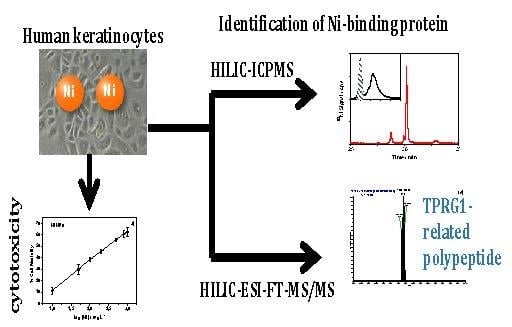
Nickel Nanoparticles Induce the Synthesis of a Tumor-Related Polypeptide in Human Epidermal Keratinocytes
Abstract
1. Introduction
2. Materials and Methods
2.1. Reagents and Chemicals
2.2. Cell Viability Assay
2.3. Preparation of Nickel Nanoparticles Suspension and Stability Test
2.4. Total Nickel Determination in Cells
2.5. Single Particle-ICPMS Analysis
2.6. Size Exclusion Chromatography-ICPMS
2.7. Hydrophilic Interaction Liquid Chromatography-ICPMS
2.8. Hydrophilic Interaction Liquid Chromatography-ESI-FT-MS/MS
2.9. Top-Down Protein Sequence Identification
3. Results and Discussion
