genotoxic properties of cerium oxide nanoparticles in a pulmonary-like cell system. Arch. Toxicol. 2016, 90, 269–278. [Google Scholar] [CrossRef]
- Wang, C.; Blough, E.; Dai, X.; Olajide, O.; Driscoll, H.; Leidy, J.W.; July, M.; Triest, W.E.; Wu, M. Protective effects of cerium oxide nanoparticles on MC3T3-E1 osteoblastic cells exposed to X-ray irradiation. Cell Physiol. Biochem. 2016, 38, 1510–1519. [Google Scholar] [CrossRef]
- Zal, Z.; Ghasemi, A.; Azizi, S.; Asgarian-Omran, H.; Montazeri, A.; Hosseinimehr, S.J. Radioprotective effect of cerium oxide nanoparticles against genotoxicity induced by ionizing radiation on human lymphocytes. Curr. Radiopharm. 2018, 11, 109–115. [Google Scholar] [CrossRef]
- Kumari, M.; Singh, S.P.; Chinde, S.; Rahman, M.F.; Mahboob, M.; Grover, P. Toxicity study of cerium oxide nanoparticles in human neuroblastoma cells. Int. J. Toxicol. 2014, 33, 86–97. [Google Scholar] [CrossRef]
- Benameur, L.; Auffan, M.; Cassien, M.; Liu, W.; Culcasi, M.; Rahmouni, H.; Stocker, P.; Tassistro, V.; Bottero, J.Y.; Rose, J.; et al. DNA damage and oxidative stress induced by CeO2 nanoparticles in human dermal fibroblasts: Evidence of a clastogenic effect as a mechanism of genotoxicity. Nanotoxicology 2015, 9, 696–705. [Google Scholar] [CrossRef] [PubMed]
- Kumari, M.; Kumari, S.I.; Kamal, S.S.; Grover, P. Genotoxicity assessment of cerium oxide nanoparticles in female Wistar rats after acute oral exposure. Mutat. Res. Genet. Toxicol. Environ. Mutagen. 2014, 775-776, 7–19. [Google Scholar] [CrossRef] [PubMed]
- Vila, L.; García-Rodríguez, A.; Cortés, C.; Velázquez, A.; Xamena, N.; Sampayo-Reyes, A.; Marcos, R.; Hernández, A. Effects of cerium oxide nanoparticles on differentiated/ undifferentiated human intestinal Caco-2cells. Chem. Biol. Interact. 2018, 283, 38–46. [Google Scholar] [CrossRef] [PubMed]
- Vila, L.; García-Rodríguez, A.; Cortés, C.; Marcos, R.; Hernández, A. Assessing the effects of silver nanoparticles on monolayers of differentiated Caco-2 cells, as a model of intestinal barrier. Food Chem. Toxicol. 2018, 116 Pt B, 1–10. [Google Scholar] [CrossRef]
- Vila, L.; Marcos, R.; Hernández, A. Long-term effects of silver nanoparticles in Caco-2 cells. Nanotoxicology 2017, 11, 771–780. [Google Scholar] [CrossRef] [PubMed]
