Printability and Cell Viability in Extrusion-Based Bioprinting from Experimental, Computational, and Machine Learning Views
Abstract
:1. Introduction
2. Experimental View
2.1. Printability
2.1.1. Evaluation of Printability
- Extrudability: the minimum extrusion pressure required to print material at the desired flow rate [19];
- Strand printability:This factor is used to compare printed strands dimeters with CAD-generated parameters strands. The expected strand diameter is [20,21]:where DS, Vn, and Qt are strands diameter, needle speed, and volumetric flow rate, respectively.So, needle speed affects the diameter of strands, and the below index is defined to compare printed strands with calculated ones:
- Integrity factor: this factor compares the thickness of printed scaffolds and designed ones
- Irregularity:In fact, this index is developed index of integrity in 3 dimensions. It compares the outer geometry of scaffolds with designed ones in X, Y, and Z directions [21]:
- Pore printability:
2.1.2. Effective Parameters on Printability
Bioink Properties
Bioprinting Parameters
Construct Design
2.1.3. Printability Improvement
2.2. Cell Viability
3. Computational View
3.1. Printability
3.1.1. Implementation
Surface Marker Techniques
Surface-Fitted Method
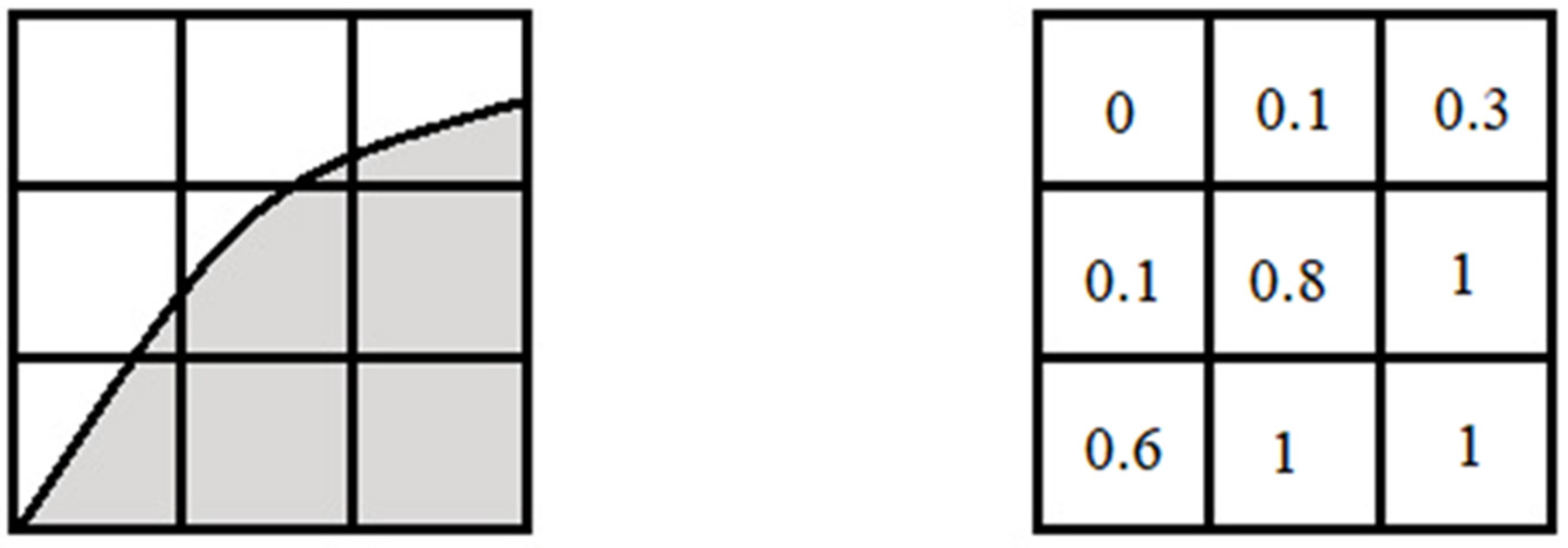
The Volume of Fluid (VOF)
Level Set Method
3.2. Cell Viability
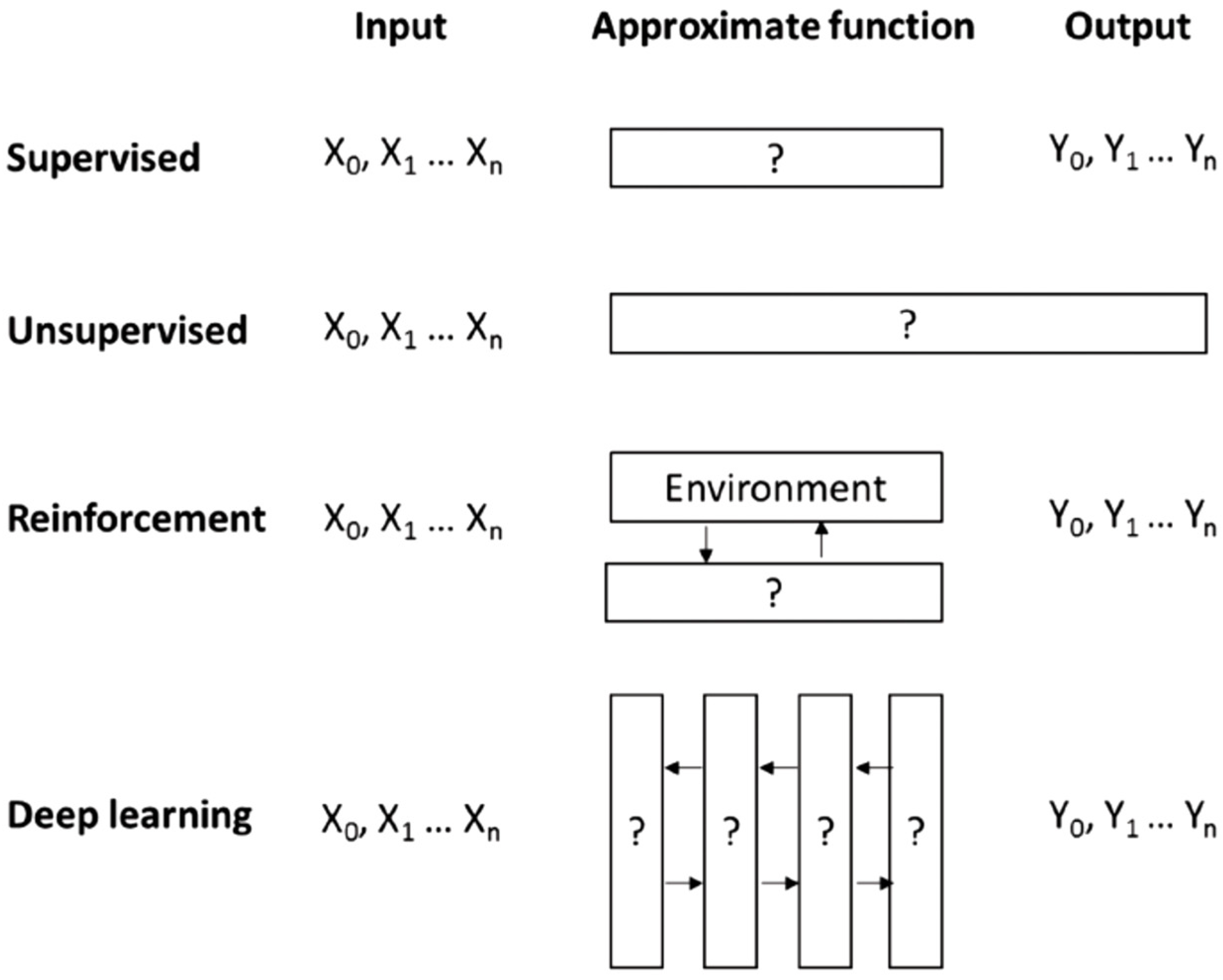
4. Machine Learning View
4.1. Printability
4.2. Cell Viability
5. Conclusions and Future Work
Author Contributions
Funding
Conflicts of Interest
References
- Abouna, G.M. Organ Shortage Crisis: Problems and Possible Solutions. Transplant. Proc. 2008, 40, 34–38. [Google Scholar] [CrossRef] [PubMed]
- Mandrycky, C.; Wang, Z.; Kim, K.; Kim, D.H. 3D bioprinting for engineering complex tissues. Biotechnol. Adv. 2016, 34, 422–434. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Khademhosseini, A.; Langer, R.; Borenstein, J.; Vacanti, J.P. Microscale technologies for tissue engineering and biology. Proc. Natl. Acad. Sci. USA 2006, 103, 2480–2487. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, X.B. Extrusion Bioprinting of Scaffolds for Tissue Engineering; Springer Nature Switzerland AG: Cham, Switzerland, 2019. [Google Scholar]
- Kozior, T.; Bochnia, J.; Gogolewski, D.; Zmarzły, P.; Rudnik, M.; Szot, W.; Szczygieł, P.; Musiałek, M. Analysis of Metrological Quality and Mechanical Properties of Models Manufactured with Photo-Curing PolyJet Matrix Technology for Medical Applications. Polymers 2022, 14, 408. [Google Scholar] [CrossRef] [PubMed]
- Betancourt, N.; Chen, X. Review of extrusion-based multi-material bioprinting processes. Bioprinting 2022, 25, e00189. [Google Scholar] [CrossRef]
- You, F.; Wu, X.; Kelly, M.; Chen, X. Bioprinting and in vitro characterization of alginate dialdehyde–Gelatin hydrogel bio-ink. Bio-Des. Manuf. 2020, 3, 48–59. [Google Scholar] [CrossRef] [Green Version]
- Delkash, Y.; Gouin, M.; Rimbeault, T.; Mohabatpour, F.; Papagerakis, P.; Maw, S.; Chen, X. Bioprinting and in vitro characterization of an eggwhite-based cell-laden patch for endothelialized tissue engineering applications. J. Funct. Biomater. 2021, 12, 45. [Google Scholar] [CrossRef]
- Sadeghianmaryan, A.; Naghieh, S.; Alizadeh Sardroud, H.; Yazdanpanah, Z.; Afzal Soltani, Y.; Sernaglia, J.; Chen, X. Extrusion-based printing of chitosan scaffolds and their in vitro characterization for cartilage tissue engineering. Int. J. Biol. Macromol. 2020, 164, 3179–3192. [Google Scholar] [CrossRef]
- Ning, L.; Zhu, N.; Smith, A.; Rajaram, A.; Hou, H.; Srinivasan, S.; Mohabatpour, F.; He, L.; McLnnes, A.; Serpooshan, V.; et al. Noninvasive Three-Dimensional in Situ and In Vivo Characterization of Bioprinted Hydrogel Scaffolds Using the X-ray Propagation-Based Imaging Technique. ACS Appl. Mater. Interfaces 2021, 13, 25611–25623. [Google Scholar] [CrossRef]
- Naghieh, S.; Sarker, M.D.; Sharma, N.K.; Barhoumi, Z.; Chen, X. Printability of 3D printed hydrogel scaffolds: Influence of hydrogel composition and printing parameters. Appl. Sci. 2020, 10, 292. [Google Scholar] [CrossRef] [Green Version]
- Zimmerling, A.; Yazdanpanah, Z.; Cooper, D.M.L.; Johnston, J.D.; Chen, X. 3D printing PCL/nHA bone scaffolds: Exploring the influence of material synthesis techniques. Biomater. Res. 2021, 25, 3. [Google Scholar] [CrossRef] [PubMed]
- Suntornnond, R.; Tan, E.Y.S.; An, J.; Chua, C.K. A Mathematical Model on the Resolution of Extrusion Bioprinting for the Development of New Bioinks. Materials 2016, 9, 756. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, Z.; Jin, Y.; Yin, J.; Xu, C.; Xiong, R.; Christensen, K.; Ringeisen, B.R.; Chrisey, D.B.; Huang, Y. Evaluation of bioink printability for bioprinting applications. Appl. Phys. Rev. 2018, 5, 041304. [Google Scholar] [CrossRef]
- Müller, M.; Becher, J.; Schnabelrauch, M.; Zenobi-Wong, M. Nanostructured Pluronic hydrogels as bioinks for 3D bioprinting. Biofabrication 2015, 7, 035006. [Google Scholar] [CrossRef]
