Abstract
The growing adoption of deep learning (DL) in early-stage cancer diagnosis has demonstrated remarkable performance across multiple imaging tasks. Yet, the lack of transparency in these models (“black-box” problem) limits their adoption in clinical environments. This study proposes a methodological framework for developing interpretable DL models to support the early histopathological diagnosis of lung cancer, with a focus on adenocarcinoma and squamous cell carcinoma. The approach leverages publicly available datasets (TCGA-LUAD, TCGA-LUSC, LC25000) and employs high-performing architectures such as EfficientNet, along with post hoc explainability techniques including Grad-CAM and SHAP. Data will be pre-processed and sampled using stratified purposeful strategies to ensure diversity and balance across subtypes and stages. Model evaluation will combine standard performance metrics with clinician feedback and the spatial alignment of visual explanations with ground-truth annotations. While implementation remains a future step, this paper proposes a methodological framework designed to guide the development of DL systems that are not only accurate but also interpretable and clinically meaningful.
1. Introduction
1.1. Contextualization
Lung cancer is the leading cause of cancer-related mortality worldwide, accounting for an incidence of approximately 2.21 million new cases in 2020. Early detection of lung cancer is crucial for improving survival rates, as earlier diagnosis allows for timely and potentially life-saving interventions [1,2]. Histopathological analysis remains the gold standard in diagnostic pathology. However, this method is dependent on pathologists’ expertise, can be time-consuming, and may result in diagnostic errors, particularly in the early stages of lung cancer, where morphological changes may be subtle [1,3].
The increasing availability of high-resolution digital scans of tissue samples, also known as whole slide images (WSIs), and the advancements in artificial intelligence (AI) allow for computational techniques to assist in cancer diagnosis. As already demonstrated in cancers like breast and prostate, deep learning (DL) models have shown impressive capabilities in identifying complex spatial patterns within tissues that may be difficult for even experienced pathologists to detect [1,4,5]. Nonetheless, one of the main barriers to the adoption of AI-driven systems in lung cancer diagnosis is explainability. The lack of clear understanding regarding how these models reach their conclusions makes clinicians hesitant to trust them fully in decision-making [6].
Recent studies have explored complementary approaches to overcome the lack of explainability of deep learning models applied to medical imaging, where the most common have been saliency-based methods like Gradient-Weighted Class Activation Mapping (Grad-CAM). These methods highlight areas in WSIs that most strongly influence a model’s prediction and are easily compatible with CNNs [7,8]. Latha et al. (2024) and Tian et al. (2024) are examples of studies that employed the Grad-CAM approach to visually assess whether the highlighted regions corresponded to cancerous tissue [9,10]. However, the efficiency of the heatmaps is limited due to low spatial resolution, susceptibility to noise, and the potential for misalignment with clinically relevant features, particularly in early-stage tumor tissue [6,8].
1.2. Aim
Following the explainability barrier, this paper aims to search for an answer to the question “Can explainable DL models enhance early-stage lung cancer diagnosis from histopathological images in a clinically meaningful and understandable manner?”.
To address this question, a methodological framework will be proposed to guide the development, training, and evaluation of explainable deep learning models in histopathological lung cancer diagnosis.
1.3. Research Paradigm
Due to the nature of the research question, this study will adopt the post-positivism paradigm. This paradigm is widely applied in empirical and computational health sciences, including biomedical AI. According to the premise of post-positivism, an objective reality exists, but the understanding of it is always partial and mediated by theory, bias, and measurement tool limitations [11]. Therefore, this paradigm can be applied to research about AI-driven histopathological diagnosis, as it supports systematic and quantitative modeling, which is essential for developing and validating DL models using datasets composed of WSIs. It also acknowledges uncertainty, reflecting the probabilistic and often ambiguous nature of early-stage cancer diagnosis and necessitating interpretable outputs that can be critically assessed rather than blindly trusted [11,12]. Other paradigms like constructivism, the transformative paradigm, and pragmatism were evaluated, but none fully aligned with the current aim of producing objective, generalizable evidence on diagnostic accuracy [11,13,14,15].
Ontologically, while a reality exists (e.g., the true presence or absence of lung cancer in tissue), the access to it is constrained by observational limitations and the used technological tools [16]. On the other side, DL models will not reveal the absolute truth but rather infer patterns based on statistical associations learnt from the training data and provide probabilities for the possible outcomes [1,17].
Regarding epistemology, knowledge about diagnostic outcomes comes from observable and measurable patterns in real-world data, such as WSIs. The outputs of DL models are interpreted as high-confidence predictions derived from learnt patterns in training data [18]. Performance metrics such as Sensitivity, Specificity, and Area Under the Curve (AUC) provide quantitative evidence of model validity [1]. Furthermore, given that model predictions can be influenced by bias and uncertainty, an additional step of critical evaluation should be integrated, involving expert clinicians to review the outputs of the explainability techniques [19]. This approach allows for an additional layer of judgment that shows why the models behave the way they do and ensures that the knowledge produced is reliable, transparent, and aligned with clinical expectations.
1.4. Research Design
In alignment with the post-positivism paradigm, a quantitative research approach will be employed, combining computational experiments and a cross-sectional clinician survey to balance empirical measurement with clinicians’ evaluations [11,20,21].