- Souza, T.A.; Franchi, L.P.; Rosa, L.R.; da Veiga, M.A.; Takahashi, C.S. Cytotoxicity and genotoxicity of silver nanoparticles of different sizes in CHO-K1 and CHO-XRS5 cell lines. Mutat. Res. Genet. Toxicol. Environ. Mutagen. 2016, 795, 70–83. [Google Scholar] [CrossRef] [PubMed]
- Jiang, X.; Foldbjerg, R.; Miclaus, T.; Wang, L.; Singh, R.; Hayashi, Y.; Sutherland, D.; Chen, C.; Autrup, H.; Beer, C. Multi-platform genotoxicity analysis of silver nanoparticles in the model cell line CHO-K1. Toxicol. Lett. 2013, 222, 55–63. [Google Scholar] [CrossRef]
- Nymark, P.; Catalán, J.; Suhonen, S.; Järventaus, H.; Birkedal, R.; Clausen, P.A.; Jensen, K.A.; Vippola, M.; Savolainen, K.; Norppa, H. Genotoxicity of polyvinylpyrrolidone-coated silver nanoparticles in BEAS-2B cells. Toxicology 2013, 313, 38–48. [Google Scholar] [CrossRef] [PubMed]
- Lebedová, J.; Hedberg, Y.S.; Odnevall Wallinder, I.; Karlsson, H.L. Size-dependent genotoxicity of silver, gold and platinum nanoparticles studied using the mini-gel comet assay and micronucleus scoring with flow cytometry. Mutagenesis 2018, 33, 77–85. [Google Scholar] [CrossRef] [PubMed]
- Louro, H.; Pinhão, M.; Santos, J.; Tavares, A.; Vital, N.; Silva, M.J. Evaluation of the cytotoxic and genotoxic effects of benchmark multi-walled carbon nanotubes in relation to their physicochemical properties. Toxicol. Lett. 2016, 262, 123–134. [Google Scholar] [CrossRef]
- Catalán, J.; Siivola, K.M.; Nymark, P.; Lindberg, H.; Suhonen, S.; Järventaus, H.; Koivisto, A.J.; Moreno, C.; Vanhala, E.; Wolff, H.; et al. In vitro and in vivo genotoxic effects of straight versus tangled multi-walled carbon nanotubes. Nanotoxicology 2016, 10, 794–806. [Google Scholar] [CrossRef] [PubMed]
- Visalli, G.; Bertuccio, M.P.; Iannazzo, D.; Piperno, A.; Pistone, A.; Di Pietro, A. Toxicological assessment of multi-walled carbon nanotubes on A549 human lung epithelial cells. Toxicol. Vitr. 2015, 29, 352–362. [Google Scholar] [CrossRef] [PubMed]
- Kato, T.; Totsuka, Y.; Ishino, K.; Matsumoto, Y.; Tada, Y.; Nakae, D.; Goto, S.; Masuda, S.; Ogo, S.; Kawanishi, M.; et al. Genotoxicity of multi-walled carbon nanotubes in both in vitro and in vivo assay systems. Nanotoxicology 2013, 7, 452–461. [Google Scholar] [CrossRef] [PubMed]
