High-Throughput Sequencing Reveals Fungal Microbiome of Apricots Grown Under Organic and Integrated Pest Management Systems
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. Extraction of Nucleic Acids
2.3. Library Preparation and Next-Generation Sequencing
2.4. Bioinformatic Analysis
2.5. Data Analyses
2.6. RT-QPCR Detection of Microbial Pathogens—‘Candidatus Phytoplasma Prunorum’, Pseudomonas Syringae, Cytospora sp.
3. Results
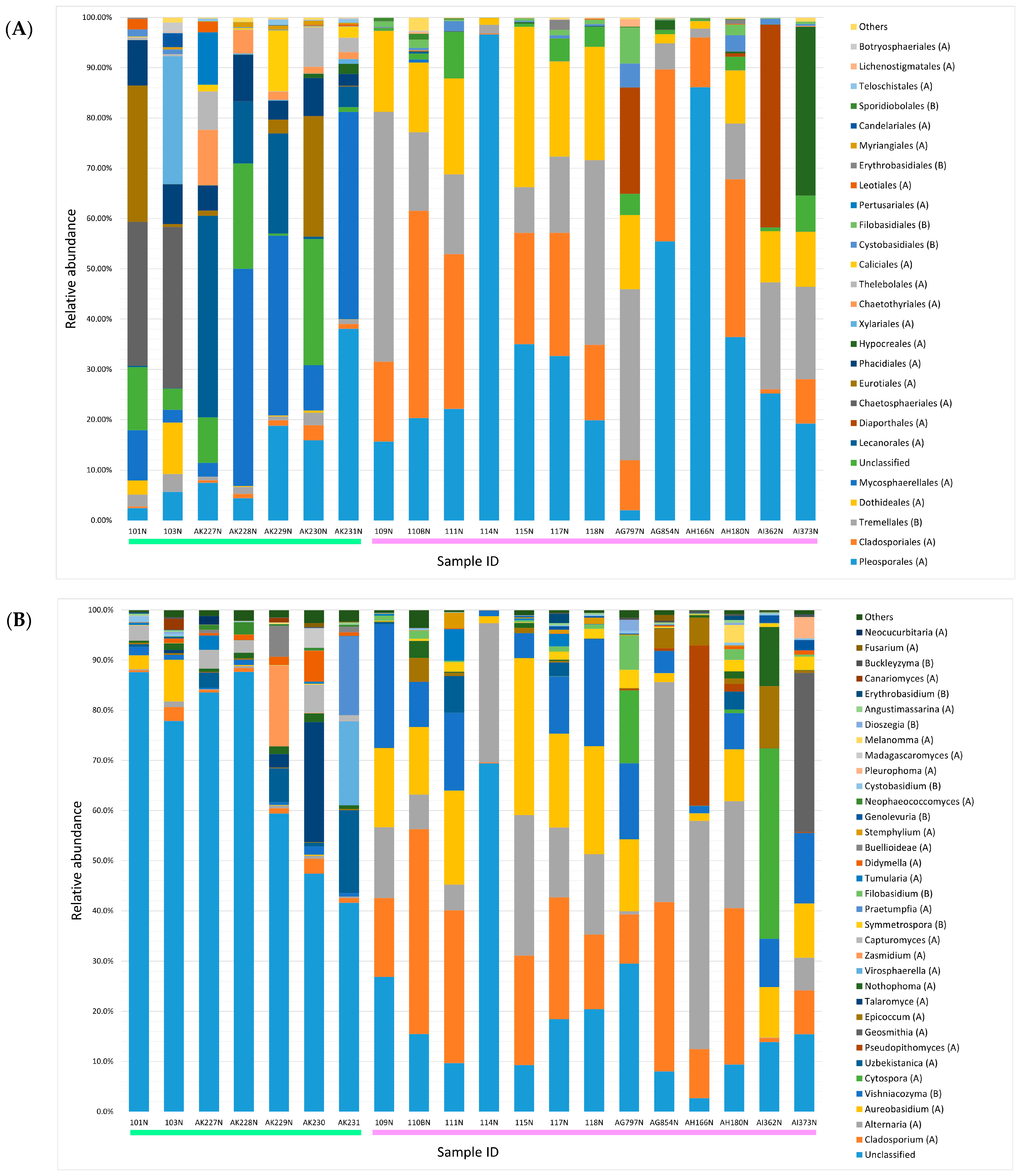
3.1. Composition of Fungal Microbiome
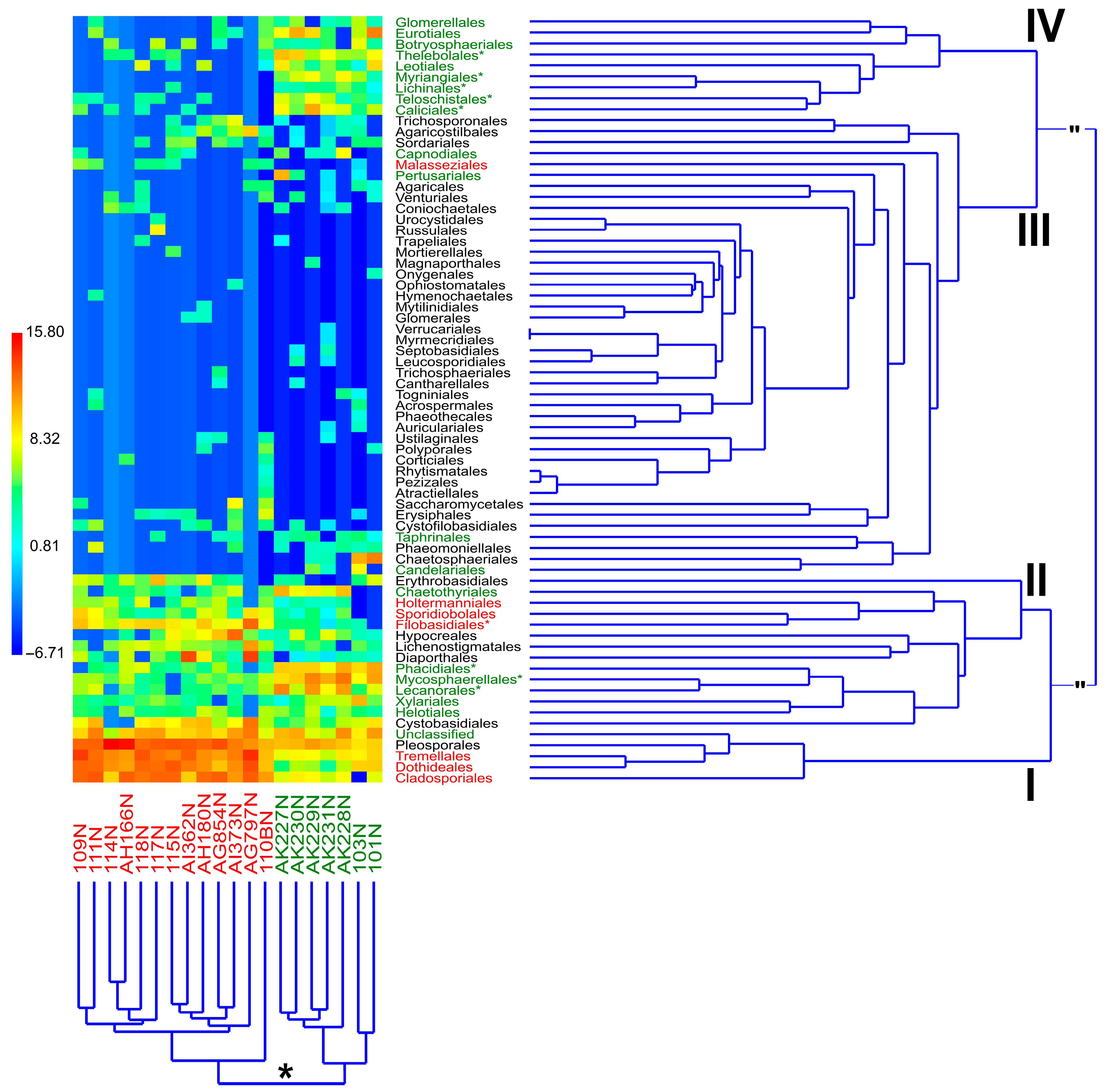
3.2. Comparison of Identified Fungal Orders and Genera Among Samples
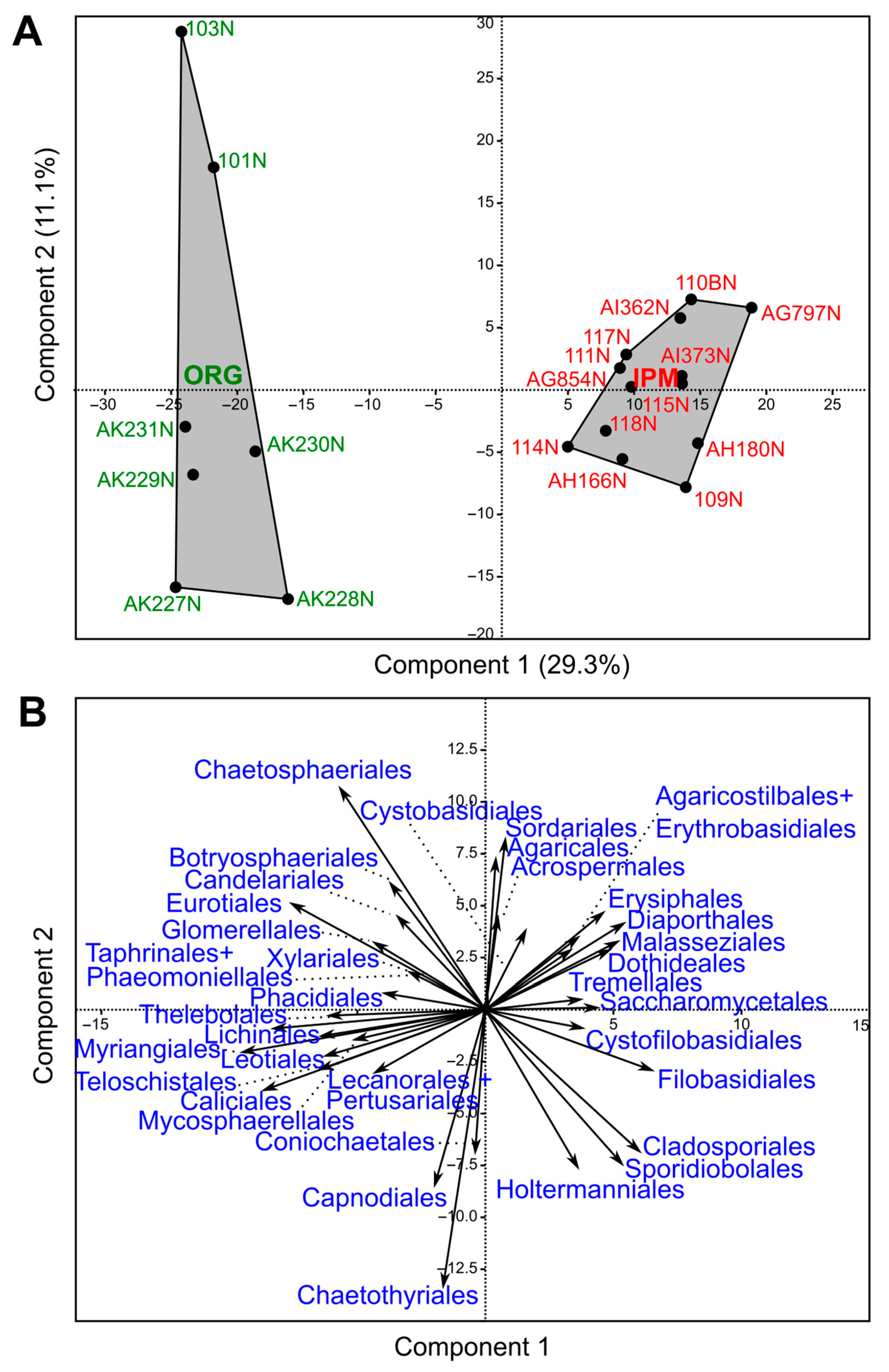
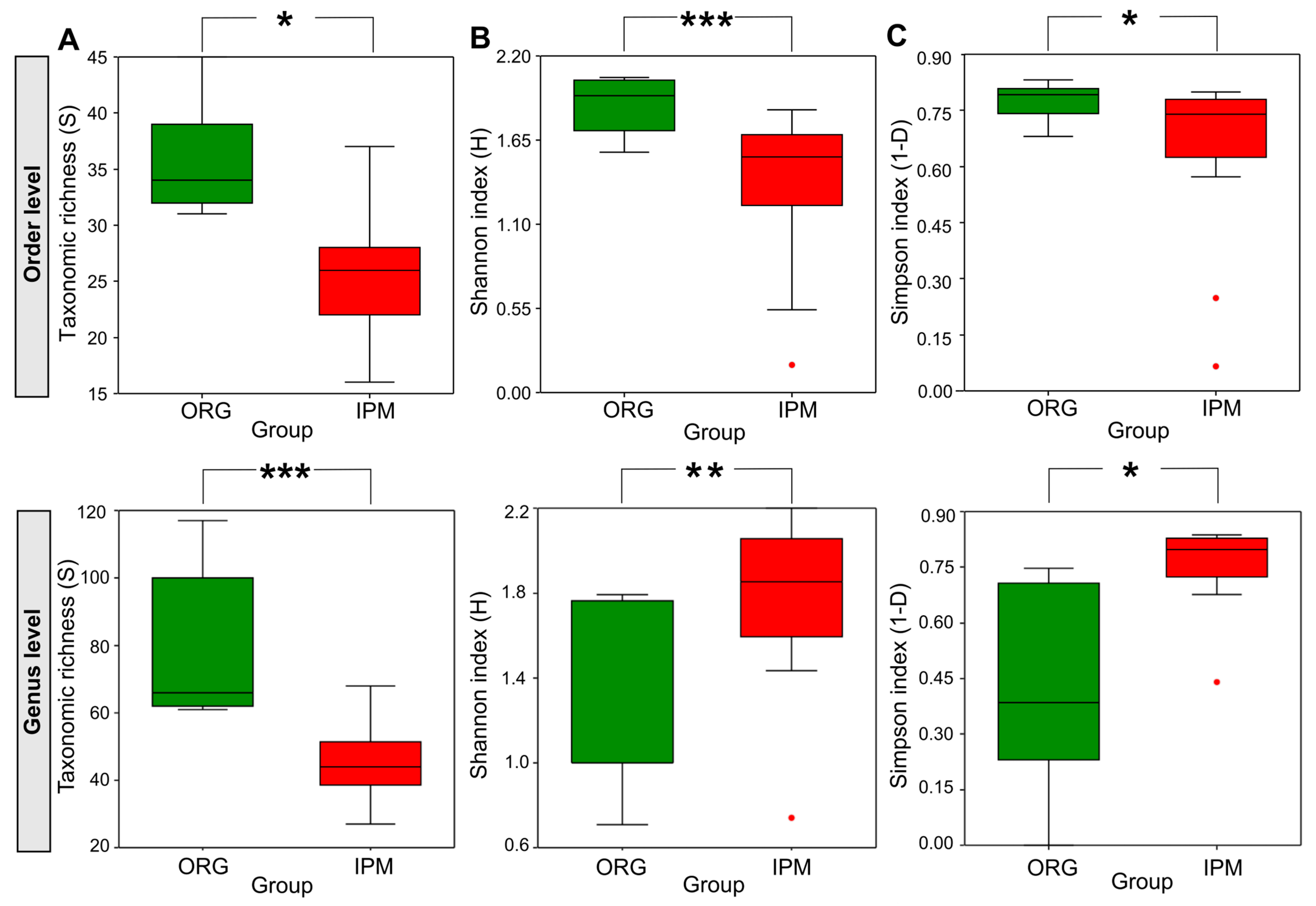