3.1. Evaluation of the Cytotoxicity of Nickel Nanoparticles and Other Nickel Compounds Towards Human Skin Cells
3.2. Determination of the Nickel Uptake
3.3. Physicochemical Form of Ni in Cell Cytosols
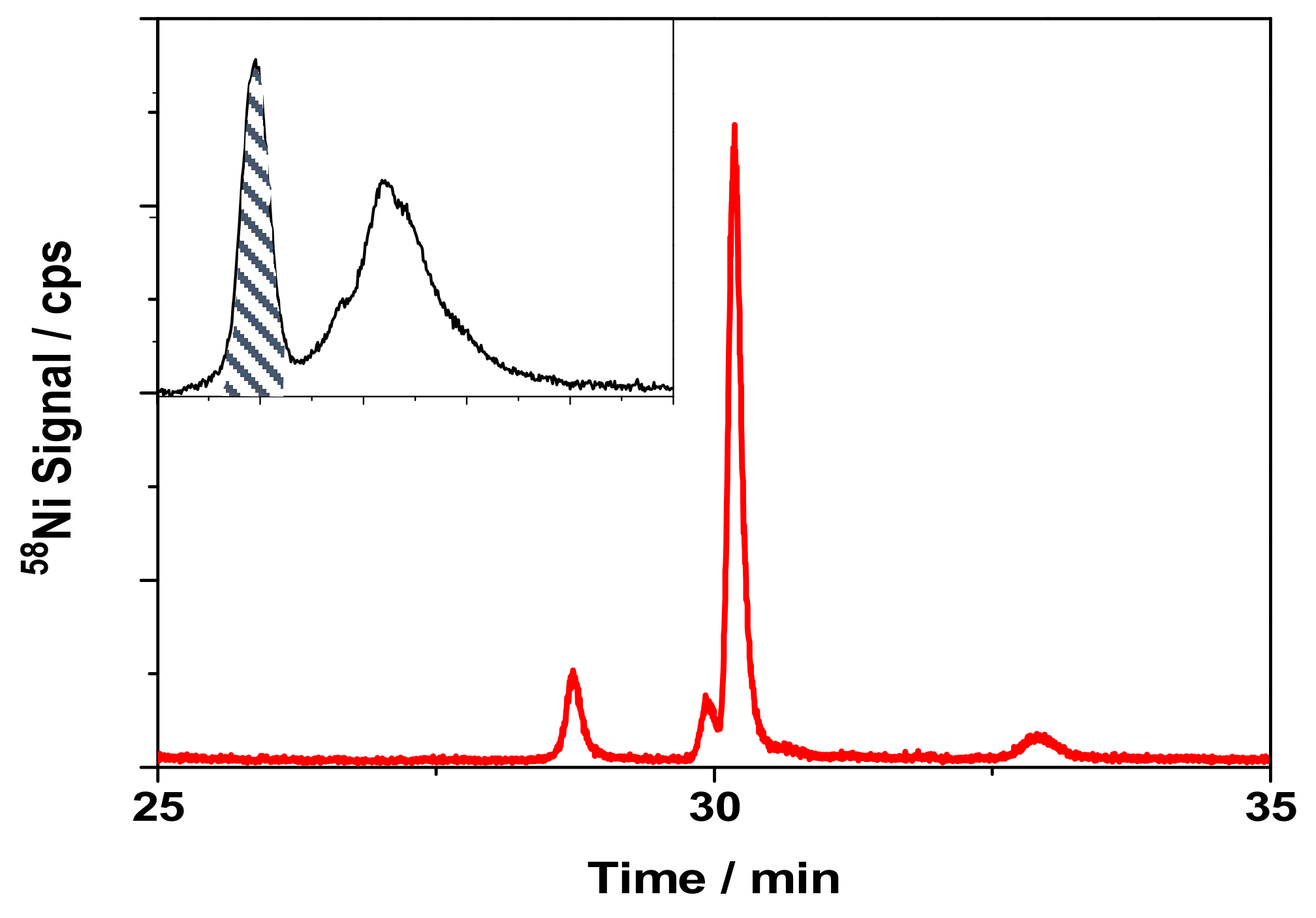
3.4. Separation of Ni-Binding Compounds
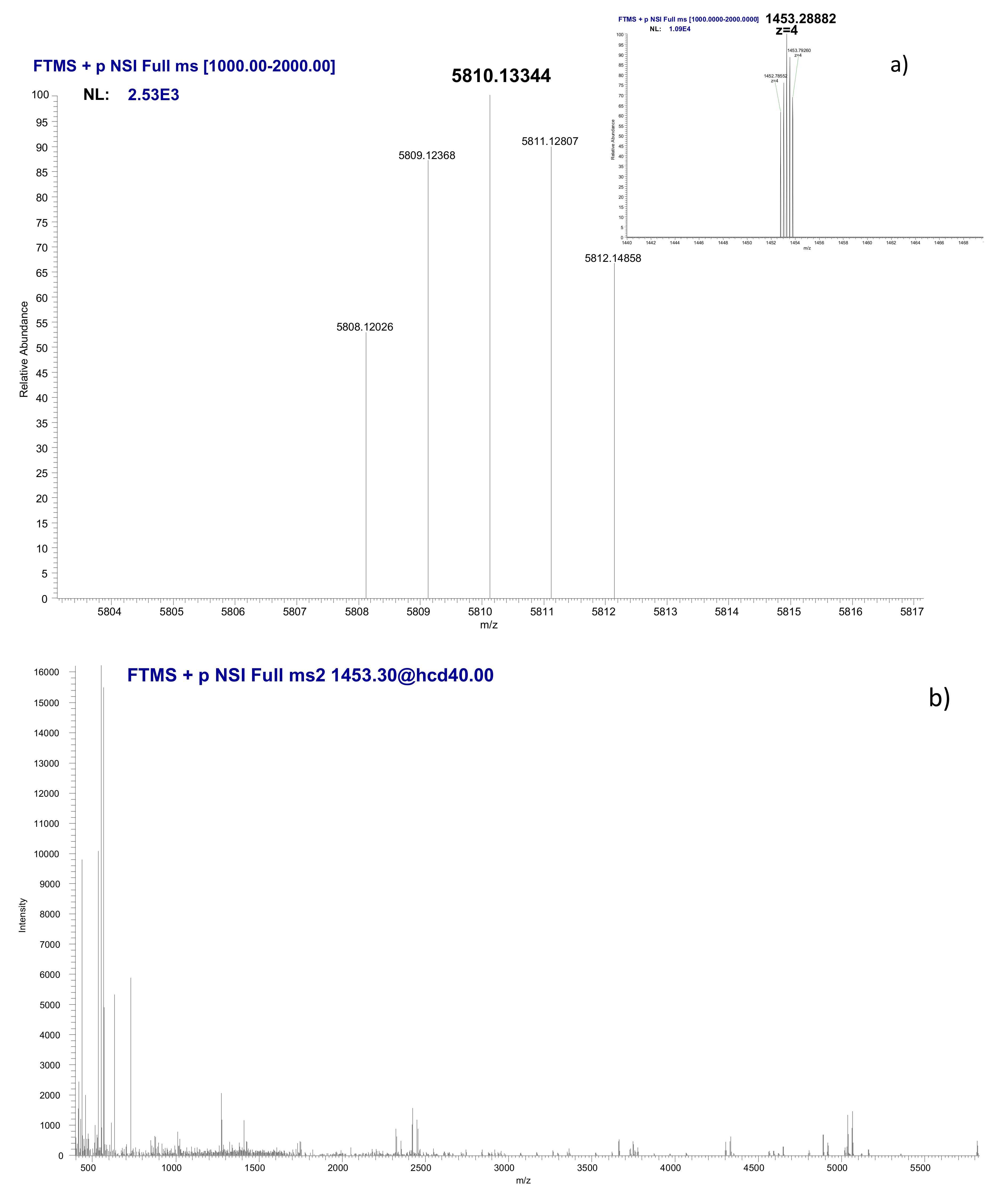
3.5. Identification of the Ni-Binding Compound Expressed by Keratinocytes
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- National Toxicology Program, Department of Health and Human Service. Nickel Compounds and Metallic Nickel; Report on Carcinogens, 14th ed.; National Toxicology Program, Department of Health and Human Service: Washington, DC, USA, 2014; pp. 1–4.
