Multimodal Approach to Assessment of Fecal Microbiota Donors based on Three Complementary Methods
Abstract
1. Introduction
2. Material and Methods
2.1. Stool Donors
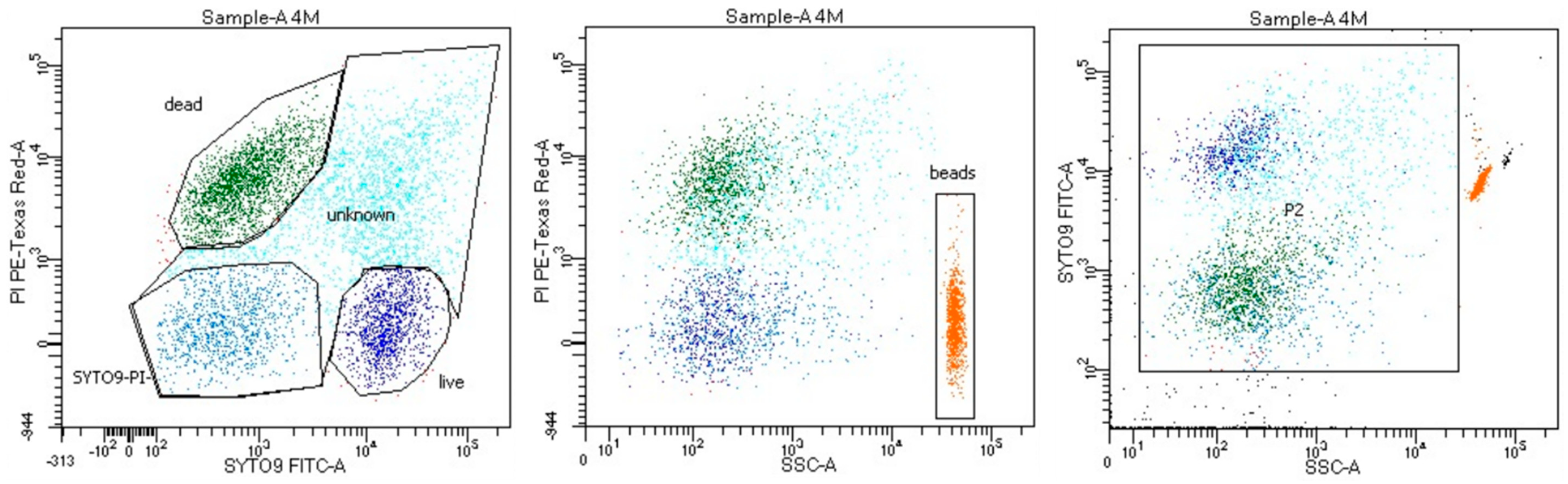
2.2. Flow Cytometry
2.3. Cultivation of Stool Microbiota
2.4. DNA Sequencing
2.5. Bioinformatic Analysis
2.6. Statistical Analysis
2.7. Sequencing Data Availability
2.8. Ethics
3. Results