Initially, the research will measure the models’ performance via statistical performance metrics, such as Sensitivity, Specificity, and F1-score. Using this approach ensures that outcomes are based on observable and reproducible patterns and supports the idea of generating empirical knowledge through experimentation [11].
Afterwards, expert clinicians will review and provide feedback on the outputs produced by explainability techniques, such as Grad-CAM, in order to assess whether the DL models’ decision-making processes align with clinical reasoning and expectations [19]. Thus, this approach guarantees that the models are not only statistically accurate but also clinically meaningful and trustworthy.
As research methodologies, the research applies the experimental methodology for model development and evaluation, and a survey methodology with a cross-sectional design to gather clinician insights. The experimental methodology involves conducting computational experiments in which DL models are trained, validated, and evaluated under varying configurations, such as different training parameters and model architectures, to measure their performance [22]. Regarding the cross-sectional survey, it will collect insights into how clinicians evaluate the relevance and clinical usefulness of the model explanations [21].
In summary, employing the quantitative research design, along with experimental and survey methodologies in this study, allows both measurable outcomes and clinicians’ insights to be integrated into the model evaluation process [11,21].
1.5. Dataset Requirements
The dataset is one of the most important elements for this study, as it will be used to train and assess the performance and explainability of DL models designed for lung cancer histopathological diagnosis. Hence, it must meet some requirements in terms of size and nature to generate clinically acceptable results.
In terms of size, large datasets are essential for training DL models effectively, reducing the risk of overfitting or underfitting, and enabling the model to learn meaningful patterns [23,24]. Since the focus of this study is on evaluating the performance of predictive DL models, rather than testing a formal hypothesis, and these models and training data are complex, sample size calculations or power analyses are not directly applicable. Instead, dataset adequacy is justified based on established practices in the field and the practical needs of training and evaluating DL models [25,26]. Based on similar studies in computational pathology, a dataset comprising several thousand WSIs or image patches is considered a reasonable target to ensure statistical robustness and reproducibility [1].
Regarding its nature, the dataset must consist of high-resolution WSIs of lung tissue stained with haematoxylin and eosin (H&E), including both malignant and benign samples. Among malignant tissues, the most common lung cancer subtypes, such as adenocarcinoma (ADC) and squamous cell carcinoma (SCC), should be represented in this initial phase of the research. Ideally, it should include the early-stage cancer samples, where histopathological features may be more subtle and difficult to detect, and reflect a demographic diversity to improve model generalizability and fairness. In addition, having WSIs already annotated according to the expected diagnostic outcome and including information related to tumor staging information and image metadata will be a plus.
The training and evaluation of the DL models will be conducted using already well-known and publicly available datasets. A stratified purposeful sampling approach will be employed to ensure that the final dataset reflects the full spectrum of histopathological subtypes, stages, and tissue morphologies in both cancerous and non-cancerous lung tissues while maintaining targeted control over sample inclusion [27]. In cases where specific strata are underrepresented, data augmentation techniques, such as patch-level resampling, color normalization, and geometric transformations, may be cautiously used to improve the class balance without compromising diagnostic validity [1,28].
In summary, the dataset requirements and sampling methods address the core challenges of histopathological DL, namely, class imbalance and tissue variability. Including early-stage cases improves its sensitivity for subtle features, while subtype diversity enhances generalizability across heterogeneous tumor types. Stratified purposeful sampling allows for the targeted inclusion of critical cases, while data augmentation supports statistical balance and robustness.
1.6. Document Structure
This paper is organized in four more sections. Section 2 presents the proposed framework, detailing the steps of dataset acquisition and how it will be prepared, the selection of DL models and their training configurations, as well as the application of the explainability techniques and evaluation of both the DL models and explainable methods employed. In Section 3, the ethical implications of using secondary clinical data, DL model transparency, potential biases, and data privacy concerns are discussed. Section 4 demonstrates the data treatment strategy across the stages of data preparation, first-order coding and evaluation, and second-order coding and modeling to ensure methodological robustness. Finally, Section 5 summarizes the main contributions of the proposed framework and describes the future steps.
2. Materials and Methods
The development of an explainable DL model for lung cancer histopathology begins with acquiring WSIs or patches that include both malignant and benign tissue (Figure 1a). Annotated regions are extracted and integrated with patch-level datasets, after which the data are balanced for population diversity and split into training and evaluation datasets (Figure 1b). The model is then trained and tested (Figure 1c), followed by the application of explainability techniques to enhance output transparency (Figure 1d). Finally, performance and explainability are assessed using predefined evaluation metrics (Figure 1e).
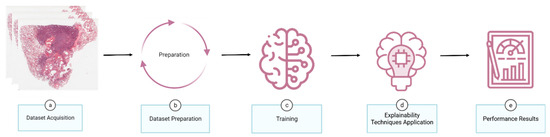
Figure 1.
Conceptual overview of a methodological framework for developing explainable deep learning models in histopathological diagnosis.
2.1. Dataset Acquisition
The model will be tackling the lung cancer subtypes with the most presence, namely, ADC and SCC. As a result, this study will employ the public and well-known datasets The Cancer Genome Atlas (TCGA) [29], particularly the two sub-cohorts TCGA-LUAD (lung ADC) [30] and TCGA-LUSC (lung SCC) [31], and the LC25000 (Lung and Colon Cancer Histopathological Image Dataset) [32].