- Kyle, S.; Jessop, Z.M.; Al-Sabah, A.; Whitaker, I.S. ‘Printability’ of Candidate Biomaterials for Extrusion Based 3D Printing: State-of-the-Art. Adv. Healthc. Mater. 2017, 6, 1700264. [Google Scholar] [CrossRef]
- Zhao, Y.; Li, Y.; Mao, S. Effect of bioink properties on printability and cell viability for 3D bioplotting of embryonic stem cells. Biofabrication 2016, 8, 035020. [Google Scholar] [CrossRef]
- Paxton, N.; Smolan, W.; Böck, T.; Melchels, F.; Groll, J.; Jungst, T. Proposal to assess printability of bioinks for extrusion-based bioprinting and evaluation of rheological properties governing bioprintability. Biofabrication 2017, 9, 044107. [Google Scholar] [CrossRef]
- Soltan, N.; Ning, L.; Mohabatpour, F.; Papagerakis, P.; Chen, X. Printability and cell viability in bioprinting alginate dialdehyde-gelatin scaffolds. ACS Biomater. Sci. Eng. 2019, 5, 2976–2987. [Google Scholar] [CrossRef]
- Sarker, M.D.; Naghieh, S.; McInnes, A.D.; Ning, L.; Schreyer, D.J.; Chen, X. Bio-fabrication of peptide-modified alginate scaffolds: Printability, mechanical stability and neurite outgrowth assessments. Bioprinting 2019, 14, e00045. [Google Scholar] [CrossRef]
- Naghieh, S.; Chen, X. Printability–A key issue in extrusion-based bioprinting. J. Pharm. Anal. 2021, 11, 564–579. [Google Scholar] [CrossRef]
- Hospodiuk, M.; Dey, M.; Sosnoski, D.; Ozbolat, I.T. The bioink: A comprehensive review on bioprintable materials. Biotechnol. Adv. 2017, 35, 217–239. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Duan, B.; Kapetanovic, E.; Hockaday, L.A.; Butcher, J.T. 3D Printed Trileaflet Valve Conduits Using Biological Hydrogels and Human Valve Interstitial Cells. Acta Biomater. 2014, 10, 1836–1846. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, Y.; Li, Y.; Mao, S.; Sun, W.; Yao, R. The influence of printing parameters on cell survival rate and printability in microextrusion-based 3D cell printing technology. Biofabrication 2015, 7, 045002. [Google Scholar] [CrossRef] [PubMed]
- Hou, X.; Chen, G.; Xing, T.; Wei, Z. Reactive ink formulated with various alcohols for improved properties and printing quality onto cotton fabrics. J. Eng. Fibers Fabr. 2019, 14, 155892501984924. [Google Scholar] [CrossRef]
- Atala, A.; Yoo, M.; Yoo, J.J.; Atala, A.; James, J.Y. Essentials of 3D Biofabrication and Translation; Academic Press: Cambridge, MA, USA, 2015. [Google Scholar]
- Chimene, D.; Lennox, K.K.; Kaunas, R.R.; Gaharwar, A.K. Advanced Bioinks for 3D Printing: A Materials Science Perspective. Ann. Biomed. Eng. 2016, 44, 2090–2102. [Google Scholar] [CrossRef]
- Chen, X.B.; Ke, H. Effects of fluid properties on dispensing processes for electronics packaging. IEEE Trans. Electron. Packag. Manuf. 2006, 29, 75–82. [Google Scholar] [CrossRef]
- Fu, Z.; Naghieh, S.; Xu, C.; Wang, C.; Sun, W.; Chen, X. Printability in extrusion bioprinting. Biofabrication 2021, 13, 033001. [Google Scholar] [CrossRef]
- Liu, W.; Zhang, Y.S.; Heinrich, M.A.; De Ferrari, F.; Jang, H.L.; Bakht, S.M.; Alvarez, M.M.; Yang, J.; Li, Y.C.; Trujillo-de Santiago, G.; et al. Rapid Continuous Multimaterial Extrusion Bioprinting. Adv. Mater. 2017, 29, 1604630. [Google Scholar] [CrossRef]
- Lee, J.M.; Ng, W.L.; Yeong, W.Y. Resolution and shape in bioprinting: Strategizing towards complex tissue and organ printing. Appl. Phys. Rev. 2019, 6, 011307. [Google Scholar] [CrossRef]
- Chen, X.B.; Li, M.G.; Ke, H. Modeling of the flow rate in the dispensing-based process for fabricating tissue scaffolds. J. Manuf. Sci. Eng. Trans. ASME 2008, 130, 021003. [Google Scholar] [CrossRef]
- Li, M.G.; Tian, X.Y.; Chen, X.B. Modeling of flow rate, pore size, and porosity for the dispensing-based tissue scaffolds fabrication. J. Manuf. Sci. Eng. Trans. ASME 2009, 131, 034501. [Google Scholar] [CrossRef]
- Sarker, M.; Chen, X.B. Modeling the Flow Behavior and Flow Rate of Medium Viscosity Alginate for Scaffold Fabrication with a Three-Dimensional Bioplotter. J. Manuf. Sci. Eng. Trans. ASME 2017, 139, 081002. [Google Scholar] [CrossRef]
- Chang, C.C.; Boland, E.D.; Williams, S.K.; Hoying, J.B. Direct-write bioprinting three-dimensional biohybrid systems for future regenerative therapies. J. Biomed. Mater. Res.-Part B Appl. Biomater. 2011, 98B, 160–170. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Billiet, T.; Gevaert, E.; De Schryver, T.; Cornelissen, M.; Dubruel, P. The 3D printing of gelatin methacrylamide cell-laden tissue-engineered constructs with high cell viability. Biomaterials 2014, 35, 49–62. [Google Scholar] [CrossRef] [PubMed]
- Naghieh, S.; Karamooz-Ravari, M.R.; Sarker, M.D.; Karki, E.; Chen, X. Influence of crosslinking on the mechanical behavior of 3D printed alginate scaffolds: Experimental and numerical approaches. J. Mech. Behav. Biomed. Mater. 2018, 80, 111–118. [Google Scholar] [CrossRef] [Green Version]