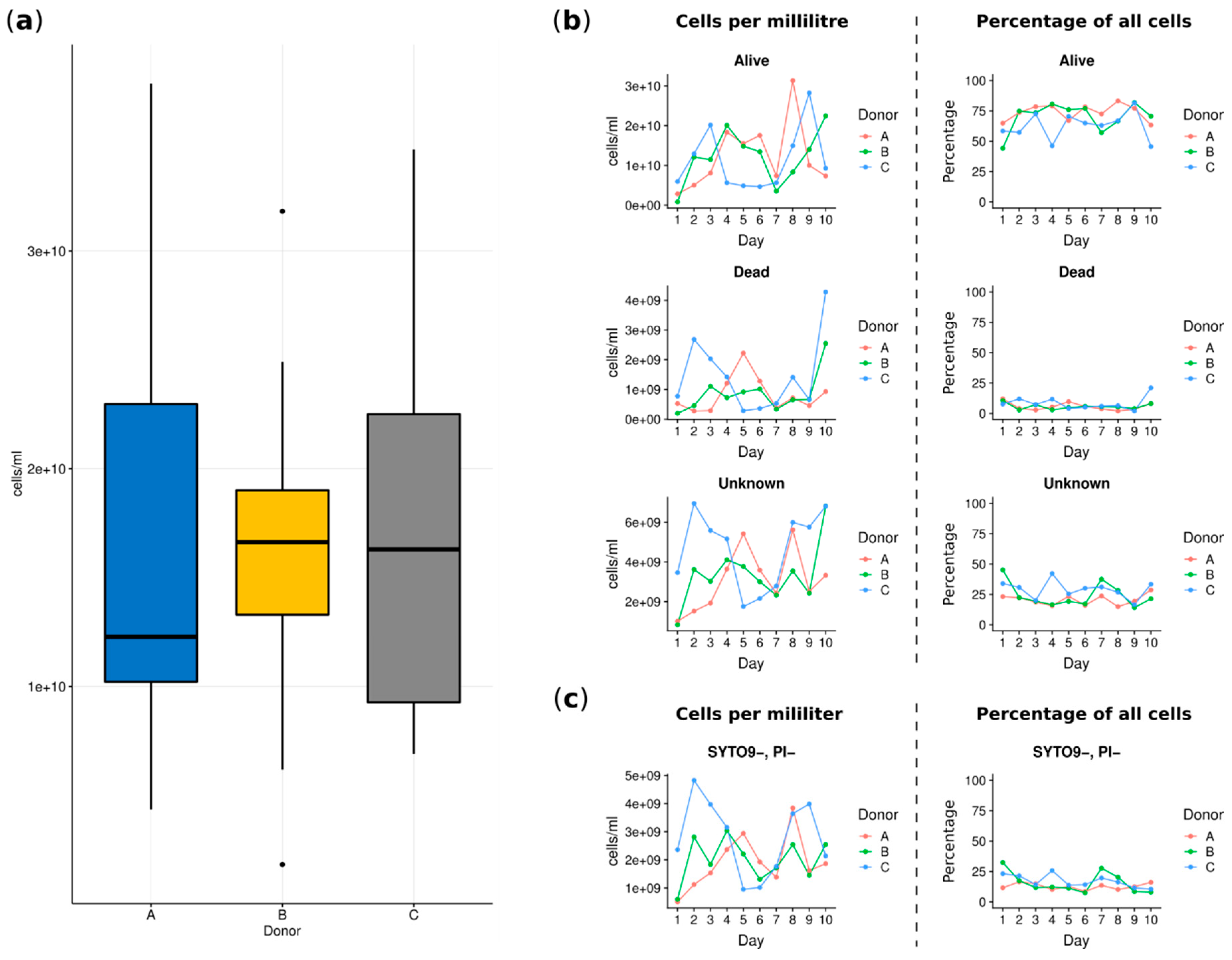
3.1. Analyses Applying the Flow Cytometry
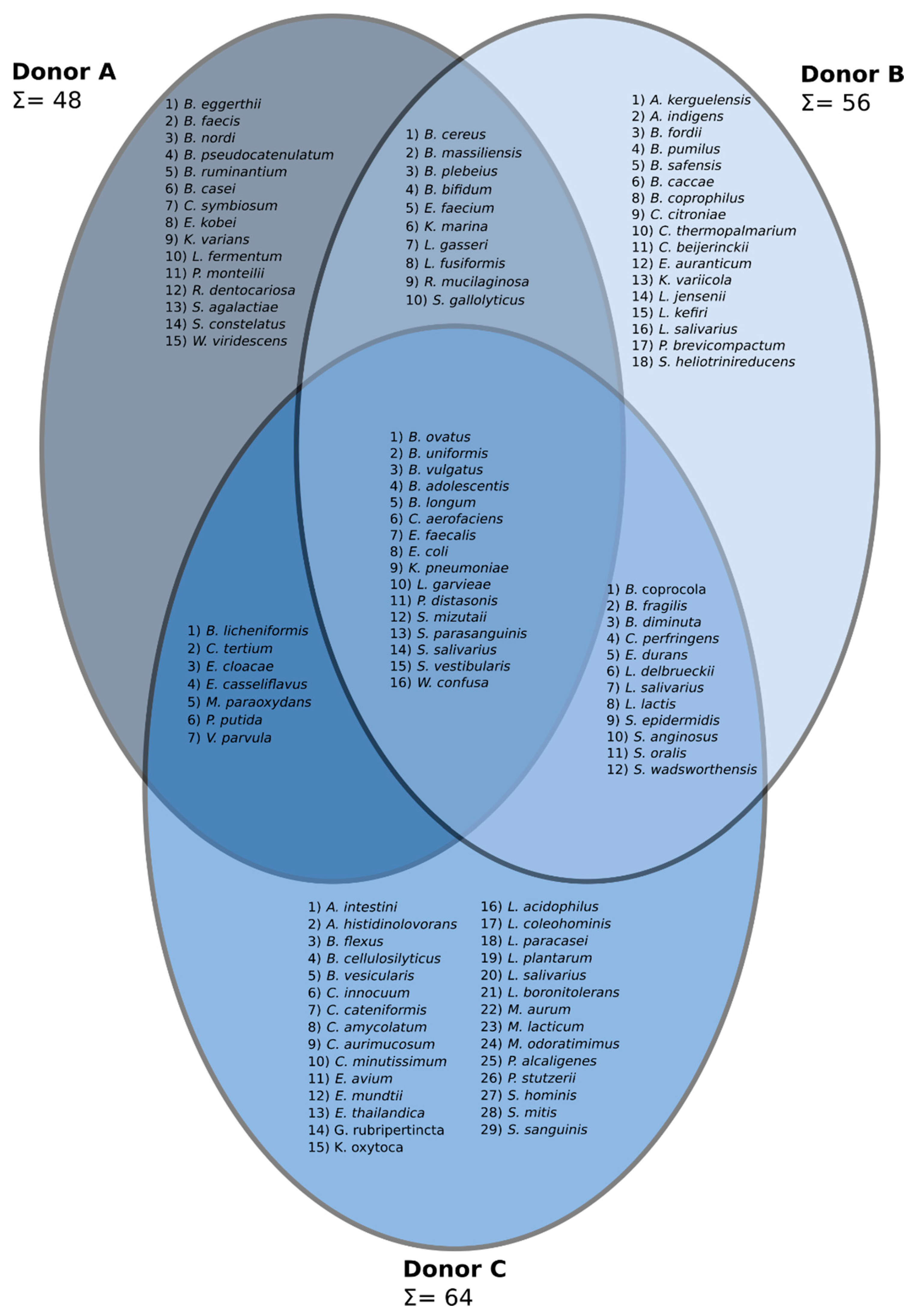
3.2. Cultivation of Stool Microbiota—Classical Microbiological Evaluation
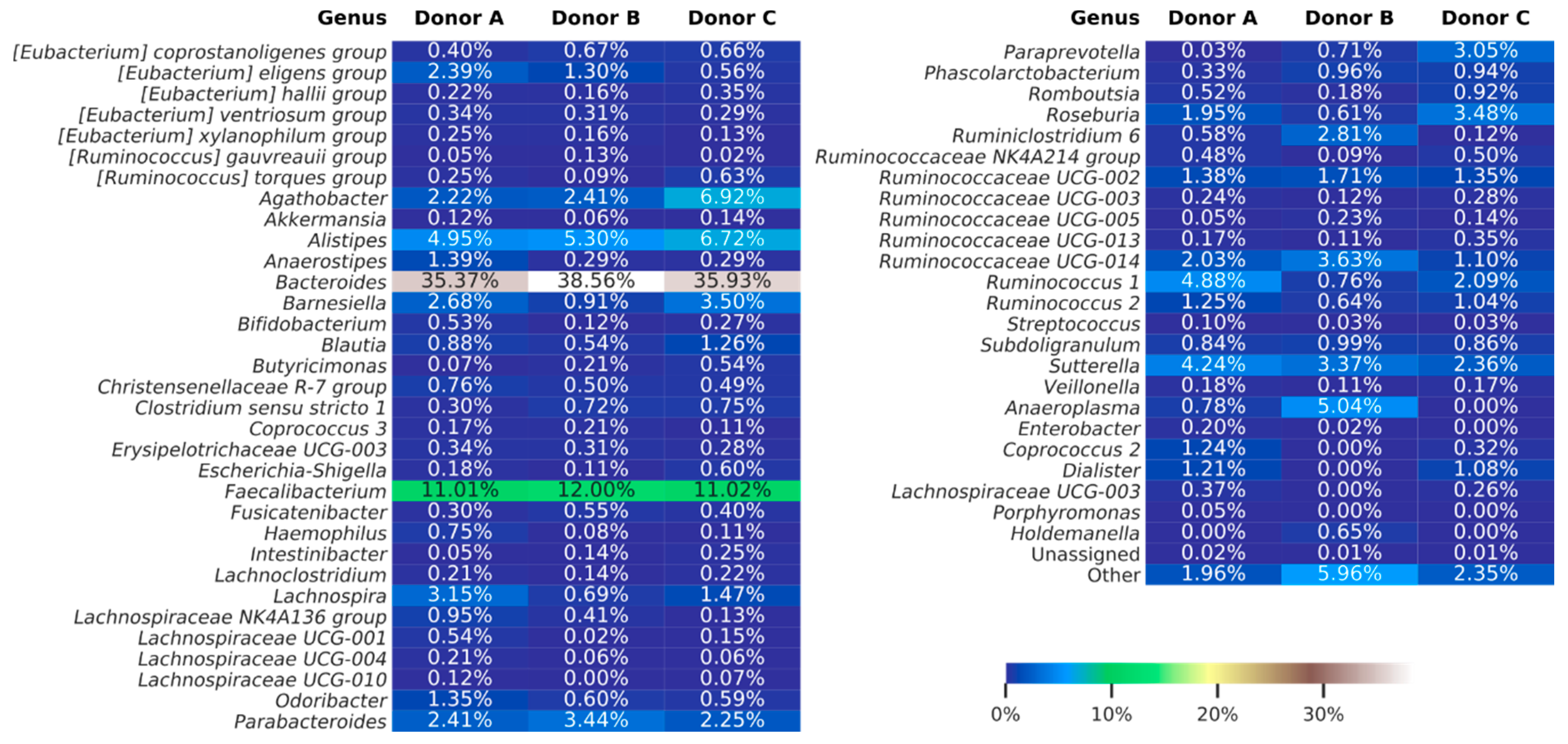
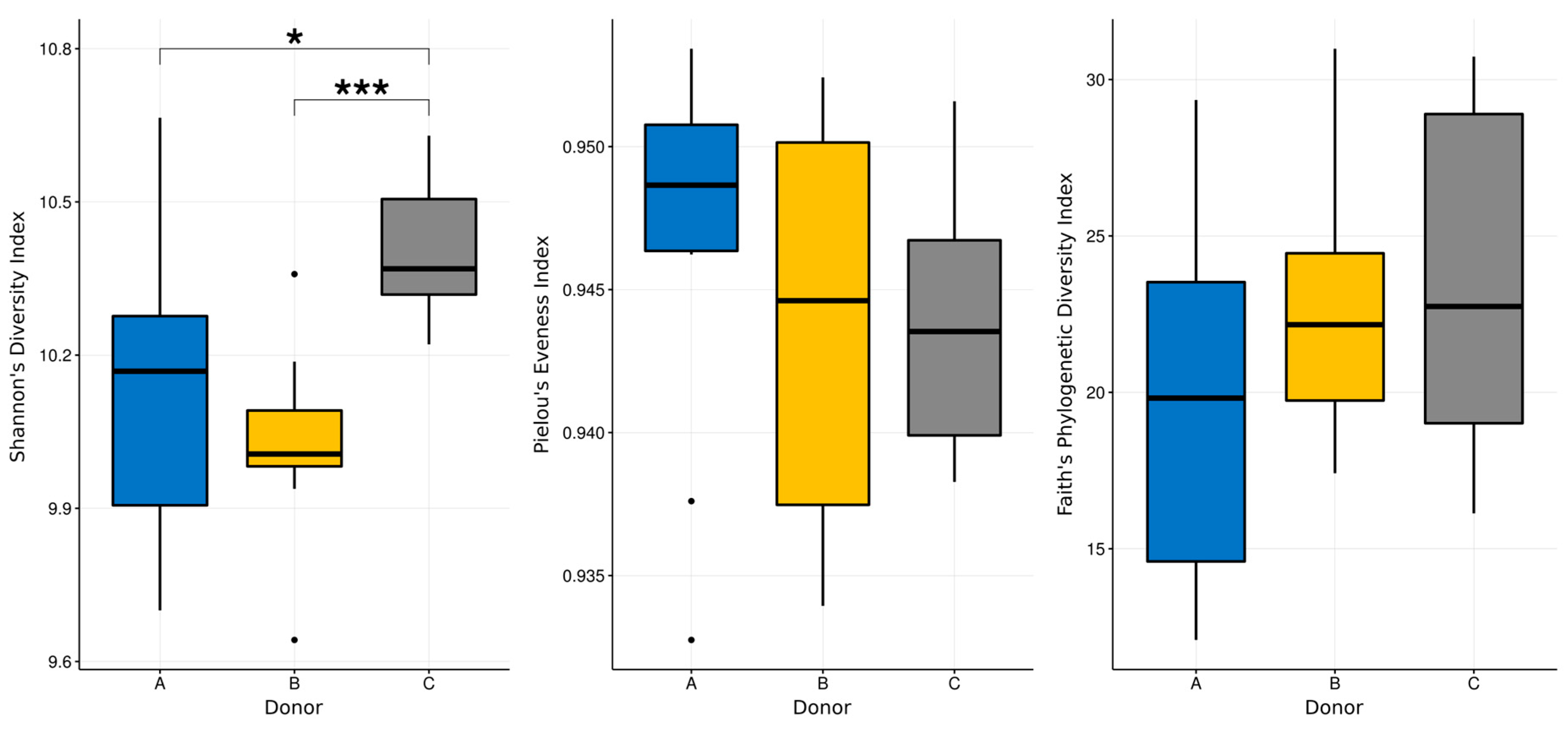
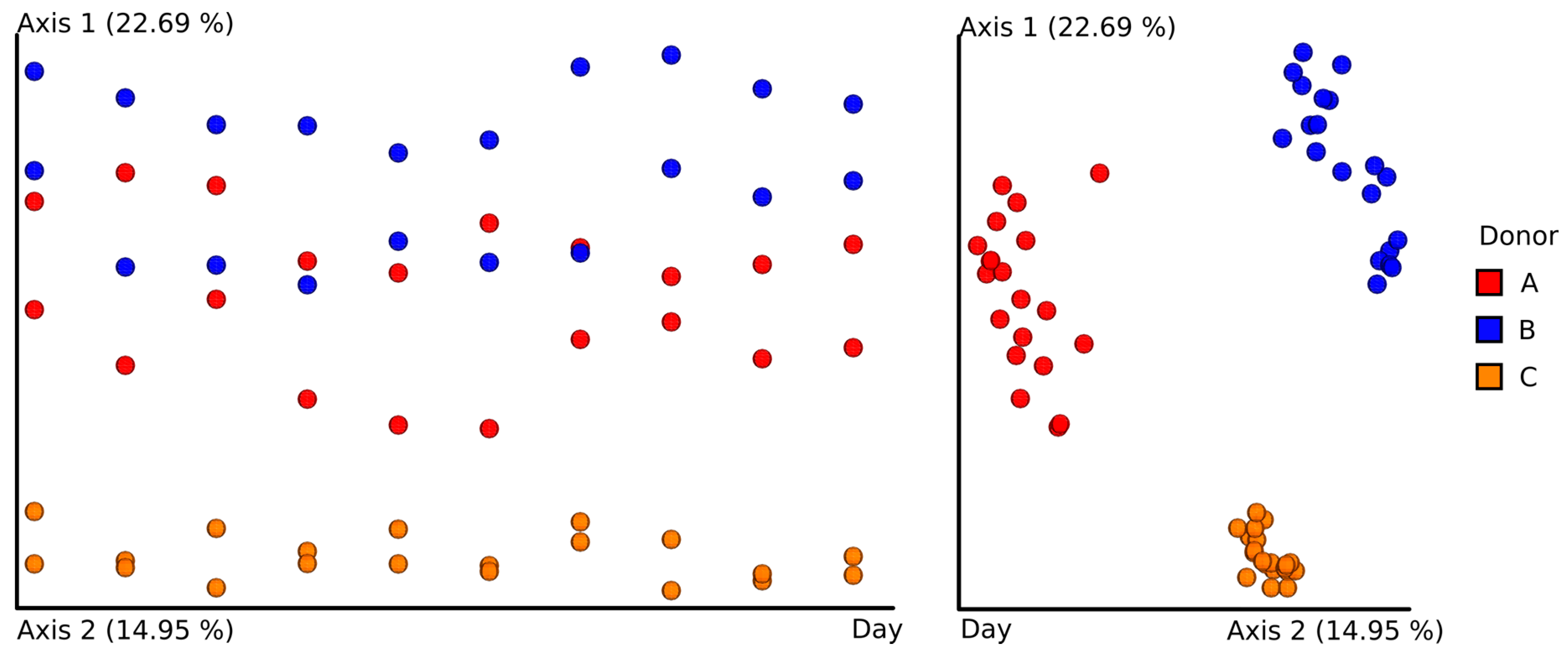
3.3. Next-Generation Sequencing
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Costello, S.P.; Soo, W.; Bryant, R.V.; Jairath, V.; Hart, A.L.; Andrews, J.M. Systematic review with meta-analysis: Faecal microbiota transplantation for the induction of remission for active ulcerative colitis. Aliment. Pharmacol. Ther. 2017, 46, 213–224. [Google Scholar] [CrossRef] [PubMed]
- Ianiro, G.; Murri, R.; Sciumè, G.D.; Impagnatiello, M.; Masucci, L.; Ford, A.C.; Law, G.R.; Tilg, H.; Sanguinetti, M.; Cauda, R.; et al. Incidence of Bloodstream Infections, Length of Hospital Stay, and Survival in Patients With Recurrent Clostridioides difficile Infection Treated With Fecal Microbiota Transplantation or Antibiotics: A Prospective Cohort Study. Ann. Intern. Med. 2019, 171, 695–702. [Google Scholar] [CrossRef] [PubMed]
- Hocquart, M.; Lagier, J.-C.; Cassir, N.; Saidani, N.; Eldin, C.; Kerbaj, J.; Delord, M.; Valles, C.; Brouqui, P.; Raoult, D.; et al. Early Fecal Microbiota Transplantation Improves Survival in Severe Clostridium difficile Infections. Clin. Infect. Dis. 2018, 66, 645–650. [Google Scholar] [CrossRef] [PubMed]
- Map of Clinical Trials Involving Fecal Microbiota Transplantation. Available online: https://www.clinicaltrials.gov/ct2/results/map?term=fecal+microbiota+transplantation&map (accessed on 9 January 2020).
