A Mortality Prediction Rule for Hematology Patients with Invasive Aspergillosis Based on Serum Galactomannan Kinetics
Abstract
1. Introduction
2. Experimental Section
2.1. Study Design
2.1.1. Data Collection
2.1.2. Definitions
2.1.3. Outcome Measurements
2.1.4. Galactomannan Testing
2.2. Statistical Mmethods
3. Results
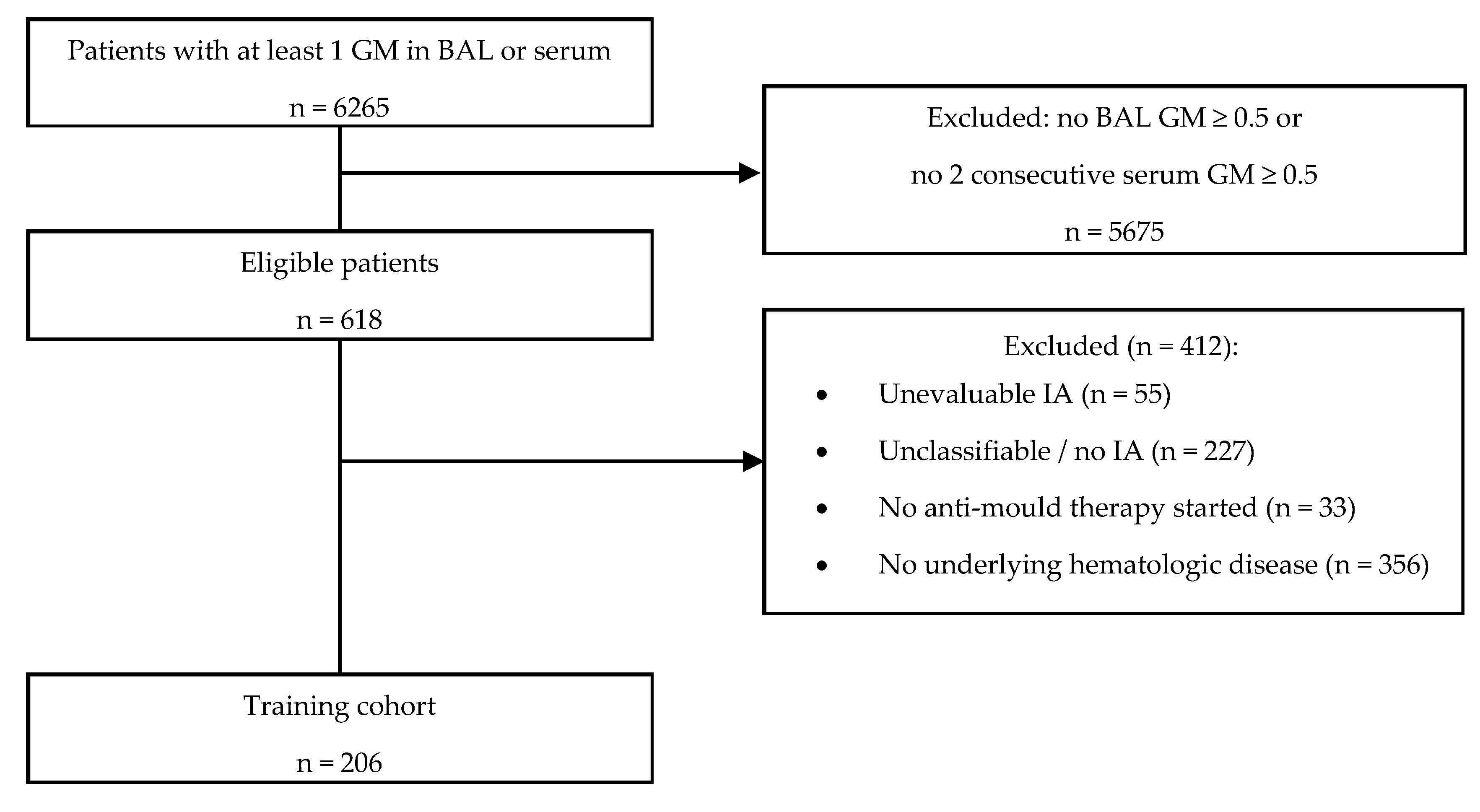
3.1. Patient Disposition and Baseline Characteristics
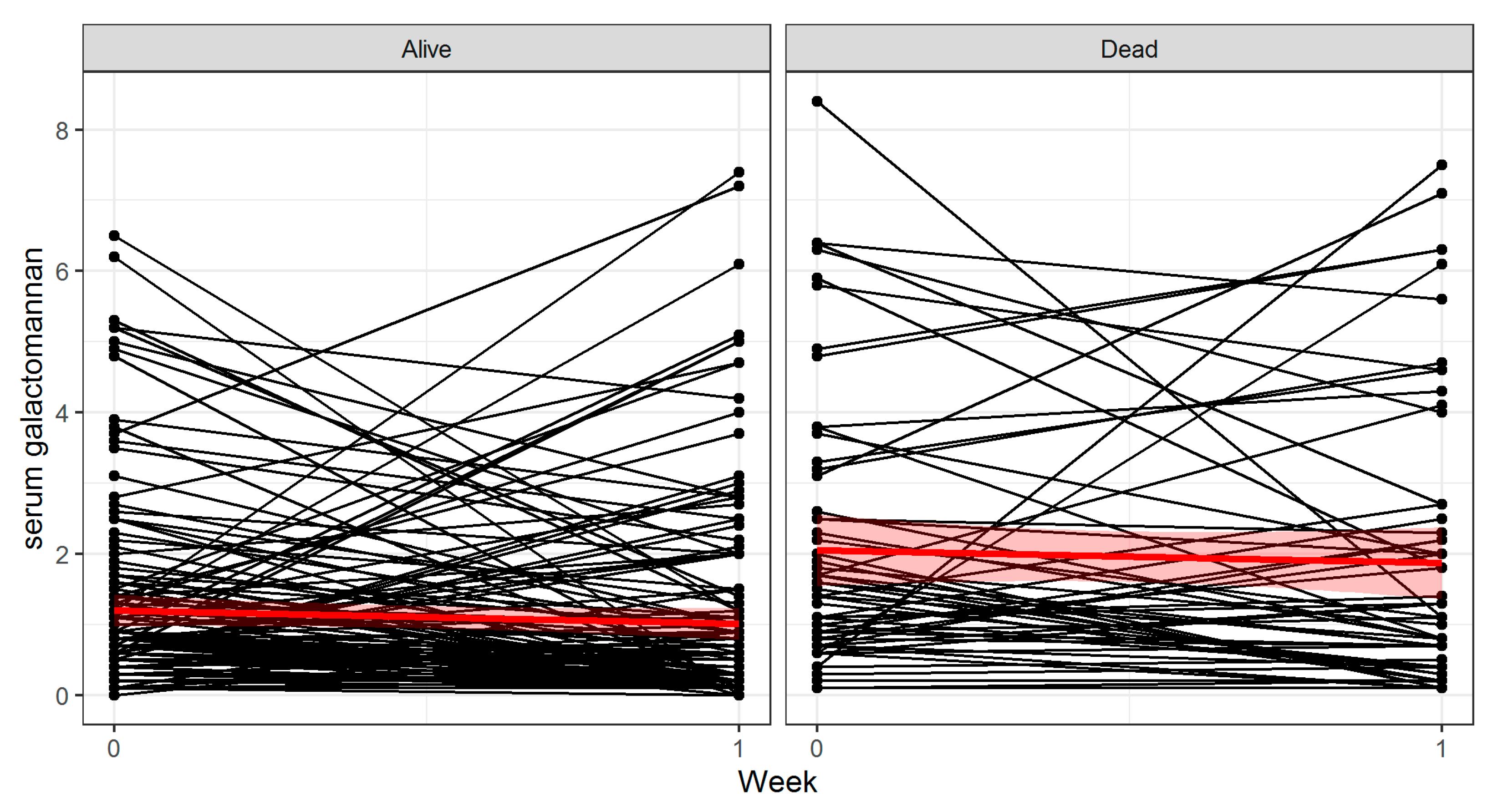
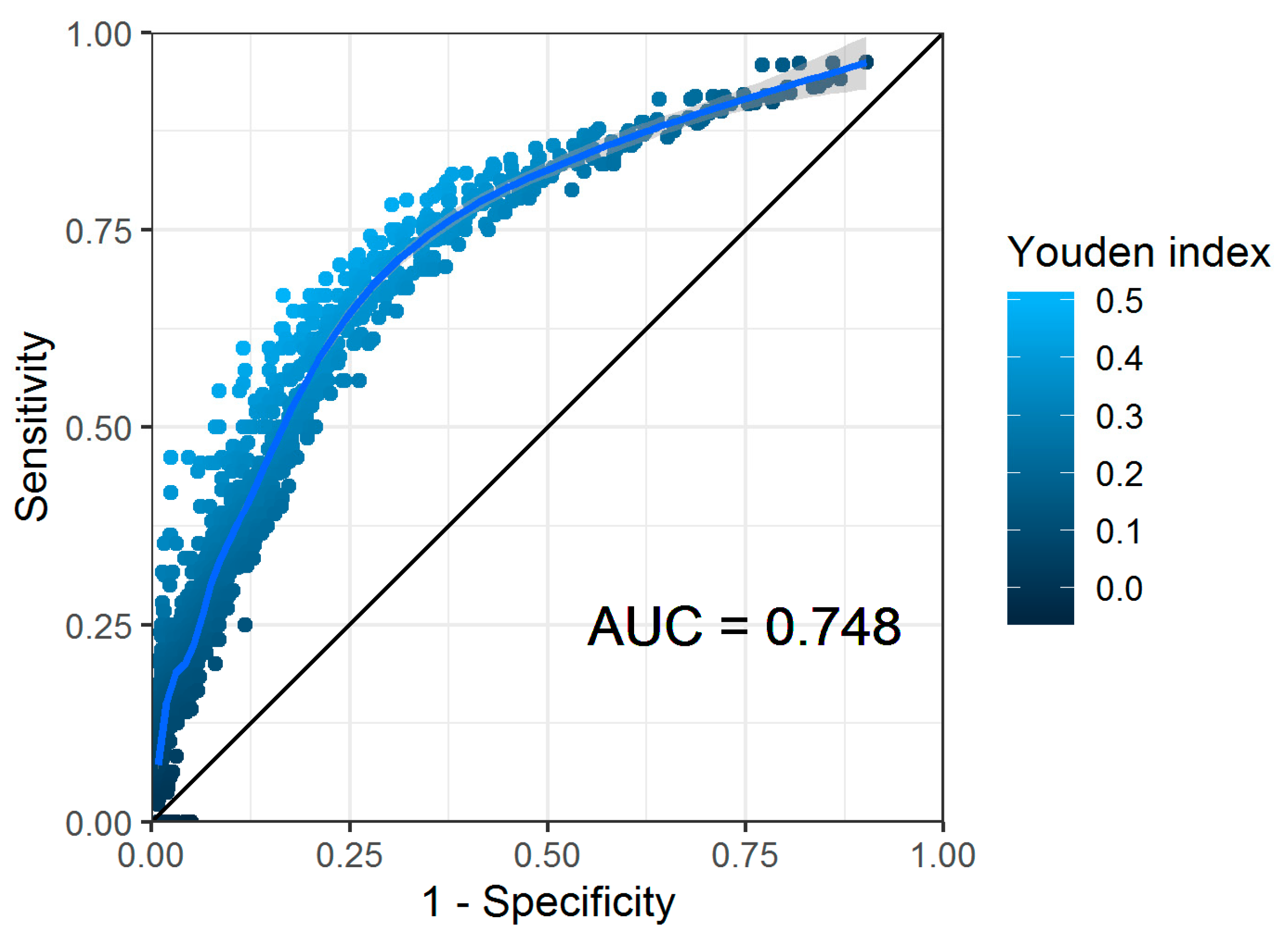
3.2. Model Creation
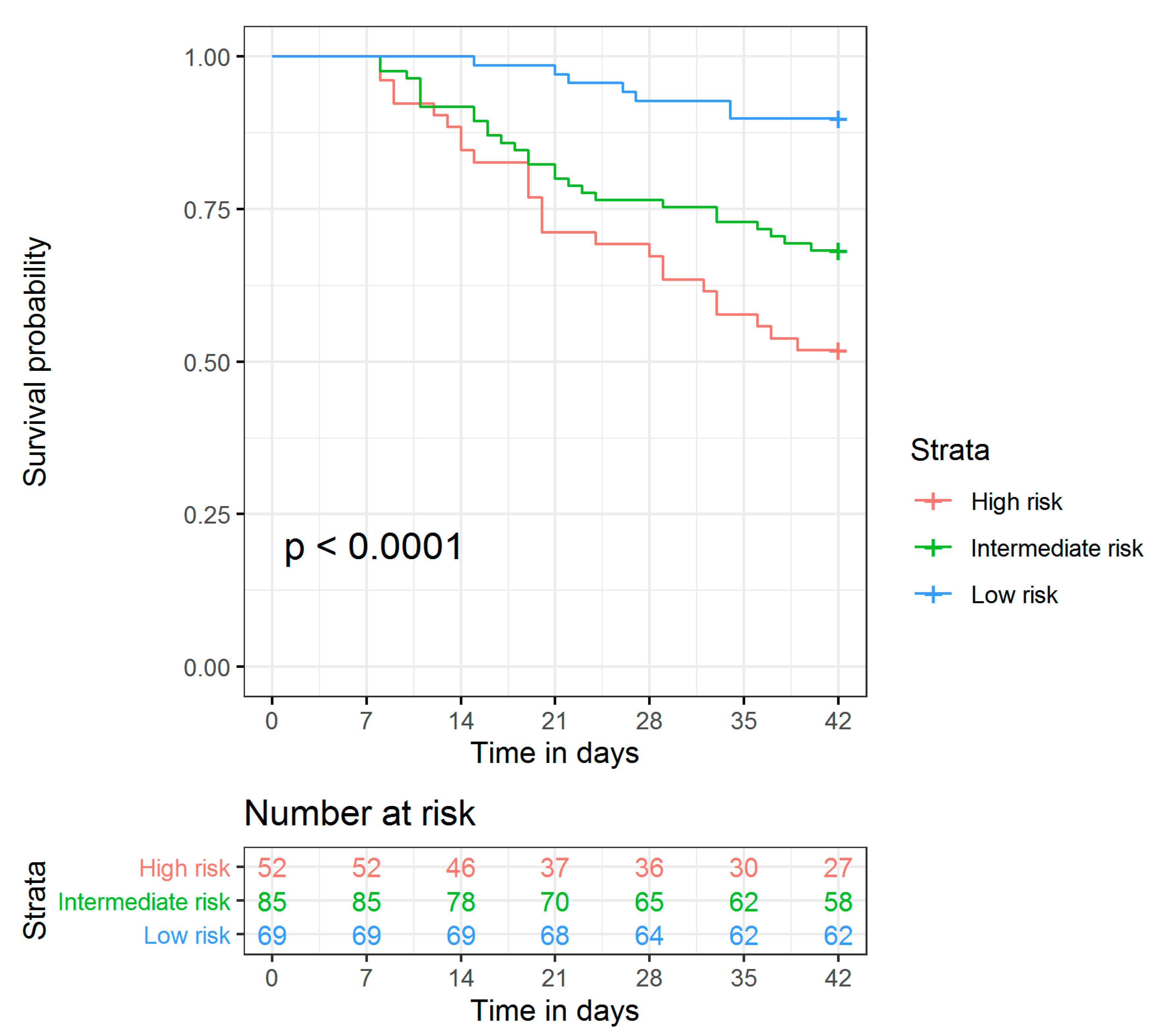
3.3. Model Validation
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Lamoth, F.; Calandra, T. Early diagnosis of invasive mould infections and disease. J. Antimicrob. Chemother. 2017, 72, i19–i28. [Google Scholar] [CrossRef]
- von Eiff, M.; Roos, N.; Schulten, R.; Hesse, M.; Zühlsdorf, M.; van de Loo, J. Pulmonary aspergillosis: Early diagnosis improves survival. Respir. Int. Rev. Thorac. Dis. 1995, 62, 341–347. [Google Scholar] [CrossRef]
- Hope, W.W.; Petraitis, V.; Petraitiene, R.; Aghamolla, T.; Bacher, J.; Walsh, T.J. The Initial 96 Hours of Invasive Pulmonary Aspergillosis: Histopathology, Comparative Kinetics of Galactomannan and (1→3)-β-d-Glucan, and Consequences of Delayed Antifungal Therapy. Antimicrob. Agents Chemother. 2010, 54, 4879–4886. [Google Scholar] [CrossRef] [PubMed]
- de Pauw, B.; Walsh, T.J.; Donnelly, J.P.; Stevens, D.A.; Edwards, J.E.; Calandra, T.; Pappas, P.G.; Maertens, J.; Lortholary, O.; Kauffman, C.A.; et al. Revised Definitions of Invasive Fungal Disease from the European Organization for Research