- Lippmann, M.; Ito, K.; Hwang, J.S.; Maciejczyk, P.; Chen, L.C. Cardiovascular effects of nickel in ambient air. Environ. Health Perspect. 2006, 114, 1662–1669. [Google Scholar] [CrossRef] [PubMed]
- Morgan, L.G.; Usher, V. Health problems associated with nickel refining and use. Ann. Occup. Hyg. 1994, 38, 189–198. [Google Scholar] [CrossRef] [PubMed]
- Garner, L.A. Contact dermatitis to metaks. Dermatol. Ther. 2004, 17, 321–327. [Google Scholar] [CrossRef] [PubMed]
- Schaumlöffel, D. Speciation of Nickel. In Handbook of Elemental Speciation II: Species in the Environment, Food, Medicine and Occupational Health; Cornelis, R., Caruso, J., Crews, H., Heumann, K., Eds.; John Wiley & Sons: Chichester, UK, 2005; ISBN 0946-672X. [Google Scholar]
- Cameron, K.S.; Buchner, V.; Tchounwou, P.B. Exploring the molecular mechanisms of nickel-induced genotoxicity and carcinogenicity: A literature review. Rev. Environ. Health 2011, 26, 81–92. [Google Scholar] [CrossRef] [PubMed]
- Costa, M.; Yan, Y.; Zhao, D.; Salnikow, K. Molecular mechanisms of nickel carcinogenesis: Gene silencing by nickel delivery to the nucleus and gene activation/inactivation by nickel-induced cell signaling. J. Environ. Monit. 2003, 5, 222–223. [Google Scholar] [CrossRef]
- Report of the Nickel Ion Bioavailability Workshop Volume I. Available online: https://www.tera.org/Peer/NiBioavailability/Nickel%20Workshop%202.10%20Meeting%20Report%20FINAL.pdf (accessed on 20 March 2020).
- Gliga, A.R.; Di Bucchianico, S.; Åkerlund, E.; Karlsson, H.L. Transcriptome Profiling and Toxicity Following Long-Term, Low Dose Exposure of Human Lung Cells to Ni and NiO Nanoparticles—Comparison with NiCl2. Nanomaterials 2020, 10, 649. [Google Scholar] [CrossRef] [PubMed]
- Oller, A.R. Respiratory Carcinogenicity Assessment of Soluble Nickel Compounds. Environ. Health Perspect. 2002, 110, 841–844. [Google Scholar] [CrossRef]
- Magaye, R.; Zhao, J. Recent progress in studies of metallic nickel and nickel-based nanoparticles’ genotoxicity and carcinogenicity. Environ. Toxicol. Pharmacol. 2012, 34, 644–650. [Google Scholar] [CrossRef]
- Muñoz, A.; Costa, M. Elucidating the mechanisms of nickel compound uptake: A review of particulate and nano-nickel endocytosis and toxicity. Toxicol. Appl. Pharmacol. 2012, 260, 1–16. [Google Scholar] [CrossRef]