- Sarker, M.; Izadifar, M.; Schreyer, D.; Chen, X. Influence of ionic crosslinkers (Ca2+ /Ba2+ /Zn2+) on the mechanical and biological properties of 3D Bioplotted Hydrogel Scaffolds. J. Biomater. Sci. Polym. Ed. 2018, 29, 1126–1154. [Google Scholar] [CrossRef]
- Lee, J.M.; Yeong, W.Y. Design and Printing Strategies in 3D Bioprinting of Cell-Hydrogels: A Review. Adv. Healthc. Mater. 2016, 5, 2856–2865. [Google Scholar] [CrossRef]
- Kesti, M.; Eberhardt, C.; Pagliccia, G.; Kenkel, D.; Grande, D.; Boss, A.; Zenobi-Wong, M. Bioprinting Complex Cartilaginous Structures with Clinically Compliant Biomaterials. Adv. Funct. Mater. 2015, 25, 7406–7417. [Google Scholar] [CrossRef] [Green Version]
- Dang, Q.; Liu, K.; Zhang, Z.; Liu, C.; Liu, X.; Xin, Y.; Cheng, X.; Xu, T.; Cha, D.; Fan, B. Fabrication and evaluation of thermosensitive chitosan/collagen/α, β-glycerophosphate hydrogels for tissue regeneration. Carbohydr. Polym. 2017, 167, 145–157. [Google Scholar] [CrossRef]
- Wüst, S.; Godla, M.E.; Müller, R.; Hofmann, S. Tunable hydrogel composite with two-step processing in combination with innovative hardware upgrade for cell-based three-dimensional bioprinting. Acta Biomater. 2014, 10, 630–640. [Google Scholar] [CrossRef]
- He, Y.; Tuck, C.J.; Prina, E.; Kilsby, S.; Christie, S.D.R.; Edmondson, S.; Hague, R.J.M.; Rose, F.R.A.J.; Wildman, R.D. A new photocrosslinkable polycaprolactone-based ink for three-dimensional inkjet printing. J. Biomed. Mater. Res.-Part B Appl. Biomater. 2017, 105, 1645–1657. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- He, Y.; Yang, F.; Zhao, H.; Gao, Q.; Xia, B.; Fu, J. Research on the printability of hydrogels in 3D bioprinting. Sci. Rep. 2016, 6, 29977. [Google Scholar] [CrossRef] [PubMed]
- Jammalamadaka, U.; Tappa, K. Recent advances in biomaterials for 3D printing and tissue engineering. J. Funct. Biomater. 2018, 9, 22. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kim, Y.; Kang, K.; Jeong, J.; Paik, S.S.; Kim, J.S.; Park, S.A.; Kim, W.D.; Park, J.; Choi, D. Three-dimensional (3D) printing of mouse primary hepatocytes to generate 3D hepatic structure. Ann. Surg. Treat. Res. 2017, 92, 67–72. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Naghieh, S.; Sarker, M.D.; Abelseth, E.; Chen, X. Indirect 3D bioprinting and characterization of alginate scaffolds for potential nerve tissue engineering applications. J. Mech. Behav. Biomed. Mater. 2019, 93, 183–193. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Markstedt, K.; Mantas, A.; Tournier, I.; Ha, D.; Gatenholm, P. 3D Bioprinting Human Chondrocytes with Nanocellulose−Alginate Bioink for Cartilage Tissue Engineering Applications. Biomacromolecules 2015, 16, 1489–1496. [Google Scholar] [CrossRef] [PubMed]
- Fu, Z.; Ouyang, L.; Xu, R.; Yang, Y.; Sun, W. Responsive biomaterials for 3D bioprinting: A review. Mater. Today 2022, 52, 112–132. [Google Scholar] [CrossRef]
- López-Marcial, G.R.; Zeng, A.Y.; Osuna, C.; Dennis, J.; García, J.M.; O’Connell, G.D. Agarose-Based Hydrogels as Suitable Bioprinting Materials for Tissue Engineering. ACS Biomater. Sci. Eng. 2018, 4, 3610–3616. [Google Scholar] [CrossRef]
- Lee, K.Y.; Mooney, D.J. Hydrogels for Tissue Engineering. Chem. Rev. 2001, 101, 1869–1880. [Google Scholar] [CrossRef]
- Gong, C.; Kong, Z.; Wang, X. The effect of agarose on 3d bioprinting. Polymers 2021, 13, 4028. [Google Scholar] [CrossRef]
- Nadernezhad, A.; Caliskan, O.S.; Topuz, F.; Afghah, F.; Erman, B.; Koc, B. Nanocomposite Bioinks Based on Agarose and 2D Nanosilicates with Tunable Flow Properties and Bioactivity for 3D Bioprinting. ACS Appl. Bio. Mater. 2019, 2, 796–806. [Google Scholar] [CrossRef] [PubMed]
- Tako, M.; Nakamura, S. Gelation mechanism of agarose. Carbohydr. Res. 1988, 180, 277–284. [Google Scholar] [CrossRef]
- Laronda, M.M.; Rutz, A.L.; Xiao, S.; Whelan, K.A.; Duncan, F.E.; Roth, E.W.; Woodruff, T.K.; Shah, R.N. A bioprosthetic ovary created using 3D printed microporous scaffolds restores ovarian function in sterilized mice. Nat. Commun. 2017, 8, 15261. [Google Scholar] [CrossRef] [PubMed]
- Du, J.; Dai, H.; Wang, H.; Yu, Y.; Zhu, H.; Fu, Y.; Ma, L.; Peng, L.; Li, L.; Wang, Q.; et al. Preparation of high thermal stability gelatin emulsion and its application in 3D printing. Food Hydrocoll. 2021, 113, 106536. [Google Scholar] [CrossRef]
- Ye, W.; Li, H.; Yu, K.; Xie, C.; Wang, P.; Zheng, Y.; Zhang, P.; Xiu, J.; Yang, Y.; Zhang, F.; et al. 3D printing of gelatin methacrylate-based nerve guidance conduits with multiple channels. Mater. Des. 2020, 192, 108757. [Google Scholar] [CrossRef]
- Dong, L.; Wang, S.J.; Zhao, X.R.; Zhu, Y.F.; Yu, J.K. 3D-printed poly (ϵ-caprolactone) scaffold integrated with cell-laden chitosan hydrogels for bone tissue engineering. Sci. Rep. 2017, 7, 4–12. [Google Scholar] [CrossRef] [Green Version]