- Mukhopadhya, I.; Segal, J.P.; Carding, S.R.; Hart, A.L.; Hold, G.L. The gut virome: The missing link between gut bacteria and host immunity? Therap. Adv. Gastroenterol. 2019, 12. [Google Scholar] [CrossRef] [PubMed]
- Ianiro, G.; Bruno, G.; Lopetuso, L.; Beghella, F.; Laterza, L.; D’Aversa, F.; Gigante, G.; Cammarota, G.; Gasbarrini, A. Role of Yeasts in Healthy and Impaired Gut Microbiota: The Gut Mycome. Curr. Pharm. Des. 2014, 20, 4565–4569. [Google Scholar] [CrossRef]
- Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature 2012, 486, 207–214. [Google Scholar] [CrossRef]
- Claesson, M.J.; O’Toole, P.W. Evaluating the latest high-throughput molecular techniques for the exploration of microbial gut communities. Gut Microbes 2010, 1, 277–278. [Google Scholar] [CrossRef]
- D’Amore, R.; Ijaz, U.Z.; Schirmer, M.; Kenny, J.G.; Gregory, R.; Darby, A.C.; Shakya, M.; Podar, M.; Quince, C.; Hall, N. A comprehensive benchmarking study of protocols and sequencing platforms for 16S rRNA community profiling. BMC Genomics 2016, 17, 55. [Google Scholar] [CrossRef]
- Lau, J.T.; Whelan, F.J.; Herath, I.; Lee, C.H.; Collins, S.M.; Bercik, P.; Surette, M.G. Capturing the diversity of the human gut microbiota through culture-enriched molecular profiling. Genome Med. 2016, 8, 72. [Google Scholar] [CrossRef]
- Hiergeist, A.; Glasner, J.; Reischl, U.; Gessner, A. Analyses of Intestinal Microbiota: Culture versus Sequencing. ILAR J. 2015, 56, 228–240. [Google Scholar] [CrossRef]
- Fraher, M.H.; O’Toole, P.W.; Quigley, E.M.M. Techniques used to characterize the gut microbiota: A guide for the clinician. Nat. Rev. Gastroenterol. Hepatol. 2012, 9, 312–322. [Google Scholar] [CrossRef] [PubMed]
- Gupta, S.; Mortensen, M.S.; Schjorring, S.; Trivedi, U.; Vestergaard, G.; Stokholm, J.; Bisgaard, H.; Krogfelt, K.A.; Sorensen, S.J. Amplicon sequencing provides more accurate microbiome information in healthy children compared to culturing. Commun. Biol. 2019, 2, 291. [Google Scholar] [CrossRef]
- Wilson, B.C.; Vatanen, T.; Cutfield, W.S.; O’Sullivan, J.M. The Super-Donor Phenomenon in Fecal Microbiota Transplantation. Front. Cell. Infect. Microbiol. 2019, 9, 2. [Google Scholar] [CrossRef] [PubMed]
- Olesen, S.W.; Gerardin, Y. Re-evaluating the evidence for fecal microbiota transplantation “super-donors” in inflammatory bowel disease. medRxiv 2020, 19011635. [Google Scholar] [CrossRef]
- Ng, S.C.; Kamm, M.A.; Yeoh, Y.K.; Chan, P.K.S.; Zuo, T.; Tang, W.; Sood, A.; Andoh, A.; Ohmiya, N.; Zhou, Y.; et al. Scientific frontiers in faecal microbiota transplantation: Joint document of Asia-Pacific Association of Gastroenterology (APAGE) and Asia-Pacific Society for Digestive Endoscopy (APSDE). Gut 2020, 69, 83–91. [Google Scholar] [CrossRef]
- El-Salhy, M.; Hatlebakk, J.G.; Gilja, O.H.; Bråthen Kristoffersen, A.; Hausken, T. Efficacy of faecal microbiota transplantation for patients with irritable bowel syndrome in a randomised, double-blind, placebo-controlled study. Gut 2020, 69, 859–867. [Google Scholar] [CrossRef]
- Bilinski, J.; Grzesiowski, P.; Sorensen, N.; Madry, K.; Muszynski, J.; Robak, K.; Wroblewska, M.; Dzieciatkowski, T.; Dulny, G.; Dwilewicz-Trojaczek, J.; et al. Microbiota Transplantation in Patients With Blood Disorders Inhibits Gut Colonization With Antibiotic-Resistant Bacteria: Results of a Prospective, Single-Center Study. Clin. Infect. Dis. 2017, 65, 364–370. [Google Scholar] [CrossRef]
- Cammarota, G.; Ianiro, G.; Kelly, C.R.; Mullish, B.H.; Allegretti, J.R.; Kassam, Z.; Putignani, L.; Fischer, M.; Keller, J.J.; Costello, S.P.; et al. International consensus conference on stool banking for faecal microbiota transplantation in clinical practice. Gut 2019, 68, 2111–2121. [Google Scholar] [CrossRef]
- Molecular Probes LIVE/DEAD BacLight Bacterial Viability and Counting Kit (L34856)—Protocol. Available online: https://assets.thermofisher.com/TFS-Assets/LSG/manuals/mp34856.pdf (accessed on 31 May 2020).
