Analysis of Nanotoxicity with Integrated Omics and Mechanobiology
Abstract
:1. Introduction
2. Omics Approaches for the Assessment of Nanotoxicity
2.1. Transcriptomics
2.2. Proteomics
2.3. Phosphoproteomics
2.4. Metabolomics
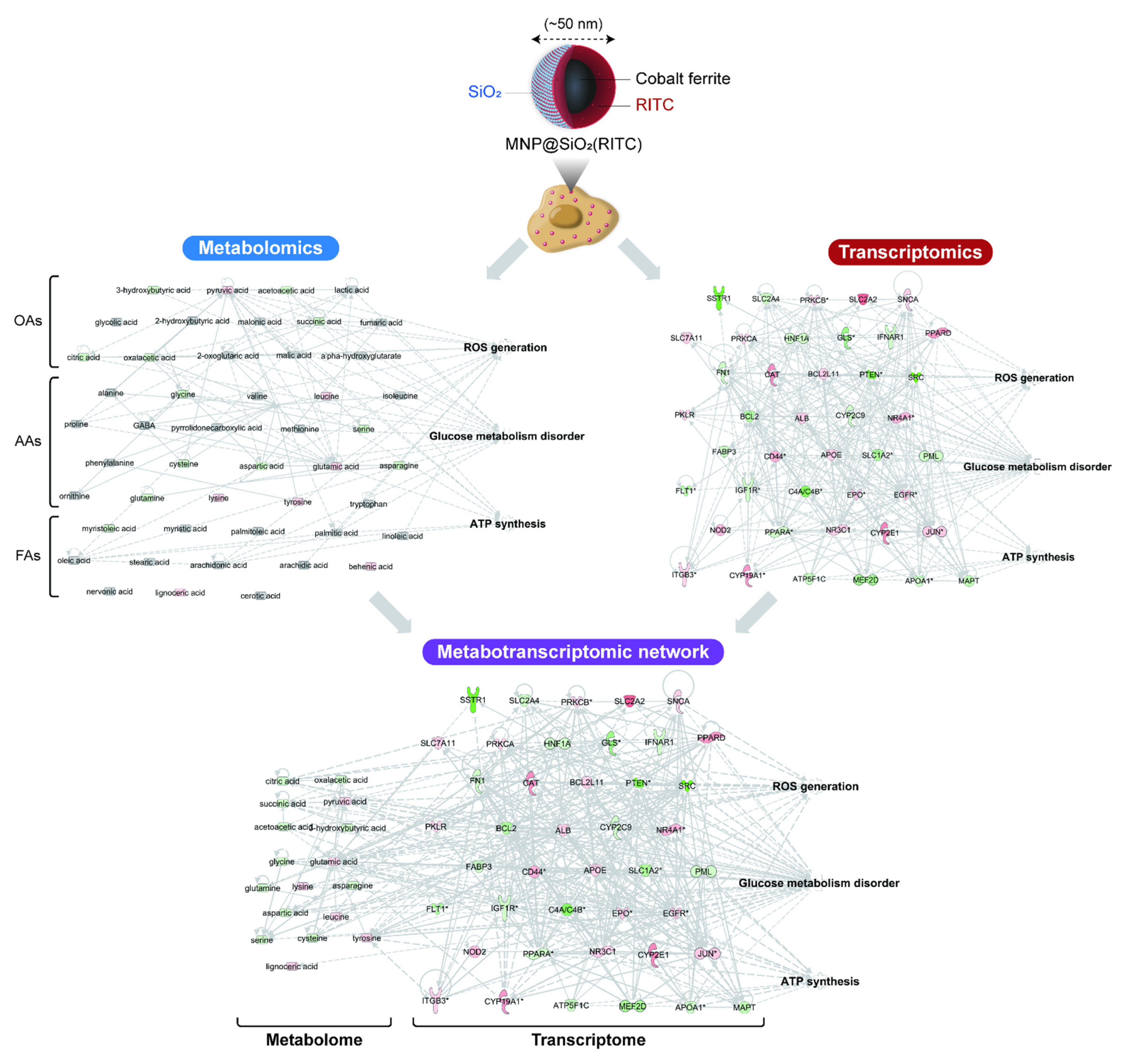
3. Integrated Omics for the Analysis of Nanotoxicity
4. ML for the Integration of Omics
5. Mechanobiology for the Analysis of Nanotoxicity
6. Summary
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Stark, W.J. Nanoparticles in Biological Systems. Angew. Chem. Int. Ed. 2011, 50, 1242–1258. [Google Scholar] [CrossRef] [PubMed]
- Sarma, A.; Bania, R.; Devi, J.R.; Deka, S. Therapeutic nanostructures and nanotoxicity. J. Appl. Toxicol. 2021, 1–24. [Google Scholar] [CrossRef]
- Cucci, L.; Trapani, G.; Hansson, Ö.; La Mendola, D.; Satriano, C. Gold Nanoparticles Functionalized with Angiogenin for Wound Care Application. Nanomaterials 2021, 11, 201. [Google Scholar] [CrossRef]
- Cucci, L.M.; Munzone, A.; Naletova, I.; Magrì, A.; La Mendola, D.; Satriano, C. Gold nanoparticles functionalized with angiogenin-mimicking peptides modulate cell membrane interactions. Biointerphases 2018, 13, 03C401. [Google Scholar] [CrossRef]
- Bouallegui, Y.; Ben Younes, R.; Bellamine, H.; Oueslati, R. Histopathology and analyses of inflammation intensity in the gills of mussels exposed to silver nanoparticles: Role of nanoparticle size, exposure time, and uptake pathways. Toxicol. Mech. Methods 2017, 27, 582–591. [Google Scholar] [CrossRef]
- Boyes, W.K.; Thornton, B.L.; Al-Abed, S.R.; Andersen, C.P.; Bouchard, D.C.; Burgess, R.M.; Hubal, E.A.C.; Ho, K.T.; Hughes, M.F.; Kitchin, K.; et al. A comprehensive framework for evaluating the environmental health and safety implications of engineered nanomaterials. Crit. Rev. Toxicol. 2017, 47, 771–814. [Google Scholar] [CrossRef]
- Ray, P.C.; Yu, H.; Fu, P.P. Toxicity and Environmental Risks of Nanomaterials: Challenges and Future Needs. J. Environ. Sci. Health Part C 2009, 27, 1–35. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zielińska, A.; Costa, B.; Ferreira, M.; Miguéis, D.; Louros, J.; Durazzo, A.; Lucarini, M.; Eder, P.; Chaud, M.; Morsink, M.; et al. Nanotoxicology and Nanosafety: Safety-By-Design and Testing at a Glance. Int. J. Environ. Res. Public Health 2020, 17, 4657. [Google Scholar] [CrossRef] [PubMed]
- Ma, C.; White, J.C.; Dhankher, O.P.; Xing, B. Metal-Based Nanotoxicity and Detoxification Pathways in Higher Plants. Environ. Sci. Technol. 2015, 49, 7109–7122. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Ji, Z.; Xia, T.; Meng, H.; Low-Kam, C.; Liu, R.; Pokhrel, S.; Lin, S.; Wang, X.; Liao, Y.-P.; et al. Use of Metal Oxide Nanoparticle Band Gap to Develop a Predictive Paradigm for Oxidative Stress and Acute Pulmonary Inflammation. ACS Nano 2012, 6, 4349–4368. [Google Scholar] [CrossRef]
- Sturla, S.J.; Boobis, A.; Fitzgerald, R.E.; Hoeng, J.; Kavlock, R.J.; Schirmer, K.; Whelan, M.; Wilks, M.; Peitsch, M.C. Systems Toxicology: From Basic Research to Risk Assessment. Chem. Res. Toxicol. 2013, 27, 314–329. [Google Scholar] [CrossRef]
- Tuncbag, N.; Gosline, S.; Kedaigle, A.J.; Soltis, A.R.; Gitter, A.; Fraenkel, E. Network-Based Interpretation of Diverse High-Throughput Datasets through the Omics Integrator Software Package. PLoS Comput. Biol. 2016, 12, e1004879. [Google Scholar] [CrossRef] [Green Version]
- Hasin, Y.; Seldin, M.; Lusis, A. Multi-omics approaches to disease. Genome Biol. 2017, 18, 1–15. [Google Scholar] [CrossRef]
- Krug, H.F.; Wick, P. Nanotoxicology: An Interdisciplinary Challenge. Angew. Chem. Int. Ed. 2011, 50, 1260–1278. [Google Scholar] [CrossRef]
- Qi, W.-Y.; Li, Q.; Chen, H.; Liu, J.; Xing, S.-F.; Xu, M.; Yan, Z.; Song, C.; Wang, S.-G. Selenium nanoparticles ameliorate Brassica napus L. cadmium toxicity by inhibiting the respiratory burst and scavenging reactive oxygen species. J. Hazard. Mater. 2021, 417, 125900. [Google Scholar] [CrossRef] [PubMed]
