Using Deep Learning for Image-Based Different Degrees of Ginkgo Leaf Disease Classification
Abstract
1. Introduction
- To classify the different degrees of disease in Ginkgo biloba leaves using a deep learning model under laboratory and field conditions that takes into account sunshine, temperature, weather, and other factors.
2. Materials and Methods
2.1. Data Set
- Ginkgo leaves have a more complete shape.
- Ginkgo leaves are flat and easy to photograph.
- Ginkgo leaves’ surfaces are clean.
- Ginkgo leaves’ periodic disease characteristics are clearly distinguishable.
2.2. Image Preprocessing and Labeling
2.3. Data Augmentation
2.4. Convolutional Neural Network Models
2.5. Training Data Sets
3. Results and Discussion
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Dekosky, S.T.; Williamson, J.D.; Fitzpatrick, A.L.; Kronmal, R.A.; Ives, D.G.; Saxton, J.A. Ginkgo biloba for prevention of dementia: A randomized controlled trial. JAMA. 2008, 300, 2253–2262. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, C.M.H.; Wolffram, S.; Ader, P.; Rimbach, G.; Packer, L.; Maguire, J.J. The in vivo neuromodulatory effects of the herbal medicine ginkgo biloba. Proc. Natl. Acad. Sci. USA 2001, 98, 6577–6580. [Google Scholar] [CrossRef] [PubMed]
- Sally, A.M.; Beed, F.D.; Harmon, C.L. Plant disease diagnostic capabilities and networks. Annu. Rev. Phytopathol. 2009, 47, 15–38. [Google Scholar]
- Sankaran, S.; Mishra, A.; Ehsani, R. A review of advanced techniques for detecting plant diseases. Comput. Electron. Agric. 2010, 72, 1–13. [Google Scholar] [CrossRef]
- Rumpf, T.; Mahlein, A.K.; Steiner, U.; Oerke, E.C.; Dehne, H.W.; Plumer, L. Early detection and classification of plant diseases with Support Vector Machines based on hyperspectral reflectance. Comput. Electron. Agric. 2010, 74, 91–99. [Google Scholar] [CrossRef]
- Tian, Y.; Zhao, C.J.; Lu, S.L.; Guo, X.Y. Multiple Classifier Combination For Recognition Of Wheat Leaf Diseases. Intell. Autom. Soft Comput. 2011, 17, 519–529. [Google Scholar] [CrossRef]
- Wilf, P.; Zhang, S.; Chikkerur, S.; Little, S.A.; Wing, S.L.; Serre, T. Computer vision cracks the leaf code. Proc. Natl. Acad. Sci. USA 2016, 113, 3305–3310. [Google Scholar] [CrossRef]
