Diversity of Epiphytic Bacterial Communities Associated with Adriatic Fucacean Species Phycosphere
Abstract
1. Introduction
2. Materials and Methods
2.1. Biological Sampling
2.2. Lipid Content and Fatty Acid Composition of Fucales
2.3. Bacterial Identification and Phycosphere Community Structure Determination
2.4. Statistical Analysis
3. Results
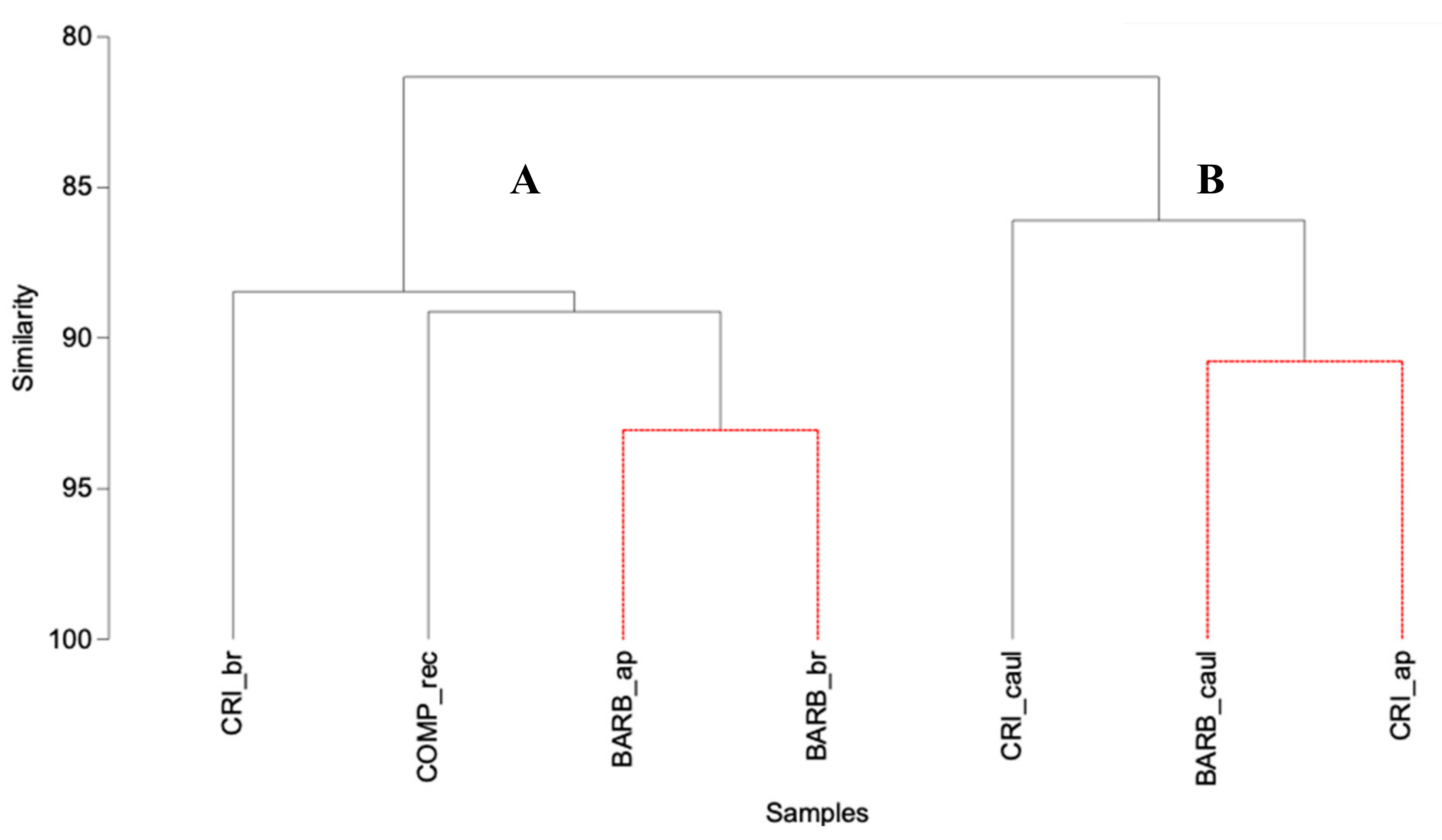
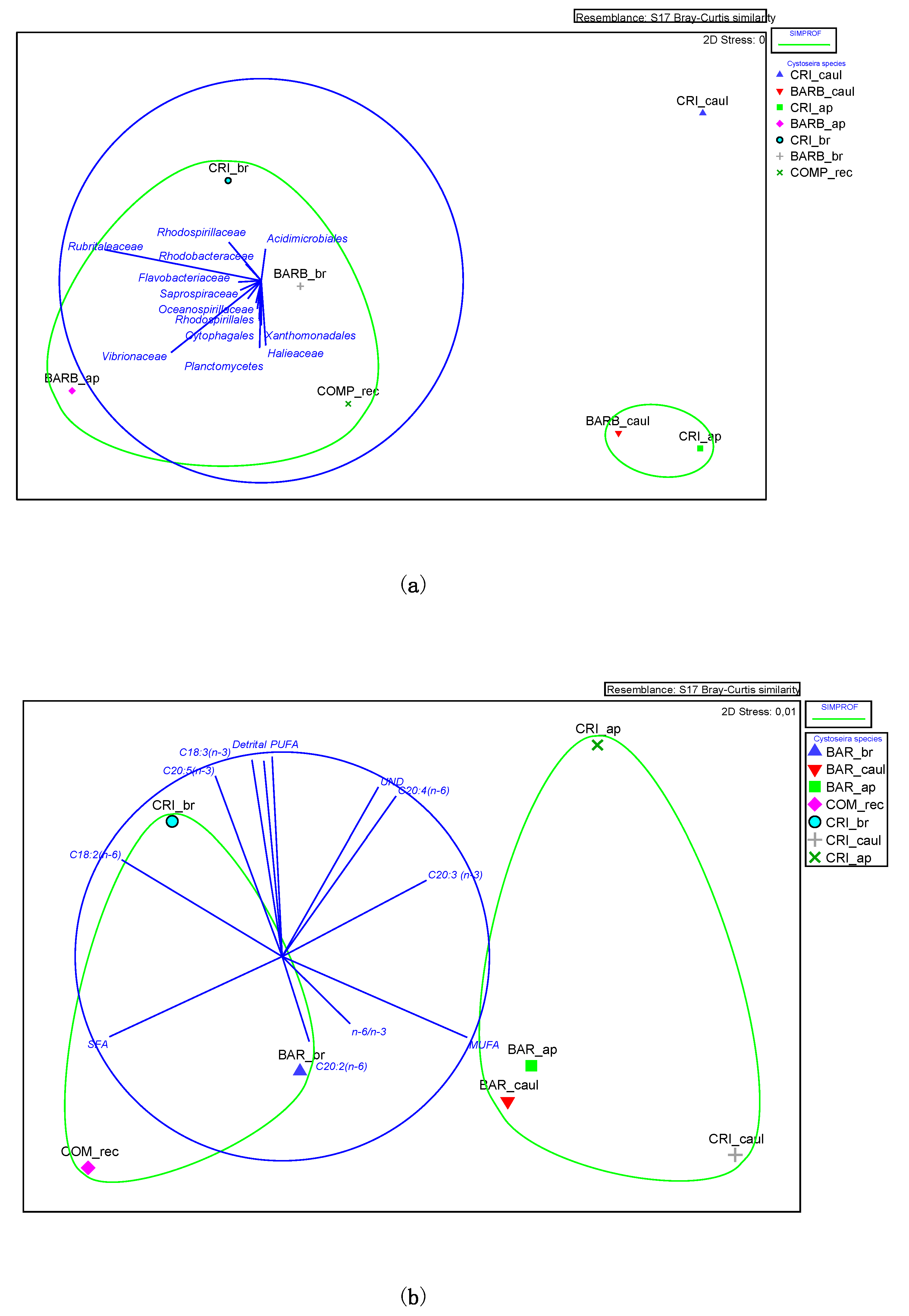
3.1. Lipid Composition and Fatty Acid Profile of the Fucales Thalli
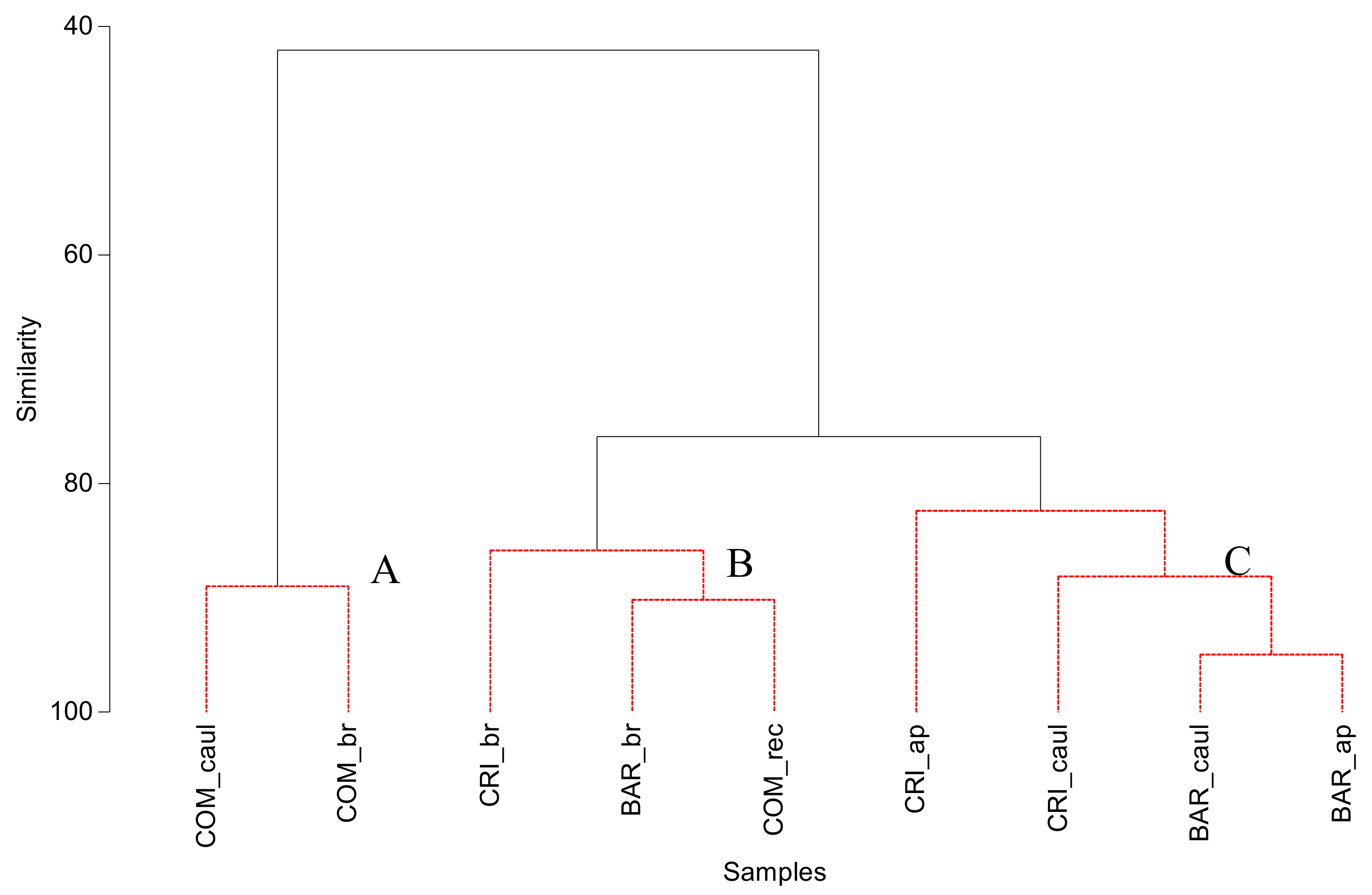
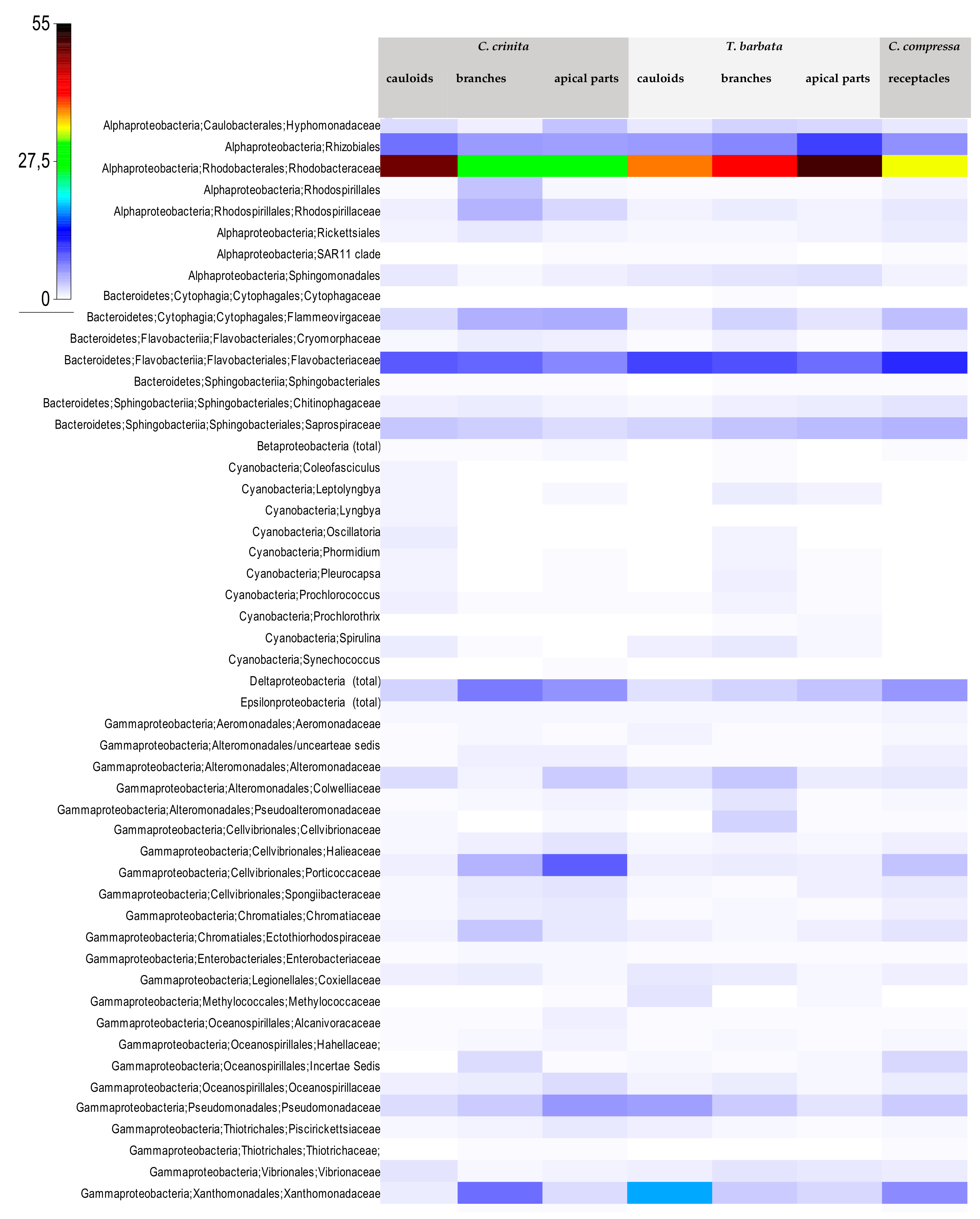
3.2. Bacterial Community Structure of the Fucacean Phycosphere
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Mangialajo, L.; Gianni, F.; Airoldi, L.; Bartolini, F.; Francour, P.; Meinesz, A.; Thiabaut, T.; Ballesteros, E. Conservation and restoration of Cystoseira forests in the Mediteranean Sea: The role of marine protected areas. Rapp. Comm. Int. Mer. Medit. 2013, 40, 621. [Google Scholar]
- Iveša, L.; Đakovac, T.; Devescovi, M. Long–term fluctuations in Cystoseira population along the west Istrian Coast (Croatia) related to eutrophication patterns in the northern Adriatic Sea. Mar. Pollut. Bull. 2016, 106, 162–173. [Google Scholar] [CrossRef]
- Coll, M.; Piroddi, C.; Steenbeek, J.; Kaschner, K.; Lasram, F.B.R.; Aguzzi, J.; Ballesteros, E.; Bianchi, C.N.; Corbera, J.; Dailianis, T.; et al. The Biodiversity of the Mediterranean Sea: Estimates, Patterns, and Threats. PLoS ONE 2010, 5, e11842. [Google Scholar] [CrossRef]
- Nasrolahi, A.; Stratil, S.B.; Jacob, K.J.; Wahl, M. A protective coat of microorganisms on macroalgae: Inhibitory effects of bacterial biofilms and epibiotic microbial assemblages on barnacle attachment. FEMS Microbiol. Ecol. 2012, 81, 583–595. [Google Scholar] [CrossRef]
- Wahl, M.; Shahnaz, L.; Dobrestsov, S.; Saha, M.; Symanowski, F.; David, K.; Lachnit, T.; Vasel, M.; Weinberger, F. Ecology and antifouling resistance in the bladder wrack Fucusvesiculosus: Patters of micriofouling and antimicrobial protection. Mar. Ecol. Prog. Ser. 2010, 411, 33–48. [Google Scholar] [CrossRef]