- Fröhlich, E. Role of omics techniques in the toxicity testing of nanoparticles. J. Nanobiotechnol. 2017, 15, 1–22. [Google Scholar] [CrossRef]
- Reyes, V.C.; Li, M.; Hoek, E.M.V.; Mahendra, S.; Damoiseaux, R. Genome-Wide Assessment in Escherichia coli Reveals Time-Dependent Nanotoxicity Paradigms. ACS Nano 2012, 6, 9402–9415. [Google Scholar] [CrossRef]
- Shin, T.H.; Seo, C.; Lee, D.Y.; Ji, M.; Manavalan, B.; Basith, S.; Chakkarapani, S.K.; Kang, S.H.; Lee, G.; Paik, M.J.; et al. Silica-coated magnetic nanoparticles induce glucose metabolic dysfunction in vitro via the generation of reactive oxygen species. Arch. Toxicol. 2019, 93, 1201–1212. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Verano-Braga, T.; Miethling-Graff, R.; Wojdyla, K.I.; Rogowska-Wrzesinska, A.; Brewer, J.; Erdmann, H.; Kjeldsen, F. Insights into the Cellular Response Triggered by Silver Nanoparticles Using Quantitative Proteomics. ACS Nano 2014, 8, 2161–2175. [Google Scholar] [CrossRef] [PubMed]
- Biola-Clier, M.; Gaillard, J.-C.; Rabilloud, T.; Armengaud, J.; Carriere, M. Titanium Dioxide Nanoparticles Alter the Cellular Phosphoproteome in A549 Cells. Nanomaterials 2020, 10, 185. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shin, T.H.; Lee, D.; Lee, H.Y.; Park, H.J.; Jin, M.S.; Paik, M.A.; Manavalan, B.; Mo, J.U.; Lee, G. Integration of metabolomics and transcriptomics in nanotoxicity studies. BMB Rep. 2018, 51, 14–20. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sun, Y.V.; Hu, Y.-J. Integrative Analysis of Multi-omics Data for Discovery and Functional Studies of Complex Human Diseases. Adv. Genet. 2016, 93, 147–190. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sun, J.; Zhou, Q.; Hu, X. Integrating multi-omics and regular analyses identifies the molecular responses of zebrafish brains to graphene oxide: Perspectives in environmental criteria. Ecotoxicol. Environ. Saf. 2019, 180, 269–279. [Google Scholar] [CrossRef]
- Hood, L. Systems biology: Integrating technology, biology, and computation. Mech. Ageing Dev. 2002, 124, 9–16. [Google Scholar] [CrossRef]
- Kim, J.S.; Yoon, T.-J.; Yu, K.N.; Kim, B.G.; Park, S.; Kim, H.W.; Lee, K.H.; Park, S.B.; Lee, J.-K.; Cho, M.H. Toxicity and Tissue Distribution of Magnetic Nanoparticles in Mice. Toxicol. Sci. 2005, 89, 338–347. [Google Scholar] [CrossRef]
- Beck, G.R.; Ha, S.-W.; Camalier, C.E.; Yamaguchi, M.; Li, Y.; Lee, J.-K.; Weitzmann, M.N. Bioactive silica-based nanoparticles stimulate bone-forming osteoblasts, suppress bone-resorbing osteoclasts, and enhance bone mineral density in vivo. Nanomed. Nanotechnol. Biol. Med. 2012, 8, 793–803. [Google Scholar] [CrossRef] [Green Version]
- Park, K.-S.; Tae, J.; Choi, B.; Kim, Y.-S.; Moon, C.; Kim, S.-H.; Lee, H.-S.; Kim, J.; Kim, J.; Park, J.; et al. Characterization, in vitro cytotoxicity assessment, and in vivo visualization of multimodal, RITC-labeled, silica-coated magnetic nanoparticles for labeling human cord blood–derived mesenchymal stem cells. Nanomed. Nanotechnol. Biol. Med. 2010, 6, 263–276. [Google Scholar] [CrossRef]
- Shim, W.; Paik, M.J.; Nguyen, D.T.; Lee, J.-K.; Lee, Y.; Kim, J.-H.; Shin, E.-H.; Kang, J.S.; Jung, H.-S.; Choi, S.; et al. Analysis of Changes in Gene Expression and Metabolic Profiles Induced by Silica-Coated Magnetic Nanoparticles. ACS Nano 2012, 6, 7665–7680. [Google Scholar] [CrossRef]
- Peng, T.; Wei, C.; Yu, F.; Xu, J.; Zhou, Q.; Shi, T.; Hu, X. Predicting nanotoxicity by an integrated machine learning and metabolomics approach. Environ. Pollut. 2020, 267, 115434. [Google Scholar] [CrossRef]