- Wäldchen, J.; Mäder, P. Plant species identification using computer vision techniques: A systematic literature review. Arch Comput. Method E. 2018, 25, 507–543. [Google Scholar] [CrossRef]
- Kumar, N.; Belhumeur, P.N.; Biswas, A.; Jacobs, D.W.; Kress, W.J. Leafsnap: A computer vision system for automatic plant species identification. In European Conference on Computer Vision; Springer: Berlin/Heidelberg, Germany, 2012; pp. 502–516. [Google Scholar]
- Wu, Q.F.; Lin, K.H.; Zhou, C.G. Feature extraction and automatic recognition of plant leaf using artificial neural network. Adv. Artif. Intell. 2007, 3, 5–12. [Google Scholar]
- Hong, F. Extraction of Leaf Vein Features Based on Artificial Neural Network-Studies on the Living Plant Identification Ⅰ. Chin. Bull. Bot. 2004, 21, 429–436. (In Chinese) [Google Scholar]
- Rastogi, A.; Arora, R.; Sharma, S. Leaf disease detection and grading using computer vision technology and fuzzy logic. In Proceedings of the 2nd International Conference on Signal Processing and Integrated Networks (SPIN), Noida, India, 19–20 February 2015; pp. 500–505. [Google Scholar]
- Mehrotra, K.; Mohan, C.K.; Ranka, S. Elements of Artificial Neural Networks. A Bradford Book; The MIT Press: Cambridge, MA, USA; London, UK, 1997. [Google Scholar]
- Hati, S.; Sajeevan, G. Plant recognition from leaf image through artificial neural network. IJCA 2013, 62, 15–18. [Google Scholar] [CrossRef]
- LeCun, Y.; Bottou, L.; Bengio, Y.; Haffner, P. Gradient-based learning applied to document Recognition. Proc. IEEE 1998, 86, 2278–2324. [Google Scholar] [CrossRef]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. Imagenet Classification with Deep Convolutional Neural Networks. Adv. Neural Inf. Process. Syst. 2012, 25, 1097–1105. [Google Scholar] [CrossRef]
- Grinblat, G.L.; Uzal, L.C.; Larese, M.G.; Granitto, P.M. Deep learning for plant identification using vein morphological patterns. Comput. Electron. Agric. 2016, 127, 418–424. [Google Scholar] [CrossRef]