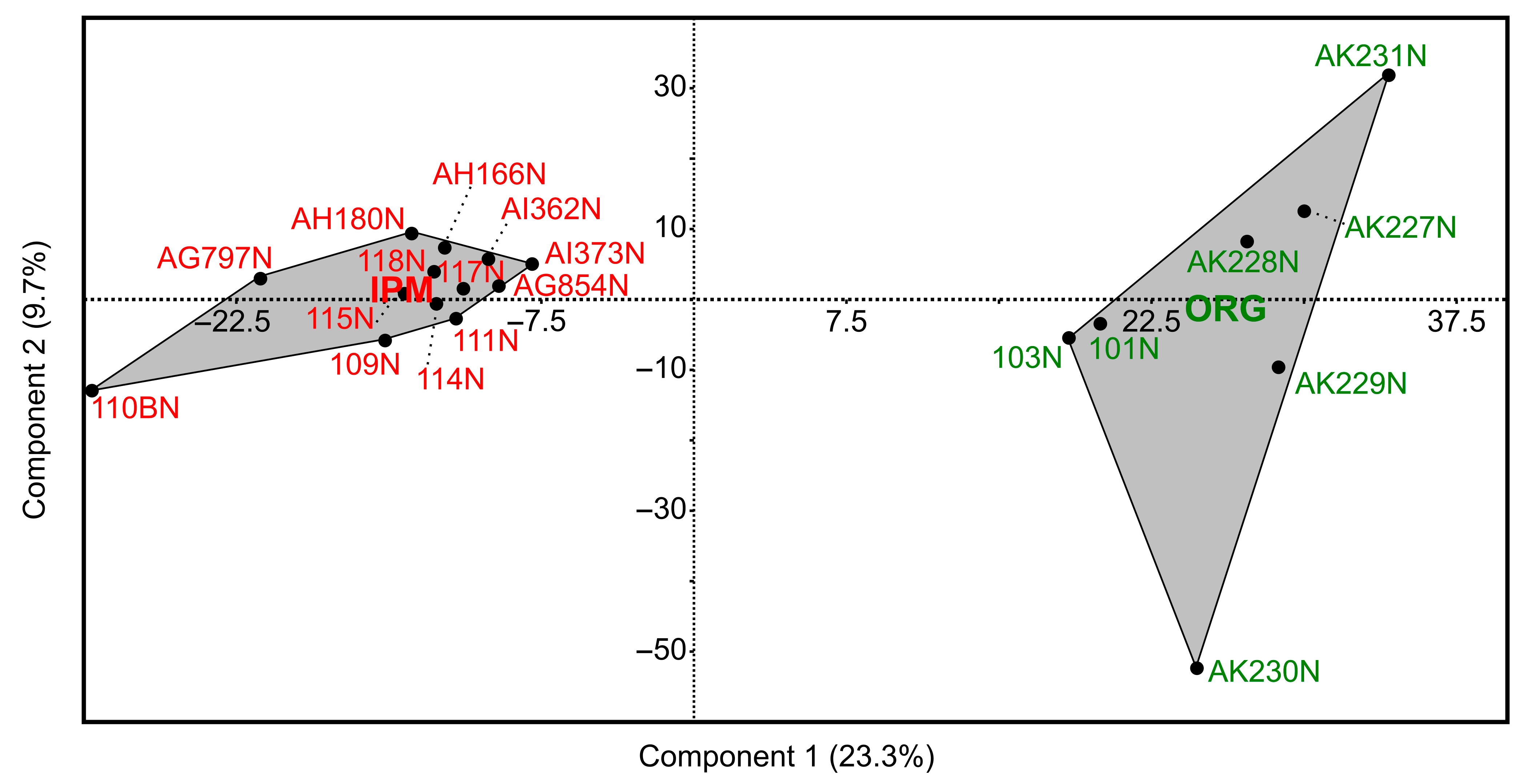
3.3. Comparison of Fungal Communities Between Organic and Integrated Pest Management Orchards
3.4. PCR Detection of Pathogenic ‘Candidatus Phytoplasma Prunorum’, Pseudomonas Syringae and Cytospora sp.
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- CIRAD. 2025. Available online: https://www.cirad.fr/en (accessed on 30 March 2025).