and Treatment of Cancer/Invasive Fungal Infections Cooperative Group and the National Institute of Allergy and Infectious Diseases Mycoses Study Group (EORTC/MSG) Consensus Group. Clin. Infect. Dis. 2008, 46, 1813–1821. [Google Scholar] [PubMed]
- Pfeiffer, C.D.; Fine, J.P.; Safdar, N. Diagnosis of Invasive Aspergillosis Using a Galactomannan Assay: A Meta-Analysis. Clin. Infect. Dis. 2006, 42, 1417–1727. [Google Scholar] [CrossRef] [PubMed]
- Lehrnbecher, T.; Robinson, P.D.; Fisher, B.T.; Castagnola, E.; Groll, A.H.; Steinbach, W.J.; Zaoutis, T.E.; Negeri, Z.F.; Beyene, J.; Phillips, B.; et al. Galactomannan, β-D-Glucan, and Polymerase Chain Reaction–Based Assays for the Diagnosis of Invasive Fungal Disease in Pediatric Cancer and Hematopoietic Stem Cell Transplantation: A Systematic Review and Meta-Analysis. Clin. Infect. Dis. 2016, 63, 1340–1348. [Google Scholar] [CrossRef] [PubMed]
- Arvanitis, M.; Anagnostou, T.; Mylonakis, E. Galactomannan and Polymerase Chain Reaction–Based Screening for Invasive Aspergillosis Among High-Risk Hematology Patients: A Diagnostic Meta-analysis. Clin. Infect. Dis. 2015, 61, 1263–1272. [Google Scholar] [CrossRef] [PubMed]
- Leeflang, M.M.G.; Debets-Ossenkopp, Y.J.; Wang, J.; Visser, C.E.; Scholten, R.J.P.M.; Hooft, L.; Bijlmer, H.A.; Reitsma, J.B.; Zhang, M.; Bossuyt, P.M.M.; et al. Galactomannan detection for invasive aspergillosis in immunocompromised patients. Cochrane Database Syst. Rev. 2015, 2015, CD007394. [Google Scholar] [CrossRef]