TCGA provides a collection of WSIs across multiple cancer types, including the two sub-cohorts TCGA-LUAD and TCGA-LUSC. These WSIs are scanned at diagnostic resolution (typically 40× magnification) and accompanied by pathological annotations and clinical metadata, including tumor staging and molecular subtyping [29].
The TCGA-LUAD cohort comprises 585 cases and approximately 1146 diagnostics, with a median patient age of 66 years and a balanced gender distribution (52% female, 48% male). The racial composition is predominantly White (>85%), with limited representation of African American, Asian, and other ethnic groups. Most cases are early-stage tumors (Stage I, ~59%), and 91% report a history of tobacco use [33,34].
Similarly, the TCGA-LUSC cohort includes 504 cases and around 1081 WSIs. This cohort exhibits a male predominance (~71%) and a slightly higher median age of 68 years. White patients represent 86% of the dataset, and smoking history is observed in more than 95% of participants. While early-stage tumors (Stages I and II) are most common, stages III and IV cases are also well represented [31,34,35].
Additionally, the LC25000 dataset is a collection of 25,000 WSI patches evenly divided across five classes, including 15,000 lung-related samples (ADC, SCC, and normal). The patches are generated from 250 H&E-stained images and augmented using generative adversarial networks to enhance morphological diversity and ensure class balance. Though LC25000 lacks associated clinical metadata or patient demographics, its accessibility, class balance, and patch-level annotation make it a widely adopted benchmark for research related to computational pathology [32].
The selection of these datasets is based on their open-access status, adherence to the Health Insurance Portability and Accountability Act (HIPAA) standards for de-identification [36], broad adoption in the computational pathology literature, and thorough documentation, all of which foster transparency and reproducibility [30,31,32].
2.2. Dataset Preparation
Having collected lung WSIs and patches, the dataset preparation process begins by gathering the WSIs without diagnostic annotations and screening them for artefacts and technical inconsistencies. Then, they are submitted to pathologists for manual annotation using digital pathology software. The pathologists will annotate regions of interest (ROIs) according to the lung cancer subtypes (ADC, SCC, benign) and stages (I–IV).
Subsequent to annotation, patches from the annotated ROIs are extracted. Using the sliding window strategy, patches are cropped with a dimension of 512 × 512 pixels, and background-only tiles are filtered out. After that, a standardized pre-processing pipeline is applied, which includes stain normalization to minimize variability due to scanner hardware or laboratory staining protocols. Patches are further subjected to quality filters, removing those containing artefacts such as blurring or excessive whitespace.
Now that the dataset is composed of lung patches, a stratified purposeful sampling strategy will be employed to construct balanced training and evaluation datasets. This approach ensures adequate representation across the lung cancer subtypes, tumor stages, and demographic attributes (age, gender, ethnicity) where metadata are available.
Even after these processes, there may exist a class imbalance in the dataset, particularly among under-represented tumor stages or demographic subgroups. Therefore, augmentation and resampling techniques will be applied to address this issue:
- •
- Patch resampling from minority groups to increase intra-class diversity.
- •
- Geometric data augmentation will include normalization by rescaling pixel values by a factor of 1/255, the application of shear transformations with an intensity of 0.2, random rotations up to an angle of 0.2 radians, zooming in or out within a 0.5 range, and horizontal flipping. These transformations are designed to expand the variability of tissue presentation.
- •
- Stain normalization to reduce color domain shifts.
All the techniques are chosen to preserve histological fidelity, ensuring that diagnostic features remain intact. As a result of applying these techniques, the variability and complexity of the dataset will increase, leading to better model generalization and performance.
2.3. Training
Following the dataset preparation, the training of the DL models represents the next phase in the pipeline of developing an explainable DL model for lung cancer histopathological diagnosis.
2.3.1. Model Selection
The selection of deep learning architecture for this study is grounded in empirical evidence from recent advancements in computational pathology and their effectiveness in cancer histopathology tasks [1,34,37,38]. From all the possible models, the following three are selected:
- •
- EfficientNet-B7: Combines depth, width, and resolution scaling to achieve high accuracy while remaining computationally feasible. In histopathology, B7 has been applied in different tasks and demonstrates high accuracy while maintaining computational feasibility [9,39].
- •
- Dense-Net 121: Features dense connectivity patterns that promote feature reuse, reduce the number of parameters, and promote gradient flow. It has achieved a high-classification performance when distinguishing cancerous from non-cancerous tissues [37,40].
- •
- ResNet-50: Incorporates residual learning, allowing for the effective training of deeper networks. It remains one of the most widely adopted models in medical imaging, including cancer histopathology [1,34,41].
All models will be pre-trained on ImageNet and subsequently fine-tuned on the prepared dataset to leverage general image features while facilitating domain adaptation to histopathological patterns.
2.3.2. Training Configuration
All selected DL models will be trained following the configuration presented in Table 1.

Table 1.
Training configuration for deep learning models.
Image patches will be resized according to the model input size, and to further optimize the model’s performance, hyperparameters such as learning rate, batch size, and augmentation parameters will be tuned using a grid search combined with validation loss monitoring for early stopping. Potential pitfalls, such as overfitting due to model complexity and dataset imbalance affecting class predictions, will be mitigated through stratified sampling, augmentation, and rigorous validation design.