- Maxwell, M. European Apricot Volumes Unchanged. 2024. Available online: https://www.fruitnet.com/eurofruit/european-apricot-volumes-unchanged/259975.article (accessed on 30 March 2025).
- Němcová, V.; Buchtová, I. Situační a výhledová zpráva Ovoce 2024 [Situation and Foresight Report Fruits 2024]; Ministry of Agriculture of the Czech Republic: Praque, Czech Republic, 2024. [Google Scholar]
- Lawrence, D.P.; Holland, L.A.; Nouri, M.T.; Travadon, R.; Abramians, A.; Michailides, T.J.; Trouillas, F.P. Molecular phylogeny of Cytospora species associated with canker diseases of fruit and nut crops in California, with the descriptions of ten new species and one new combination. IMA Fungus 2018, 9, 333–370. [Google Scholar] [CrossRef]
- Zhu, H.; Pan, M.; Bezerra, J.D.P.; Tian, C.; Fan, X. Discovery of Cytospora species associated with canker disease of tree hosts from Mount Dongling of China. MycoKeys 2020, 62, 97–121. [Google Scholar] [CrossRef] [PubMed]
- CABI. Eutypa Lata (Eutypa Dieback). 2021. Available online: https://www.cabidigitallibrary.org/doi/10.1079/cabicompendium.23591 (accessed on 30 March 2025).
- Hrustic, J.; Delibasic, G.; Stankovic, I.; Grahovac, M.; Krstic, B.; Bulajic, A.; Tanovic, B. Monilinia spp. causing brown rot of stone fruit in Serbia. Plant Dis. 2015, 99, 709–717. [Google Scholar] [CrossRef]
- Grígel, J.; Cerny, K.; Mrázková, M.; Havrdová, L.; Zahradník, D.; Jílková, B.; Hrabetová, M. Phytophthora root and collar rots in fruit orchards in the Czech Republic. Phytopathol. Mediterr. 2019, 58, 261–275. [Google Scholar] [CrossRef]
- Hiemstra, J. A compendium of Verticillium wilts in tree species. J. Reprod. Fertil. 1998. Available online: https://www.researchgate.net/publication/254834373_A_Compendium_of_Verticillium_Wilts_in_Tree_Species (accessed on 30 March 2025).
- OEPP/EPPO. Stone fruits. PP2/33-Good plant protection practice. Bull. OEPP/EPPO Bull. 2004, 34, 427–438. [Google Scholar]
- Regulation_2019/2072. Document 32019R2072 Commission Implementing Regulation (EU) 2019/2072 of 28 November 2019 Establishing Uniform Conditions for the Implementation of Regulation (EU) 2016/2031 of the European Parliament and the Council, as Regards Protective Measures Against Pests of Plants, and Repealing Commission Regulation (EC) No 690/2008 and Amending Commission Implementing Regulation (EU) 2018/2019. 2019. Available online: https://eur-lex.europa.eu/eli/reg_impl/2019/2072/oj/eng (accessed on 30 March 2025).
- Ye, S.; Jia, H.; Cai, G.; Tian, C.; Ma, R. Morphology, DNA phylogeny, and pathogenicity of Wilsonomyces carpophilus isolate causing shot-hole disease of Prunus divaricata and Prunus armeniaca in wild-fruit forest of Western Tianshan mountains, China. Forests 2020, 11, 319. [Google Scholar] [CrossRef]
- Morvan, G. Pathogenic organisms as causes of apricot decline. Acta Hortic. 1995, 384, 521–532. [Google Scholar] [CrossRef]
- Krška, B.; Gogolková¡, K.; Ondrášek, I.; Nečas, T. Evaluation of precocious decline of young apricot orchards in the Czech Republic. Acta Hortic. 2012, 966, 175–182. [Google Scholar] [CrossRef]
- Pečenka, J.; Peňázová, E.; Tekielska, D.; Ondrášek, I.; Nečas, T.; Eichmeier, A. Fungi detected in trunk of stone fruits in the Czech Republic. Acta Agrar. Debr. 2019, 72, 121–127. [Google Scholar] [CrossRef]
- Zezlina, I.; Rot, M.; Kac, M.; Trdan, S. Causal agents of stone fruit diseases in Slovenia and the potential for diminishing their economic impact—A review. Plant Protect. Sci. 2016, 52, 149–157. [Google Scholar] [CrossRef]
- Lamichhane, J.R.; Varvaro, L.; Parisi, L.; Audergon, J.M.; Morris, C.E. Disease and frost damage of woody plants caused by Pseudomonas syringae: Seeing the forest for the trees. Adv. Agron. 2014, 126, 235–295. [Google Scholar] [CrossRef]
- Kennelly, M.M.; Cazorla, F.M.; de Vicente, A.; Ramos, C.; Sundin, G.W. Pseudomonas syringae diseases of fruit trees: Progress toward understanding and control. Plant Dis. 2007, 91, 4–17. [Google Scholar] [CrossRef]
- Bultreys, A.; Kaluzna, M. Bacterial cankers caused by Pseudomonas syringae on stone fruit species with special emphasis on the pathovars and race 1 and race 2. J. Plant Pathol. 2010, 92, S21–S33. [Google Scholar]
- Ogawa, J.M.; Zehr, E.I.; Bird, G.W.; Ritchie, D.F.; Uriu, K.; Uyemoto, J.K. Compendium of Stone Fruit Diseases; American Phytopathological Society: St. Paul, MN, USA, 1995. [Google Scholar]
- Soylu, S.; Kara, M.; Kurt, Å.; Soylu, E.M.; Uysal, A. Determination of fungal and bacterial disease agents of apricot trees growing in Hatay province. Acta Hortic. 2020, 1290, 111–114. [Google Scholar] [CrossRef]