- Asakura, M.; Sasaki, T.; Sugiyama, T.; Takaya, M.; Koda, S.; Nagano, K.; Arito, H.; Fukushima, S. Genotoxicity and cytotoxicity of multi-wall carbon nanotubes in cultured Chinese hamster lung cells in comparison with chrysotile A fibers. J. Occup. Health 2010, 52, 155–166. [Google Scholar] [CrossRef] [PubMed]
- Patlolla, A.K.; Hussain, S.M.; Schlager, J.J.; Patlolla, S.; Tchounwou, P.B. Comparative study of the clastogenicity of functionalized and nonfunctionalized multiwalled carbon nanotubes in bone marrow cells of Swiss-Webster mice. Environ. Toxicol. 2010, 25, 608–621. [Google Scholar] [CrossRef] [PubMed]
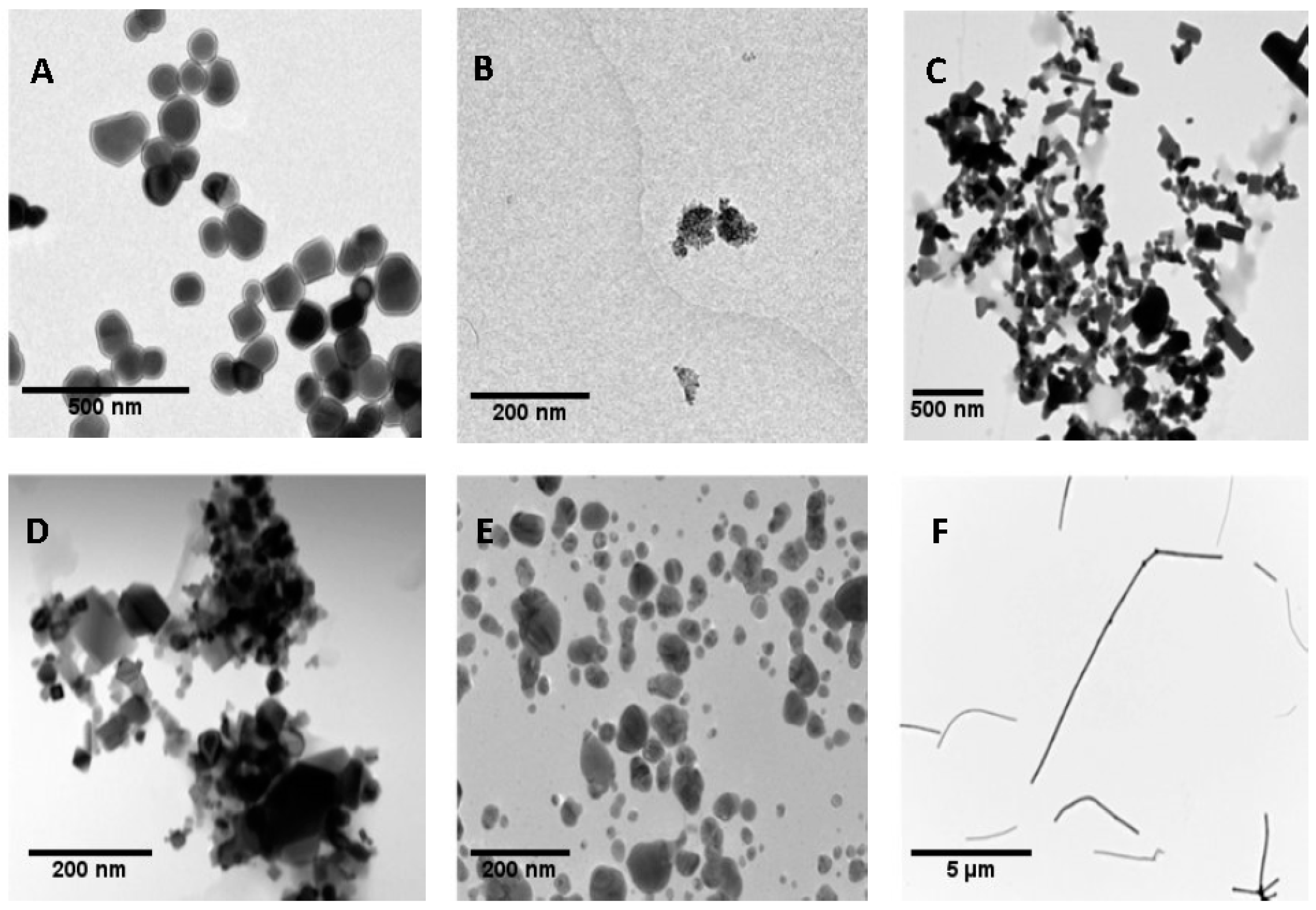
| NP | TiO2 NM100 | TiO2 NM101 | ZnO NM110 | CeO2 NM212 | Ag NM300K | MWCNT * NM401 |
|---|---|---|---|---|---|---|
| TEM | 104.01 ± 39.42 | 54.69 ± 35.39 | 132.37 ± 69.53 | 70.33 ± 49.61 | 7.75 ± 2.48 | 6012.09 ± 4091.45 |
| DLS (10 μg/mL) | 195.3 ± 2.50 | 152.2 ± 62.28 | 171.17 ± 13.61 | 172.2 ± 45.24 | 19.02 ± 1.78 | - |
| Nanomaterial | FCMN Assay | CBMN Assay |
|---|---|---|
| NM100 | − | − |
| NM101 | − (+ *) | − |
| NM110 | − | +/− |
| NM212 | − | nd |
| NM300K | +/− | − |
| NM401 | +/− | − |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
García-Rodríguez, A.; Kazantseva, L.; Vila, L.; Rubio, L.; Velázquez, A.; Ramírez, M.J.; Marcos, R.; Hernández, A. Micronuclei Detection by Flow Cytometry as a High-Throughput Approach for the Genotoxicity Testing of Nanomaterials. Nanomaterials 2019, 9, 1677. https://doi.org/10.3390/nano9121677
García-Rodríguez A, Kazantseva L, Vila L, Rubio L, Velázquez A, Ramírez MJ, Marcos R, Hernández A. Micronuclei Detection by Flow Cytometry as a High-Throughput Approach for the Genotoxicity Testing of Nanomaterials. Nanomaterials. 2019; 9(12):1677. https://doi.org/10.3390/nano9121677
Chicago/Turabian StyleGarcía-Rodríguez, Alba, Liliya Kazantseva, Laura Vila, Laura Rubio, Antonia Velázquez, María José Ramírez, Ricard Marcos, and Alba Hernández. 2019. "Micronuclei Detection by Flow Cytometry as a High-Throughput Approach for the Genotoxicity Testing of Nanomaterials" Nanomaterials 9, no. 12: 1677. https://doi.org/10.3390/nano9121677
APA StyleGarcía-Rodríguez, A., Kazantseva, L., Vila, L., Rubio, L., Velázquez, A., Ramírez, M. J., Marcos, R., & Hernández, A. (2019). Micronuclei Detection by Flow Cytometry as a High-Throughput Approach for the Genotoxicity Testing of Nanomaterials. Nanomaterials, 9(12), 1677. https://doi.org/10.3390/nano9121677