- Lachnit, T.; Wahl, M.; Harder, T. Isolated thallus-associated compounds from the macroalgae Fucusvesiculosus mediate bacterial surface colonization in the field similar to that on the natural algae. Biofouling 2010, 26, 247–255. [Google Scholar] [CrossRef] [PubMed]
- Lachnit, T.; Meske, D.; Wahl, M.; Harder, T.; Schmitz, R. Epibacterial community patterns on marine macroalgae are host—Specific but temporally variable. Environ. Microbiol. 2011, 13, 655–665. [Google Scholar] [CrossRef] [PubMed]
- Iveša, L.; Lyons, D.M.; Devescovi, M. Assessment of the ecological status of north-eastern Adriatic coastal waters (Istria, Croatia) using macroalgal assemblages for the European Union Water Framework Directive. Aquat. Conserv. Mar. Freshw. Ecosyst. 2009, 19, 14–23. [Google Scholar] [CrossRef]
- Valdazo, J.; Ascenisión, V.R.; Espino, F.; Haroun, R.; Tuya, F. Massive decline of Cystoseiraabies—Marina forests in Gran Canaria Island (Canary Islands, eastern Atlantic). Sci. Mar. 2017, 81, 499–507. [Google Scholar] [CrossRef]
- Mačić, V.; Svirčev, Z. Macroepiphytes on Cystoseira species (Phaeophyceae) on the coast of Montenegro. Fresenius Environ. Bull. 2014, 23, 29–34. [Google Scholar]
- Egan, S.; Harder, T.; Burke, C.; Steinberg, P.; Kjelleberg, S.; Thomas, T. The seaweed holobiont: Understanding seaweed-bacteria interactions. FEMS Microbiol. Rev. 2013, 37, 462–476. [Google Scholar] [CrossRef] [PubMed]
- Huggett, M.J.; McMahon, K.; Bernasconi, R. Future warming and acidification result in multiple ecological impacts to a temperate coralline alga. Environ. Microbiol. 2018, 20, 2769–2782. [Google Scholar] [CrossRef] [PubMed]
- Dobretsov, S.; Teplitski, M.; Paul, V.J. Quorum sensing in the marine environment and its relationship to biofouling. Biofouling 2009, 25, 413–427. [Google Scholar] [CrossRef] [PubMed]
- Piazza, V.; Roussis, V.; Garaventa, F.; Greco, G.; Smyrniotopoulos, V.; Vagias, C.; Faimali, M. Terpenes from the red alga Sphaerococcuscoronopifolius inhibit the settlement of barnacles. Mar. Biotechnol. 2011, 13, 764–772. [Google Scholar] [CrossRef]
- Potin, P.; Bouarab, K.; Salaün, J.P.; Pohnert, G.; Kloareg, B. Biotic interations of marine algae. Curr. Opin. Plant. Biol. 2002, 5, 308–317. [Google Scholar] [CrossRef]
