ARTD10/PARP10 Induces ADP-Ribosylation of GAPDH and Recruits GAPDH into Cytosolic Membrane-Free Cell Bodies When Overexpressed in Mammalian Cells
Abstract
1. Introduction
2. Material and Methods
2.1. Cell Culture, Transfections and Fractionation
2.2. ADP-Ribosylation Assays and GAPDH Activity Assay
2.3. Western Blotting and Antibodies
2.4. Co-Immunoprecipitation, Immunoprecipitation, and GST-Pull Down
2.5. Proteomic Analysis
2.6. Indirect Immunofluorescence and Antibodies
2.7. Electron Microscopy
2.8. Statistical Analysis
3. Results and Discussion
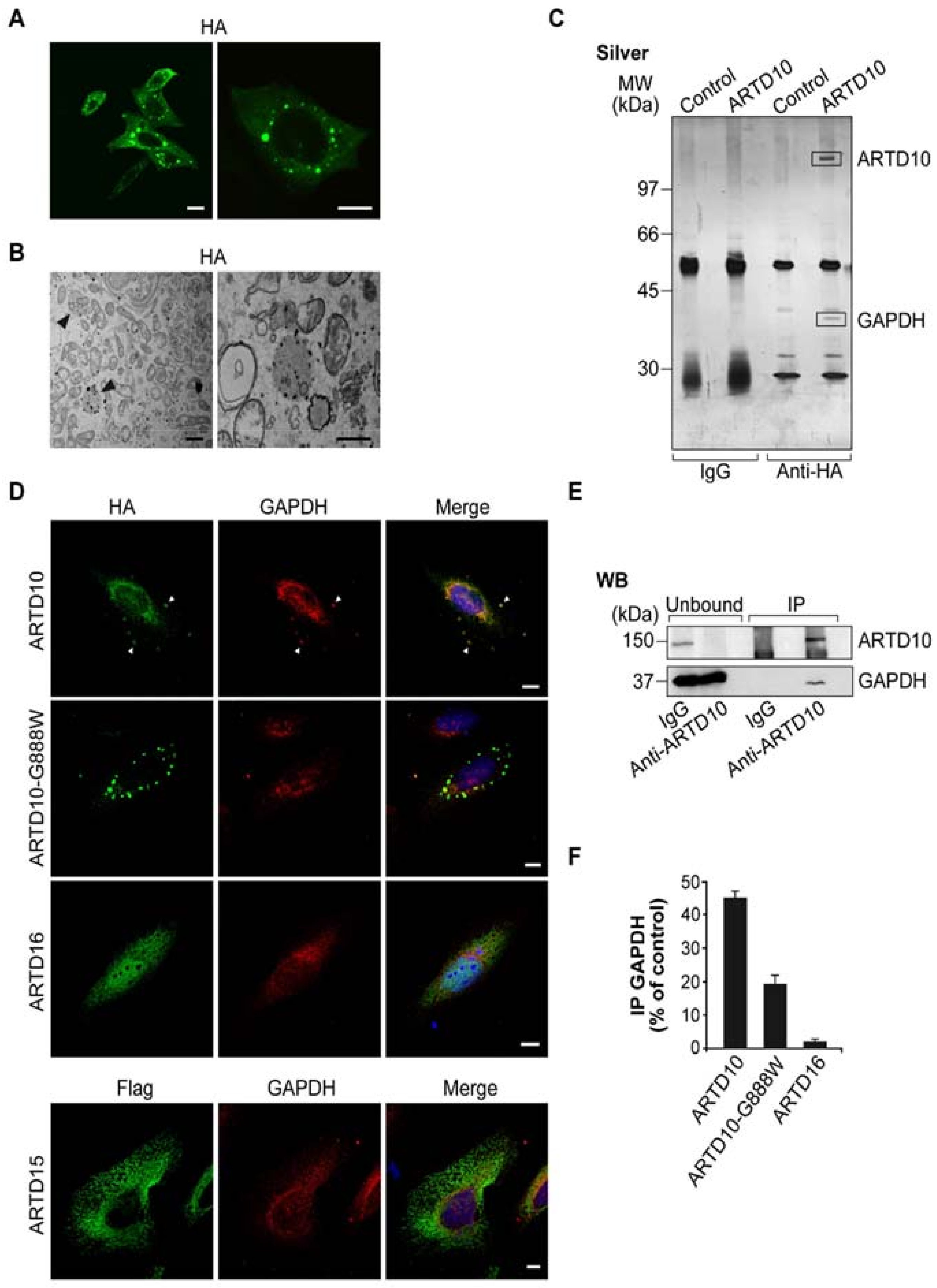
3.1. ARTD10 Localises in Membrane-Free Cell Bodies
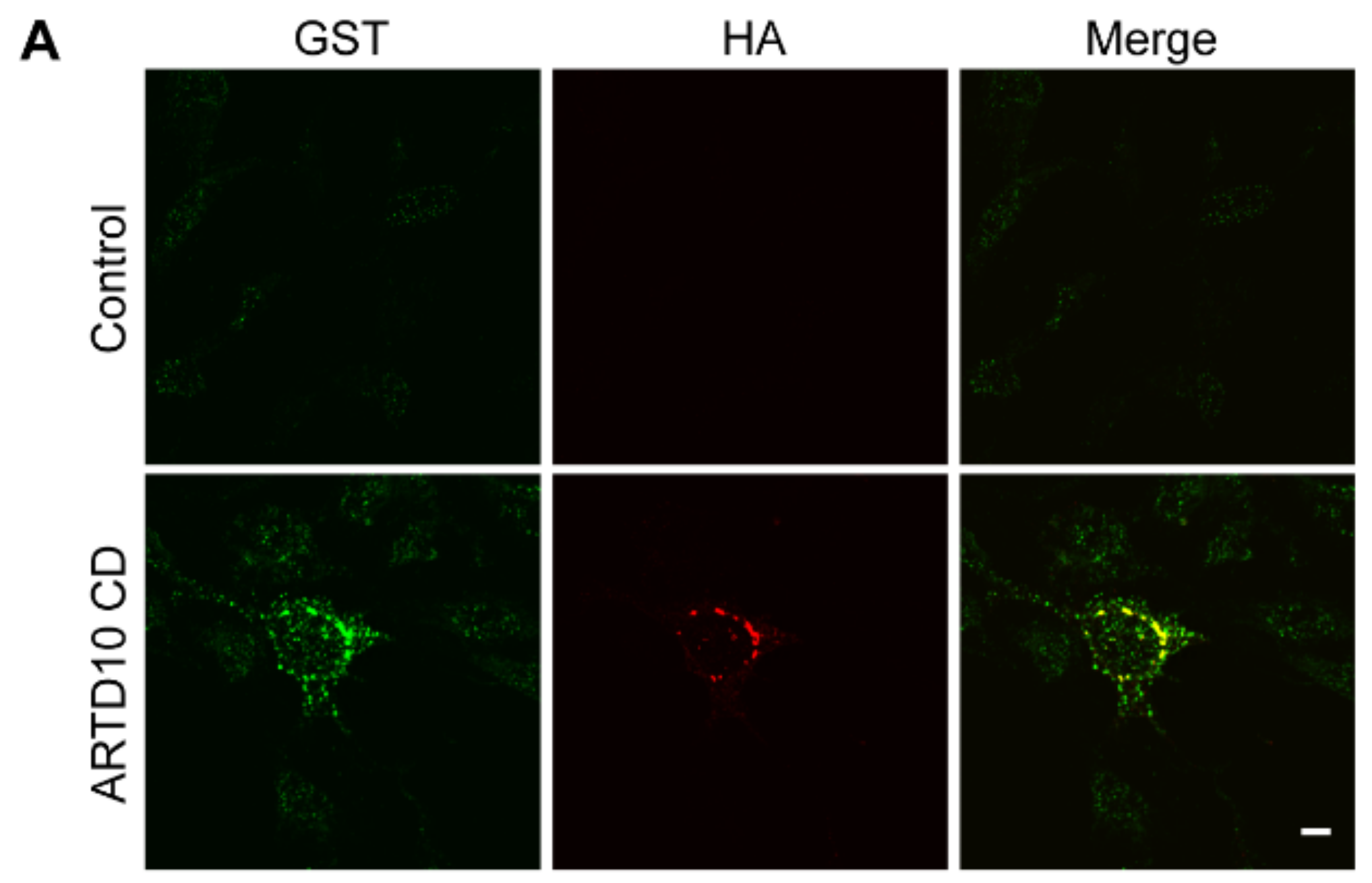
3.2. ARTD10 Interacts and Co-Localises with GAPDH in Membrane-Free Cell Bodies
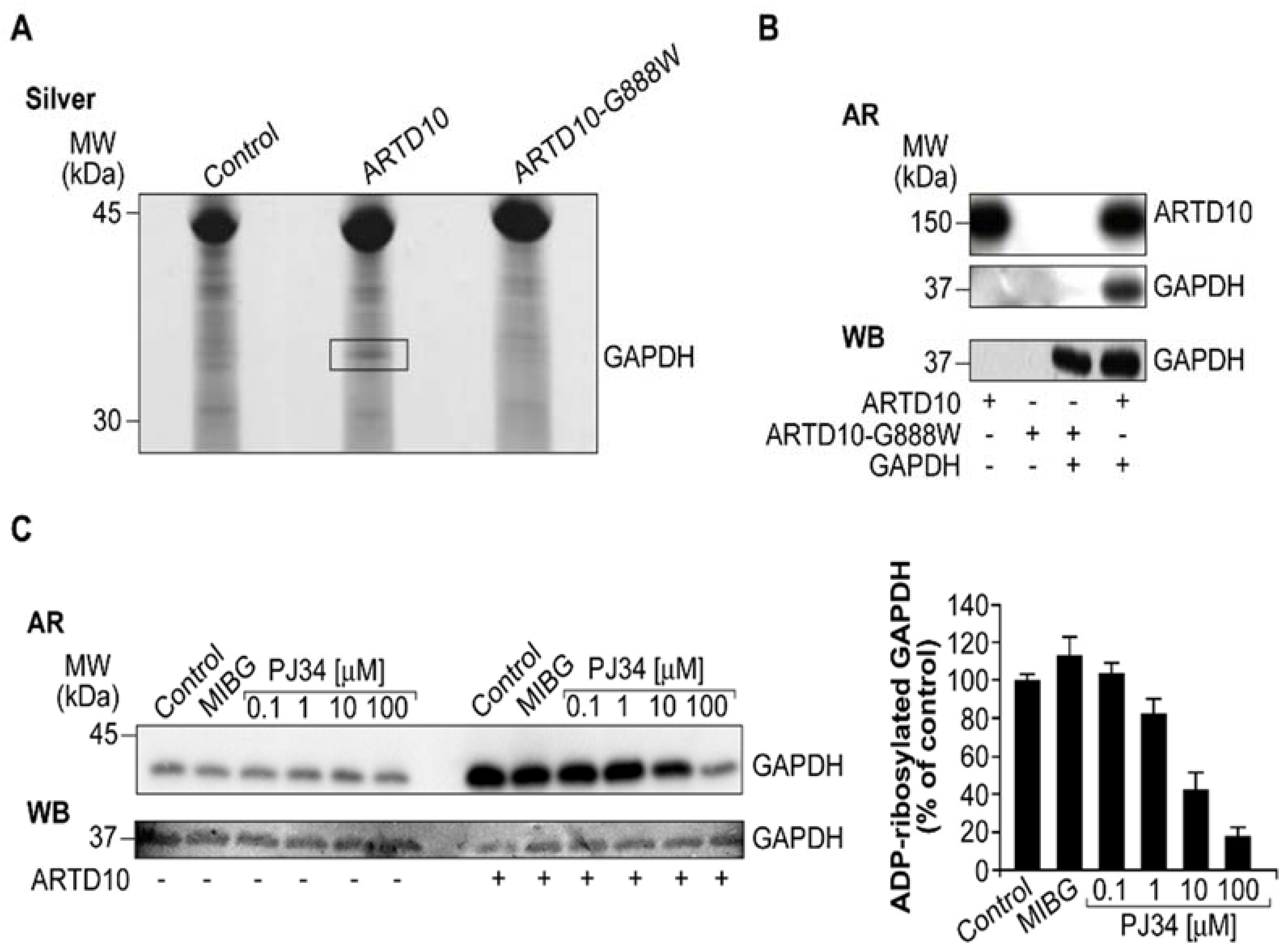
3.3. ARTD10 Catalyses Mono-ADP-Ribosylation of GAPDH
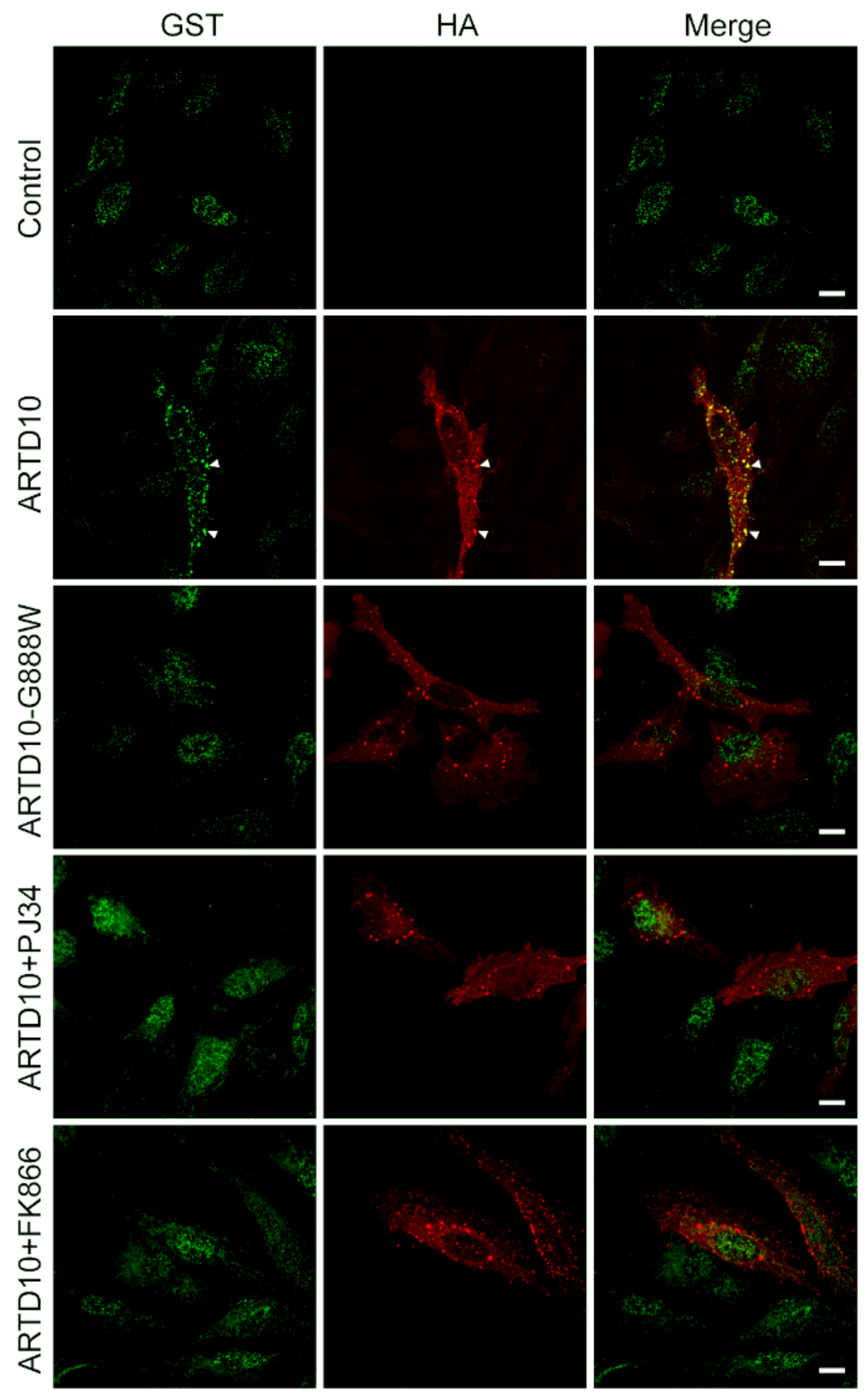
3.4. mAf1521 Allows Visualisation of the ARTD10 Membrane-Free Cell Bodies
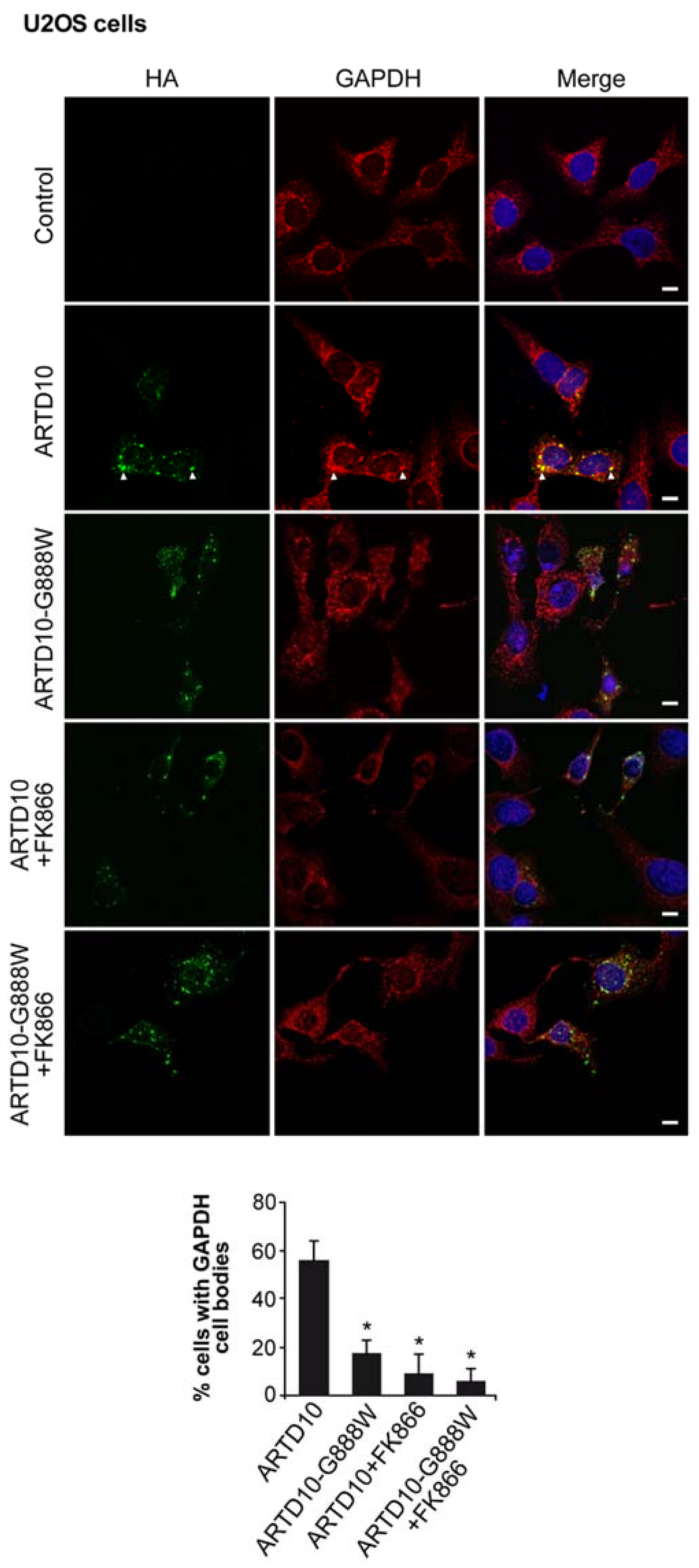
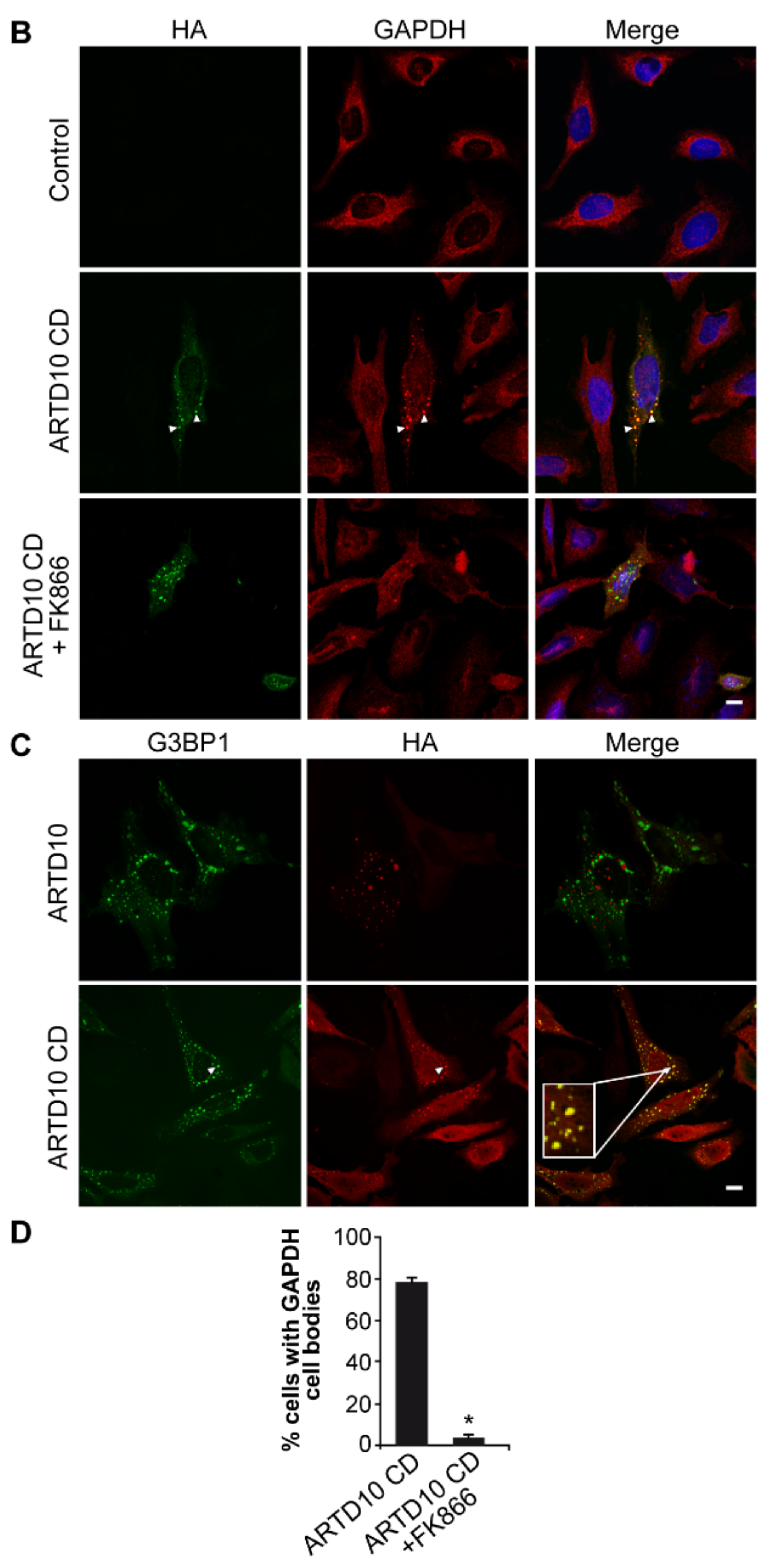
3.5. ARTD10-Dependent GAPDH Localisation in Membrane-Free Cell Bodies
3.6. The Catalytic Domain of ARTD10 Sequesters GAPDH into Stress Granules
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| ART | ADP-ribosyltransferase |
| GAPDH | glyceraldehyde-3-phosphate dehydrogenase |
| CHO cells | Chinese hamster ovary cells |
| HeLa cells | human cervix adenocarcinoma cells |
| GST | glutatione S-transferase |
| PAR | poly-ADP-ribose |
| PARP | PAR polymerase |
| PARG | poly (ADP-ribose) glycohydrolase |
| mART | mono-ADP-ribosyltransferase |
| ARH | ADP-ribosyl hydrolase |
References
- Di Girolamo, M.; Dani, N.; Stilla, A.; Corda, D. Physiological relevance of the endogenous mono(ADP-ribosyl)ation of cellular proteins. FEBS J. 2005, 272, 4565–4575. [Google Scholar] [CrossRef] [PubMed]