- Herlemann, D.P.; Labrenz, M.; Jurgens, K.; Bertilsson, S.; Waniek, J.J.; Andersson, A.F. Transitions in bacterial communities along the 2000 km salinity gradient of the Baltic Sea. ISME J. 2011, 5, 1571–1579. [Google Scholar] [CrossRef]
- Klindworth, A.; Pruesse, E.; Schweer, T.; Peplies, J.; Quast, C.; Horn, M.; Glockner, F.O. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 2013, 41, e1. [Google Scholar] [CrossRef]
- Rodriguez-R, L.M.; Gunturu, S.; Tiedje, J.M.; Cole, J.R.; Konstantinidis, K.T. Nonpareil 3: Fast Estimation of Metagenomic Coverage and Sequence Diversity. mSystems 2018, 3. [Google Scholar] [CrossRef]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A.; Abnet, C.C.; Al-Ghalith, G.A.; Alexander, H.; Alm, E.J.; Arumugam, M.; Asnicar, F.; et al. Reproducible, interactive, scalable and extensible microbiome data science usin QIIME 2. Nat. Biotechnol. 2019, 37, 852–857. [Google Scholar] [CrossRef]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.A.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef] [PubMed]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glöckner, F.O. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 2012, 41, D590–D596. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef]
- Price, M.N.; Dehal, P.S.; Arkin, A.P. FastTree 2—Approximately Maximum-Likelihood Trees for Large Alignments. PLoS ONE 2010, 5. [Google Scholar] [CrossRef] [PubMed]
- Vazquez-Baeza, Y.; Pirrung, M.; Gonzalez, A.; Knight, R. EMPeror: A tool for visualizing high-throughput microbial community data. Gigascience 2013, 2, 16. [Google Scholar] [CrossRef]
- Mandal, S.; Van Treuren, W.; White, R.A.; Eggesbo, M.; Knight, R.; Peddada, S.D. Analysis of composition of microbiomes: A novel method for studying microbial composition. Microb. Ecol. Health Dis. 2015, 26, 27663. [Google Scholar] [CrossRef]
- Wickham, H. ggplot2, 2nd ed.; Use R! Springer International Publishing: Berlin/Heidelberg, Germany, 2016; ISBN 978-3-319-24275-0. [Google Scholar]
- Wilke, C. Cowplot—Streamlined Plot Theme and Plot Annotations for Ggplot2. Available online: https://github.com/wilkelab/cowplot (accessed on 7 January 2020).
- Kassambara, A. Ggpubr: ‘Ggplot2’ Based Publication Ready Plots. Available online: https://github.com/kassambara/ggpubr (accessed on 7 January 2020).
- Barb, J.J.; Oler, A.J.; Kim, H.-S.; Chalmers, N.; Wallen, G.R.; Cashion, A.; Munson, P.J.; Ames, N.J. Development of an Analysis Pipeline Characterizing Multiple Hypervariable Regions of 16S rRNA Using Mock Samples. PLoS ONE 2016, 11, e0148047. [Google Scholar] [CrossRef]
- David, L.A.; Maurice, C.F.; Carmody, R.N.; Gootenberg, D.B.; Button, J.E.; Wolfe, B.E.; Ling, A.V.; Devlin, A.S.; Varma, Y.; Fischbach, M.A.; et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature 2014, 505, 559–563. [Google Scholar] [CrossRef]
- Ott, S.J.; Waetzig, G.H.; Rehman, A.; Moltzau-Anderson, J.; Bharti, R.; Grasis, J.A.; Cassidy, L.; Tholey, A.; Fickenscher, H.; Seegert, D.; et al. Efficacy of Sterile Fecal Filtrate Transfer for Treating Patients With Clostridium difficile Infection. Gastroenterology 2017, 152, 799–811.e7. [Google Scholar] [CrossRef] [PubMed]
- Papanicolas, L.E.; Choo, J.M.; Wang, Y.; Leong, L.E.X.; Costello, S.P.; Gordon, D.L.; Wesselingh, S.L.; Rogers, G.B. Bacterial viability in faecal transplants: Which bacteria survive? EBioMedicine 2019, 41, 509–516. [Google Scholar] [CrossRef] [PubMed]
- Rajan, S.K.; Lindqvist, M.; Brummer, R.J.; Schoultz, I.; Repsilber, D. Phylogenetic microbiota profiling in fecal samples depends on combination of sequencing depth and choice of NGS analysis method. PLoS ONE 2019, 14, e0222171. [Google Scholar] [CrossRef]
- Lloyd-Price, J.; Abu-Ali, G.; Huttenhower, C. The healthy human microbiome. Genome Med. 2016, 8, 51. [Google Scholar] [CrossRef] [PubMed]
- Peled, J.U.; Gomes, A.L.C.; Devlin, S.M.; Littmann, E.R.; Taur, Y.; Sung, A.D.; Weber, D.; Hashimoto, D.; Slingerland, A.E.; Slingerland, J.B.; et al. Microbiota as Predictor of Mortality in Allogeneic Hematopoietic-Cell Transplantation. N. Engl. J. Med. 2020, 382, 822–834. [Google Scholar] [CrossRef]