- Bouarab, K.; Adas, F.; Gaquerel, E.; Kloareg, B.; Salaün, J.P.; Potin, P. The innate immunity of a marine red alga involves oxylipins from both the eicosanoid and octadecanoid pathways. Plant Physiol. 2004, 135, 1838–1848. [Google Scholar] [CrossRef]
- Küpper, F.C.; Gaquerel, E.; Cosse, A.; Adas, F.; Peters, A.F.; Müller, D.G.; Kloareg, B.; Salaün, J.-P.; Potin, P. Free fatty acids and methyl jasmonate trigger defense reactions in Laminaria dignitata. Plant. Cell. Physiol. 2009, 50, 789–800. [Google Scholar] [CrossRef]
- Duarte, B.; Martins, I.; Rosa, R.; Matos, A.R.; Roleda, M.Y.; Reusch, T.B.H.; Engelen, A.H.; Serrao, E.A.; Pearson, G.A.; Marques, J.C.; et al. Climate change Impacts on Seagrass Meadows and Macroalgal Forests: An Integrative Perspective on Acclimation and Adaptation Potential. Front. Mar. Sci. 2018. [Google Scholar] [CrossRef]
- Vizetto-Duarte, C.; Pereira, H.; Bruno de Suosa, C.; Rauter, A.P.; Albericio, F.; Custódio, L.; Barreira, L.; Varela, J. Fatty acid profile of different species of algae of the Cystoseira genus: A nutraceutical perspective. Nat. Prod. Res. 2015, 29, 13. [Google Scholar] [CrossRef]
- Burke, C.; Kjelleberg, S.; Thomas, T. Selective Extraction of Bacterial DNA from the Surfaces of Macroalgae. Appl. Environ. Microbiol. 2008, 75, 252–256. [Google Scholar] [CrossRef]
- De Rosa, M.; Gambacorta, A. The lipids of archaebacteria. Prog. Lipid Res. 1988, 27, 153–175. [Google Scholar] [CrossRef]
- Klindworth, A.; Pruesse, E.; Schweer, T.; Peplies, J.; Quast, C.; Horn, M.; Glöckner, F., O. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 2013, 41, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Schloss, P.D.; Westcott, S.L.; Ryabin, T.; Hall, J.R.; Hartmann, M.; Hollister, E.B.; Lesniewski, R.A.; Oakley, B.B.; Parks, D.H.; Robinson, C.J.; et al. Introducing mothur: Open-Source, Platform-Independent, Community-Supported Software for Describing and Comparing Microbial Communities. Appl. Environ. Microbiol. 2009, 75, 7537–7541. [Google Scholar] [CrossRef] [PubMed]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glöckner, F.O. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 2013, 41, 590–596. [Google Scholar] [CrossRef] [PubMed]
- Pruesse, E.; Peplies, J.; Glöckner, F.O. SINA: Accurate high-throughput multiple sequence alignment of ribosomal RNA genes. Bioinformatics 2012, 28, 1823–1829. [Google Scholar] [CrossRef]