- Dani, N.; Jorge Moura Barbosa, A.; Del Rio, A.; Di Girolamo, M. ADP-Ribosylated Proteins as Old and New Drug Targets for Anticancer Therapy: The Example of ARF6. Curr. Pharm. Des. 2013, 19, 624–633. [Google Scholar] [CrossRef] [PubMed]
- Koch-Nolte, F.; Kernstock, S.; Mueller-Dieckmann, C.; Weiss, M.S.; Haag, F. Mammalian ADP-ribosyltransferases and ADP-ribosylhydrolases. Front. Biosci. 2008, 13, 6716–6729. [Google Scholar] [CrossRef] [PubMed]
- Hassa, P.O.; Hottiger, M.O. The diverse biological roles of mammalian PARPS, a small but powerful family of poly-ADP-ribose polymerases. Front. Biosci. 2008, 13, 3046–3082. [Google Scholar] [CrossRef] [PubMed]
- Gibson, B.A.; Kraus, W.L. New insights into the molecular and cellular functions of poly(ADP-ribose) and PARPs. Nat. Rev. Mol. Cell Biol. 2012, 13, 411–424. [Google Scholar] [CrossRef] [PubMed]
- Mangerich, A.; Bürkle, A. Pleiotropic cellular functions of PARP1 in longevity and aging: Genome maintenance meets inflammation. Oxid. Med. Cell. Longev. 2012, 2012, 321653. [Google Scholar] [CrossRef] [PubMed]
- Amé, J.C.; Spenlehauer, C.; de Murcia, G. The PARP superfamily. Bioessays 2004, 26, 882–893. [Google Scholar] [CrossRef] [PubMed]
- Corda, D.; di Girolamo, M. Functional aspects of protein mono-ADP-ribosylation. EMBO J. 2003, 22, 1953–1958. [Google Scholar] [CrossRef] [PubMed]
- Fabrizio, G.; Scarpa, E.S.; Di Girolamo, M. State of the art of protein mono-ADP-ribosylation: Biological role and therapeutic potential. Front. Biosci. 2015, 20, 405–430. [Google Scholar]
- Eno, G.H.; Ledford, B.E. ADP-ribosylation of the 78-kDa glucose-regulated protein during nutritional stress. Eur. J. Biochem. 1989, 186, 205–211. [Google Scholar]
- Ledford, B.E.; Leno, G.H. ADP-ribosylation of the molecular chaperone GRP78/BiP. Mol. Cell. Biochem. 1994, 138, 141–148. [Google Scholar] [CrossRef] [PubMed]