- Rajabi, M.; McConnell, M.; Cabral, J.; Ali, M.A. Chitosan hydrogels in 3D printing for biomedical applications. Carbohydr. Polym. 2021, 260, 117768. [Google Scholar] [CrossRef]
- Wu, Q.; Therriault, D.; Heuzey, M.-C. Processing and Properties of Chitosan Inks for 3D Printing of Hydrogel Microstructures. ACS Biomater. Sci. Eng. 2018, 4, 2643–2652. [Google Scholar] [CrossRef]
- Wu, Q.; Maire, M.; Lerouge, S.; Therriault, D.; Heuzey, M.-C. 3D Printing of Microstructured and Stretchable Chitosan Hydrogel for Guided Cell Growth. Adv. Biosyst. 2017, 1, 1700058. [Google Scholar] [CrossRef]
- Lee, V.; Singh, G.; Trasatti, J.P.; Bjornsson, C.; Xu, X.; Tran, T.N.; Yoo, S.S.; Dai, G.; Karande, P. Design and fabrication of human skin by three-dimensional bioprinting. Tissue Eng.-Part C Methods 2014, 20, 473–484. [Google Scholar] [CrossRef] [Green Version]
- Fu, F.; Qin, Z.; Xu, C.; Chen, X.Y.; Li, R.X.; Wang, L.N.; Peng, D.W.; Sun, H.T.; Tu, Y.; Chen, C.; et al. Magnetic resonance imaging-three-dimensional printing technology fabricates customized scaffolds for brain tissue engineering. Neural Regen. Res. 2017, 12, 614–622. [Google Scholar] [CrossRef] [PubMed]
- Rhee, S.; Puetzer, J.L.; Mason, B.N.; Reinhart-King, C.A.; Bonassar, L.J. 3D Bioprinting of Spatially Heterogeneous Collagen Constructs for Cartilage Tissue Engineering. ACS Biomater. Sci. Eng. 2016, 2, 1800–1805. [Google Scholar] [CrossRef] [PubMed]
- Abelseth, E.; Abelseth, L.; De La Vega, L.; Beyer, S.T.; Wadsworth, S.J.; Willerth, S.M. 3D Printing of Neural Tissues Derived from Human Induced Pluripotent Stem Cells Using a Fibrin-Based Bioink. ACS Biomater. Sci. Eng. 2019, 5, 234–243. [Google Scholar] [CrossRef] [PubMed]
- England, S.; Rajaram, A.; Schreyer, D.J.; Chen, X. Bioprinted fibrin-factor XIII-hyaluronate hydrogel scaffolds with encapsulated Schwann cells and their in vitro characterization for use in nerve regeneration. Bioprinting 2017, 5, 1–9. [Google Scholar] [CrossRef]
- Zhao, D.; Huang, Y.; Ao, Y.; Han, C.; Wang, Q.; Li, Y.; Liu, J.; Wei, Q.; Zhang, Z. Effect of pore geometry on the fatigue properties and cell affinity of porous titanium scaffolds fabricated by selective laser melting. J. Mech. Behav. Biomed. Mater. 2018, 88, 478–487. [Google Scholar] [CrossRef]
- Li, H.; Yin, Y.; Xiang, Y.; Liu, H.; Guo, R. A novel 3D printing PCL/GelMA scaffold containing USPIO for MRI-guided bile duct repair. Biomed. Mater. 2020, 15, 45004. [Google Scholar] [CrossRef] [PubMed]
- Ouyang, L.; Highley, C.B.; Rodell, C.B.; Sun, W.; Burdick, J.A. 3D Printing of Shear-Thinning Hyaluronic Acid Hydrogels with Secondary Cross-Linking. ACS Biomater. Sci. Eng. 2016, 2, 1743–1751. [Google Scholar] [CrossRef]
- Highley, C.B.; Rodell, C.B.; Burdick, J.A. Direct 3D Printing of Shear-Thinning Hydrogels into Self-Healing Hydrogels. Adv. Mater. 2015, 27, 5075–5079. [Google Scholar] [CrossRef]
- Petta, D.; D’Amora, U.; Ambrosio, L.; Grijpma, D.W.; Eglin, D.; D’Este, M. Hyaluronic acid as a bioink for extrusion-based 3D printing. Biofabrication 2020, 12, 032001. [Google Scholar] [CrossRef]
- Shie, M.Y.; Chang, W.C.; Wei, L.J.; Huang, Y.H.; Chen, C.H.; Shih, C.T.; Chen, Y.W.; Shen, Y.F. 3D printing of cytocompatible water-based light-cured polyurethane with hyaluronic acid for cartilage tissue engineering applications. Materials 2017, 10, 136. [Google Scholar] [CrossRef]
- Ouyang, L.; Highley, C.B.; Sun, W.; Burdick, J.A. A Generalizable Strategy for the 3D Bioprinting of Hydrogels from Nonviscous Photo-crosslinkable Inks. Adv. Mater. 2017, 29, 1604983. [Google Scholar] [CrossRef]
- Joas, S.; Tovar, G.E.M.; Celik, O.; Bonten, C.; Southan, A. Extrusion-based 3d printing of poly(Ethylene glycol) diacrylate hydrogels containing positively and negatively charged groups. Gels 2018, 4, 69. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bai, J.; Wang, H.; Gao, W.; Liang, F.; Wang, Z.; Zhou, Y.; Lan, X.; Chen, X.; Cai, N.; Huang, W.; et al. Melt electrohydrodynamic 3D printed poly (ε-caprolactone)/polyethylene glycol/roxithromycin scaffold as a potential anti-infective implant in bone repair. Int. J. Pharm. 2020, 576, 118941. [Google Scholar] [CrossRef] [PubMed]
- Khan, F.; Valere, S.; Fuhrmann, S.; Arrighi, V.; Bradley, M. Synthesis and cellular compatibility of multi-block biodegradable poly(ε-caprolactone)-based polyurethanes. J. Mater. Chem. B 2013, 1, 2590–2600. [Google Scholar] [CrossRef] [PubMed]
- Hsiao, S.H.; Hsu, S.H. Synthesis and Characterization of Dual Stimuli-Sensitive Biodegradable Polyurethane Soft Hydrogels for 3D Cell-Laden Bioprinting. ACS Appl. Mater. Interfaces 2018, 10, 29273–29287. [Google Scholar] [CrossRef]
- Hung, K.C.; Tseng, C.S.; Hsu, S.H. Synthesis and 3D Printing of biodegradable polyurethane elastomer by a water-based process for cartilage tissue engineering applications. Adv. Healthc. Mater. 2014, 3, 1578–1587. [Google Scholar] [CrossRef]