- Jiménez-Lamana, J.; Szpunar, J. Analytical approaches for the characterization of nickel proteome. Metallomics 2017, 9, 1014–1027. [Google Scholar] [CrossRef] [PubMed]
- Mosmann, T. Rapid Colorimetric Assay for Cellular Growth and Survival: Application to Proliferation and Cytotoxicity Assays. J. Immunol. Methods 1983, 65, 55–63. [Google Scholar] [CrossRef]
- International Organization for Standardization. ISO 10993-5 Biological Evaluation of Medical Devices—Part 5: Tests for In Vitro Cytotoxicity; ISO: Geneva, Switzerland, 2009; Volume 2007, pp. 1–11. [Google Scholar] [CrossRef]
- Cox, B.; Emili, A. Tissue subcellular fractionation and protein extraction for use in mass-spectrometry-based proteomics. Nat. Protoc. 2006, 1, 1872–1878. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Sirotkin, Y.; Shen, Y.; Anderson, G.; Tsai, Y.S.; Ting, Y.S.; Goodlett, D.R.; Smith, R.D.; Bafna, V.; Pevzner, P.A. Protein Identification Using Top-Down Spectra. Mol. Cell. Proteomics 2011, 11, M111.008524. [Google Scholar] [CrossRef]
- Costa, A.; Eberlin, S.; Polettini, A.; Pereira, A.; Pereira, C.; Ferreira, N.; Dolis, E.; Torloni, L. Neuromodulatory and Anti-Inflammatory Ingredient for Sensitive Skin: In Vitro Assessment. Inflamm. Allergy-Drug Targets 2014, 13, 191–198. [Google Scholar] [CrossRef]
- Fletcher, G.G.; Rossetto, F.E.; Turnbull, J.D.; Nieboer, E. Toxicity, Uptake, and Mutagenicity of Particulate and Soluble Nickel Compounds. Environ. Health Perspect. 1994, 102, 69–79. [Google Scholar]
- Landolph, J.R., Jr.; Xue, W.; Warshawsky, D. Whole animal carcinogenicity bioassays. In Molecular Carcinogenesis and the Molecular Biology of Human Cancer; Warshawsky, D., Landolph, J.R., Jr., Eds.; CRC Press: Boca Raton, FL, USA, 2015; pp. 25–44. ISBN 9781479978007. [Google Scholar]
- Bevan, R.J.; Harrison, P.T.C. Threshold and non-threshold chemical carcinogens: A survey of the present regulatory landscape. Regul. Toxicol. Pharmacol. 2017, 88, 291–302. [Google Scholar] [CrossRef]
- Neumann, H.-G. Risk assessment of chemical carcinogens and thresholds. Crit. Rev. Toxicol. 2009, 39, 449–461. [Google Scholar] [CrossRef]