- Segal, B.H.; Herbrecht, R.; Stevens, D.A.; Ostrosky-Zeichner, L.; Sobel, J.; Viscoli, C.; Walsh, T.J.; Maertens, J.; Patterson, T.F.; Perfect, J.R.; et al. Defining Responses to Therapy and Study Outcomes in Clinical Trials of Invasive Fungal Diseases: Mycoses Study Group and European Organization for Research and Treatment of Cancer Consensus Criteria. Clin. Infect. Dis. 2008, 47, 674–683. [Google Scholar] [CrossRef]
- Miceli, M.H.; Maertens, J.; Buvé, K.; Grazziutti, M.; Woods, G.; Rahman, M.; Barlogie, B.; Anaissie, E.J. Immune reconstitution inflammatory syndrome in cancer patients with pulmonary aspergillosis recovering from neutropenia: Proof of principle, description, and clinical and research implications. Cancer 2007, 110, 112–120. [Google Scholar] [CrossRef]
- Sheppard, D.C.; Marr, K.A.; Fredricks, D.N.; Chiang, L.Y.; Doedt, T.; Filler, S.G. Comparison of three methodologies for the determination of pulmonary fungal burden in experimental murine aspergillosis. Clin. Microbiol. Infect. 2006, 12, 376–380. [Google Scholar] [CrossRef] [PubMed]
- Kovanda, L.L.; Petraitiene, R.; Petraitis, V.; Walsh, T.J.; Desai, A.; Bonate, P.; Hope, W.W. Pharmacodynamics of isavuconazole in experimental invasive pulmonary aspergillosis: Implications for clinical breakpoints. J. Antimicrob. Chemother. 2016, 71, 1885–1891. [Google Scholar] [CrossRef] [PubMed]
- Petraitiene, R.; Petraitis, V.; Groll, A.H.; Sein, T.; Schaufele, R.L.; Francesconi, A.; Bacher, J.; Avila, N.A.; Walsh, T.J. Antifungal Efficacy of Caspofungin (MK-0991) in Experimental Pulmonary Aspergillosis in Persistently Neutropenic Rabbits: Pharmacokinetics, Drug Disposition, and Relationship to Galactomannan Antigenemia. Antimicrob. Agents Chemother. 2002, 46, 12–23. [Google Scholar] [CrossRef] [PubMed]