- Domínguez-Robles, J.; Mancinelli, C.; Mancuso, E.; García-Romero, I.; Gilmore, B.F.; Casettari, L.; Larrañeta, E.; Lamprou, D.A. 3D printing of drug-loaded thermoplastic polyurethane meshes: A potential material for soft tissue reinforcement in vaginal surgery. Pharmaceutics 2020, 12, 63. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Murphy, S.V.; Skardal, A.; Atala, A. Evaluation of hydrogels for bio-printing applications. J. Biomed. Mater. Res.-Part A 2013, 101A, 272–284. [Google Scholar] [CrossRef]
- Rajaram, A.; Schreyer, D.J.; Chen, D.X.B. Use of the polycation polyethyleneimine to improve the physical properties of alginate-hyaluronic acid hydrogel during fabrication of tissue repair scaffolds. J. Biomater. Sci. Polym. Ed. 2015, 26, 433–445. [Google Scholar] [CrossRef]
- Panwar, A.; Tan, L.P. Current Status of Bioinks for Micro-Extrusion-Based. Molecules 2016, 21, 685. [Google Scholar] [CrossRef]
- Chung, J.H.Y.; Naficy, S.; Yue, Z.; Kapsa, R.; Quigley, A.; Moulton, S.E.; Wallace, G.G. Bio-ink properties and printability for extrusion printing living cells. Biomater. Sci. 2013, 1, 763–773. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mouser, V.H.M.; Melchels, F.P.W.; Visser, J.; Dhert, W.J.A.; Gawlitta, D.; Malda, J. Yield stress determines bioprintability of hydrogels based on gelatin-methacryloyl and gellan gum for cartilage bioprinting. Biofabrication 2016, 8, 35003. [Google Scholar] [CrossRef] [PubMed]
- Bertassoni, L.E.; Cardoso, J.C.; Manoharan, V.; Cristino, A.L.; Bhise, N.S.; Araujo, W.A.; Zorlutuna, P.; Vrana, N.E.; Ghaemmaghami, A.M.; Dokmeci, M.R.; et al. Direct-write bioprinting of cell-laden methacrylated gelatin hydrogels. Biofabrication 2014, 6, 024105. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, W.; Heinrich, M.A.; Zhou, Y.; Akpek, A.; Hu, N.; Liu, X.; Guan, X.; Zhong, Z.; Jin, X.; Khademhosseini, A.; et al. Extrusion Bioprinting of Shear-Thinning Gelatin Methacryloyl Bioinks. Adv. Healthc. Mater. 2017, 6, 1601451. [Google Scholar] [CrossRef]
- Izadifar, Z.; Chang, T.; Kulyk, W.; Chen, X.; Eames, B.F. Analyzing biological performance of 3D-printed, cell-impregnated hybrid constructs for cartilage tissue engineering. Tissue Eng.-Part C Methods 2016, 22, 173–188. [Google Scholar] [CrossRef] [Green Version]
- Olubamiji, A.D.; Zhu, N.; Chang, T.; Nwankwo, C.K.; Izadifar, Z.; Honaramooz, A.; Chen, X.; Eames, B.F. Traditional Invasive and Synchrotron-Based Noninvasive Assessments of Three-Dimensional-Printed Hybrid Cartilage Constructs in Situ. Tissue Eng.-Part C Methods 2017, 23, 156–168. [Google Scholar] [CrossRef]
- Kundu, J.; Shim, J.-H.; Jang, J.; Kim, S.-W.; Cho, D.-W. An additive manufacturing-based PCL–alginate–chondrocyte bioprinted scaffold for cartilage tissue engineering. J. Tissue Eng. Regen. Med. 2015, 9, 1286–1297. [Google Scholar] [CrossRef]
- Pati, F.; Shim, J.-H.; Lee, J.-S.; Cho, D.-W. 3D printing of cell-laden constructs for heterogeneous tissue regeneration. Manuf. Lett. 2013, 1, 49–53. [Google Scholar] [CrossRef]
- Lee, J.-S.; Hong, J.M.; Jung, J.W.; Shim, J.-H.; Oh, J.-H.; Cho, D.-W. 3D printing of composite tissue with complex shape applied to ear regeneration. Biofabrication 2014, 6, 024103. [Google Scholar] [CrossRef]
- Hinton, T.J.; Hudson, A.; Pusch, K.; Lee, A.; Feinberg, A.W. 3D Printing PDMS Elastomer in a Hydrophilic Support Bath via Freeform Reversible Embedding. 3D Printers 2016, 2, 1781–1786. [Google Scholar] [CrossRef]
- Ding, H.; Chang, R.C. Printability Study of Bioprinted Tubular Structures Using Liquid Hydrogel Precursors in a Support Bath. Appl. Sci. 2018, 8, 403. [Google Scholar] [CrossRef] [Green Version]
- Sodupe-Ortega, E.; Sanz-Garcia, A.; Pernia-Espinoza, A.; Escobedo-Lucea, C. Accurate Calibration in Multi-Material 3D Bioprinting for Tissue Engineering. Materials 2018, 11, 1402. [Google Scholar] [CrossRef] [Green Version]
- Wong, K.H.K.; Chan, J.M.; Kamm, R.D.; Tien, J. Microfluidic Models of Vascular Functions. Annu. Rev. Biomed. Eng. 2012, 14, 205–230. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Potter, C.M.F.; Lao, K.H.; Zeng, L.; Xu, Q. Role of Biomechanical Forces in Stem Cell Vascular Lineage Differentiation. Arterioscler. Thromb. Vasc. Biol. 2014, 34, 2184–2190. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Boularaoui, S.; Al Hussein, G.; Khan, K.A.; Christoforou, N.; Stefanini, C. An overview of extrusion-based bioprinting with a focus on induced shear stress and its effect on cell viability. Bioprinting 2020, 20, e00093. [Google Scholar] [CrossRef]
- Rajaram, A.; Schreyer, D.; Chen, D. Bioplotting Alginate/Hyaluronic Acid Hydrogel Scaffolds with Structural Integrity and Preserved Schwann Cell Viability. 3D Print. Addit. Manuf. 2014, 1, 194–203. [Google Scholar] [CrossRef]
- Ning, L.; Xu, Y.; Chen, X.; Schreyer, D.J. Influence of mechanical properties of alginate-based substrates on the performance of Schwann cells in culture. J. Biomater. Sci. Polym. Ed. 2016, 27, 898–915. [Google Scholar] [CrossRef]