- Lee, S.H.; Chan, C.S.; Wilkin, P.; Remagnino, P. Deep-plant: Plant identification with convolutional neural networks. In Proceedings of the IEEE International Conference on Image Processing (ICIP), Quebec City, QC, Canada, 27–30 September 2015; pp. 452–456. [Google Scholar]
- Carranza-Rojas, J.; Goeau, H.; Bonnet, P.; Mata-Montero, E.; Joly, A. Going deeper in the automated identification of Herbarium specimens. BMC Evol. Biol. 2017, 17, 181. [Google Scholar] [CrossRef]
- Ferentinos, K.P. Deep learning models for plant disease detection and diagnosis. Comput. Electron. Agric. 2018, 145, 311–318. [Google Scholar] [CrossRef]
- Sladojevic, S.; Arsenovic, M.; Anderla, A.; Culibrk, D.; Stefanovic, D. Deep Neural Networks Based Recognition of Plant Diseases by Leaf Image Classification. Comput. Intell. Neurosc. 2016, 2016, 3289801. [Google Scholar] [CrossRef]
- Mohanty, S.P.; Hughes, D.P.; Salathé, M. Using Deep Learning for Image-Based Plant Disease Detection. Front Plant Sci. 2016, 17, 1419. [Google Scholar] [CrossRef]
- Brahimi, M.; Boukhalfa, K.; Moussaoui, A. Deep learning for tomato diseases: Classification and symptoms visualization. Appl. Artif. Intell. 2017, 31, 299–315. [Google Scholar] [CrossRef]
- Lu, Y.; Yi, S.J.; Zeng, N.Y.; Liu, Y.R.; Zhang, Y. Identification of rice diseases using deep convolutional neural networks. Neurocomputing 2017, 267, 378–384. [Google Scholar] [CrossRef]
- Hawkins, D.M. The problem of overfitting. J. Chem. Inf. Comput. Sci. 2004, 35, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Simonyan, K.; Zisserman, A. Very deep convolutional networks for large-scale image recognition. arXiv 2014, arXiv:1409.1556. [Google Scholar]
- Szegedy, C.; Vanhoucke, V.; Ioffe, S.; Shlens, J.; Wojna, Z. Rethinking the inception architecture for computer vision. In Proceedings of the Computer Vision and Pattern Recognition 2016, Las Vegas, NV, USA, 27 June–28 July 2016; pp. 2818–2826. [Google Scholar]
- Schmidhuber, J. Deep Learning in neural networks: An overview. Neural Netw. 2015, 61, 85–117. [Google Scholar] [CrossRef] [PubMed]
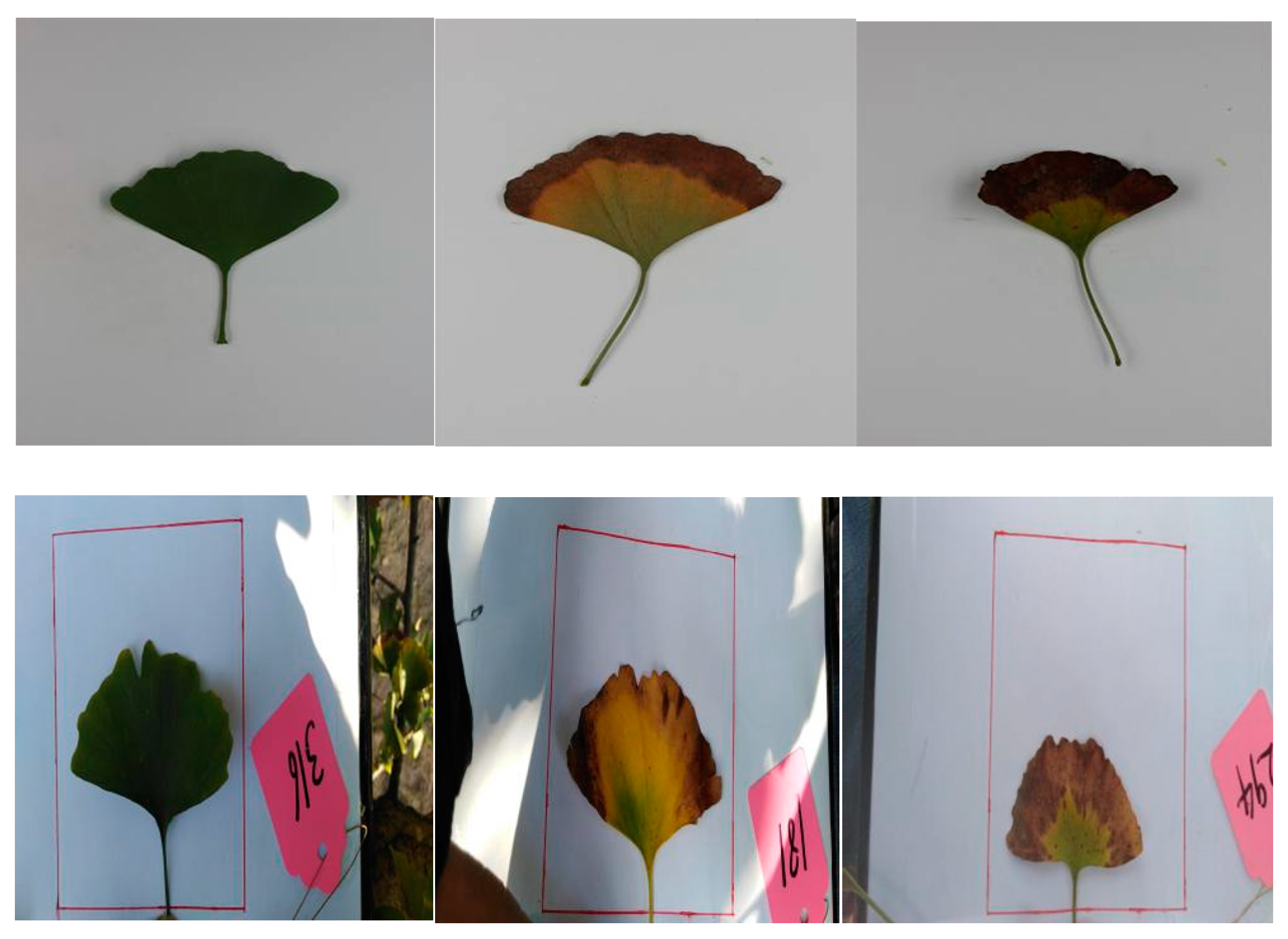








| Disease Degree | Laboratory Conditions | Field Conditions |
|---|---|---|
| Healthy | 326 | 87 |
| Mild | 671 | 1945 |
| Severe | 322 | 376 |