- Tennekes, H.A.; Pletz, J.; Sanchez-Bayo, F. Development of a Dose-Response Model for Risk Assessment of Receptor-Mediated Effects Henk A. Tennekes. Intern. Med. Rev. 2016, 1, 1–13. [Google Scholar]
- Alarifi, S.; Ali, D.; Alakhtani, S.; Suhaibani, E.S.; Al-qahtani, A.A. Reactive Oxygen Species-Mediated DNA Damage and Apoptosis in Human Skin Epidermal Cells After Exposure to Nickel Nanoparticles. Biol. Trace Element Res. 2014, 84–93. [Google Scholar] [CrossRef] [PubMed]
- Magaye, R.; Zhou, Q.; Bowman, L.; Zou, B.; Mao, G.; Xu, J.; Castranova, V.; Zhao, J.; Ding, M. Metallic Nickel Nanoparticles May Exhibit Higher Carcinogenic Potential than Fine Particles in JB6 Cells. PLoS ONE 2014, 9, e92418. [Google Scholar] [CrossRef] [PubMed]
- Jin, C.; Zhu, B.; Wang, X.; Lu, Q. Cytotoxicity of Titanium Dioxide Nanoparticles in Mouse Fibroblast Cells. Chem. Res. Toxicol. 2008, 21, 1871–1877. [Google Scholar] [CrossRef] [PubMed]
- Sahu, D.; Kannan, G.M.; Tailang, M.; Vijayaraghavan, R. In Vitro Cytotoxicity of Nanoparticles: A Comparison between Particle Size and Cell Type. J. Nanosci. 2016, 2016, 1–9. [Google Scholar] [CrossRef]
- Chusuei, C.C.; Wu, C.; Mallavarapu, S.; Yao, F.; Hou, S.; Hsu, C.; Winiarz, J.G.; Aronstam, R.S.; Huang, Y. Chemico-Biological Interactions Cytotoxicity in the age of nano: The role of fourth period transition metal oxide nanoparticle physicochemical properties. Chem. Biol. Interact. 2013, 206, 319–326. [Google Scholar] [CrossRef]
- Acevedo, F.; Serra, M.Á.; Ermolli, M.; Clerici, L.; Vesterberg, O. Nickel-induced proteins in human HaCaT keratinocytes: Annexin II and phosphoglycerate kinase. Toxicology 2001, 159, 33–41. [Google Scholar] [CrossRef]
- Ermolli, M.; Menné, C.; Pozzi, G.; Serra, M.Á.; Clerici, L.A. Nickel, cobalt and chromium-induced cytotoxicity and intracellular accumulation in human hacat keratinocytes. Toxicology 2001, 159, 23–31. [Google Scholar] [CrossRef]
- Laborda, F.; Bolea, E.; Jiménez-Lamana, J. Single particle inductively coupled plasma mass spectrometry: A powerful tool for nanoanalysis. Anal. Chem. 2014, 86, 2270–2278. [Google Scholar] [CrossRef]