- Ning, L.; Guillemot, A.; Zhao, J.; Kipouros, G.; Chen, X. Influence of Flow Behavior of Alginate-Cell Suspensions on Cell Viability and Proliferation. Tissue Eng. Part C Methods 2016, 22, 652–662. [Google Scholar] [CrossRef]
- Li, M.; Tian, X.; Kozinski, J.A.; Chen, X.; Hwang, D.K. Modeling Mechanical Cell Damage in the Bioprinting Process Employing a Conical Needle. J. Mech. Med. Biol. 2015, 15, 1550073. [Google Scholar] [CrossRef]
- Ning, L.; Yang, B.; Mohabatpour, F.; Betancourt, N.; Sarker, M.D.; Papagerakis, P.; Chen, X. Process-induced cell damage: Pneumatic versus screw-driven bioprinting. Biofabrication 2020, 12, 025011. [Google Scholar] [CrossRef]
- Blaeser, A.; Filipa, D.; Campos, D.; Puster, U.; Richtering, W.; Stevens, M.M. Controlling Shear Stress in 3D Bioprinting is a Key Factor to Balance Printing Resolution and Stem Cell Integrity. Adv. Healthc. Mater. 2016, 5, 326–333. [Google Scholar] [CrossRef] [PubMed]
- Hull, S.M.; Brunel, L.G.; Heilshorn, S.C. 3D Bioprinting of Cell-Laden Hydrogels for Improved Biological Functionality. Adv. Mater. 2022, 34, 2103691. [Google Scholar] [CrossRef] [PubMed]
- Nair, K.; Gandhi, M.; Khalil, S.; Yan, K.C.; Marcolongo, M.; Barbee, K.; Sun, W. Characterization of cell viability during bioprinting processes. Biotechnol. J. 2009, 4, 1168–1177. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Tian, X.; Schreyer, D.J.; Chen, X. Effect of needle geometry on flow rate and cell damage in the dispensing-based biofabrication process. Biotechnol. Prog. 2011, 27, 1777–1784. [Google Scholar] [CrossRef]
- Lucas, L.; Aravind, A.; Emma, P.; Christophe, M.; Edwin-Joffrey, C. Rheology, simulation and data analysis toward bioprinting cell viability awareness. Bioprinting 2021, 21, e00119. [Google Scholar] [CrossRef]
- Kingsley, D.M.; Ouyang, L.; Yao, R.; Mao, S.; Chen, X.; Na, J.; Sun, W. Three-dimensional bioprinting of embryonic stem cells directs highly uniform embryoid body formation Three-dimensional bioprinting of embryonic stem cells directs highly uniform embryoid body formation. Biofabrication 2015, 7, 044101. [Google Scholar]
- Murphy, S.V.; Atala, A. 3D bioprinting of tissues and organs. Nat. Biotechnol. 2014, 32, 773–785. [Google Scholar] [CrossRef]
- Baena, J.M.; Jiménez, G.; López-Ruiz, E.; Antich, C.; Griñán-Lisón, C.; Perán, M.; Gálvez-Martín, P.; Marchal, J.A. Volume-by-volume bioprinting of chondrocytes-alginate bioinks in high temperature thermoplastic scaffolds for cartilage regeneration. Exp. Biol. Med. 2019, 244, 13–21. [Google Scholar] [CrossRef]
- Mohd Bohari, S.P.; Hukins, D.; Grover, L. Effect of calcium alginate concentration on viability and proliferation of encapsulated fibroblasts. Biomed. Mater. Eng. 2011, 21, 159–170. [Google Scholar] [CrossRef]
- Li, M.; Tian, X.; Zhu, N.; Schreyer, D.J.; Chen, X. Modeling process-induced cell damage in the biodispensing process. Tissue Eng. Part C Methods 2010, 16, 533–542. [Google Scholar] [CrossRef]
- Ning, L.; Betancourt, N.; Schreyer, D.J.; Chen, X. Characterization of Cell Damage and Proliferative Ability during and after Bioprinting. ACS Biomater. Sci. Eng. 2018, 4, 3906–3918. [Google Scholar] [CrossRef]
- Göhl, J.; Markstedt, K.; Mark, A.; Håkansson, K.; Gatenholm, P.; Edelvik, F. Simulations of 3D bioprinting: Predicting bioprintability of nanofibrillar inks. Biofabrication 2018, 10, 34105. [Google Scholar] [CrossRef] [PubMed]
- Stiehm, M.; Wüstenhagen, C.; Siewert, S.; Grabow, N.; Schmitz, K.-P. Numerical simulation of pulsatile flow through a coronary nozzle model based on FDA’s benchmark geometry. Curr. Dir. Biomed. Eng. 2017, 3, 775–778. [Google Scholar] [CrossRef] [Green Version]
- Brindha, J.; Privita Edwina, R.A.G.; Rajesh, P.K.; Rani, P. Influence of rheological properties of protein bio-inks on printability: A simulation and validation study. Mater. Today Proc. 2016, 3, 3285–3295. [Google Scholar] [CrossRef]
- Wu, H.-C.; Lin, H.-J. Effects of Actuating Pressure Waveforms on the Droplet Behavior in a Piezoelectric Inkjet. Mater. Trans. 2010, 51, 2269–2276. [Google Scholar] [CrossRef] [Green Version]
- Leppiniemi, J.; Lahtinen, P.; Paajanen, A.; Mahlberg, R.; Metsä-Kortelainen, S.; Pinomaa, T.; Pajari, H.; Vikholm-Lundin, I.; Pursula, P.; Hytönen, V.P. 3D-Printable Bioactivated Nanocellulose-Alginate Hydrogels. ACS Appl. Mater. Interfaces 2017, 9, 21959–21970. [Google Scholar] [CrossRef] [Green Version]
- Brackbill, J.U.; Kothe, D.B.; Zemach, C. A continuum Method for Modeling Surface Tension. Comput. Phys. 1992, 100, 335–354. [Google Scholar] [CrossRef]
- Furlani, E.P. Fluid Mechanics for Inkjet Printing. In Fundamentals of Inkjet Printing; Wiley-VCH Verlag GmbH & Co. KGaA: Weinheim, Germany, 2015; pp. 13–56. [Google Scholar]
- Caboussat, A. Numerical Simulation of Two-Phase Free Surface Flows. Arch. Comput. Methods Eng. 2005, 12, 165–224. [Google Scholar] [CrossRef]