- Tan, J.L.; Tien, J.; Pirone, D.M.; Gray, D.S.; Bhadriraju, K.; Chen, C.S. Cells lying on a bed of microneedles: An approach to isolate mechanical force. Proc. Natl. Acad. Sci. USA 2003, 100, 1484–1489. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- DU Roure, O.; Saez, A.; Buguin, A.; Austin, R.H.; Chavrier, P.; Silberzan, P.; Ladoux, B. Force mapping in epithelial cell migration. Proc. Natl. Acad. Sci. USA 2005, 102, 2390–2395. [Google Scholar] [CrossRef] [Green Version]
- Shin, T.H.; Ketebo, A.A.; Lee, D.Y.; Lee, S.; Kang, S.H.; Basith, S.; Manavalan, B.; Kwon, D.H.; Park, S.; Lee, G. Decrease in membrane fluidity and traction force induced by silica-coated magnetic nanoparticles. J. Nanobiotechnol. 2021, 19, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Ketebo, A.A.; Shin, T.H.; Jun, M.; Lee, G.; Park, S. Effect of silica-coated magnetic nanoparticles on rigidity sensing of human embryonic kidney cells. J. Nanobiotechnol. 2020, 18, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Fan, T.W.-M.; Higashi, R.M.; Lane, A.N. Integrating Metabolomics and Transcriptomics for Probing Se Anticancer Mechanisms. Drug Metab. Rev. 2006, 38, 707–732. [Google Scholar] [CrossRef]
- Alsagaby, S.A.; Vijayakumar, R.; Premanathan, M.; Mickymaray, S.; Alturaiki, W.; Al-Baradie, R.S.; AlGhamdi, S.; Aziz, M.A.; Alhumaydhi, F.A.; Alzahrani, F.A.; et al. Transcriptomics-Based Characterization of the Toxicity of ZnO Nanoparticles Against Chronic Myeloid Leukemia Cells. Int. J. Nanomed. 2020, ume 15, 7901–7921. [Google Scholar] [CrossRef]
- Panahi, Y.; Fattahi, A.; Zarei, F.; Ghasemzadeh, N.; Mohammadpoor, A.; Abroon, S.; Nojadeh, J.N.; Khojastefard, M.; Akbarzadeh, A.; Ghasemnejad, T. Next-generation sequencing approaches for the study of genome and epigenome toxicity induced by sulfur mustard. Arch. Toxicol. 2018, 92, 3443–3457. [Google Scholar] [CrossRef] [PubMed]
- Federico, A.; Serra, A.; Ha, M.K.; Kohonen, P.; Choi, J.-S.; Liampa, I.; Nymark, P.; Sanabria, N.; Cattelani, L.; Fratello, M.; et al. Transcriptomics in Toxicogenomics, Part II: Preprocessing and Differential Expression Analysis for High Quality Data. Nanomaterials 2020, 10, 903. [Google Scholar] [CrossRef]
- Simon, D.F.; Domingos, R.; Hauser, C.; Hutchins, C.M.; Zerges, W.; Wilkinson, K.J. Transcriptome Sequencing (RNA-seq) Analysis of the Effects of Metal Nanoparticle Exposure on the Transcriptome of Chlamydomonas reinhardtii. Appl. Environ. Microbiol. 2013, 79, 4774–4785. [Google Scholar] [CrossRef] [Green Version]
- Chen, Z.; Gao, S.-H.; Jin, M.; Sun, S.; Lu, J.; Yang, P.; Bond, P.; Yuan, Z.; Guo, J. Physiological and transcriptomic analyses reveal CuO nanoparticle inhibition of anabolic and catabolic activities of sulfate-reducing bacterium. Environ. Int. 2019, 125, 65–74. [Google Scholar] [CrossRef] [PubMed]
- Yoon, T.-J.; Kim, J.S.; Kim, B.G.; Yu, K.N.; Cho, M.-H.; Lee, J.-K. Multifunctional Nanoparticles Possessing A? Magnetic Motor Effect? for Drug or Gene Delivery. Angew. Chem. Int. Ed. 2005, 44, 1068–1071. [Google Scholar] [CrossRef]
- Bartel, D.P. MicroRNAs: Target Recognition and Regulatory Functions. Cell 2009, 136, 215–233. [Google Scholar] [CrossRef] [Green Version]
- Griffiths-Jones, S.; Saini, H.K.; van Dongen, S.; Enright, A.J. miRBase: Tools for microRNA genomics. Nucleic Acids Res. 2007, 36, D154–D158. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, L.; Wan, H.; Liu, Q.; Wang, D. Multi-walled carbon nanotubes-induced alterations in microRNA let-7 and its targets activate a protection mechanism by conferring a developmental timing control. Part. Fibre Toxicol. 2017, 14, 1–11. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, R.; Ren, M.; Rui, Q.; Wang, D. A mir-231-Regulated Protection Mechanism against the Toxicity of Graphene Oxide in Nematode Caenorhabditis elegans. Sci. Rep. 2016, 6, 32214. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Wu, Q.; Li, Y.; Nouara, A.; Jia, R.; Wang, D. In vivo translocation and toxicity of multi-walled carbon nanotubes are regulated by microRNAs. Nanoscale 2014, 6, 4275–4284. [Google Scholar] [CrossRef]