2.3.3. Hardware and Software Configuration
Each selected DL model will be implemented using the TensorFlow library [42] and trained on high-performance computing infrastructure with the following specifications:
- •
- Central Process Unit(s): 2 × AMD EPYC ROME 7452 (32 cores, 64 threads, 2.35 GHz).
- •
- Random Access Memory: 16 × 32 GB DDR4 RAM @ 3200 MHz (ECC Registered).
- •
- Graphical Process Unit(s): 4 × NVIDIA TESLA A100 40 GB.
- •
- Physical Memory: 2 × 3.84 TB NVMe PCIe SSDs (Western Digital DC SN640).
- •
- Operating System: Ubuntu 20.04 LTS with CUDA 11.6 and cuDNN 8.2.
2.4. Explainability Techniques’ Application
To enhance the transparency and clinical utility of the DL models used in this study, several interpretability techniques will be applied to the trained networks [8]:
- •
- Saliency and Gradient-Based Visualization: This highlights the spatial regions of input histopathological patches that contribute most to the model’s classification. Grad-CAM will be used to generate class-discriminative heatmaps by projecting gradients onto the final convolutional layer.
- •
- Occlusion Sensitivity: This involves masking parts of the image and observing changes in the output prediction. This helps to identify regions where the model is most sensitive.
- •
- Shapley Additive Explanations (SHAP): Values will be computed to quantify the contribution of each input feature toward the final decision.
These explainability techniques will enable pathologists to interactively explore and interpret model decisions and to improve user trust and model accountability, supporting the integration of AI into real-world clinical workflows. Figure 2 shows how the Grad-CAM technique can be applied to a lung cancer WSI [1]. The overlaid heatmap highlights regions that contribute most to the model’s prediction, helping pathologists interpret model behavior.
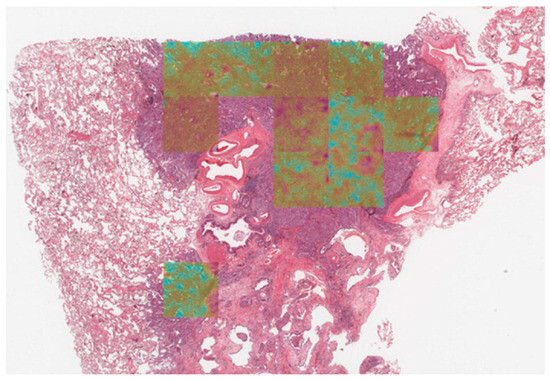
Figure 2.
Example of Grad-CAM application on a whole slide image of lung cancer. The heatmap highlights tumor regions, with lighter to darker shades indicating stronger tumor likelihood, while non-tumoral regions appear without overlay, showing only the lung tissue (adapted from [1]).
2.5. Evaluation
The evaluation of this study consists of two phases: (1) Model Validation and (2) Explainability Methods’ Validation.
2.5.1. Model Validation
In the evaluation of the models, the most used performance metrics in the literature will be employed [43]:
- •
- Accuracy: Represents the overall correctness of the model by calculating the proportion of correctly classified instances (both positive and negative) over the total number of instances.
- •
- Sensitivity: Measures the model’s ability to correctly identify positive tumoral cases. It is defined as the ratio of true positives to the sum of true positives and false negatives.
- •
- Specificity: Assesses the model’s ability to correctly identify negative cases (benign tissue). It is calculated as the ratio of true negatives to the sum of true negatives and false positives.
- •
- F1-score: Measures a weighted mean of Specificity and Sensitivity, balancing the trade-off between over-detection and under-detection.
- •
- AUC: Measures the model’s ability to distinguish between classes across varying classification thresholds. It plots the true positive rate against the false positive rate, with values closer to 1.0 indicating superior discriminative capability.
- •
- Cohen’s Kappa: Quantifies the inter-rater agreement between the predicted labels and the true labels, adjusting for agreement that could occur by chance.
These metrics will provide a comprehensive view of the model’s performance, balancing overall accuracy, the ability to diagnose lung cancer early, minimizing false positives, and ensuring reliable and reproducible predictions.
In a hypothetical scenario with values similar to those presented in Table 2, an accuracy of 0.92 and an AUC of 0.95 would indicate that the model can reliably distinguish between tumoral and non-tumoral samples. A Cohen’s Kappa of 0.87 demonstrates strong agreement with ground truth beyond chance, reinforcing the model’s consistency. Therefore, results such as these will help clinicians understand the reliability and robustness of the model in early lung cancer diagnosis, thereby increasing their confidence in its application in a clinical workflow.

Table 2.
Synthetic results from a hypothetical experiment of lung cancer diagnosis.
Besides the performance metrics, a structured survey will be made available to pathologists where, after testing, they will select on a Likert scale how the model behaved and if the results were satisfactory.
2.5.2. Explainability Methods’ Validation
The explainability methods of the models will be evaluated not only by visual inspection but also through systematic, measurable criteria [8]:
- •
- Localization Accuracy: Assesses whether saliency-based methods such as Grad-CAM correctly localize cancerous tissue. Metrics such as the Intersection Over Union (IoU) will be used to quantify the alignment between the highlighted heatmaps and annotated tumor regions.