- Yeoh, G.H.; Tu, J. Chapter 3-Solution Methods for Multi-Phase Flows. In Computational Techniques for Multiphase Flows; Yeoh, G.H., Tu, J., Eds.; Butterworth-Heinemann: Oxford, UK, 2010; pp. 95–242. ISBN 978-0-08-046733-7. [Google Scholar]
- Chan, K.S.; Pericleous, K.; Cross, M. Numerical simulation of flows encountered during mold-filling. Top. Catal. 1991, 15, 624–631. [Google Scholar] [CrossRef]
- Kothe, D.B. Volume Tracking of Interfaces Having Surface The University of Memphis; American Institute of Aeronautics and Astronautics: Reston, VA, USA, 1999; p. 859. [Google Scholar] [CrossRef] [Green Version]
- Noh, W.F.; Woodward, P. SLIC (Simple Line Interface Calculation); Springer: Berlin/Heidelberg, Germany, 1976. [Google Scholar]
- Youngs, D. Time-Dependent Multi-material Flow with Large Fluid Distortion. Numer. Methods Fluid Dyn. 1982, 273–285. Available online: https://ci.nii.ac.jp/naid/10029507464/ (accessed on 10 February 2022).
- Lee, M.S.; Aute, V.; Riaz, A.; Radermacher, R. A Review on Direct Two-Phase, Phase Change Flow Simulation Methods and their Applications. In Proceedings of the International Refrigeration and Air Conditioning Conference, Lafayette, LA, USA, 16 July 2012; pp. 1–9. [Google Scholar]
- Shepel, S.V.; Smith, B.L. On surface tension modelling using the level set method. Numer. Methods Fluids 2009, 59, 147–171. [Google Scholar] [CrossRef]
- Hutchings, I.M.; Martin, G.D.; Hoath, S.D. Introductory Remarks. In Fundamentals of Inkjet Printing; Wiley-VCH Verlag GmbH & Co. KGaA: Weinheim, Germany, 2015; pp. 1–12. ISBN 9783527684724. [Google Scholar]
- Landau, L.D.; Lifshitz, E.M. Chapter VII-Surface Phenomena. In Fluid Mechanics (Second Edition); Landau, L.D., Lifshitz, E.M., Eds.; Pergamon: Oxford, UK, 1987; pp. 238–250. ISBN 978-0-08-033933-7. [Google Scholar]
- Shirani, E.; Ashgriz, N.; Mostaghimi, J. Interface pressure calculation based on conservation of momentum for front capturing methods. J. Comput. Phys. 2005, 203, 154–175. [Google Scholar] [CrossRef]
- Lafaurie, B.; Nardoni, C.; Scardovelei, R.; Zaleski, S.; Zanetti, G. Modeling Merging and Fragmentation in Multiphase flows with SURFER. Comput. Phys. 1994, 113, 147. [Google Scholar] [CrossRef]
- Lepowsky, E.; Muradoglu, M.; Tasoglu, S. Bioprinting Towards preserving post-printing cell viability and improving the resolution: Past, present, and future of 3D bioprinting theory. Bioprinting 2018, 11, e00034. [Google Scholar] [CrossRef]
- Hendriks, J.; Visser, C.W.; Henke, S.; Leijten, J.; Saris, D.B.F. Optimizing cell viability in droplet- based cell deposition. Nat. Publ. Gr. 2015, 5, 11304. [Google Scholar] [CrossRef]
- Lee, S.S.; Yim, Y.; Ahn, K.H.; Lee, S.J. Extensional flow-based assessment of red blood cell deformability using hyperbolic converging microchannel. Biomed. Microdevices 2009, 11, 1021–1027. [Google Scholar] [CrossRef]
- Tanzeglock, T.; Soos, M.; Stephanopoulos, G.; Morbidelli, M. Induction of Mammalian Cell Death by Simple Shear and Extensional Flows. Biotechnol. Bioeng. 2009, 104, 360–370. [Google Scholar] [CrossRef]
- Aguado, B.A.; Mulyasasmita, W.; Su, J.; Lampe, K.J.; Heilshorn, S.C. Improving Viability of Stem Cells During Syringe Needle Flow Through the Design of Hydrogel Cell Carriers. Tissue Eng. Part A 2012, 18, 806–815. [Google Scholar] [CrossRef] [Green Version]
- Reina-Romo, E.; Mandal, S.; Amorim, P.; Bloemen, V.; Ferraris, E.; Geris, L. Towards the Experimentally-Informed In Silico Nozzle Design Optimization for Extrusion-Based Bioprinting of Shear-Thinning Hydrogels. Front. Bioeng. Biotechnol. 2021, 9, 701778. [Google Scholar] [CrossRef]
- Yu, C.; Jiang, J. A perspective on using machine learning in 3D bioprinting. Int. J. Bioprint. 2020, 6, 4–11. [Google Scholar] [CrossRef]
- An, J.; Chua, C.K.; Mironov, V. Application of Machine Learning in 3D Bioprinting: Focus on Development of Big Data and Digital Twin. Int. J. Bioprint. 2021, 7, 342. [Google Scholar] [CrossRef]
- Ng, W.L.; Chan, A.; Ong, Y.S.; Chua, C.K. Deep learning for fabrication and maturation of 3D bioprinted tissues and organs. Virtual Phys. Prototyp. 2020, 15, 340–358. [Google Scholar] [CrossRef]
- Costa, J.B.; Silva-Correia, J.; Reis, R.L.; Oliveira, J.M. Deep learning in bioengineering and biofabrication: A powerful technology boosting translation from research to clinics. J. 3D Print. Med. 2021, 5, 191–211. [Google Scholar] [CrossRef]
- Ruberu, K.; Senadeera, M.; Rana, S.; Gupta, S.; Chung, J.; Yue, Z.; Venkatesh, S.; Wallace, G. Coupling machine learning with 3D bioprinting to fast track optimisation of extrusion printing. Appl. Mater. Today 2021, 22, 100914. [Google Scholar] [CrossRef]
- Jin, Z.; Zhang, Z.; Shao, X.; Gu, G.X. Monitoring Anomalies in 3D Bioprinting with Deep Neural Networks. ACS Biomater. Sci. Eng. 2021. [Google Scholar] [CrossRef] [PubMed]
- Shi, J.; Song, J.; Song, B.; Lu, W.F. Multi-Objective Optimization Design through Machine Learning for Drop-on-Demand Bioprinting. Engineering 2019, 5, 586–593. [Google Scholar] [CrossRef]