- O’Farrell, P. High resolution two-dimensional electrophoresis of proteins. J. Biol. Chem. 1975, 250, 4007–4021. [Google Scholar] [CrossRef]
- Ma, T.; Zhang, A. Integrate multi-omics data with biological interaction networks using Multi-view Factorization AutoEncoder (MAE). BMC Genom. 2019, 20, 1–11. [Google Scholar] [CrossRef]
- Gonzalez-Gonzalez, M.; Acevedo, R.J.; Matarraz, S.; Jara-Acevedo, M.; Paradinas, S.; Sayagües, J.; Orfao, A.; Fuentes, M. Nanotechniques in proteomics: Protein microarrays and novel detection platforms. Eur. J. Pharm. Sci. 2012, 45, 499–506. [Google Scholar] [CrossRef]
- Agrawal, G.K.; Timperio, A.M.; Zolla, L.; Bansal, V.; Shukla, R.; Rakwal, R. Biomarker discovery and applications for foods and beverages: Proteomics to nanoproteomics. J. Proteom. 2013, 93, 74–92. [Google Scholar] [CrossRef]
- Karkossa, I.; Raps, S.; Von Bergen, M.; Schubert, K. Systematic Review of Multi-Omics Approaches to Investigate Toxicological Effects in Macrophages. Int. J. Mol. Sci. 2020, 21, 9371. [Google Scholar] [CrossRef]
- Nienhaus, K.; Nienhaus, G.U. Towards a molecular-level understanding of the protein corona around nanoparticles—Recent advances and persisting challenges. Curr. Opin. Biomed. Eng. 2019, 10, 11–22. [Google Scholar] [CrossRef]
- Qin, M.; Zhang, J.; Li, M.; Yang, D.; Liu, D.; Song, S.; Fu, J.; Zhang, H.; Dai, W.; Wang, X.; et al. Proteomic analysis of intracellular protein corona of nanoparticles elucidates nano-trafficking network and nano-bio interactions. Theranostics 2020, 10, 1213–1229. [Google Scholar] [CrossRef] [PubMed]
- Blume, J.E.; Manning, W.C.; Troiano, G.; Hornburg, D.; Figa, M.; Hesterberg, L.; Platt, T.L.; Zhao, X.; Cuaresma, R.A.; Everley, P.A.; et al. Rapid, deep and precise profiling of the plasma proteome with multi-nanoparticle protein corona. Nat. Commun. 2020, 11, 1–14. [Google Scholar] [CrossRef]
- Roy, D.N.; Goswami, R.; Pal, A. Nanomaterial and toxicity: What can proteomics tell us about the nanotoxicology? Xenobiotica 2016, 47, 632–643. [Google Scholar] [CrossRef]
- Matysiak, M.; Kapka-Skrzypczak, L.; Brzóska, K.; Gutleb, A.C.; Kruszewski, M. Proteomic approach to nanotoxicity. J. Proteom. 2016, 137, 35–44. [Google Scholar] [CrossRef]
- Zhang, T.; Gaffrey, M.J.; Thrall, B.D.; Qian, W.-J. Mass spectrometry-based proteomics for system-level characterization of biological responses to engineered nanomaterials. Anal. Bioanal. Chem. 2018, 410, 6067–6077. [Google Scholar] [CrossRef]
- Duan, G.; Walther, D. The Roles of Post-translational Modifications in the Context of Protein Interaction Networks. PLoS Comput. Biol. 2015, 11, e1004049. [Google Scholar] [CrossRef]
- Deribe, Y.L.; Pawson, T.; Dikic, I. Post-translational modifications in signal integration. Nat. Struct. Mol. Biol. 2010, 17, 666–672. [Google Scholar] [CrossRef] [PubMed]
- Lu, C.; Thompson, C.B. Metabolic Regulation of Epigenetics. Cell Metab. 2012, 16, 9–17. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fiehn, O. Metabolomics—The link between genotypes and phenotypes. Plant Mol. Biol. 2002, 48, 155–171. [Google Scholar] [CrossRef]
- Zlatkis, A.; Brazell, R.S.; Poole, C.F. The role of organic volatile profiles in clinical diagnosis. Clin. Chem. 1981, 27, 789–797. [Google Scholar] [CrossRef]
- Zabala-Letona, A.; Arruabarrena-Aristorena, A.; Martín-Martín, N.; Fernandez-Ruiz, S.; Sutherland, J.D.; Clasquin, M.; Tomas-Cortazar, J.; Jimenez, J.; Torres, I.; Quang, P.; et al. mTORC1-dependent AMD1 regulation sustains polyamine metabolism in prostate cancer. Nature 2017, 547, 109–113. [Google Scholar] [CrossRef] [PubMed]
- Emwas, A.-H.; Roy, R.; McKay, R.T.; Tenori, L.; Saccenti, E.; Gowda, G.A.N.; Raftery, D.; AlAhmari, F.; Jaremko, L.; Jaremko, M.; et al. NMR Spectroscopy for Metabolomics Research. Metabolites 2019, 9, 123. [Google Scholar] [CrossRef] [Green Version]