- Shafquat, A.; Joice, R.; Simmons, S.L.; Huttenhower, C. Functional and phylogenetic assembly of microbial communities in the human microbiome. Trends Microbiol. 2014, 22, 261–266. [Google Scholar] [CrossRef]
- Jumas-Bilak, E.; Carlier, J.-P.; Jean-Pierre, H.; Mory, F.; Teyssier, C.; Gay, B.; Campos, J.; Marchandin, H. Acidaminococcus intestini sp. nov., isolated from human clinical samples. Int. J. Syst. Evol. Microbiol. 2007, 57, 2314–2319. [Google Scholar] [CrossRef]
- Cook, G.M.; Rainey, F.A.; Chen, G.; Stackebrandt, E.; Russell, J.B. Emendation of the description of Acidaminococcus fermentans, a trans-aconitate- and citrate-oxidizing bacterium. Int. J. Syst. Bacteriol. 1994, 44, 576–578. [Google Scholar] [CrossRef]
- Rajilic-Stojanovic, M.; Smidt, H.; de Vos, W.M. Diversity of the human gastrointestinal tract microbiota revisited. Environ. Microbiol. 2007, 9, 2125–2136. [Google Scholar] [CrossRef]
- Stanislawski, M.A.; Lozupone, C.A.; Wagner, B.D.; Eggesbo, M.; Sontag, M.K.; Nusbacher, N.M.; Martinez, M.; Dabelea, D. Gut microbiota in adolescents and the association with fatty liver: The EPOCH study. Pediatr. Res. 2018, 84, 219–227. [Google Scholar] [CrossRef]
- Lun, H.; Yang, W.; Zhao, S.; Jiang, M.; Xu, M.; Liu, F.; Wang, Y. Altered gut microbiota and microbial biomarkers associated with chronic kidney disease. Microbiologyopen 2019, 8, e00678. [Google Scholar] [CrossRef] [PubMed]
- Meehan, C.J.; Beiko, R.G. A phylogenomic view of ecological specialization in the Lachnospiraceae, a family of digestive tract-associated bacteria. Genome Biol. Evol. 2014, 6, 703–713. [Google Scholar] [CrossRef] [PubMed]
- Rocas, I.N.; Siqueira, J.F.J. Characterization of Dialister species in infected root canals. J. Endod. 2006, 32, 1057–1061. [Google Scholar] [CrossRef] [PubMed]
- Hiranmayi, K.V.; Sirisha, K.; Ramoji Rao, M.V.; Sudhakar, P. Novel Pathogens in Periodontal Microbiology. J. Pharm. Bioallied Sci. 2017, 9, 155–163. [Google Scholar] [CrossRef]
- Vermeire, S.; Joossens, M.; Verbeke, K.; Wang, J.; Machiels, K.; Sabino, J.; Ferrante, M.; Van Assche, G.; Rutgeerts, P.; Raes, J. Donor Species Richness Determines Faecal Microbiota Transplantation Success in Inflammatory Bowel Disease. J. Crohns. Colitis 2016, 10, 387–394. [Google Scholar] [CrossRef]
- Kump, P.; Wurm, P.; Grochenig, H.P.; Wenzl, H.; Petritsch, W.; Halwachs, B.; Wagner, M.; Stadlbauer, V.; Eherer, A.; Hoffmann, K.M.; et al. The taxonomic composition of the donor intestinal microbiota is a major factor influencing the efficacy of faecal microbiota transplantation in therapy refractory ulcerative colitis. Aliment. Pharmacol. Ther. 2018, 47, 67–77. [Google Scholar] [CrossRef] [PubMed]

© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bilinski, J.; Dziurzynski, M.; Grzesiowski, P.; Podsiadly, E.; Stelmaszczyk-Emmel, A.; Dzieciatkowski, T.; Dziewit, L.; Basak, G.W. Multimodal Approach to Assessment of Fecal Microbiota Donors based on Three Complementary Methods. J. Clin. Med. 2020, 9, 2036. https://doi.org/10.3390/jcm9072036
Bilinski J, Dziurzynski M, Grzesiowski P, Podsiadly E, Stelmaszczyk-Emmel A, Dzieciatkowski T, Dziewit L, Basak GW. Multimodal Approach to Assessment of Fecal Microbiota Donors based on Three Complementary Methods. Journal of Clinical Medicine. 2020; 9(7):2036. https://doi.org/10.3390/jcm9072036
Chicago/Turabian StyleBilinski, Jaroslaw, Mikolaj Dziurzynski, Pawel Grzesiowski, Edyta Podsiadly, Anna Stelmaszczyk-Emmel, Tomasz Dzieciatkowski, Lukasz Dziewit, and Grzegorz W. Basak. 2020. "Multimodal Approach to Assessment of Fecal Microbiota Donors based on Three Complementary Methods" Journal of Clinical Medicine 9, no. 7: 2036. https://doi.org/10.3390/jcm9072036
APA StyleBilinski, J., Dziurzynski, M., Grzesiowski, P., Podsiadly, E., Stelmaszczyk-Emmel, A., Dzieciatkowski, T., Dziewit, L., & Basak, G. W. (2020). Multimodal Approach to Assessment of Fecal Microbiota Donors based on Three Complementary Methods. Journal of Clinical Medicine, 9(7), 2036. https://doi.org/10.3390/jcm9072036