- Li, W.; Godzik, A. Cd-hit: A fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinfomatics Appl. Note 2006, 22, 1658–1659. [Google Scholar] [CrossRef]
- Camacho, C.; Coulouris, G.; Avagyan, V.; Ma, N.; Papadopoulos, J.S.; Bealer, K.; Madden, T.L. BLAST+: Architecture and applications. BMC Bioinform. 2009, 10, 421. [Google Scholar] [CrossRef]
- Kumari, P.; Kumar, M.; Gupta, V.; Reddy, C.R.K.; Jha, B. Tropical marine macroalgae as potential sources of nutritionally important PUFAs. Food Chem. 2010, 120, 749–757. [Google Scholar] [CrossRef]
- Pereira, H.; Barreira, L.; Figueiredo, F.; Custodio, L.; Vizetto-Durte, C.; Polo, C.; Rešek, E.; Engelen, A.; Varela, J. Polyunsaturated Fatty Acids of Marine Macroalgae: Potential for Nutritional and Pharmaceutical Applications. Mar. Drugs 2012, 10, 1920. [Google Scholar] [CrossRef]
- Selvarajan, R.; Sibanda, T.; Venkatachalam, S.; Ogola, H.J.O.; Obieze, C.C.; Msagati, T.A. Distribution, Interaction and Functional Profiles of Epiphytic Bacterial Communities from the Rocky Intertidal Seaweeds, South Africa. Sci. Rep. 2019, 9, 1–13. [Google Scholar] [CrossRef]
- Silva, G.; Pereira, R.B.; Valentão, P.; Andrade, P.B.; Sousa, C. Distinct fatty acid profile of ten brown macroalgae. Rev. Bras. Farm. 2013, 23, 608–613. [Google Scholar] [CrossRef]
- White, D.C.; Bobbie, R.J.; Nickels, J.S.; Fazio, S.D.; Davis, W.M. Nonselective biochemical methods for the determination of fungal mass and community structure in estuarine detrital microflora. Bot. Mar. 1980, 23, 239–250. [Google Scholar]
- Simon, M.; Scheuner, C.; Meier-Kolthoff, J.P.; Brinkhoff, T.; Wagner-Döbler, I.; Ulbrich, M.; Klenk, H.-P.; Schomburg, D.; Petersen, J.; Göker, M. Phylogenomics of Rhodobacteraceae reveals evolutionary adaptation to marine and non-marine habitats. ISME J. 2017, 11, 1483–1499. [Google Scholar] [CrossRef] [PubMed]
- Elifantz, H.; Horn, G.; Ayon, M.; Cohen, Y.; Minz, D. Rhodobacteraceaeare the key members of the microbial community of the initial biofilm formed in Eastern Mediterranean coastal seawater. FEMS Microbiol. Ecol. 2013, 85, 348–357. [Google Scholar] [CrossRef]