- Chambers, J.E.; Petrova, K.; Tomba, G.; Vendruscolo, M.; Ron, D. ADP ribosylation adapts an ER chaperone response to short-term fluctuations in unfolded protein load. J. Cell Biol. 2012, 198, 371–385. [Google Scholar] [CrossRef] [PubMed]
- Fabrizio, G.; Di Paola, S.; Stilla, A.; Giannotta, M.; Ruggiero, C.; Menzel, S.; Koch-Nolte, F.; Sallese, M.; Di Girolamo, M. ARTC1-mediated ADP-ribosylation of GRP78/BiP: A new player in endoplasmic-reticulum stress responses. Cell. Mol. Life Sci. 2015, 72, 1209–1225. [Google Scholar] [CrossRef] [PubMed]
- Lupi, R.; Corda, D.; Di Girolamo, M. Endogenous ADP-ribosylation of the G protein beta subunit prevents the inhibition of type 1 adenylyl cyclase. J. Biol. Chem. 2000, 275, 9418–9424. [Google Scholar] [CrossRef] [PubMed]
- Lupi, R.; Dani, N.; Dietrich, A.; Marchegiani, A.; Turacchio, S.; Berrie, C.P.; Moss, J.; Gierschik, P.; Corda, D.; Di Girolamo, M. Endogenous mono-ADP-ribosylation of the free Gbetagamma prevents stimulation of phosphoinositide 3 kinase-gamma and phospholipase C- beta2 and Is activated by G-protein-coupled receptors. Biochem. J. 2002, 367, 825–832. [Google Scholar] [CrossRef] [PubMed]
- Dani, N.; Mayo, E.; Stilla, A.; Marchegiani, A.; Di Paola, S.; Corda, D.; Di Girolamo, M. Mono-ADP-ribosylation of the G Protein βγ Dimer Is Modulated by Hormones and Inhibited by Arf6. J. Biol. Chem. 2011, 286, 5995–6005. [Google Scholar] [CrossRef] [PubMed]
- Herrero-Yraola, A.; Bakhit, S.M.; Franke, P.; Weise, C.; Schweiger, M.; Jorcke, D.; Ziegler, M. Regulation of glutamate dehydrogenase by reversible ADP-ribosylation in mitochondria. EMBO J. 2001, 20, 2404–2412. [Google Scholar] [CrossRef] [PubMed]
- Haigis, M.C.; Mostoslavsky, R.; Haigis, K.M.; Fahie, K.; Christodoulou, D.C.; Murphy, A.J.; Valenzuela, D.M.; Yancopoulos, G.D.; Karow, M.; Blander, G.; et al. SIRT4 inhibits glutamate dehydrogenase and opposes the effects of calorie restriction in pancreatic beta cells. Cell 2006, 126, 941–954. [Google Scholar] [CrossRef] [PubMed]