| Laboratory Conditions | Field Conditions | |
|---|---|---|
| Original picture size (KB) | 1887–6770 | 1270–2847 |
| Original picture size (pixels) | 5184 × 3456 | 4160 × 2080 |
| Experimental picture size (KB) | 157–540 | 298–643 |
| Experimental picture size (pixels) | 1800 × 1200 | 4160 × 2080 |
| Disease Degree | Laboratory Conditions | Field Conditions |
|---|---|---|
| Healthy | 5569 | 5569 |
| Mild | 5964 | 5964 |
| Severe | 4137 | 4137 |
| Total | 15,670 | 15,670 |
| Parameters | VGG-16 | Inception V3 |
|---|---|---|
| Batch-size | 64 | 64 |
| Step | 2000 | 4000 |
| Input-width | 224 | 299 |
| Input-height | 224 | 299 |
| Learning | rate | 0.01–0.0001 |
| Learning Rate | VGG16 | |||
|---|---|---|---|---|
| Laboratory Conditions | Field Conditions | |||
| Accuracy | Loss | Accuracy | Loss | |
| 0.01 | 93.75% | 0.15 | 81.25% | 0.51 |
| 0.005 | 98.44% | 0.05 | 87.50% | 0.26 |
| 0.001 | 98.44% | 0.02 | 92.19% | 0.17 |
| 0.0005 | 98.44% | 0.03 | 89.06% | 0.24 |
| 0.0001 | 98.44% | 0.05 | 85.94% | 0.37 |
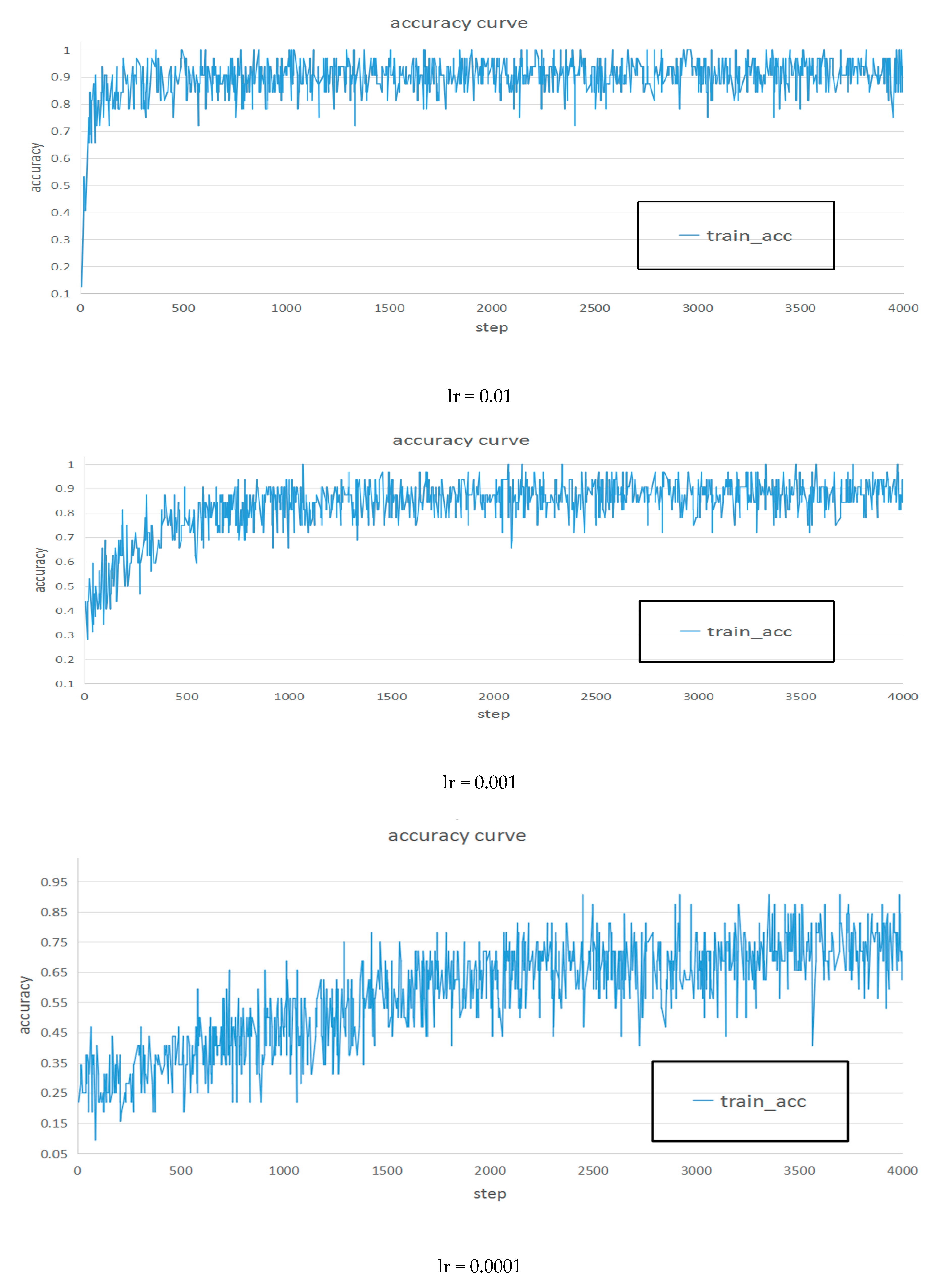
| Learning Rate | Inception V3 | |
|---|---|---|
| Laboratory Conditions | Field Conditions | |
| Accuracy | Accuracy | |
| 0.01 | 92.30% | 93.20% |
| 0.001 | 88.60% | 89.00% |
| 0.0001 | 73.40% | 73.20% |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, K.; Lin, J.; Liu, J.; Zhao, Y. Using Deep Learning for Image-Based Different Degrees of Ginkgo Leaf Disease Classification. Information 2020, 11, 95. https://doi.org/10.3390/info11020095
Li K, Lin J, Liu J, Zhao Y. Using Deep Learning for Image-Based Different Degrees of Ginkgo Leaf Disease Classification. Information. 2020; 11(2):95. https://doi.org/10.3390/info11020095
Chicago/Turabian StyleLi, Kaizhou, Jianhui Lin, Jinrong Liu, and Yandong Zhao. 2020. "Using Deep Learning for Image-Based Different Degrees of Ginkgo Leaf Disease Classification" Information 11, no. 2: 95. https://doi.org/10.3390/info11020095
APA StyleLi, K., Lin, J., Liu, J., & Zhao, Y. (2020). Using Deep Learning for Image-Based Different Degrees of Ginkgo Leaf Disease Classification. Information, 11(2), 95. https://doi.org/10.3390/info11020095