- Bennett, J.E.; Friedman, M.M.; Dupont, B. Receptor-mediated clearance of Aspergillus galactomannan. J. Infect. Dis. 1987, 155, 1005–1010. [Google Scholar] [CrossRef]
- Cai, X.; Ni, W.; Wei, C.; Cui, J. Diagnostic value of the serum galactomannan and (1, 3)-β-D-glucan assays for invasive pulmonary aspergillosis in non-neutropenic patients. Intern. Med. Tokyo Jpn. 2014, 53, 2433–2437. [Google Scholar] [CrossRef]
- Marr, K.A.; Schlamm, H.T.; Herbrecht, R.; Rottinghaus, S.T.; Bow, E.J.; Cornely, O.A.; Heinz, W.J.; Jagannatha, S.; Koh, L.P.; Kontoyiannis, D.P.; et al. Combination Antifungal Therapy for Invasive Aspergillosis: A Randomized Trial. Ann. Intern. Med. 2015, 162, 81–89. [Google Scholar] [CrossRef]
- Wingard, J.R.; Ribaud, P.; Schlamm, H.T.; Herbrecht, R. Changes in causes of death over time after treatment for invasive aspergillosis. Cancer 2008, 112, 2309–2312. [Google Scholar] [CrossRef]
- Azur, M.J.; Stuart, E.A.; Frangakis, C.; Leaf, P.J. Multiple Imputation by Chained Equations: What is it and how does it work? Int. J. Methods Psychiatr. Res. 2011, 20, 40–49. [Google Scholar] [CrossRef]
- Youden, W.J. Index for rating diagnostic tests. Cancer 1950, 3, 32–35. [Google Scholar] [CrossRef]
- Herbrecht, R.; Denning, D.W.; Patterson, T.F.; Bennett, J.E.; Greene, R.E.; Oestmann, J.-W.; Kern, W.V.; Marr, K.A.; Ribaud, P.; Lortholary, O.; et al. Voriconazole versus Amphotericin B for Primary Therapy of Invasive Aspergillosis. N. Engl. J. Med. 2002, 347, 408–415. [Google Scholar] [CrossRef]