- Bien, S.; Kraus, C.; Damm, U. Novel collophorina-like genera and species from Prunus trees and vineyards in Germany. Persoonia 2020, 45, 46–67. [Google Scholar] [CrossRef]
- Damm, U.; Crous, P.W.; Fourie, P.H. Botryosphaeriaceae as potential pathogens of Prunus species in South Africa, with descriptions of Diplodia africana and Lasiodiplodia plurivora. Mycologia 2007, 99, 664–680. [Google Scholar] [CrossRef] [PubMed]
- Damm, U.; Crous, P.F.; Fourie, P.H. A fissitunicate ascus mechanism in the Calosphaeriaceae, and novel species of Jattaea and Calosphaeria on Prunus wood. Persoonia 2008, 20, 39–52. [Google Scholar] [CrossRef] [PubMed]
- Damm, U.; Fourie, P.F.; Crous, P.W. Coniochaeta (Lecythophora), Collophora gen. nov. and Phaeomoniella species associated with wood necroses of Prunus trees. Persoonia 2010, 24, 60–80. [Google Scholar] [CrossRef]
- Damm, U.; Mostert, L.; Crous, P.W.; Fourie, P.H. Novel Phaeoacremonium species associated with necrotic wood of Prunus trees. Persoonia 2008, 20, 87–102. [Google Scholar] [CrossRef]
- Garcia, J.F.; Lawrence, D.P.; Morales-Cruz, A.; Travadon, R.; Minio, A.; Hernandez-Martinez, R.; Rolshausen, P.E.; Baumgartner, K.; Cantu, D. Phylogenomics of plant-associated Botryosphaeriaceae species. Front. Microbiol. 2021, 12, 652802. [Google Scholar] [CrossRef]
- Moyo, P.; Damm, U.; Mostert, L.; Halleen, F. Eutypa, Eutypella, and Cryptovalsa species (Diatrypaceae) associated with Prunus species in South Africa. Plant Dis. 2018, 102, 1402–1409. [Google Scholar] [CrossRef] [PubMed]
- Oprea, M.; Jinga, V. Lignivorous fungi evolution attack on apricot tree in Romania. Acta Hortic. 2010, 862, 515–518. [Google Scholar] [CrossRef]
- Król, E.D.; Abramczyk, B.A.; Zalewska, E.D.; Zimowska, B. Fungi inhabiting fruit tree shoots with special reference to the Diaporthe (Phomopsis) genus. Acta Sci. Pol. Hortorum Cultus 2017, 16, 113–126. [Google Scholar] [CrossRef]
- Garić, R.; Arsenijević, M. Phomopsis—uzročnik nove bolesti kajsije [Phomopsis—the causal agent of a new disease of apricots]. Zast. Bilja 1990, 193, 269–274. [Google Scholar]
- Elbeaino, T.; Giampetruzzi, A.; De Stradis, A.; Digiaro, M. Deep-sequencing analysis of an apricot tree with vein clearing symptoms reveals the presence of a novel betaflexivirus. Virus Res. 2014, 181, 1–5. [Google Scholar] [CrossRef]
- Marais, A.; Faure, C.; Mustafayev, E.; Barone, M.; Alioto, D.; Candresse, T. Characterization by deep sequencing of Prunus virus T, a novel Tepovirus infecting Prunus species. Phytopathology 2015, 105, 135–140. [Google Scholar] [CrossRef]
- Maliogka, V.I.; Minafra, A.; Saldarelli, P.; Ruiz-Garcia, A.B.; Glasa, M.; Katis, N.; Olmos, A. Recent advances on detection and characterization of fruit tree viruses using high-throughput sequencing technologies. Viruses 2018, 10, 436. [Google Scholar] [CrossRef]
- Liang, S.X.; Liu, H.; Wu, S.H.; Xu, S.J.; Jin, D.C.; Faiola, F.; Zhuang, X.L.; Zhuang, G.Q.; Qu, D.; Fan, H.Y.; et al. Genetic diversity of diazotrophs and total bacteria in the phyllosphere of Pyrus serotina, Prunus armeniaca, Prunus avium, and Vitis vinifera. Can. J. Microbiol. 2019, 65, 642–652. [Google Scholar] [CrossRef] [PubMed]
- ELI. Directive 2009/128/EC of the European Parliament and of the Council of 21 October 2009 Establishing a Framework for Community Action to Achieve the Sustainable Use of Pesticides. 2019. Available online: https://eur-lex.europa.eu/eli/dir/2009/128/2019-07-26 (accessed on 1 July 2025).
- MACZ. The National Action Plan to Reduce the Use of Pesticides in the Czech Republic; Ministry of Agriculture; Ministry of Agriculture of the Czech Republic: Prague, Czech Republic, 2013; Available online: https://food.ec.europa.eu/document/download/681292de-c674-45a9-aeff-958d2454edce_en (accessed on 1 July 2025).
- SISPO. Směrnice Pro Pěstování Peckovin V Systému Integrované Produkce Ovoce. Svaz Pro Integrované Systémy Produkce Ovoce. 2016. Available online: https://www.sispo.cz/wp-content/uploads/2024/07/Peckoviny_smernice_IP.pdf (accessed on 1 July 2025).
- ELI. Regulation (EU) 2018/848 of the European Parliament and of the Council of 30 May 2018 on Organic Production and Labelling of Organic Products 2018. 2018. Available online: https://eur-lex.europa.eu/legal-content/EN/TXT/?uri=CELEX:32018R0848 (accessed on 1 July 2025).
- Anonymous. Organic Farming Act No. 242/2000. Parliament of the Czech Republic. 2000. Available online: https://www.fao.org/faolex/results/details/en/c/LEX-FAOC182674/ (accessed on 1 July 2025).