- Jwa, M.; Chang, P. PARP16 is a tail-anchored endoplasmic reticulum protein required for the PERK- and IRE1alpha-mediated unfolded protein response. Nat. Cell Biol. 2012, 14, 1223–1230. [Google Scholar] [CrossRef] [PubMed]
- Di Paola, S.; Micaroni, M.; Di Tullio, G.; Buccione, R.; Di Girolamo, M. PARP16/ARTD15 Is a Novel Endoplasmic-Reticulum-Associated Mono-ADP-ribosyltransferase that interacts with, and modifies Karyopherin-β1. PLoS ONE 2012, 7, e37352. [Google Scholar] [CrossRef] [PubMed]
- Di Girolamo, M.; Fabrizio, G.; Salvatore Scarpa, E.; Di Paola, S. NAD+-Dependent Enzymes at the Endoplasmic Reticulum. Curr. Top. Med. Chem. 2014, 13, 3001–3010. [Google Scholar] [CrossRef]
- Feijs, K.L.; Kleine, H.; Braczynski, A.; Forst, A.H.; Herzog, N.; Verheugd, P.; Linzen, U.; Kremmer, E.; Lüscher, B. ARTD10 substrate identification on protein microarrays: Regulation of GSK3beta by mono-ADP-ribosylation. Cell Commun. Signal. 2013, 11, 5. [Google Scholar] [CrossRef] [PubMed]
- Rosenthal, F.; Feijs, K.L.; Frugier, E.; Bonalli, M.; Forst, A.H.; Imhof, R.; Winkler, H.C.; Fischer, D.; Caflisch, A.; Hassa, P.O.; Lüscher, B. Macrodomain-containing proteins are new mono-ADP-ribosylhydrolases. Nat. Struct. Mol. Biol. 2013, 20, 502–507. [Google Scholar] [CrossRef] [PubMed]
- Yu, M.; Schreek, S.; Cerni, C.; Schamberger, C.; Lesniewicz, K.; Poreba, E.; Vervoorts, J.; Walsemann, G.; Grötzinger, J.; Kremmer, E.; et al. PARP-10, a novel Myc-interacting protein with poly(ADP-ribose) polymerase activity, inhibits transformation. Oncogene 2005, 24, 1982–1993. [Google Scholar] [CrossRef] [PubMed]