- Maertens, J.A.; Raad, I.I.; Marr, K.A.; Patterson, T.F.; Kontoyiannis, D.P.; Cornely, O.A.; Bow, E.J.; Rahav, G.; Neofytos, D.; Aoun, M.; et al. Isavuconazole versus voriconazole for primary treatment of invasive mould disease caused by Aspergillus and other filamentous fungi (SECURE): A phase 3, randomised-controlled, non-inferiority trial. Lancet Lond. Engl. 2016, 387, 760–769. [Google Scholar] [CrossRef]
- Donnelly, J.P.; Chen, S.C.; Kauffman, C.A.; Steinbach, W.J.; Baddley, J.W.; Verweij, P.E.; Clancy, C.J.; Wingard, J.R.; Lockhart, S.R.; Groll, A.H.; et al. Revision and Update of the Consensus Definitions of Invasive Fungal Disease From the European Organization for Research and Treatment of Cancer and the Mycoses Study Group Education and Research Consortium. Clin. Infect. Dis. 2019. (epub ahead of print). [Google Scholar] [CrossRef] [PubMed]
- Mercier, T.; Guldentops, E.; Lagrou, K.; Maertens, J. Galactomannan, a Surrogate Marker for Outcome in Invasive Aspergillosis: Finally Coming of Age. Front. Microbiol. 2018, 9, 661. [Google Scholar] [CrossRef] [PubMed]
- Chai, L.Y.A.; Kullberg, B.-J.; Johnson, E.M.; Teerenstra, S.; Khin, L.W.; Vonk, A.G.; Maertens, J.; Lortholary, O.; Donnelly, P.J.; Schlamm, H.T.; et al. Early Serum Galactomannan Trend as a Predictor of Outcome of Invasive Aspergillosis. J. Clin. Microbiol. 2012, 50, 2330–2336. [Google Scholar] [CrossRef]
- Kovanda, L.L.; Kolamunnage-Dona, R.; Neely, M.; Maertens, J.; Lee, M.; Hope, W.W. Pharmacodynamics of Isavuconazole for Invasive Mold Disease: Role of Galactomannan for Real-Time Monitoring of Therapeutic Response. Clin. Infect. Dis. 2017, 64, 1557–1563. [Google Scholar] [CrossRef]