- Pan, Z.; Raftery, D. Comparing and combining NMR spectroscopy and mass spectrometry in metabolomics. Anal. Bioanal. Chem. 2006, 387, 525–527. [Google Scholar] [CrossRef] [PubMed]
- Phukan, G.; Shin, T.H.; Shim, J.S.; Paik, M.J.; Lee, J.-K.; Choi, S.; Kim, Y.M.; Kang, S.H.; Kim, H.S.; Kang, Y.; et al. Silica-coated magnetic nanoparticles impair proteasome activity and increase the formation of cytoplasmic inclusion bodies in vitro. Sci. Rep. 2016, 6, 29095. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Misra, B.; Langefeld, C.D.; Olivier, M.; Cox, L.A. Integrated omics: Tools, advances and future approaches. J. Mol. Endocrinol. 2019, 62, R21–R45. [Google Scholar] [CrossRef] [Green Version]
- Hartiala, J.A.; Tang, W.H.W.; Wang, Z.; Crow, A.L.; Stewart, A.; Roberts, R.; McPherson, R.; Erdmann, J.; Willenborg, C.; Hazen, S.L.; et al. Genome-wide association study and targeted metabolomics identifies sex-specific association of CPS1 with coronary artery disease. Nat. Commun. 2016, 7, 10558. [Google Scholar] [CrossRef] [Green Version]
- Shah, S.H.; Newgard, C.B. Integrated metabolomics and genomics: Systems approaches to biomarkers and mechanisms of cardiovascular disease. Circ. Cardiovasc. Genet. 2015, 8, 410–419. [Google Scholar] [CrossRef] [Green Version]
- Berg, K.C.G.; Eide, P.W.; Eilertsen, I.A.; Johannessen, B.; Bruun, J.; Danielsen, S.A.; Bjørnslett, M.; Meza-Zepeda, L.A.; Eknæs, M.; Lind, G.E.; et al. Multi-omics of 34 colorectal cancer cell lines—A resource for biomedical studies. Mol. Cancer 2017, 16, 1–16. [Google Scholar] [CrossRef]
- Rebollar, E.A.; Antwis, R.; Becker, M.H.; Belden, L.; Bletz, M.C.; Brucker, R.M.; Harrison, X.; Hughey, M.C.; Kueneman, J.G.; Loudon, A.H.; et al. Using “Omics” and Integrated Multi-Omics Approaches to Guide Probiotic Selection to Mitigate Chytridiomycosis and Other Emerging Infectious Diseases. Front. Microbiol. 2016, 7, 68. [Google Scholar] [CrossRef]
- Kim, J.; Woo, H.R.; Gil Nam, H. Toward Systems Understanding of Leaf Senescence: An Integrated Multi-Omics Perspective on Leaf Senescence Research. Mol. Plant 2016, 9, 813–825. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kang, W.; Li, X.; Sun, A.; Yu, F.; Hu, X. Study of the Persistence of the Phytotoxicity Induced by Graphene Oxide Quantum Dots and of the Specific Molecular Mechanisms by Integrating Omics and Regular Analyses. Environ. Sci. Technol. 2019, 53, 3791–3801. [Google Scholar] [CrossRef]
- Ebrahim, A.; Brunk, E.; Tan, J.; O’Brien, E.J.; Kim, D.; Szubin, R.; Lerman, J.A.; Lechner, A.; Sastry, A.; Bordbar, A.; et al. Multi-omic data integration enables discovery of hidden biological regularities. Nat. Commun. 2016, 7, 13091. [Google Scholar] [CrossRef] [Green Version]
- Braakhuis, H.M.; Park, M.V.D.Z.; Gosens, I.; De Jong, W.H.; Cassee, F.R. Physicochemical characteristics of nanomaterials that affect pulmonary inflammation. Part. Fibre Toxicol. 2014, 11, 18. [Google Scholar] [CrossRef] [Green Version]
- Gatoo, M.A.; Naseem, S.; Arfat, M.Y.; Dar, A.M.; Qasim, K.; Zubair, S. Physicochemical Properties of Nanomaterials: Implication in Associated Toxic Manifestations. BioMed Res. Int. 2014, 2014, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Alcantar, N.A.; Aydil, E.; Israelachvili, J.N. Polyethylene glycol-coated biocompatible surfaces. J. Biomed. Mater. Res. 2000, 51, 343–351. [Google Scholar] [CrossRef]
- Silva, M.M.; Calado, R.; Marto, J.; Bettencourt, A.; Almeida, A.J.; Gonçalves, L.M.D. Chitosan Nanoparticles as a Mucoadhesive Drug Delivery System for Ocular Administration. Mar. Drugs 2017, 15, 370. [Google Scholar] [CrossRef] [Green Version]
- Ding, Y.-F.; Li, S.; Liang, L.; Huang, Q.; Yuwen, L.; Yang, W.J.; Wang, R.; Wang, L.-H. Highly Biocompatible Chlorin e6-Loaded Chitosan Nanoparticles for Improved Photodynamic Cancer Therapy. ACS Appl. Mater. Interfaces 2018, 10, 9980–9987. [Google Scholar] [CrossRef] [PubMed]