- Montoro Bustos, A.R.; Purushotham, K.P.; Possolo, A.; Farkas, N.; Vladár, A.E.; Murphy, K.E.; Winchester, M.R. Validation of Single Particle ICP-MS for Routine Measurements of Nanoparticle Size and Number Size Distribution. Anal. Chem. 2018, 90, 14376–14386. [Google Scholar] [CrossRef]
- Laborda, F.; Jiménez-Lamana, J.; Bolea, E.; Castillo, J.R. Selective identification, characterization and determination of dissolved silver(i) and silver nanoparticles based on single particle detection by inductively coupled plasma mass spectrometry. J. Anal. Atomic Spectrom. 2011, 26, 1362. [Google Scholar] [CrossRef]
- Jiménez-Lamana, J.; Wojcieszek, J.; Jakubiak, M.; Asztemborska, M.; Szpunar, J. Single particle ICP-MS characterization of platinum nanoparticles uptake and bioaccumulation by Lepidium sativum and Sinapis alba plants. J. Anal. Atom. Spectrom. 2016, 31, 2321–2329. [Google Scholar] [CrossRef]
- Kińska, K.; Jiménez-Lamana, J.; Kowalska, J.; Krasnodębska-Ostręga, B.; Szpunar, J. Study of the uptake and bioaccumulation of palladium nanoparticles by Sinapis alba using single particle ICP-MS. Sci. Total Environ. 2018, 615, 1078–1085. [Google Scholar] [CrossRef] [PubMed]
- Zook, J.M.; Long, S.E.; Cleveland, D.; Geronimo, C.L.A.; MacCuspie, R.I. Measuring silver nanoparticle dissolution in complex biological and environmental matrices using UV-visible absorbance. Anal. Bioanal. Chem. 2011, 401, 1993–2002. [Google Scholar] [CrossRef] [PubMed]
- Bolea, E.; Jiménez-Lamana, J.; Laborda, F.; Abad-Álvaro, I.; Bladé, C.; Arola, L.; Castillo, J.R. Detection and characterization of silver nanoparticles and dissolved species of silver in culture medium and cells by AsFlFFF-UV-Vis-ICPMS: Application to nanotoxicity tests. Analyst 2014, 139, 914–922. [Google Scholar] [CrossRef]
- Hansen, U.; Thünemann, A.F. Characterization of Silver Nanoparticles in Cell Culture Medium Containing Fetal Bovine Serum. Langmuir 2015, 31, 6842–6852. [Google Scholar] [CrossRef]
- Munusamy, P.; Wang, C.; Engelhard, M.H.; Baer, D.R.; Smith, J.N.; Kodali, V.; Thrall, B.D.; Chen, S.; Porter, A.E.; Ryan, M.P.; et al. Comparison of 20 nm silver nanoparticles synthesized with and without a gold core: Structure, dissolution in cell culture media, and biological impact on macrophages. Biointerphases 2015, 10, 1–16. [Google Scholar] [CrossRef]