- Kovanda, L.L.; Desai, A.V.; Hope, W.W. Prognostic value of galactomannan: Current evidence for monitoring response to antifungal therapy in patients with invasive aspergillosis. J. Pharmacokinet. Pharmacodyn. 2017, 44, 143–151. [Google Scholar] [CrossRef]
- Maertens, J.; Van Eldere, J.; Verhaegen, J.; Verbeken, E.; Verschakelen, J.; Boogaerts, M. Use of Circulating Galactomannan Screening for Early Diagnosis of Invasive Aspergillosis in Allogeneic Stem Cell Transplant Recipients. J. Infect. Dis. 2002, 186, 1297–1306. [Google Scholar] [CrossRef]
- Fisher, C.E.; Stevens, A.M.; Leisenring, W.; Pergam, S.A.; Boeckh, M.; Hohl, T.M. The Serum Galactomannan Index Predicts Mortality in Hematopoietic Stem Cell Transplant Recipients With Invasive Aspergillosis. Clin. Infect. Dis. 2013, 57, 1001–1004. [Google Scholar] [CrossRef]
- Jung, J.; Kim, M.Y.; Chong, Y.P.; Lee, S.-O.; Choi, S.-H.; Kim, Y.S.; Woo, J.H.; Kim, S.-H. Clinical characteristics, radiologic findings, risk factors and outcomes of serum galactomannan-negative invasive pulmonary aspergillosis. J. Microbiol. Immunol. Infect. Wei Mian Yu Gan Ran Za Zhi 2017, 51, 802–809. [Google Scholar] [CrossRef]
- Neofytos, D.; Railkar, R.; Mullane, K.M.; Fredricks, D.N.; Granwehr, B.; Marr, K.A.; Almyroudis, N.G.; Kontoyiannis, D.P.; Maertens, J.; Fox, R.; et al. Correlation between Circulating Fungal Biomarkers and Clinical Outcome in Invasive Aspergillosis. PLoS ONE 2015, 10, e0129022. [Google Scholar] [CrossRef]