- Kleine, H.; Herrmann, A.; Lamark, T.; Forst, A.H.; Verheugd, P.; Lüscher-Firzlaff, J.; Lippok, B.; Feijs, K.L.; Herzog, N.; Kremmer, E.; et al. Dynamic subcellular localization of the mono-ADP-ribosyltransferase ARTD10 and interaction with the ubiquitin receptor p62. Cell Commun. Signal. 2012, 10, 28. [Google Scholar] [CrossRef] [PubMed]
- Cohen, P.; Frame, S. The renaissance of GSK3. Nat. Rev. Mol. Cell Biol. 2001, 2, 769–776. [Google Scholar] [CrossRef] [PubMed]
- Wu, D.; Pan, W. GSK3: A multifaceted kinase in Wnt signaling. Trends Biochem. Sci. 2010, 35, 161–168. [Google Scholar] [CrossRef] [PubMed]
- Verheugd, P.; Forst, A.H.; Milke, L.; Herzog, N.; Feijs, K.L.; Kremmer, E.; Kleine, H.; Lüscher, B. Regulation of NF-kappaB signalling by the mono-ADP-ribosyltransferase ARTD10. Nat. Commun. 2013, 4, 1683. [Google Scholar] [CrossRef] [PubMed]
- Hayden, M.S.; Ghosh, S. Signaling to NF-κB. Genes Dev. 2004, 18, 2195–2224. [Google Scholar] [CrossRef] [PubMed]
- Bruns, G.A.; Gerald, P.S. Human glyceraldehyde-3-phosphate dehydrogenase in man-rodent somatic cell hybrids. Science 1976, 192, 54–56. [Google Scholar] [CrossRef] [PubMed]
- Sirover, M.A. New insights into an old protein: The functional diversity of mammalian glyceraldehyde-3-phosphate dehydrogenase. Biochim. Biophys. Acta 1999, 1432, 159–184. [Google Scholar] [CrossRef]
- Seidler, N.W. Functional diversity. Adv. Exp. Med. Biol. 2013, 985, 103–147. [Google Scholar] [PubMed]
- Du, X.; Matsumura, T.; Edelstein, D.; Rossetti, L.; Zsengellér, Z.; Szabó, C.; Brownlee, M. Inhibition of GAPDH activity by poly(ADP-ribose) polymerase activates three major pathways of hyperglycemic damage in endothelial cells. J. Clin. Investig. 2003, 112, 1049–1057. [Google Scholar] [CrossRef] [PubMed]
- Long, C.A.; Boloum, V.; Albadawi, H.; Tsai, S.; Yoo, H.J.; Oklu, R.; Goldman, M.H.; Watkins, M.T. Poly-ADP-ribose-polymerase inhibition ameliorates hind limb ischemia reperfusion injury in a murine model of type 2 diabetes. Ann. Surg. 2013, 258, 1087–1095. [Google Scholar] [CrossRef] [PubMed]