- •
- Sensitivity to Input Perturbation: Measures how significantly the prediction changes with systematic masking.
- •
- Reliability: Each interpretability method will be measured across multiple augmentations (e.g., flipped or rotated patches). A stable explanation method should generate consistent attribution maps regardless of minor transformations.
- •
- Pathologists’ Evaluation: Pathologists will be asked to rate the clinical acceptability and diagnostic utility of the model explanations.
This combined evaluation strategy ensures that the DL models are not only accurate but also transparent, trustworthy, and aligned with clinical reasoning, facilitating their adoption into clinical workflows.
3. Ethical Considerations
Given the sensitive nature of clinical data and the increasing scrutiny of AI in healthcare, the ethical considerations in this study are significant and complex. Although this study involves the secondary use of publicly available data, it still requires careful attention to privacy, fairness, transparency, and future clinical implications.
The chosen datasets (TCGA and LC25000) are explicitly designed for research use. All data are publicly accessible and anonymized, as they do not provide information that allows the identification of the patient, and they are HIPAA compliant. As such, the use of these datasets falls under the secondary use of public domain data and does not require ethical approval from a research ethics committee under most institutional guidelines. Nevertheless, all dataset usage will adhere to the associated data use agreements and licensing terms, and full source attribution will be provided in line with open science standards.
Despite the use of public data, the risk of algorithmic bias remains. These datasets may under-represent minority demographic groups and rare tumor types, which could limit the generalizability of the model. As such, dataset diversity will be reported, and mitigation measures should be implemented. These measures include the application of the stratified purposeful sampling approach to promote representation across all clinical sub-groups, and the implementation of data augmentation techniques for a better class balance. The feedback from pathologists will also help to identify biases or limitations in model interpretability across different case types.
One of the main aspects of this research is the integration of explainability techniques to address the “black box” nature of DL models. Through these techniques, pathologists will be able to understand and evaluate the model’s reasoning process, supporting an informed decision-making process and improving transparency.
Details of the used model architecture, training configurations, and evaluation results will be published in an open-access format so that other researchers can reproduce them and perform peer validations.
As this study does not involve new data collection or direct interaction with patients, formal ethical approval is not required at this stage. However, when ready for deployment to a clinician environment, the model will require an ethical and regulatory review. This review may include trials, patient consent protocols, and integration with clinical decision support systems that ensure pathologists remain in control of the final diagnostic decisions.
4. Data Analysis
The analytical strategy in this study is organized into three sequential stages: data preparation, first-order coding and evaluation, and second-order coding and modeling (Figure 3). This framework is designed to operate directly on real-world histopathology datasets (e.g., TCGA-LUAD, LC25000), combining quantitative outputs (model predictions, Grad-CAM heatmaps) with ground-truth annotations from pathologists. This structure enables a rigorous and complete assessment of the DL models, addressing both predictive accuracy and explainability.
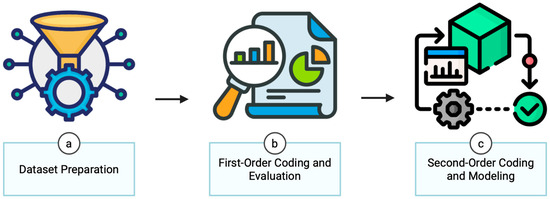
Figure 3.
An analytical strategy to develop an explainable deep learning algorithm for lung cancer histopathological diagnosis.
4.1. Data Preparation
As outlined in the Data Preparation section of Materials, Methods, and Procedural Design, the acquired low-quality WSIs or patches (obtained from real-world datasets such as TCGA-LUAD and LC25000) will be excluded to maintain data integrity, and the dataset will be standardized in terms of image format, label nomenclature, and patch-level resolution, followed by normalization and augmentation. Ground-truth annotations are completed by pathologists and aligned spatially with the image data. For interpretability validation, the annotations indicating diagnostically relevant regions of interest are stored to later compare with visual attribution maps (e.g., Grad-CAM heatmaps). The output of the model, such as predictions and Grad-CAM heatmaps, is systematically logged and formatted to serve as quantitative data for analysis.
4.2. First-Order Coding and Evaluation
In the second phase, model performance on the real datasets is quantitatively assessed using common metrics, and assumptions of distributional normality are examined to guide statistical comparisons across model architectures. Additionally, first-cycle qualitative coding is conducted on the explainability outputs. Descriptive coding is used to annotate salient features highlighted in attribution maps (e.g., “nuclear focus”, “stromal activation”), while attribute coding captures contextual metadata, such as cancer subtype and confidence level [44].
4.3. Second-Order Coding and Modeling
The third phase involves comparing the model’s performance using measures, such as an analysis of variance or non-parametric equivalents, depending on data distribution characteristics. The alignment between Grad-CAM visualizations and annotated regions is evaluated, while saliency variability is tested through controlled input perturbations [8,45].
Regarding second-cycle coding, focused coding identifies recurrent or clinically significant first-cycle codes, and pattern coding aggregates them into broader themes such as “clinically coherent focus” or “non-diagnostic attribution”. Axial coding is employed to explore the relationships between these themes across different models, cancer subtypes, and prediction outcomes, revealing latent dynamics in model behavior and interpretability [44].