- Liu, J.; Pennell, K.G.; Hurt, R.H. Kinetics and mechanisms of nanosilver oxysulfidation. Environ. Sci. Technol. 2011, 45, 7345–7353. [Google Scholar] [CrossRef]
- Loza, K.; Diendorf, J.; Sengstock, C.; Ruiz-Gonzalez, L.; Gonzalez-Calbet, J.M.; Vallet-Regi, M.; Köller, M.; Epple, M. The dissolution and biological effects of silver nanoparticles in biological media. J. Mater. Chem. B 2014, 2, 1634–1643. [Google Scholar] [CrossRef]
- Perez, J.E.; Alsharif, N.; Martínez-Banderas, A.I.; Othman, B.; Merzaban, J.; Ravasi, T.; Kosel, J. Additional Review of In Vitro Toxicity of Nanoparticles and Nanorods. In Cytotoxicity; IntechOpen: London, UK, 2018; pp. 234–265. [Google Scholar]
- Buszewski, B.; Noga, S. Hydrophilic interaction liquid chromatography (HILIC)-a powerful separation technique. Anal. Bioanal. Chem. 2012, 402, 231–247. [Google Scholar] [CrossRef]
- Mounicou, S.; Ouerdane, L.; Passagne, I. Identification of Metallothionein Subisoforms in HPLC Using Accurate Mass and Online Sequencing by Electrospray Hybrid Linear Ion Trap-Orbital Ion Trap Mass Spectrometry. Anal. Chem. 2010, 82, 6947–6957. [Google Scholar] [CrossRef]
- Antonini, D.; Dentice, M.; Mahtani, P.; De Rosa, L.; Della Gatta, G.; Mandinova, A.; Salvatore, D.; Stupka, E.; Missero, C. Tprg, a gene predominantly expressed in skin, is a direct target of the transcription factor p63. J. Investig. Dermatol. 2008, 128, 1676–1685. [Google Scholar] [CrossRef] [PubMed]
- Peana, M.; Medici, S.; Nurchi, V.M.; Crisponi, G.; Zoroddu, M.A. Nickel binding sites in histone proteins: Spectroscopic and structural characterization. Coord. Chem. Rev. 2013, 257, 2737–2751. [Google Scholar] [CrossRef]
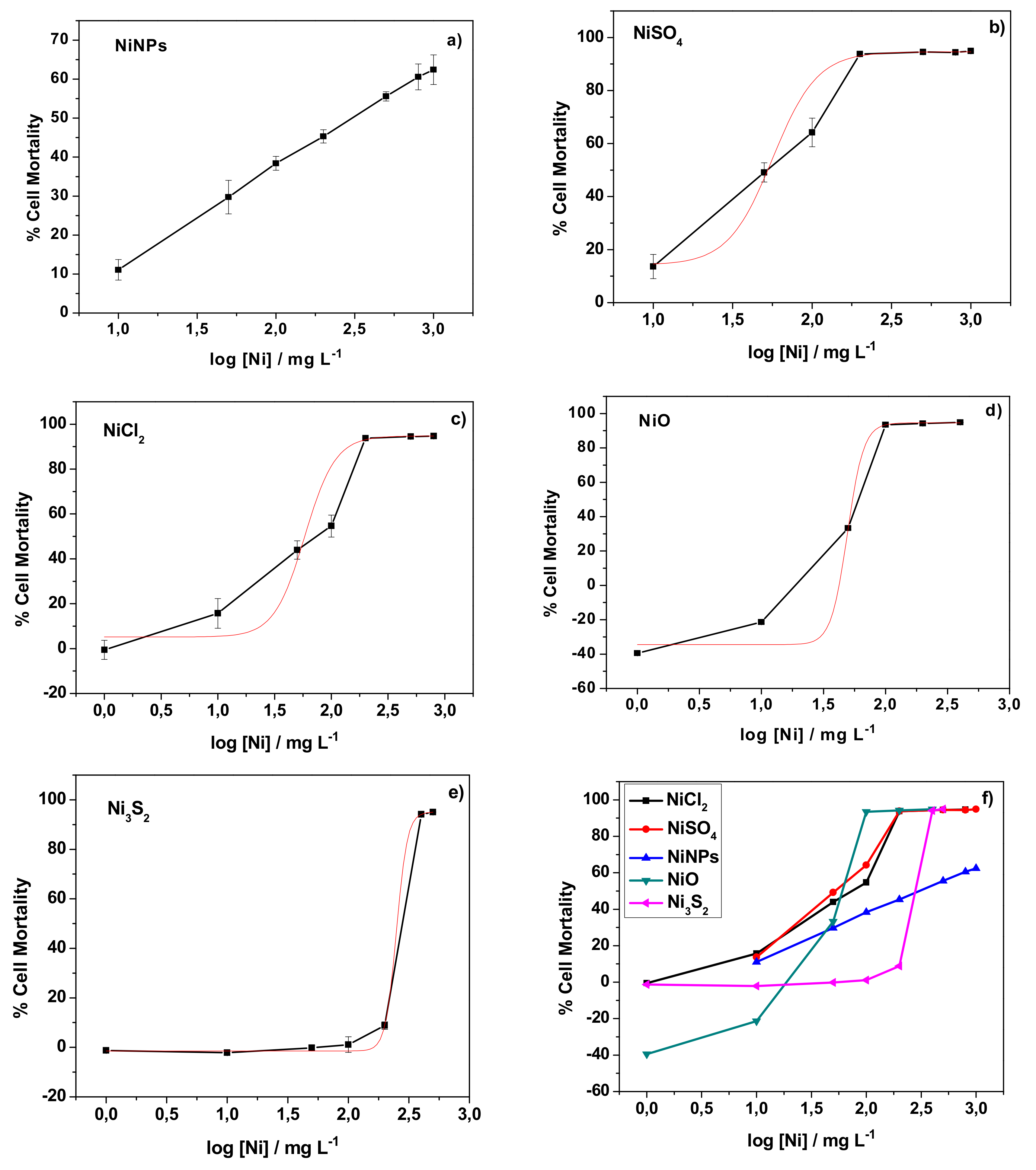
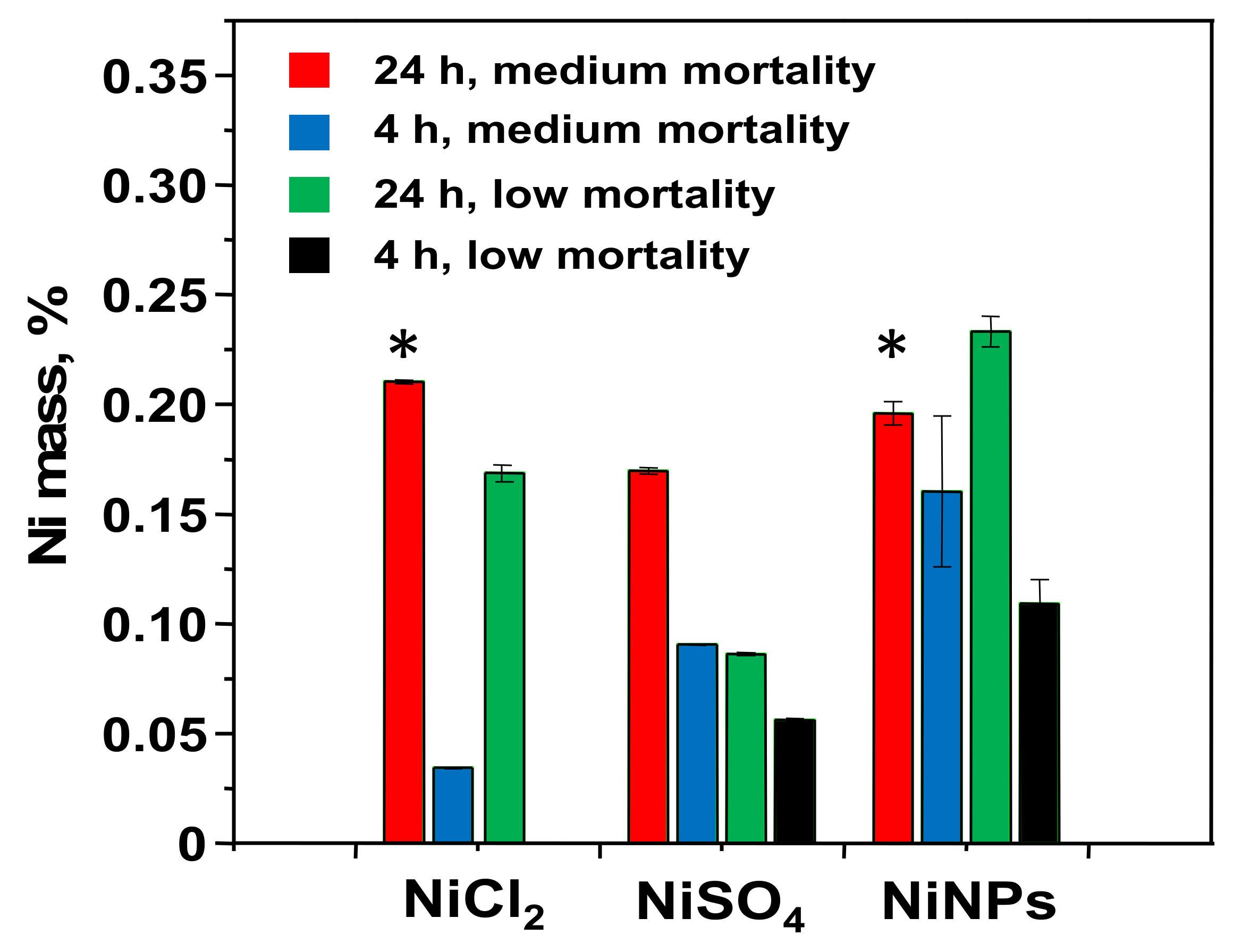

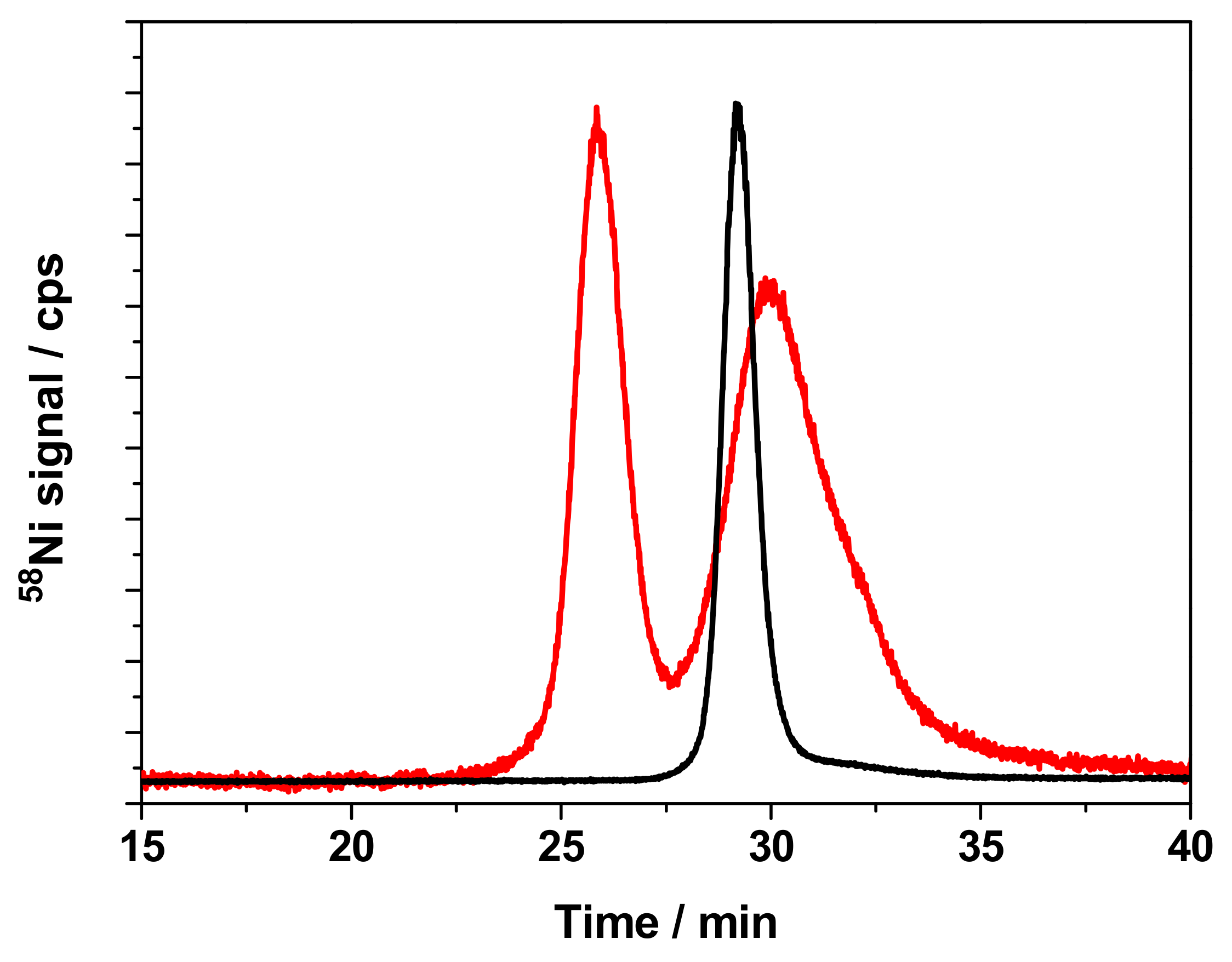


| Compound | NiNPs | NiSO4 | NiCl2 | NiO | Ni3S2 |
|---|---|---|---|---|---|
| LD50, mg L−1 | - | 56.4 | 58.3 | 49.2 | 249.5 |
| Compound | Ni Mass Added, µg | Ni Mass in Medium, µg | Ni Mass in Cells, µg (%) | Rec, % | Ni Mass in Cytosol, µg (%) |
|---|---|---|---|---|---|
| NiNPs | 1000 | 42.6 ± 0.1 | 37.1 ± 0.2 (3.71 ± 0.02) | 8 ± 1 | 1.96 ± 0.05 (0.20 ± 0.01) * |