- Alvarado, P.; Huang, Y.; Wang, J.; Garrido, I.; Leiva, S. Phylogeny and bioactivity of epiphytic Gram-positive bacteria isolated from three co-occurring antarctic macroalgae. Antonie van Leeuwenhoek 2018, 111, 1543–1555. [Google Scholar] [CrossRef]
- Alamsjah, M.A.; Hirao, S.; Ishibashi, F.; Oda, T.; Fujita, Y. Algicidal activity of polyunsaturated fatty acids derived from Ulva fasciata and U. pertusa (Ulvaceae, Chlorophyta) on phytoplankton. J. Appl. Phycol. 2008, 20, 713–720. [Google Scholar] [CrossRef]
- Raphael, L. Quorum Sensing in Marine Biofilms and Environments; Quorum Sensing; Academic Press: Cambridge, MA, USA, 2019; pp. 55–96. [Google Scholar]
- Mancuso, P.; D’Hondt, S.; Willems, A.; Airoldi, L.; De Clerck, O. Diversity and Temporal Dynamics of the Epiphytic Bacterial Communities Associated with the Canopy-Forming Seaweed Cystoseira compressa (Esper) Gerloff and Nizamuddin. Front. Microbiol. 2016, 7, 476. [Google Scholar] [CrossRef]
- Fernandes, N.; Steinberg, P.; Rusch, D.; Kjelleberg, S.; Thomas, T. Community Structure and Functional Gene Profile of Bacteria on Healthy and Diseased Thalli of the Red Seaweed Delisea pulchra. PLoS ONE 2012, 7, e50854. [Google Scholar] [CrossRef]
- Zozaya-Valdes, E.; Egan, S.; Thomas, T. A comprehensive analysis of the microbial communities of healthy and diseased marine macroalgae and the detection of known and potential bacterial pathogens. Front. Microbiol. 2015, 6, 146. [Google Scholar] [CrossRef]
- Bulleri, F.; Eriksson, B.K.; Queirós, A.; Airoldi, L.; Arenas, F.; Arvanitidis, C.; Bouma, T.J.; Crowe, T.P.; Davoult, D.; Guizien, K.; et al. Harnessing positive species interactions as a tool against climate-driven loss of coastal biodiversity. PLoS Biol. 2018, 16, e2006852. [Google Scholar] [CrossRef]
- Albakosh, M.A.; Naidoo, R.K.; Kirby, B.; Bauer, R. Identification of epiphytic bacterial communities associated with the brown alga Splachnidium rugosum. Environ. Boil. Fishes 2015, 28, 1891–1901. [Google Scholar] [CrossRef]
- Muñoz, C.; Hidalgo, C.; Zapata, M.; Jeison, D.; Riquelme, C.; Rivas, M. Use of Cellulolytic Marine Bacteria for Enzymatic Pretreatment in Microalgal Biogas Production. Appl. Environ. Microbiol. 2014, 80, 4199–4206. [Google Scholar] [CrossRef] [PubMed]