- Kim, S.-J.; Cheong, J.-W.; Min, Y.H.; Choi, Y.J.; Lee, D.-G.; Lee, J.-H.; Yang, D.-H.; Lee, S.M.; Kim, S.-H.; Kim, Y.S.; et al. Success rate and risk factors for failure of empirical antifungal therapy with itraconazole in patients with hematological malignancies: A multicenter, prospective, open-label, observational study in Korea. J. Korean Med. Sci. 2014, 29, 61–68. [Google Scholar] [CrossRef] [PubMed]
| Training Cohort | Validation Cohort | p-Value | |
|---|---|---|---|
| n | 206 | 285 | |
| Age, years (median [IQR]) | 58.66 [49.30, 67.22] | 55.00 [42.00, 63.00] | <0.001 |
| BMI, kg/m2 (median [IQR]) | 23.61 [20.80, 27.12] | 23.30 [20.60, 26.35] | 0.699 |
| Male gender (%) | 129 (62.6) | 161 (56.5) | 0.204 |
| Proven IA (%) | 32 (15.5) | 13 (4.6) | <0.001 |
| Bilirubin, mg/dL (median [IQR]) | 0.66 [0.47, 0.97] | ||
| Creatinine, mg/dL (median [IQR]) | 0.70 [0.54, 1.03] | ||
| Neutrophils/µL (median [IQR]) | 0 [0, 845] | ||
| Monocytes/µL (median [IQR]) | 0 [0, 100] | ||
| Platelets/µL (median [IQR]) | 39,000 [20,250, 71,500] | ||
| Active GvHD (%) | 28 (13.7) | ||
| High dose corticosteroids * (%) | 46 (22.4) | ||
| Initial antifungal therapy (%) | |||
| Azole | 113 (54.9) | ||
| Echinocandin | 48 (23.3) | ||
| Polyene | 32 (15.5) | ||
| Combination | 11 (5.3) | ||
| Other | 2 (1.0) |
| Test | Baseline Cut-off | Week 1 Cut-off | Sensitivity (95% CI) | Specificity (95% CI) | Youden Index |
|---|---|---|---|---|---|
| Baseline sGM | 1.0 | - | 0.64 (0.51–0.76) | 0.65 (0.56–0.72) | 0.29 |
| Relative sGM decline by W1 | - | −1.5 | 0.07 (0.02–0.16) | 0.88 (0.81–0.93) | −0.05 |
| Absolute sGM decline by W1 | - | 0.8 | 0.78 (0.71–0.85) | 0.32 (0.21–0.46) | 0.10 |