- Deveze-Alvarez, M.; García-Soto, J.; Martínez-Cadena, G. Glyceraldehyde-3-phosphate dehydrogenase is negatively regulated by ADP-ribosylation in the fungus Phycomyces blakesleeanus. Microbiology 2001, 147, 2579–2584. [Google Scholar] [CrossRef] [PubMed][Green Version]
- De Matteis, M.A.; Di Girolamo, M.; Colanzi, A.; Pallas, M.; Di Tullio, G.; McDonald, L.J.; Moss, J.; Santini, G.; Bannykh, S.; Corda, D. Stimulation of endogenous ADP-ribosylation by brefeldin A. Proc. Natl. Acad. Sci. USA 1994, 91, 1114–1118. [Google Scholar] [CrossRef] [PubMed]
- Di Girolamo, M.; Silletta, M.G.; De Matteis, M.A.; Braca, A.; Colanzi, A.; Pawlak, D.; Rasenick, M.M.; LuINI, A.L.; Corda, D. Evidence that the 50-kDa substrate of brefeldin A-dependent ADP- ribosylation binds GTP and is modulated by the G-protein beta gamma subunit complex. Proc. Natl. Acad. Sci. USA 1995, 92, 7065–7069. [Google Scholar] [CrossRef] [PubMed]
- Herzog, N.; Hartkamp, J.D.; Verheugd, P.; Treude, F.; Forst, A.H.; Feijs, K.L.; Lippok, B.E.; Kremmer, E.; Kleine, H.; Lüscher, B. Caspase-dependent cleavage of the mono-ADP-ribosyltransferase ARTD10 interferes with its pro-apoptotic function. FEBS J. 2013, 280, 1330–1343. [Google Scholar] [CrossRef] [PubMed]
- Stilla, A.; Di Paola, S.; Dani, N.; Krebs, C.; Arrizza, A.; Corda, D.; Haag, F.; Koch-Nolte, F.; Di Girolamo, M. Characterisation of a novel glycosylphosphatidylinositol-anchored mono-ADP-ribosyltransferase isoform in ovary cells. Eur. J. Cell Biol. 2011, 90, 665–677. [Google Scholar] [CrossRef] [PubMed]
- Dani, N.; Stilla, A.; Marchegiani, A.; Tamburro, A.; Till, S.; Ladurner, A.G.; Corda, D.; Di Girolamo, M. Combining affinity purification by ADP-ribose-binding macro domains with mass spectrometry to define the mammalian ADP-ribosyl proteome. Proc. Natl. Acad. Sci. USA 2009, 106, 4243–4248. [Google Scholar] [CrossRef] [PubMed]