- Tuomisto, H.L.; Hodge, I.D.; Riordan, P.; Macdonald, D.W. Does organic farming reduce environmental impacts?—A meta-analysis of European research. J. Environ. Manag. 2012, 112, 309–320. [Google Scholar] [CrossRef]
- Legendre, P.; Legendre, L. Numerical Ecology; Elsevier: Amsterdam, The Netherlands, 1998. [Google Scholar]
- Jackson, D.A. Stopping Rules in Principal Components-Analysis—A Comparison of Heuristic and Statistical Approaches. Ecology 1993, 74, 2204–2214. [Google Scholar] [CrossRef]
- Hammer, O.; Harper, D.; Ryan, P. PAST: Paleontological Statistics Software package for education and data analysis. Palaeontol. Electron. 2001, 4, 1–9. [Google Scholar]
- Rejlová, M.; Valentová, L.; Navrátil, M.; Šafářová, D.; Čmejla, R. Prevalence of pathogens associated with apricot apoplexy in commercial orchards in the Czech Republic. VPO 2021, 27, 53–69. (In Czech) [Google Scholar]
- Longa, C.M.O.; Antonielli, L.; Bozza, E.; Sicher, C.; Pertot, I.; Perazzolli, M. Plant organ and sampling time point determine the taxonomic structure of microbial communities associated to apple plants in the orchard environment. Microbiol. Res. 2022, 258, 126991. [Google Scholar] [CrossRef] [PubMed]
- Abdelfattah, A.; Freilich, S.; Bartuv, R.; Zhimo, V.Y.; Kumar, A.; Biasi, A.; Salim, S.; Feygenberg, O.; Burchard, E.; Dardick, C.; et al. Global analysis of the apple fruit microbiome: Are all apples the same? Environ. Microbiol. 2021, 23, 6038–6055. [Google Scholar] [CrossRef] [PubMed]
- Al Riachy, R.; Strub, C.; Durand, N.; Chochois, V.; Lopez-Lauri, F.; Fontana, A.; Schorr-Galindo, S. The influence of long-term storage on the epiphytic microbiome of postharvest apples and on Penicillium expansum occurrence and patulin accumulation. Toxins 2024, 16, 102. [Google Scholar] [CrossRef] [PubMed]
- Dugan, F.M. Etiology of preharvest colonization of Bing cherry fruit by fungi. Phytopathology 1994, 84, 1031–1036. [Google Scholar] [CrossRef]
- Nekrasov, E.V.; Shumilova, L.P.; Gomzhina, M.M.; Aleksandrova, A.V.; Kokaeva, L.Y.; Pavlova, L.M. Diversity of endophytic fungi in annual shoots of Prunus mandshurica (Rosaceae) in the south of Amur region, Russia. Diversity 2022, 14, 1124. [Google Scholar] [CrossRef]
- Galli, V.; Romboli, Y.; Barbato, D.; Mari, E.; Venturi, M.; Guerrini, S.; Granchi, L. Indigenous Aureobasidium pullulans strains as biocontrol agents of Botrytis cinerea on grape berries. Sustainability 2021, 13, 9389. [Google Scholar] [CrossRef]
- Bellamy, S.; Shaw, M.; Xu, X. Field application of Bacillus subtilis and Aureobasidium pullulans to reduce Monilinia laxa post-harvest rot on cherry. Eur. J. Plant Pathol. 2022, 163, 761–766. [Google Scholar] [CrossRef]
- Zeng, Q.; Johnson, K.B.; Mukhtar, S.; Nason, S.; Huntley, R.; Millet, F.; Yang, C.-H.; Hassani, M.A.; Zuverza-Mena, N.; Sundin, G.W. Aureobasidium pullulans from the fire blight biocontrol product, blossom protect, induces host resistance in apple flowers. Phytopathology 2023, 113, 1192–1201. [Google Scholar] [CrossRef] [PubMed]
- Ippolito, A.; Ghaouth, A.; Wilson, C.; Wisniewski, M. Control of postharvest decay of apple fruit by Aureobasidium pullulans and induction of defense responses. Postharvest Biol. Tec. 2000, 19, 265–272. [Google Scholar] [CrossRef]
- Cignola, R.; Boato, A.; Sadallah, A.; Firrao, G.; Di Francesco, A. Molecular characterization of Aureobasium spp. strains isolated during the cold season. A preliminary efficacy evaluation as novel potential biocontrol agents against postharvest pathogens. Eur. J. Plant Pathol. 2023, 167, 221–233. [Google Scholar] [CrossRef]
- Di Francesco, A.; Di Foggia, M.; Baraldi, E. Aureobasidium pullulans volatile organic compounds as alternative postharvest method to control brown rot of stone fruits. Food Microbiol. 2020, 87, 103395. [Google Scholar] [CrossRef] [PubMed]
- Oztekin, S.; Dikmetas, D.N.; Devecioglu, D.; Acar, E.G.; Karbancioglu-Guler, F. Recent insights into the use of antagonistic yeasts for sustainable biomanagement of postharvest pathogenic and mycotoxigenic fungi in fruits with their prevention strategies against mycotoxins. J. Agric. Food Chem. 2023, 71, 9923–9950. [Google Scholar] [CrossRef]
- Arzanlou, M. Molecular characterization of Aureobasidium species in Iran. Res. Mol. Med. 2014, 2, 28–33. [Google Scholar] [CrossRef]
- Pinto, C.; Custódio, V.; Nunes, M.; Songy, A.; Rabenoelina, F.; Courteaux, B.; Clément, C.; Gomes, A.C.; Fontaine, F. Understand the potential role of Aureobasidium pullulans, a resident microorganism from grapevine, to prevent the infection caused by Diplodia seriata. Front. Microbiol. 2018, 9, 3047. [Google Scholar] [CrossRef]
- Dimakopoulou, M.; Tjamos, S.E.; Antoniou, P.P.; Pietri, A.; Battilani, P.; Avramidis, N.; Markakis, E.A.; Tjamos, E.C. Phyllosphere grapevine yeast Aureobasidium pullulans reduces Aspergillus carbonarius (sour rot) incidence in wine-producing vineyards in Greece. Biol. Control 2008, 46, 158–165. [Google Scholar] [CrossRef]
- Pose, G.N.; Ludemann, V.; Fernandez, D.; Segura, J.A.; Pinto, V.F. Alternaria species associated with ‘moldy heart’ on peaches in Argentina. Trop. Plant Pathol. 2010, 35, 174–177. Available online: https://www.scielo.br/j/tpp/a/4xyjqcfXRXNVb9YJbvz6Wqf/?lang=en (accessed on 1 July 2025).