- Ito, A.; Honda, H.; Kobayashi, T. Cancer immunotherapy based on intracellular hyperthermia using magnetite nanoparticles: A novel concept of “heat-controlled necrosis” with heat shock protein expression. Cancer Immunol. Immunother. 2005, 55, 320–328. [Google Scholar] [CrossRef] [PubMed]
- Cavill, R.; Jennen, D.; Kleinjans, J.; Briedé, J.J. Transcriptomic and metabolomic data integration. Briefings Bioinform. 2015, 17, 891–901. [Google Scholar] [CrossRef] [Green Version]
- Hu, M.; Li, J.; Hou, M.; Liu, X.; Cui, S.; Yang, X.; Liu, L.; Jiang, X.; Mu, G. Transcriptomic and metabolomic joint analysis reveals distinct flavonoid biosynthesis regulation for variegated testa color development in peanut (Arachis hypogaea L.). Sci. Rep. 2021, 11, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Evans, T.G. Considerations for the use of transcriptomics in identifying the ‘genes that matter’ for environmental adaptation. J. Exp. Biol. 2015, 218, 1925–1935. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shin, T.H.; Lee, D.Y.; Manavalan, B.; Basith, S.; Na, Y.-C.; Yoon, C.; Lee, H.-S.; Paik, M.J.; Lee, G. Silica-coated magnetic nanoparticles activate microglia and induce neurotoxic d-serine secretion. Part. Fibre Toxicol. 2021, 18, 1–18. [Google Scholar] [CrossRef]
- Basith, S.; Manavalan, B.; Shin, T.H.; Lee, G. Machine intelligence in peptide therapeutics: A next-generation tool for rapid disease screening. Med. Res. Rev. 2020, 40, 1276–1314. [Google Scholar] [CrossRef] [PubMed]
- Assawamakin, A.; Prueksaaroon, S.; Kulawonganunchai, S.; Shaw, P.J.; Varavithya, V.; Ruangrajitpakorn, T.; Tongsima, S. Biomarker Selection and Classification of “-Omics” Data Using a Two-Step Bayes Classification Framework. BioMed Res. Int. 2013, 2013, 1–9. [Google Scholar] [CrossRef] [Green Version]
- Kim, S.; Jhong, J.-H.; Lee, J.-J.; Koo, J.-Y. Meta-analytic support vector machine for integrating multiple omics data. BioData Min. 2017, 10, 1–14. [Google Scholar] [CrossRef] [Green Version]
- Lummertz da Rocha, E.; Malleshaiah, M. Trajectory Algorithms to Infer Stem Cell Fate Decisions. In Computational Stem Cell Biology: Methods and Protocols; Cahan, P., Ed.; Springer: New York, NY, USA, 2019; pp. 193–209. [Google Scholar]
- Karim, R.; Beyan, O.; Zappa, A.; Costa, I.G.; Rebholz-Schuhmann, D.; Cochez, M.; Decker, S. Deep learning-based clustering approaches for bioinformatics. Briefings Bioinform. 2020, 22, 393–415. [Google Scholar] [CrossRef] [Green Version]
- Martorell-Marugán, J.; Tabik, S.; Benhammou, Y.; del Val, C.; Zwir, I.; Herrera, F.; Carmona-Sáez, P. Deep Learning in Omics Data Analysis and Precision Medicine. In Computational Biology; Husi, H., Ed.; Codon Publications: Brisbane, Australia, 2019. [Google Scholar]
- Furxhi, I.; Murphy, F. Predicting In Vitro Neurotoxicity Induced by Nanoparticles Using Machine Learning. Int. J. Mol. Sci. 2020, 21, 5280. [Google Scholar] [CrossRef] [PubMed]
- Winkler, D.A. Role of Artificial Intelligence and Machine Learning in Nanosafety. Small 2020, 16, e2001883. [Google Scholar] [CrossRef]
- Oh, E.; Liu, R.; Nel, A.; Gemill, K.B.; Bilal, M.; Cohen, R.L.A.N.M.B.Y.; Medintz, K.B.G.I.L. Meta-analysis of cellular toxicity for cadmium-containing quantum dots. Nat. Nanotechnol. 2016, 11, 479–486. [Google Scholar] [CrossRef] [PubMed]
- Vogel, V.; Sheetz, M.P. Local force and geometry sensing regulate cell functions. Nat. Rev. Mol. Cell Biol. 2006, 7, 265–275. [Google Scholar] [CrossRef]
- Huang, Y.; Schell, C.; Huber, T.B.; Simsek, A.N.; Hersch, N.; Merkel, R.; Gompper, G.; Sabass, B. Traction force microscopy with optimized regularization and automated Bayesian parameter selection for comparing cells. Sci. Rep. 2019, 9, 1–16. [Google Scholar] [CrossRef] [Green Version]