| Carpodesmia crinita | ||||||
|---|---|---|---|---|---|---|
| Fatty Acid Methyl Ester | Cauloid | Branches | Apical Parts | |||
| AV | ± sd | AV | ± sd | AV | ± sd | |
| C14:1 | - | - | - | - | - | - |
| C14:0 | 16.73 | 0.05 | 8.97 | 2.74 | 8.56 | 0.78 |
| C15:0 iso | 0.11 | 0.17 | 0.32 | 0.07 | 0.34 | 0.12 |
| C15:0 ante | 0.10 | |||||
| C15:0 | 0.79 | 0.09 | 1.00 | 0.16 | 0.75 | 0.01 |
| C16:1 | 5.03 | 1.10 | 3.63 | 0.28 | 3.03 | 0.31 |
| C16:0 | 24.75 | 0.42 | 35.90 | 9.25 | 23.37 | 9.91 |
| C17:0 iso | - | - | 0.22 | - | 0.31 | 0.12 |
| C17:0 ante | - | - | 0.09 | - | 0.05 | - |
| C17:1 | 0.14 | 0.11 | 0.19 | 0.07 | 0.54 | 0.66 |
| C17:0 | 0.38 | 0.13 | 0.47 | 0.07 | 0.53 | 0.21 |
| C18:3(n-3) | 0.48 | 0.36 | 2.70 | 2.20 | 3.53 | 1.07 |
| C18:2(n-6) | 0.74 | 0.56 | 6.62 | 9.53 | 2.20 | 1.29 |
| C18:1 | 30.40 | 3.23 | 14.08 | 12.13 | 22.58 | 4.95 |
| C18:0 | 1.85 | 0.78 | 3.66 | 0.81 | 3.75 | 3.55 |
| C20:4(n-6) | 8.38 | 1.46 | 8.51 | 3.98 | 13.82 | 5.57 |
| C20:5(n-3) | 2.00 | 0.60 | 5.46 | 3.63 | 5.39 | 2.17 |
| C20:3(n-3) | 2.83 | 0.78 | 1.88 | 1.01 | 3.25 | 1.58 |
| C20:2(n-6) | 0.66 | 0.29 | 0.45 | 0.04 | 0.74 | 0.11 |
| C20:1 | 0.09 | 0.14 | 0.03 | 0.05 | 0.29 | 0.15 |
| C20:0 | 1.38 | 1.28 | 1.01 | 0.48 | 0.95 | 0.39 |
| C22:4(n-6) | - | - | - | - | 0.09 | - |
| C22:6(n-3) | - | - | 0.13 | 0.20 | 0.27 | 0.37 |
| C22:4(n-3) | 0.09 | - | - | - | 0.10 | 0.09 |
| C22:2(n-6) | - | - | - | - | 0.10 | - |
| C22:1 | 0.32 | 0.49 | - | - | - | - |
| C22:0 | 0.22 | 0.28 | 0.64 | 0.22 | 0.54 | 0.33 |
| SFA | 46.21 | 0.52 | 52.37 | 13.66 | 39.14 | 12.41 |
| MUFA | 35.99 | 3.59 | 17.93 | 11.74 | 26.44 | 4.45 |
| PUFA | 15.18 | 4.04 | 25.75 | 1.52 | 29.49 | 9.67 |
| Bacterial | 1.42 | 0.27 | 2.38 | 0.23 | 2.52 | 1.11 |
| n-6/n-3 | 3.34 | - | 2.67 | 1.74 | 1.91 | 0.26 |
| UND | 2.73 | 0.19 | 2.92 | 2.22 | 4.32 | 2.93 |
| Total lipids (mg/g d.w.) | 11.60 | 2.03 | 10.49 | 1.90 | 10.22 | 3.20 |
| Treptacantha barbata | ||||||
|---|---|---|---|---|---|---|
| Fatty Acid Methyl Ester | Cauloid | Branches | Apical Parts | |||
| AV | ± sd | AV | ± sd | AV | ± sd | |
| C14:1 | - | - | - | - | - | - |
| C14:0 | 9.44 | 2.49 | 6.98 | 0.68 | 8.80 | 1.70 |
| C15:0 iso | 0.07 | 0.07 | 0.11 | 0.15 | 0.16 | 0.05 |