- Thomidis, T.; Michailides, T.; Exadaktylou, E. Contribution of pathogens to peach fruit rot in Northern Greece and their sensitivity to iprodione, carbendazim, thiophanate-methyl and tebuconazole fungicides. J. Phytopathol. 2009, 157, 194–200. [Google Scholar] [CrossRef]
- Valiuskaite, A. Micromycetes infecting stone fruit trees. Biologija 2002, 1, 18–21. [Google Scholar]
- Fancello, F.; Bianco, A.; Niccolai, M.; Zara, G.; Coronas, R.; Serra, E.; D’Hallewin, G.; Valentoni, A.; Santoru, A.; Pretti, L.; et al. Fruit microbial communities of the Bisucciu sardinian apricot cultivar (Prunus armeniaca L.) as a reservoir of new brewing starter strains. Fermentation 2022, 8, 364. [Google Scholar] [CrossRef]
- Gierl, L.; Fischer, M. Grapevine trunk disease in German viticulture II. Associated fungi occurring on non-hosts, and first report of Phaeoacremonium angustius. Vitis 2017, 56, 103–110. [Google Scholar] [CrossRef]
- Barata, A.; Malfeito-Ferreira, M.; Loureiro, V. The microbial ecology of wine grape berries. Int. J. Food Microbiol. 2012, 153, 243–259. [Google Scholar] [CrossRef]
- Camatti-Sartori, V.; Silva-Ribeiro, R.; Valdebenito- Sanhueza, R.; Pagnocca, F.; Echeverrigaray, S.; Azevedo, J. Endophytic yeasts and filamentous fungi associated with southern Brazilian apple (Malus domestica) orchards subjected to conventional, integrated or organic cultivation. J. Basic Microbiol. 2005, 45, 397–402. [Google Scholar] [CrossRef]
- Ucar, K.; Saner, G.; Engindeniz, S. The comparative economic analysis of organic and conventional dried apricot production: A case study for Turkey. Fresen. Environ. Bull. 2017, 26, 4555–4560. [Google Scholar]
- Purin, S.; Klauberg, O.; Stürmer, S.L. Mycorrhizae activity and diversity in conventional and organic apple orchards from Brazil. Soil Biol. Biochem. 2006, 38, 1831–1839. [Google Scholar] [CrossRef]
- Ciccoritti, R.; Ciorba, R.; Mitrano, F.; Cutuli, M.; Amoriello, T.; Ciaccia, C.; Testani, E.; Ceccarelli, D. Diversification and soil management effects on the quality of organic apricots. Agronomy 2021, 11, 1791. [Google Scholar] [CrossRef]
- Novotný, D.; Lukas, J.; Brozova, J.; Růžičková, P. Comparison of the occurrence of fungi causing postharvest diseases of apples grown in organic and integrated production systems in orchards in the Czech Republic. Czech Mycol. 2019, 71, 99–121. [Google Scholar] [CrossRef]
- Khwantongyim, P.; Wansee, S.; Lu, X.; Zhang, W.; Sun, G.Y. Variations in the community structure of fungal microbiota associated with apple fruit shaped by fruit bagging—Based practice. J. Fungi 2021, 7, 764. [Google Scholar] [CrossRef] [PubMed]
- Janakiev, T.; Berić, T.; Stević, T.; Stanković, S.; Bačić, J.; Majstorović, H.; Fira, D.; Dimkić, I. The microbiome of the ‘Williams’ pear variety grown in the organic orchard and antifungal activity by the autochthonous bacterial and yeast isolates. Microorganisms 2022, 10, 1282. [Google Scholar] [CrossRef]
- Varanda, C.; Oliveira, M.; Materatski, P.; Landum, M.; Clara, M.; Félix, M. Fungal endophytic communities associated to the phyllosphere of grapevine cultivars under different types of management. Fungal Biol. 2016, 120, 1525–1536. [Google Scholar] [CrossRef] [PubMed]
- Pancher, M.; Ceol, M.; Corneo, P.; Longa, C.; Yousaf, S.; Pertot, I.; Campisano, A. Fungal endophytic communities in grapevines (Vitis vinifera L.) respond to crop management. Appl. Environ. Microbiol. 2012, 78, 4308–4317. [Google Scholar] [CrossRef]
- Conti, M.; Cinget, B.; Labbé, C.; Asselin, Y.; Bélanger, R.R. New insights into the fungal diversity of cranberry fruit rot in Quebec farms through a large-scale molecular analysis. Plant Dis. 2022, 106, 215–222. [Google Scholar] [CrossRef] [PubMed]
- Granado, J.; Thürig, B.; Kieffer, E.; Petrini, L.; Fliessbach, A.; Tamm, L.; Weibel, F.P.; Wyss, G.S. Culturable fungi of stored ‘Golden Delicious’ apple fruits: A one-season comparison study of organic and integrated production systems in Switzerland. Microb. Ecol. 2008, 56, 720–732. [Google Scholar] [CrossRef]
- Schaeffer, R.N.; Pfeiffer, V.W.; Basu, S.; Brousil, M.; Strohm, C.; DuPont, S.T.; Vannette, R.L.; Crowder, D.W. Orchard management and landscape context mediate the pear floral microbiome. Appl. Environ. Microbiol. 2021, 87, 101138. [Google Scholar] [CrossRef]
- De Corato, U.; Trupo, M. Apricot trees gummosis caused by Botryosphaeria dothidea (Moug.: Fr.) Ces. et De Not. in Southern Italy (Basilicata). PETRIA 2008, 18, 43–48. [Google Scholar]
- Guarnaccia, V.; Kraus, C.; Markakis, E.; Alves, A.; Armengol, J.; Eichmeier, A.; Compant, S.; Gramaje, D. Fungal trunk diseases of fruit trees in Europe: Pathogens, spread and future directions. Phytopathol. Mediterr. 2022, 61, 563–599. [Google Scholar] [CrossRef]
- Mitrev, S.; Kungulovski, D.; Arsov, E.; Kovacevik, B. First report of Calosphaeria pulchella causing canker and branch dieback of sour cherry trees (Prunus cerasus) in North Macedonia. Phytopathol. Mediterr. 2025, 64, 3–8. [Google Scholar] [CrossRef]
- Oprea, M.; Şesan, T.; Bălan, V. Schizophyllum commune–canker and dieback disease of apricot trees in orchards of southeastern Romania. Rev. Roum. Biol. Biol. 1994, 39, 35–40. [Google Scholar]
- Spetik, M.; Pecenka, J.; Stuskova, K.; Stepanova, B.; Eichmeier, A.; Kiss, T. Fungal trunk diseases causing decline of apricot and plum trees in the Czech Republic. Plant Dis. 2024, 108, 1425–1436. [Google Scholar] [CrossRef] [PubMed]
- Rejlová, M.; Valentová, L.; Čmejla, R. Zdravotní stav a patogeny v experimentální výsadbě meruněk: Vývoj v čase [Health status and pathogens in an experimental apricot orchard: Develeopment in time]. VPO 2021, 27, 119–132. [Google Scholar]
- Lacok, P. Fungi and apricot cultures in Slovakia (Czechoslovakia) at present. Acta Hortic. 1986, 192, 205–212. [Google Scholar] [CrossRef]
- Dosba, F.; Lansac, M.; Mazy, K.; Garnier, M.; Eyquard, J.P. Incidence of different disease associated with mycoplasma like organisms in different species of Prunus. Acta Hortic. 1990, 283, 311–320. [Google Scholar] [CrossRef]
- Marcone, C.; Jarausch, B.; Jarausch, W. Candidatus Phytoplasma prunorum, the causal agent of European stone fruit yellows: An overview. J. Plant Pathol. 2010, 92, 19–34. Available online: https://www.researchgate.net/publication/267030640_Candidatus_Phytoplasma_prunorum_the_causal_agent_Of_European_stone_fruit_yellows_An_overview (accessed on 30 March 2025).