- Polishchuk, R.S.; Mironov, A.A. Correlative video light/electron microscopy. Curr. Protoc. Cell Biol. 2001. [Google Scholar] [CrossRef] [PubMed]
- Polishchuk, V.P.; Tyvonchuk, T.P.; Rengevich, I.; Beketov, G.V.; Budzanivskaia, I.G.; Boĭko, A.L. Use of atomic force microscopy to study the morphology and structure of viruses. Mikrobiol. Zhurnal 2000, 62, 40–43. [Google Scholar]
- Kleine, H.; Poreba, E.; Lesniewicz, K.; Hassa, P.O.; Hottiger, M.O.; Litchfield, D.W.; Shilton, B.H.; Lüscher, B. Substrate-assisted catalysis by PARP10 limits its activity to mono-ADP-ribosylation. Mol. Cell 2008, 32, 57–69. [Google Scholar] [CrossRef] [PubMed]
- Loesberg, C.; Smets, L.A. Meta-iodobenzylguanidine (MIBG), a novel high-affinity substrate for cholera toxin that interferes with cellular mono(ADP-ribosylation). Biochim. Biophys. Acta 1990, 1037, 92–99. [Google Scholar] [CrossRef]
- Yates, S.P.; Taylor, P.L.; Jørgensen, R.; Ferraris, D.; Zhang, J.; Andersen, G.R.; Merrill, A.R. Structure-function analysis of water-soluble inhibitors of the catalytic domain of exotoxin A from Pseudomonas aeruginosa. Biochem. J. 2005, 385, 667–675. [Google Scholar] [CrossRef] [PubMed]
- Forst, A.H.; Karlberg, T.; Herzog, N.; Thorsell, A.G.; Gross, A.; Feijs, K.L.; Verheugd, P.; Kursula, P.; Nijmeijer, B.; Kremmer, E.; et al. Recognition of mono-ADP-ribosylated ARTD10 substrates by ARTD8 macrodomains. Structure 2013, 21, 462–475. [Google Scholar] [CrossRef] [PubMed]
- Leung, A.K.; Todorova, T.; Ando, Y.; Chang, P. Poly(ADP-ribose) regulates post-transcriptional gene regulation in the cytoplasm. RNA Biol. 2012, 9, 542–548. [Google Scholar] [CrossRef] [PubMed]
- Leung, A.K.; Vyas, S.; Rood, J.E.; Bhutkar, A.; Sharp, P.A.; Chang, P. Poly(ADP-ribose) regulates stress responses and microRNA activity in the cytoplasm. Mol. Cell 2011, 42, 489–499. [Google Scholar] [CrossRef] [PubMed]