- Shi, J.; Wu, B.; Song, B.; Song, J.; Li, S.; Trau, D.; Lu, W.F. Learning-Based Cell Injection Control for Precise Drop-on-Demand Cell Printing. Ann. Biomed. Eng. 2018, 46, 1267–1279. [Google Scholar] [CrossRef] [PubMed]
- Menon, A.; Póczos, B.; Feinberg, A.W.; Washburn, N.R. Optimization of Silicone 3D Printing with Hierarchical Machine Learning. 3D Print. Addit. Manuf. 2019, 6, 181–189. [Google Scholar] [CrossRef]
- Conev, A.; Litsa, E.E.; Perez, M.R.; Diba, M.; Mikos, A.G.; Kavraki, L.E. Machine learning-guided three-dimensional printing of tissue engineering scaffolds. Tissue Eng.-Part A 2020, 26, 1359–1368. [Google Scholar] [CrossRef]
- Lee, J.; Oh, S.J.; An, S.H.; Kim, W.D.; Kim, S.H.; Kim, S.H. Machine learning-based design strategy for 3D printable bioink: Elastic modulus and yield stress determine printability. Biofabrication 2020, 12, 035018. [Google Scholar] [CrossRef]
- Tian, S.; Stevens, R.; McInnes, B.T.; Lewinski, N.A. Machine assisted experimentation of extrusion-based bioprinting systems. Micromachines 2021, 12, 780. [Google Scholar] [CrossRef]
- Specht, D.F. A general regression neural network. IEEE Trans. Neural Netw. 1991, 2, 568–576. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Imandoust, S.B.; Bolandraftar, M. Application of K-Nearest Neighbor (KNN) Approach for Predicting Economic Events: Theoretical Background. Int. J. Eng. Res. Appl. 2013, 3, 605–610. [Google Scholar]
- Hastie, T.; Tibshirani, R.; James, G.; Witten, D. An Introduction to Statistical Learning, Springer Texts; Springer: Berlin/Heidelberg, Germany, 2006; Volume 102, ISBN 9780387781884. [Google Scholar]
- Breiman, L. Random Forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef] [Green Version]
- Xu, H.; Liu, Q.; Casillas, J.; Mcanally, M.; Mubtasim, N.; Gollahon, L.S.; Wu, D.; Xu, C. Prediction of cell viability in dynamic optical projection stereolithography-based bioprinting using machine learning. J. Intell. Manuf. 2020, 33, 995–1005. [Google Scholar] [CrossRef]



| Biomaterials | Advantages | Limits | Target Tissue/Application | Ref. |
|---|---|---|---|---|
| Alginate | • Water-soluble • High water-absorbing • Highly biocompatible • Rapid cross-linking | • Low viscosity • Lack of molecules adhesion | • Liver • Nerve • cartilage | [4,37,45,46,47,48] |
| Agarose | • Water-soluble • Responsive to temperature • Rapid gelation | • Poor cell attachment | • Cartilage | [46,49,50,51,52,53,54] |
| Gelatin | • Highly bioactive • Highly biocompatible • Responsive to temperature | • Poor mechanical properties | • Ovary • Nerve | [4,45,55,56,57] |
| Chitosan | • Antibacterial | • Slow gelation • Poor water solubility | • Bone • Cartilage • Drug delivery • Wound dressing | [58,59,60,61] |
| Collagen | • High cell attachment • Responsive to PH and temperature | • Slow gelation • Poor mechanical properties | • Skin • Nerve • Cartilage | [49,62,63,64] |
| Fibrin | • High cell adhesion • Highly bioactive | • Rapid degradation • Poor mechanical stability | • Nerve | [65,66] |
| Polycaprolactone (PCL) | • Low cost • Biodegradable polyester • Excellent rheological and viscoelastic properties upon heating | • Just for hard tissue • Extended degradation | • Bone • Drug delivery | [67,68] |
| Hyaluronic acid (HA) | • Water-soluble • Highly biocompatible • Good shear thinning properties | • Rapid degradation rate • Poor mechanical properties • Required modification for stable cross-linking | • Wound healing • Bone • Cartilage • Hearth • Nerve | [49,69,70,71,72] |
| Polyethylene glycol (PEG) | • Water-soluble High capacity for chemical modification | • Poor biodegradability • Poor cell attachment | • Would dressing • Bone | [73,74,75] |
| Polyurethane | • Highly biocompatible • Tunable dol–gel transition temperature | • Slow biodegradability • Poor cell attachment | • Cartilage • Drug delivery | [76,77,78,79] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Malekpour, A.; Chen, X. Printability and Cell Viability in Extrusion-Based Bioprinting from Experimental, Computational, and Machine Learning Views. J. Funct. Biomater. 2022, 13, 40. https://doi.org/10.3390/jfb13020040
Malekpour A, Chen X. Printability and Cell Viability in Extrusion-Based Bioprinting from Experimental, Computational, and Machine Learning Views. Journal of Functional Biomaterials. 2022; 13(2):40. https://doi.org/10.3390/jfb13020040
Chicago/Turabian StyleMalekpour, Ali, and Xiongbiao Chen. 2022. "Printability and Cell Viability in Extrusion-Based Bioprinting from Experimental, Computational, and Machine Learning Views" Journal of Functional Biomaterials 13, no. 2: 40. https://doi.org/10.3390/jfb13020040
APA StyleMalekpour, A., & Chen, X. (2022). Printability and Cell Viability in Extrusion-Based Bioprinting from Experimental, Computational, and Machine Learning Views. Journal of Functional Biomaterials, 13(2), 40. https://doi.org/10.3390/jfb13020040