| C15:0 ante | 0.02 | 0.03 | - | - | - | - |
| C15:0 | 0.75 | 0.29 | 0.76 | 0.21 | 0.54 | 0.13 |
| C16:1 | 2.39 | 0.39 | 2.25 | 0.30 | 1.47 | 0.46 |
| C16:0 | 32.01 | 10.35 | 39.82 | 1.91 | 29.36 | 4.89 |
| C17:0 iso | 0.14 | 0.13 | - | - | - | - |
| C17:0 ante | 0.03 | 0.03 | - | - | - | - |
| C17:1 | 0.19 | 0.10 | 0.59 | 0.49 | 0.35 | 0.18 |
| C17:0 | 0.39 | 0.18 | 0.21 | 0.14 | 0.25 | 0.08 |
| C18:3(n-3) | 0.72 | 0.48 | 1.61 | 0.22 | 1.19 | 0.79 |
| C18:2(n-6) | 1.96 | 2.20 | 2.84 | 0.82 | 0.93 | 0.48 |
| C18:1 | 25.25 | 11.11 | 22.75 | 2.67 | 25.12 | 9.16 |
| C18:0 | 2.21 | 1.07 | 1.60 | 0.36 | 1.05 | 0.29 |
| C20:4(n-6) | 7.92 | 4.54 | 7.33 | 0.54 | 9.13 | 5.51 |
| C20:5(n-3) | 1.53 | 1.53 | 3.45 | 1.01 | 1.79 | 1.19 |
| C20:3(n-3) | 2.32 | 1.88 | 1.08 | 0.72 | 1.85 | 0.94 |
| C20:2(n-6) | 0.69 | 0.35 | 0.67 | 0.17 | 0.67 | 0.28 |
| C20:1 | - | - | - | - | 0.17 | 0.05 |
| C20:0 | 0.75 | 0.24 | 0.82 | 0.28 | 0.65 | 0.34 |
| C22:4(n-6) | - | - | - | - | - | - |
| C22:6(n-3) | - | - | - | - | - | - |
| C22:4(n-3) | 0.03 | 0.05 | - | - | - | - |
| C22:2(n-6) | - | - | - | - | - | - |
| C22:1 | - | - | - | - | - | - |
| C22:0 | 0.43 | 0.14 | 0.29 | 0.19 | 0.19 | 0.06 |
| SFA | 46.23 | 11.09 | 50.58 | 3.18 | 41.00 | 7.52 |
| MUFA | 27.83 | 11.57 | 25.59 | 2.60 | 27.11 | 9.69 |
| PUFA | 15.17 | 6.60 | 16.98 | 0.99 | 15.57 | 9.20 |
| Bacterial | 1.59 | 0.44 | 1.67 | 0.49 | 1.30 | 0.26 |
| n-6/n-3 | 2.21 | 0.19 | 2.06 | 0.75 | 2.47 | 0.37 |
| UND | 2.04 | 1.19 | 1.74 | 0.19 | 2.37 | 1.35 |
| Total lipids (mg/g d.w.) | 6.31 | 0.42 | 7.23 | 1.35 | 7.49 | 2.59 |
| Cystoseira compressa | ||||||
|---|---|---|---|---|---|---|
| Fatty Acid Methyl Ester | Cauloid | Old Branches | Receptacle | |||
| AV | ± sd | AV | ± sd | AV | ± sd | |
| C14:1 | 0.28 | 0.41 | 0.25 | 0.38 | ||
| C14:0 | 11.92 | 0.99 | 9.50 | 3.78 | 6.57 | 2.90 |
| C15:0 iso | 0.20 | 0.20 | ||||
| C15:0 ante | ||||||
| C15:0 | 1.58 | 1.06 | 0.90 | 0.20 | 0.59 | 0.05 |
| C16:1 | 0.33 | 0.44 | 1.85 | 1.33 | 3.33 | 1.69 |
| C16:0 | 65.81 | 6.68 | 71.39 | 3.35 | 48.41 | 17.13 |
| C17:0 iso | ||||||
| C17:0 ante | ||||||
| C17:1 | 0.11 | 0.11 | 0.56 | 0.45 | ||
| C17:0 | 1.24 | 0.82 | 0.60 | 0.15 | 0.25 | 0.17 |
| C18:3(n-3) | 1.07 | 1.07 | ||||