- Isabelle, M.; Gagné, J.P.; Gallouzi, I.E.; Poirier, G.G. Quantitative proteomics and dynamic imaging reveal that G3BP-mediated stress granule assembly is poly(ADP-ribose)-dependent following exposure to MNNG-induced DNA alkylation. J. Cell Sci. 2012, 125, 4555–4566. [Google Scholar] [CrossRef] [PubMed]

© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mayo, E.; Fabrizio, G.; Scarpa, E.S.; Stilla, A.; Dani, N.; Chiacchiera, F.; Kleine, H.; Attanasio, F.; Lüscher, B.; Di Girolamo, M. ARTD10/PARP10 Induces ADP-Ribosylation of GAPDH and Recruits GAPDH into Cytosolic Membrane-Free Cell Bodies When Overexpressed in Mammalian Cells. Challenges 2018, 9, 22. https://doi.org/10.3390/challe9010022
Mayo E, Fabrizio G, Scarpa ES, Stilla A, Dani N, Chiacchiera F, Kleine H, Attanasio F, Lüscher B, Di Girolamo M. ARTD10/PARP10 Induces ADP-Ribosylation of GAPDH and Recruits GAPDH into Cytosolic Membrane-Free Cell Bodies When Overexpressed in Mammalian Cells. Challenges. 2018; 9(1):22. https://doi.org/10.3390/challe9010022
Chicago/Turabian StyleMayo, Emilia, Gaia Fabrizio, Emanuele Salvatore Scarpa, Annalisa Stilla, Nadia Dani, Fulvio Chiacchiera, Henning Kleine, Francesca Attanasio, Bernhard Lüscher, and Maria Di Girolamo. 2018. "ARTD10/PARP10 Induces ADP-Ribosylation of GAPDH and Recruits GAPDH into Cytosolic Membrane-Free Cell Bodies When Overexpressed in Mammalian Cells" Challenges 9, no. 1: 22. https://doi.org/10.3390/challe9010022
APA StyleMayo, E., Fabrizio, G., Scarpa, E. S., Stilla, A., Dani, N., Chiacchiera, F., Kleine, H., Attanasio, F., Lüscher, B., & Di Girolamo, M. (2018). ARTD10/PARP10 Induces ADP-Ribosylation of GAPDH and Recruits GAPDH into Cytosolic Membrane-Free Cell Bodies When Overexpressed in Mammalian Cells. Challenges, 9(1), 22. https://doi.org/10.3390/challe9010022