- Mohammed, D.; Versaevel, M.; Bruyère, C.; Alaimo, L.; Luciano, M.; Vercruysse, E.; Procès, A.; Gabriele, S. Innovative Tools for Mechanobiology: Unraveling Outside-In and Inside-Out Mechanotransduction. Front. Bioeng. Biotechnol. 2019, 7, 162. [Google Scholar] [CrossRef]
- Wang, J.H.C.; Thampatty, B.P. An Introductory Review of Cell Mechanobiology. Biomech. Model. Mechanobiol. 2006, 5, 1–16. [Google Scholar] [CrossRef]
- Ghassemi, S.; Biais, N.; Maniura, K.; Wind, S.J.; Sheetz, M.P.; Hone, J. Fabrication of elastomer pillar arrays with modulated stiffness for cellular force measurements. J. Vac. Sci. Technol. B Microelectron. Nanometer Struct. 2008, 26, 2549–2553. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ghassemi, S.; Meacci, G.; Liu, S.; Gondarenko, A.A.; Mathur, A.; Roca-Cusachs, P.; Sheetz, M.P.; Hone, J. Cells test substrate rigidity by local contractions on submicrometer pillars. Proc. Natl. Acad. Sci. USA 2012, 109, 5328–5333. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vogel, V.; Sheetz, M.P. Cell fate regulation by coupling mechanical cycles to biochemical signaling pathways. Curr. Opin. Cell Biol. 2009, 21, 38–46. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dupont, S.; Morsut, L.; Aragona, M.; Enzo, E.; Giulitti, S.; Cordenonsi, M.; Zanconato, F.; Le Digabel, J.; Forcato, M.; Bicciato, S.; et al. Role of YAP/TAZ in mechanotransduction. Nature 2011, 474, 179–183. [Google Scholar] [CrossRef]
- Schwartz, M.A. Integrins and Extracellular Matrix in Mechanotransduction. Cold Spring Harb. Perspect. Biol. 2010, 2, a005066. [Google Scholar] [CrossRef]
- Prager-Khoutorsky, M.; Lichtenstein, A.; Krishnan, R.; Rajendran, K.; Mayo, A.; Kam, Z.; Geiger, B.; Bershadsky, A. Fibroblast polarization is a matrix-rigidity-dependent process controlled by focal adhesion mechanosensing. Nature 2011, 13, 1457–1465. [Google Scholar] [CrossRef]


| Entrez Gene Name | Symbol | ID b | Location | Signal Fold Change a | |
|---|---|---|---|---|---|
| 0.1 μg/μL | 1.0 μg/μL | ||||
| albumin | ALB | 1565228_s_at | Extracellular Space | 5.50 | 4.16 |
| apolipoprotein A1 | APOA1 | 217073_x_at | Extracellular Space | −3.75 | −1.53 |
| apolipoprotein E | APOE | 203382_s_at | Extracellular Space | 5.13 | 3.38 |
| ATP synthase F1 subunit gamma | ATP5F1C | 214132_at | Cytoplasm | −6.06 | −1.28 |
| BCL2 apoptosis regulator | BCL2 | 203684_s_at | Cytoplasm | −10.46 | −7.26 |
| BCL2 like 11 | BCL2L11 | 208536_s_at | Cytoplasm | 6.37 | 1.70 |
| catalase | CAT | 215573_at | Cytoplasm | 17.41 | 13.38 |
| CD44 molecule (Indian blood group) | CD44 | 234411_x_at | Plasma Membrane | 11.38 | −4.89 |
| complement C4A (Rodgers blood group) | C4A/C4B | 208451_s_at | Extracellular Space | −20.71 | 9.05 |
| cytochrome P450 family 19 subfamily A member 1 | CYP19A1 | 1554296_at | Cytoplasm | 15.56 | −17.51 |
| cytochrome P450 family 2 subfamily C member 9 | CYP2C9 | 214419_s_at | Cytoplasm | −3.60 | 1.19 |
| cytochrome P450 family 2 subfamily E member 1 | CYP2E1 | 209976_s_at | Cytoplasm | 18.60 | 3.69 |
| epidermal growth factor receptor | EGFR | 1565484_x_at | Plasma Membrane | 9.62 | 9.52 |
| erythropoietin | EPO | 207257_at | Extracellular Space | 8.02 | 8.34 |
| fatty acid binding protein 3 | FABP3 | 214285_at | Cytoplasm | −3.13 | −4.84 |
| fibronectin 1 | FN1 | 214702_at | Extracellular Space | −4.96 | 1.89 |
| fms related receptor tyrosine kinase 1 | FLT1 | 210287_s_at | Plasma Membrane | −10.50 | −3.85 |
| glutaminase | GLS | 203158_s_at | Cytoplasm | −14.49 | −2.26 |
| HNF1 homeobox A | HNF1A | 216930_at | Nucleus | −4.21 | 1.60 |
| insulin like growth factor 1 receptor | IGF1R | 243358_at | Plasma Membrane | −4.50 | −1.44 |
| integrin subunit beta 3 | ITGB3 | 215240_at | Plasma Membrane | 5.79 | 5.93 |
| interferon alpha and beta receptor subunit 1 | IFNAR1 | 204191_at | Plasma Membrane | −5.46 | 1.37 |