- Ivanova, L. First occurrence of apricot blast disease caused by Pseudomonas syringae in the North-Eastern part of Bulgaria. Acta Hortic. 2009, 825, 149–151. [Google Scholar] [CrossRef]
- Scortichini, M. Severe outbreak of Pseudomonas syringae pv. syringae on new apricot cultivars in central Italy. J. Plant Pathol. 2006, 88, S68. [Google Scholar]
- Morvan, G. Apricot Decline Today and Tomorrow. Bull. OEPP/EPPO Bull. 1977, 7, 137–147. [Google Scholar] [CrossRef]
- Travadon, R.; Baumgartner, K. Molecular polymorphism and phenotypic diversity in the Eutypa dieback pathogen. Phytopathology 2015, 105, 255–264. [Google Scholar] [CrossRef]
- Montesinos, E.; Vilardell, P. Relationships among population-levels of Pseudomonas-syringae, amount of ice nuclei, and incidence of blast of dormant flower buds in commercial pear orchards in Catalunya, Spain. Phytopathology 1991, 81, 113–119. [Google Scholar] [CrossRef]
- Lindow, S.E.; Arny, D.C.; Upper, C.D. Bacterial ice nucleation—A factor in frost injury to plants. Plant Physiol. 1982, 70, 1084–1089. [Google Scholar] [CrossRef] [PubMed]
- Xin, X.F.; Kvitko, B.; He, S.Y. Pseudomonas syringae: What it takes to be a pathogen. Nat. Rev. Microbiol. 2018, 16, 316–328. [Google Scholar] [CrossRef] [PubMed]
- Xin, X.F.; He, S.Y. Pseudomonas syringae pv. tomato DC3000: A model pathogen for probing disease susceptibility and hormone signaling in plants. Annu. Rev. Phytopathol. 2013, 51, 473–498. [Google Scholar] [CrossRef]
- Block, A.; Alfano, J.R. Plant targets for Pseudomonas syringae type III effectors: Virulence targets or guarded decoys? Curr. Opin. Microbiol. 2011, 14, 39–46. [Google Scholar] [CrossRef] [PubMed]
| Sampling Location | Growing System | Sample ID | Sampling Date | Symptoms |
|---|---|---|---|---|
| Holovousy (50.419383 N, 15.665607 E) | IPM | AI362N | 28 June 2019 | Apoplexy |
| AI373N | Apoplexy | |||
| AG854N | 4 September 2018 | Branch dieback, leaf wilting | ||
| AH180N | 12 October 2018 | Leaf wilting | ||
| AH166N | Apoplexy | |||
| AG797N | 31 August 2018 | Stunting, premature leaf dropping | ||
| Martinice (49.0059833 N, 16.8208667 E) | IPM | 109N | 3 May 2019 | Branch dieback |
| 110BN | Branch dieback | |||
| 111N | Branch dieback | |||
| 114N | Branch dieback | |||
| 115N | Branch dieback | |||
| 117N | Branch dieback | |||
| 118N | Branch dieback | |||
| Lednice (48.7920883 N, 16.7951131 E) | ORG | AK227N | 3 August 2020 | Stunting, bark cracking |
| AK228N | Branch dieback | |||
| AK229N | Shot hole | |||
| AK230N | Leaf yellowing | |||
| AK231N | Branch dieback | |||
| Velké Pavlovice (48.9161167 N, 16.8181167 E) | ORG | 101N | 3 May 2019 | Apoplexy |
| 103N | Apoplexy |
| Identified Microorganism | Primer or Probe | Sequence (5’-3’) | Final Concentration [µM] |
|---|---|---|---|
| ‘Candidatus Phytoplasma prunorum’ | Forward | GCAGCTGCGGTAATACATGG | 0.50 |
| Reverse | GAATTCCACTTGCCTCTATCCAA | 0.50 | |
| Probe | AGTTCAACGCTTAACGTTGTGATGCTAT | 0.25 | |
| Pseudomonas syringae | Forward | TCGAGCGGCAGCACGGGT | 0.50 |
| Reverse | AGGCCCGAAGGTCCCCTG | 0.50 | |
| Probe | TTGTACCTGGTGGCGAGCGG | 0.25 | |
| Cytospora sp. | Forward | ACCCAGAAACCCTTTGTGAACTTAT | 0.50 |
| Reverse | CCGGCGGGCCTGCTGTCC | 0.50 | |
| Probe | CGTTGCCTCGGCGCTGGCTGC | 0.25 |
| Sample ID | Growing System | PCR ESFY | PCR PS | PCR Cyt | HTS Cyt |
|---|---|---|---|---|---|
| AI362N | IPM | − | − | + | + |
| AI373N | − | − | + | + | |
| AG854N | − | − | + | + | |
| AH180N | − | − | + | + | |
| AH166N | ? | ? | + | + | |
| AG797N | ? | − | + | + | |
| 109N | IPM | + | + | − | − |
| 110BN | + | + | + | + | |
| 111N | + | + | + | + | |
| 114N | + | + | − | − | |
| 115N | + | + | + | + | |
| 117N | + | + | + | + | |
| 118N | + | + | + | − | |
| AK227N | ORG | + | + | + | − |
| AK228N | + | + | + | + | |
| AK229N | + | + | + | + | |
| AK230N | − | + | − | − | |
| AK231N | + | + | + | − | |
| 101N | ORG | + | + | − | + |
| 103N | + | + | + | + |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Navrátil, M.; Šafářová, D.; Čmejla, R.; Duchoslav, M.; Sedlák, J. High-Throughput Sequencing Reveals Fungal Microbiome of Apricots Grown Under Organic and Integrated Pest Management Systems. Agriculture 2025, 15, 1825. https://doi.org/10.3390/agriculture15171825
Navrátil M, Šafářová D, Čmejla R, Duchoslav M, Sedlák J. High-Throughput Sequencing Reveals Fungal Microbiome of Apricots Grown Under Organic and Integrated Pest Management Systems. Agriculture. 2025; 15(17):1825. https://doi.org/10.3390/agriculture15171825
Chicago/Turabian StyleNavrátil, Milan, Dana Šafářová, Radek Čmejla, Martin Duchoslav, and Jiří Sedlák. 2025. "High-Throughput Sequencing Reveals Fungal Microbiome of Apricots Grown Under Organic and Integrated Pest Management Systems" Agriculture 15, no. 17: 1825. https://doi.org/10.3390/agriculture15171825
APA StyleNavrátil, M., Šafářová, D., Čmejla, R., Duchoslav, M., & Sedlák, J. (2025). High-Throughput Sequencing Reveals Fungal Microbiome of Apricots Grown Under Organic and Integrated Pest Management Systems. Agriculture, 15(17), 1825. https://doi.org/10.3390/agriculture15171825