In summary, this analysis does not only focus on model accuracy but also investigates how models make decisions, where they focus attention, and whether these rationales align with clinical reasoning [8,19].
5. Conclusions
The development of DL tools in cancer diagnostics has shown promising advances in accuracy and efficiency. However, their widespread clinical adoption remains hindered due to the “black box” problem of DL models, where decision-making processes are opaque to clinicians. Therefore, this paper proposed a methodological framework for the development and evaluation of interpretable deep learning models tailored to the early histopathological diagnosis of lung cancer. In this framework, large-scale public datasets, state-of-the-art models, and proven explainability techniques will be used to achieve clinically acceptable predictive accuracy with transparent decision-making. The performance evaluation of the models and explainability output will be made with widely used quantitative metrics while relying also on structured and measurable feedback from pathologists. The results from this work may reinforce the potential of explainable AI to enhance the reliability, transparency, and integration of AI systems into routine pathology workflows.
As this study represents a methodological proposal, future work will focus on its practical implementation and validation. The implementation will include dataset preparation, model training using DL models such as EfficientNet, and the application of explainability methods including Grad-CAM and SHAP. Besides assessing model performance, the visual explanations will also be aligned with diagnostic ground truth through both a spatial analysis and expert review. Additionally, future efforts will also include the addition of rarer lung cancer subtypes to the dataset and the deployment of the system in a real-world clinical environment.
Ultimately, the main objective is to turn this methodological framework into a practical, clinically relevant tool that not only improves diagnostic accuracy but also builds trust in how AI supports decisions in real-world medical settings.
Author Contributions
Conceptualization, N.F., S.C. and V.C.; methodology, N.F., S.C. and V.C.; software, N.F., S.C. and V.C.; validation, N.F., S.C. and V.C.; formal analysis, N.F., S.C. and V.C.; investigation, N.F., S.C. and V.C.; resources, N.F., S.C. and V.C.; data curation, N.F., S.C. and V.C.; writing—original draft preparation, N.F.; writing—review and editing, N.F., S.C. and V.C.; visualization, N.F., S.C. and V.C.; supervision, S.C. and V.C.; project administration, V.C.; funding acquisition, V.C. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by FCT/MCTES grant numbers UIDB/05549:2Ai and UIDP/05549:2Ai.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Publicly available datasets were analyzed in this study. These data can be found in https://portal.gdc.cancer.gov and in https://www.kaggle.com/datasets/javaidahmadwani/lc25000 (accessed on 24 April 2025).
Acknowledgments
The authors would like to thank The Cancer Genome Atlas, the Genomic Data Commons Data Portal, and Andrew Borkowski et al. for the lung cancer datasets’ availability. During the preparation of this manuscript/study, the authors used [GPT-4o] for the purposes of sentence refinement, based on the authors’ original wording. The authors have reviewed and edited the output and take full responsibility for the content of this publication.
Conflicts of Interest
Author Sofia Campelos was employed by the company IMP Diagnostics. The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Abbreviations
The following abbreviations are used in this manuscript:
| ADC | Adenocarcinoma |
| AI | Artificial Intelligence |
| AUC | Area Under the Curve |
| DL | Deep Learning |
| Grad-CAM | Gradient-Weighted Class Activation Mapping |
| H&E | Haematoxylin and Eosin |
| HIPAA | Health Insurance Portability and Accountability Act |
| IoU | Intersection over Union |
| LC25000 | Lung and Colon Cancer Histopathological Image Dataset |
| ROI | Region of Interest |
| SCC | Squamous Cell Carcinoma |
| SHAP | Shapley Additive Explanations |
| TCGA | The Cancer Genome Atlas |
| WSI | Whole Slide Image |
References
- Faria, N.; Campelos, S.; Carvalho, V. A Novel Convolutional Neural Network Algorithm for Histopathological Lung Cancer Detection. Appl. Sci. 2023, 13, 6571. [Google Scholar] [CrossRef]
- Bera, K.; Schalper, K.A.; Rimm, D.L.; Velcheti, V.; Madabhushi, A. Artificial Intelligence in Digital Pathology—New Tools for Diagnosis and Precision Oncology. Nat. Rev. Clin. Oncol. 2019, 16, 703–715. [Google Scholar] [CrossRef] [PubMed]
- Faria, N.; Campelos, S.; Carvalho, V. Lorenz, R., Fred, A., Gamboa, H., Eds.; Cancer Detec-Lung Cancer Diagnosis Support System: First Insights. In Bioinformatics; Proceedings of the 15th International Joint Conference on Biomedical Engineering Systems and Technologies—Vol 3: Bioinformatics, Virtual Event, February 9–11, 2022; Scitepress: Setubal, Portugal, 2021; pp. 81–88. [Google Scholar]