| C18:2(n-6) | 0.08 | 0.12 | 3.39 | 3.07 | ||
| C18:1 | 0.17 | 0.22 | 3.08 | 1.38 | 16.76 | 5.51 |
| C18:0 | 16.26 | 3.66 | 4.62 | 1.88 | 2.70 | 0.86 |
| C20:4(n-6) | 5.85 | 4.73 | ||||
| C20:5(n-3) | 2.82 | 1.64 | ||||
| C20:3(n-3) | 1.75 | 1.82 | ||||
| C20:2(n-6) | 0.81 | 0.53 | ||||
| C20:1 | 0.32 | 0.41 | 0.04 | 0.05 | ||
| C20:0 | 1.44 | 0.96 | 1.79 | 0.02 | 1.61 | 0.87 |
| C22:4(n-6) | ||||||
| C22:6(n-3) | ||||||
| C22:4(n-3) | 0.05 | 0.07 | ||||
| C22:2(n-6) | ||||||
| C22:1 | 0.09 | 0.14 | ||||
| C22:0 | 1.20 | 0.80 | 0.96 | 0.13 | 0.63 | 0.20 |
| SFA | 99.45 | 0.73 | 89.95 | 334.00 | 60.75 | 15.93 |
| MUFA | 0.50 | 0.67 | 5.72 | 3.16 | 20.93 | 5.43 |
| PUFA | 0.08 | 0.12 | 15.75 | 10.70 | ||
| Bacterial | 2.82 | 1.88 | 1.81 | 0.43 | 1.40 | 0.63 |
| n-6/n-3 | 2.25 | 0.69 | ||||
| UND | 0.01 | 0.01 | 0.07 | 0.04 | 1.67 | 1.24 |
| Total lipids (mg/g d.w.) | 6.98 | 0.98 | 9.48 | 0.38 | 7.60 | 2.73 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Blažina, M.; Iveša, L.; Matošović, K.; Pustijanac, E.; Devescovi, M.; Pritišanac, E.; Fafanđel, M.; Batel, R. Diversity of Epiphytic Bacterial Communities Associated with Adriatic Fucacean Species Phycosphere. J. Mar. Sci. Eng. 2020, 8, 967. https://doi.org/10.3390/jmse8120967
Blažina M, Iveša L, Matošović K, Pustijanac E, Devescovi M, Pritišanac E, Fafanđel M, Batel R. Diversity of Epiphytic Bacterial Communities Associated with Adriatic Fucacean Species Phycosphere. Journal of Marine Science and Engineering. 2020; 8(12):967. https://doi.org/10.3390/jmse8120967
Chicago/Turabian StyleBlažina, Maria, Ljiljana Iveša, Ksenija Matošović, Emina Pustijanac, Massimo Devescovi, Ena Pritišanac, Maja Fafanđel, and Renato Batel. 2020. "Diversity of Epiphytic Bacterial Communities Associated with Adriatic Fucacean Species Phycosphere" Journal of Marine Science and Engineering 8, no. 12: 967. https://doi.org/10.3390/jmse8120967
APA StyleBlažina, M., Iveša, L., Matošović, K., Pustijanac, E., Devescovi, M., Pritišanac, E., Fafanđel, M., & Batel, R. (2020). Diversity of Epiphytic Bacterial Communities Associated with Adriatic Fucacean Species Phycosphere. Journal of Marine Science and Engineering, 8(12), 967. https://doi.org/10.3390/jmse8120967