| NiCl2 | 250 | 248 ± 14 | 0.59 ± 0.01 (0.24 ± 0.01) | 99 ± 5 | 0.53 ± 0.01 (0.21 ± 0.01) * |
| NiSO4 | 250 | 240 ± 2 | 0.54 ± 0.01 (0.21 ± 0.01) | 96 ± 1 | 0.42 ± 0.01 (0.17 ± 0.01) |
| Ion Type | Theoretical Mass | Observed Mass | Mass Difference (Da) | Mass Difference (ppm) |
|---|---|---|---|---|
| B4 | 426.2227 | 426.2264 | 0.00368 | 8.6 |
| B8 | 876.4566 | 876.4566 | −0.00002 | −0.023 |
| B9 | 963.4886 | 963.4879 | −0.00069 | −0.72 |
| B12 | 1208.5898 | 1208.5947 | 0.00493 | 4.1 |
| B14 | 1421.7124 | 1421.7138 | 0.00147 | 1.0 |
| B15 | 1478.7338 | 1478.7388 | 0.00499 | 3.4 |
| B16 | 1606.8288 | 1606.8423 | 0.01354 | 8.4 |
| B19 | 1979.0483 | 1979.0335 | −0.01484 | −7.5 |
| B23 | 2421.2910 | 2421.2671 | −0.02396 | −9.9 |
| B45 | 4902.6000 | 4902.5637 | −0.03635 | −7.4 |
| Y10 | 1069.5556 | 1069.5532 | −0.00239 | −2.2 |
| Y11 | 1126.5771 | 1126.5867 | 0.00964 | 8.6 |
| Y20 | 2156.1490 | 2156.1305 | −0.01852 | −8.6 |
| Y33 | 3617.8915 | 3617.8719 | −0.01961 | −5.4 |
| Y34 | 3730.9756 | 3730.9461 | −0.02947 | −7.9 |
| Y47 | 5046.6786 | 5046.6357 | −0.04290 | −8.5 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jiménez-Lamana, J.; Godin, S.; Aragonès, G.; Bladé, C.; Szpunar, J.; Łobinski, R. Nickel Nanoparticles Induce the Synthesis of a Tumor-Related Polypeptide in Human Epidermal Keratinocytes. Nanomaterials 2020, 10, 992. https://doi.org/10.3390/nano10050992
Jiménez-Lamana J, Godin S, Aragonès G, Bladé C, Szpunar J, Łobinski R. Nickel Nanoparticles Induce the Synthesis of a Tumor-Related Polypeptide in Human Epidermal Keratinocytes. Nanomaterials. 2020; 10(5):992. https://doi.org/10.3390/nano10050992
Chicago/Turabian StyleJiménez-Lamana, Javier, Simon Godin, Gerard Aragonès, Cinta Bladé, Joanna Szpunar, and Ryszard Łobinski. 2020. "Nickel Nanoparticles Induce the Synthesis of a Tumor-Related Polypeptide in Human Epidermal Keratinocytes" Nanomaterials 10, no. 5: 992. https://doi.org/10.3390/nano10050992
APA StyleJiménez-Lamana, J., Godin, S., Aragonès, G., Bladé, C., Szpunar, J., & Łobinski, R. (2020). Nickel Nanoparticles Induce the Synthesis of a Tumor-Related Polypeptide in Human Epidermal Keratinocytes. Nanomaterials, 10(5), 992. https://doi.org/10.3390/nano10050992