- Gour, M.; Jain, S.; Sunil Kumar, T. Residual Learning Based CNN for Breast Cancer Histopathological Image Classification. Int. J. Imaging Syst. Tech. 2020, 30, 621–635. [Google Scholar] [CrossRef]
- Litjens, G.; Kooi, T.; Bejnordi, B.E.; Setio, A.A.A.; Ciompi, F.; Ghafoorian, M.; van der Laak, J.A.W.M.; van Ginneken, B.; Sánchez, C.I. A Survey on Deep Learning in Medical Image Analysis. Med. Image Anal. 2017, 42, 60–88. [Google Scholar] [CrossRef] [PubMed]
- Chaddad, A.; Peng, J.; Xu, J.; Bouridane, A. Survey of Explainable AI Techniques in Healthcare. Sensors 2023, 23, 634. [Google Scholar] [CrossRef] [PubMed]
- Patrício, C.; Neves, J.C.; Teixeira, L.F. Explainable Deep Learning Methods in Medical Image Classification: A Survey. ACM Comput. Surv. 2023, 56, 1–41. [Google Scholar] [CrossRef]
- Salahuddin, Z.; Woodruff, H.C.; Chatterjee, A.; Lambin, P. Transparency of Deep Neural Networks for Medical Image Analysis: A Review of Interpretability Methods. Comput. Biol. Med. 2022, 140, 105111. [Google Scholar] [CrossRef]
- Latha, M.; Kumar, P.S.; Chandrika, R.R.; Mahesh, T.R.; Kumar, V.V.; Guluwadi, S. Revolutionizing Breast Ultrasound Diagnostics with EfficientNet-B7 and Explainable AI. BMC Med. Imaging 2024, 24, 230. [Google Scholar] [CrossRef]
- Tian, L.; Wu, J.; Song, W.; Hong, Q.; Liu, D.; Ye, F.; Gao, F.; Hu, Y.; Wu, M.; Lan, Y.; et al. Precise and Automated Lung Cancer Cell Classification Using Deep Neural Network with Multiscale Features and Model Distillation. Sci. Rep. 2024, 14, 10471. [Google Scholar] [CrossRef]
- Creswell, J.W.; Creswell, J.D. Research Design: Qualitative, Quantitative, and Mixed Methods Approaches; SAGE Publications: London, UK, 2017; ISBN 978-1-5063-8669-0. [Google Scholar]
- Ghassemi, M.; Oakden-Rayner, L.; Beam, A.L. The False Hope of Current Approaches to Explainable Artificial Intelligence in Health Care. Lancet Digit. Health 2021, 3, e745–e750. [Google Scholar] [CrossRef]
- Lincoln, Y.S.; Lynham, S.A.; Guba, E.G. Paradigmatic Controversies, Contradictions, and Emerging Confluences, Revisited. Sage Handb. Qual. Res. 2011, 4, 97–128. [Google Scholar]
- Mertens, D.M. Transformative Research: Personal and Societal. Int. J. Transform. Res. 2017, 4, 18–24. [Google Scholar] [CrossRef]
- Elder-Vass, D. Pragmatism, Critical Realism and the Study of Value. J. Crit. Realism 2022, 21, 261–287. [Google Scholar] [CrossRef]
- Lawani, A. Critical Realism: What You Should Know and How to Apply It. Qual. Res. J. 2020, 21, 320–333. [Google Scholar] [CrossRef]
- Topol, E. Deep Medicine: How Artificial Intelligence Can Make Healthcare Human Again; Basic Books: New York, NY, USA, 2019; ISBN 978-1-5416-4464-9. [Google Scholar]
- Haibe-Kains, B.; Adam, G.A.; Hosny, A.; Khodakarami, F.; Waldron, L.; Wang, B.; McIntosh, C.; Goldenberg, A.; Kundaje, A.; Greene, C.S.; et al. Transparency and Reproducibility in Artificial Intelligence. Nature 2020, 586, E14–E16. [Google Scholar] [CrossRef]
- Holzinger, A.; Langs, G.; Denk, H.; Zatloukal, K.; Müller, H. Causability and Explainability of Artificial Intelligence in Medicine. WIREs Data Min. Knowl. Discov. 2019, 9, e1312. [Google Scholar] [CrossRef]
- Dawadi, S.; Shrestha, S.; Giri, R.A. Mixed-Methods Research: A Discussion on Its Types, Challenges, and Criticisms. JPSE 2021, 2, 25–36. [Google Scholar] [CrossRef]
- Cohen, L.; Manion, L.; Morrison, K. Research Methods in Education, 8th ed.; Routledge: London, UK, 2017; ISBN 978-1-315-45653-9. [Google Scholar]
- LeCun, Y.; Bengio, Y.; Hinton, G. Deep Learning. Nature 2015, 521, 436–444. [Google Scholar] [CrossRef] [PubMed]
- Goodfellow, I.; Bengio, Y.; Courville, A. Deep Learning; MIT Press: Cambridge, MA, USA, 2016. [Google Scholar]
- Yamashita, R.; Nishio, M.; Do, R.K.G.; Togashi, K. Convolutional Neural Networks: An Overview and Application in Radiology. Insights Imaging 2018, 9, 611–629. [Google Scholar] [CrossRef] [PubMed]
- Goldenholz, D.M.; Sun, H.; Ganglberger, W.; Westover, M.B. Sample Size Analysis for Machine Learning Clinical Validation Studies. Biomedicines 2023, 11, 685. [Google Scholar] [CrossRef]
- Collins, G.S.; Ogundimu, E.O.; Altman, D.G. Sample Size Considerations for the External Validation of a Multivariable Prognostic Model: A Resampling Study. Stat. Med. 2016, 35, 214–226. [Google Scholar] [CrossRef] [PubMed]
- Suri, H. Purposeful Sampling in Qualitative Research Synthesis. Qual. Res. J. 2011, 11, 63–75. [Google Scholar] [CrossRef]
- Chen, I.Y.; Joshi, S.; Ghassemi, M. Treating Health Disparities with Artificial Intelligence. Nat. Med. 2020, 26, 16–17. [Google Scholar] [CrossRef]
- National Cancer Institute. The Cancer Genome Atlas Program (TCGA). Available online: https://www.cancer.gov/ccg/research/genome-sequencing/tcga (accessed on 10 April 2025).