| Jun proto-oncogene, AP-1 transcription factor subunit | JUN | 201465_s_at | Nucleus | 7.34 | 1.43 |
| microtubule associated protein tau | MAPT | 203930_s_at | Plasma Membrane | −8.28 | −1.43 |
| myocyte enhancer factor 2D | MEF2D | 225641_at | Nucleus | −11.89 | −3.92 |
| nuclear receptor subfamily 3 group C member 1 | NR3C1 | 232431_at | Nucleus | 4.01 | −1.09 |
| nuclear receptor subfamily 4 group A member 1 | NR4A1 | 211143_x_at | Nucleus | 11.14 | 4.23 |
| nucleotide binding oligomerization domain containing 2 | NOD2 | 220066_at | Cytoplasm | 9.56 | 7.03 |
| peroxisome proliferator activated receptor alpha | PPARA | 1560981_a_at | Nucleus | −4.81 | 6.60 |
| peroxisome proliferator activated receptor delta | PPARD | 210636_at | Nucleus | 15.79 | 7.90 |
| phosphatase and tensin homolog | PTEN | 240964_at | Cytoplasm | −24.40 | 4.66 |
| promyelocytic leukemia | PML | 239582_at | Nucleus | −5.15 | 1.31 |
| protein kinase C alpha | PRKCA | 215195_at | Cytoplasm | 3.53 | 2.76 |
| protein kinase C beta | PRKCB | 227824_at | Cytoplasm | 6.41 | −10.37 |
| pyruvate kinase L/R | PKLR | 207858_s_at | Cytoplasm | 3.38 | 1.11 |
| solute carrier family 1 member 2 | SLC1A2 | 217055_x_at | Plasma Membrane | −11.34 | −8.32 |
| solute carrier family 2 member 2 | SLC2A2 | 206535_at | Plasma Membrane | 26.34 | 1.90 |
| solute carrier family 2 member 4 | SLC2A4 | 206603_at | Plasma Membrane | −5.42 | 3.00 |
| solute carrier family 7 member 11 | SLC7A11 | 217678_at | Plasma Membrane | 5.61 | 2.00 |
| somatostatin receptor 1 | SSTR1 | 235591_at | Plasma Membrane | −29.18 | −1.32 |
| SRC proto-oncogene, non-receptor tyrosine kinase | SRC | 1558210_at | Cytoplasm | −33.92 | −12.56 |
| synuclein alpha | SNCA | 236081_at | Cytoplasm | 3.22 | −1.89 |
| 3-hydroxybutyric acid | 300-85-6 | Other | −1.49 | −1.39 | |
| acetoacetic acid | 541-50-4 | Other | −1.40 | −1.40 | |
| citric acid | 77-92-9 | Other | −1.39 | −1.27 | |
| glycine | 56-40-6 | Other | −1.53 | −1.07 | |
| asparagine | 70-47-3 | Other | −1.21 | 1.01 | |
| aspartic acid | 56-84-8 | Other | −1.22 | −1.12 | |
| cysteine | 52-90-4 | Other | −1.24 | 1.09 | |
| glutamic acid | 56-86-0 | Other | 1.50 | 1.24 | |
| glutamine | 56-85-9 | Other | −1.89 | −1.51 | |
| leucine | 61-90-5 | Other | 1.38 | 1.28 | |
| lysine | 56-87-1 | Other | 1.91 | 1.25 | |
| serine | 56-45-1 | Other | −1.97 | 2.12 | |
| tyrosine | 60-18-4 | Other | 1.21 | −1.09 | |
| lignoceric acid | 557-59-5 | Other | 1.33 | 1.21 | |
| oxalacetic acid | 328-42-7 | Other | −1.47 | −1.32 | |
| pyruvic acid | 127-17-3 | Other | 1.41 | −6.13 | |
| succinic acid | 110-15-6 | Other | −2.28 | −1.21 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shin, T.H.; Nithiyanandam, S.; Lee, D.Y.; Kwon, D.H.; Hwang, J.S.; Kim, S.G.; Jang, Y.E.; Basith, S.; Park, S.; Mo, J.-S.; et al. Analysis of Nanotoxicity with Integrated Omics and Mechanobiology. Nanomaterials 2021, 11, 2385. https://doi.org/10.3390/nano11092385
Shin TH, Nithiyanandam S, Lee DY, Kwon DH, Hwang JS, Kim SG, Jang YE, Basith S, Park S, Mo J-S, et al. Analysis of Nanotoxicity with Integrated Omics and Mechanobiology. Nanomaterials. 2021; 11(9):2385. https://doi.org/10.3390/nano11092385
Chicago/Turabian StyleShin, Tae Hwan, Saraswathy Nithiyanandam, Da Yeon Lee, Do Hyeon Kwon, Ji Su Hwang, Seok Gi Kim, Yong Eun Jang, Shaherin Basith, Sungsu Park, Jung-Soon Mo, and et al. 2021. "Analysis of Nanotoxicity with Integrated Omics and Mechanobiology" Nanomaterials 11, no. 9: 2385. https://doi.org/10.3390/nano11092385
APA StyleShin, T. H., Nithiyanandam, S., Lee, D. Y., Kwon, D. H., Hwang, J. S., Kim, S. G., Jang, Y. E., Basith, S., Park, S., Mo, J.-S., & Lee, G. (2021). Analysis of Nanotoxicity with Integrated Omics and Mechanobiology. Nanomaterials, 11(9), 2385. https://doi.org/10.3390/nano11092385