- Albertina, B.; Watson, M.; Holback, C.; Jarosz, R.; Kirk, S.; Lee, Y.; Rieger-Christ, K.; Lemmerman, J. The Cancer Genome Atlas Lung Adenocarcinoma Collection (TCGA-LUAD); The Cancer Imaging Archive: Palo Alto, CA, USA, 2016. [Google Scholar]
- Kirk, S.; Lee, Y.; Kumar, P.; Filippini, J.; Albertina, B.; Watson, M.; Rieger-Christ, K.; Lemmerman, J. The Cancer Genome Atlas Lung Squamous Cell Carcinoma Collection (TCGA-LUSC); The Cancer Imaging Archive: Palo Alto, CA, USA, 2016. [Google Scholar]
- Borkowski, A.A.; Bui, M.M.; Thomas, L.B.; Wilson, C.P.; DeLand, L.A.; Mastorides, S.M. Lung and Colon Cancer Histopathological Image Dataset (LC25000). arXiv 2019, arXiv:1912.12142. [Google Scholar] [CrossRef]
- cBioPortal for Cancer Genomics—Lung Adenocarcinoma (TCGA, GDC). Available online: https://www.cbioportal.org/study/summary?id=luad_tcga_gdc (accessed on 3 May 2025).
- Coudray, N.; Ocampo, P.S.; Sakellaropoulos, T.; Narula, N.; Snuderl, M.; Fenyö, D.; Moreira, A.L.; Razavian, N.; Tsirigos, A. Classification and Mutation Prediction from Non-Small Cell Lung Cancer Histopathology Images Using Deep Learning. Nat. Med. 2018, 24, 1559–1567. [Google Scholar] [CrossRef]
- cBioPortal for Cancer Genomics—Lung Squamous Cell Carcinoma (TCGA, GDC). Available online: https://www.cbioportal.org/study/summary?id=lusc_tcga_gdc (accessed on 3 May 2025).
- CDC Health Insurance Portability and Accountability Act of 1996 (HIPAA). Available online: https://www.cdc.gov/phlp/php/resources/health-insurance-portability-and-accountability-act-of-1996-hipaa.html (accessed on 6 April 2025).
- Sumon, R.I.; Mazumdar, M.A.I.; Uddin, S.M.I.; Kim, H.-C. Exploring Deep Learning and Machine Learning Techniques for Histopathological Image Classification in Lung Cancer Diagnosis. In Proceedings of the 2024 International Conference on Electrical, Computer and Energy Technologies (ICECET), Sydney, Australia, 25–27 July 2024; pp. 1–6. [Google Scholar]
- Katar, O.; Yildirim, O.; Tan, R.-S.; Acharya, U.R. A Novel Hybrid Model for Automatic Non-Small Cell Lung Cancer Classification Using Histopathological Images. Diagnostics 2024, 14, 2497. [Google Scholar] [CrossRef]
- Tan, M.; Le, Q.V. EfficientNet: Rethinking Model Scaling for Convolutional Neural Networks. In International conference on machine learning; PMLR: Cambridge, MA, USA, 2020; pp. 6105–6114. [Google Scholar]
- Huang, G.; Liu, Z.; van der Maaten, L.; Weinberger, K.Q. Densely Connected Convolutional Networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Salt Lake City, UT, USA, 18–22 June 2018. [Google Scholar]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep Residual Learning for Image Recognition. In Proceedings of the 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, NV, USA, 27–30 June 2016; pp. 770–778. [Google Scholar]
- Abadi, M.; Agarwal, A.; Barham, P.; Brevdo, E.; Chen, Z.; Citro, C.; Corrado, G.S.; Davis, A.; Dean, J.; Devin, M.; et al. TensorFlow: Large-Scale Machine Learning on Heterogeneous Distributed Systems. arXiv 2016, arXiv:1603.04467. [Google Scholar]
- Müller, D.; Soto-Rey, I.; Kramer, F. Towards a Guideline for Evaluation Metrics in Medical Image Segmentation. BMC Res. Notes 2022, 15, 210. [Google Scholar] [CrossRef]
- Saldaña, J. The Coding Manual for Qualitative Researchers, 3rd ed.; SAGE: Los Angeles, CA, USA; London, UK; New Delhi, India; Singapore; Washington, DC, USA, 2016; ISBN 978-1-4739-0249-7. [Google Scholar]
- Adebayo, J.; Gilmer, J.; Muelly, M.; Goodfellow, I.; Hardt, M.; Kim, B. Sanity Checks for Saliency Maps. In Advances in Neural Information Processing Systems; Curran Associates, Inc.: Red Hook, NY, USA, 2018; Volume 31. [Google Scholar]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).