| Baseline + relative sGM decline by W1 | 1.0 | −1.5 | 0.00 (0.00–0.2.) | 0.94 (0.87–0.98) | −0.06 |
| Baseline + absolut sGM decline by W1 | 1.0 | 0.8 | 0.00 (0.00–0.18) | 1.00 (0.83–1.00) | 0.00 |
| Baseline and absolute sGM value at baseline and W1 | 1.4 | 0.4 | 0.78 (0.60–0.91) | 0.70 (0.59–0.79) | 0.48 |
| High Risk | Intermediate Risk | Low Risk | p | |
|---|---|---|---|---|
| n | 54 | 84 | 68 | |
| Proven IA (%) | 17 (31.5) | 12 (14.3) | 3 (4.4) | <0.001 |
| Initial Therapy (%) | 0.209 | |||
| Azole | 29 (53.7) | 39 (46.4) | 45 (66.2) | |
| Echinocandin | 14 (25.9) | 20 (23.8) | 14 (20.6) | |
| Combination | 4 (7.4) | 6 (7.1) | 1 (1.5) | |
| Polyene | 7 (13.0) | 17 (20.2) | 8 (11.8) | |
| Other | 0 (0.0) | 2 (2.4) | 0 (0.0) | |
| Prophylaxis (%) | 3 (5.6) | 3 (3.6) | 3 (4.4) | 0.856 |
| Male Gender (%) | 32 (59.3) | 56 (66.7) | 41 (60.3) | 0.605 |
| Bilirubin, mg/dL (median [IQR]) | 0.78 [0.52, 1.19] | 0.75 [0.50, 1.11] | 0.64 [0.45, 0.90] | 0.077 |
| Creatinine, mg/dL (median [IQR]) | 0.74 [0.52, 1.11] | 0.66 [0.52, 1.01] | 0.68 [0.56, 0.91] | 0.845 |
| New mycological criteria * (%) | 54 (100.0) | 76 (96.2) | 40 (74.1) | <0.001 |
| High dose corticosteroids (%) | 15 (27.8) | 18 (21.4) | 13 (19.1) | 0.505 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mercier, T.; Wera, J.; Chai, L.Y.A.; Lagrou, K.; Maertens, J. A Mortality Prediction Rule for Hematology Patients with Invasive Aspergillosis Based on Serum Galactomannan Kinetics. J. Clin. Med. 2020, 9, 610. https://doi.org/10.3390/jcm9020610
Mercier T, Wera J, Chai LYA, Lagrou K, Maertens J. A Mortality Prediction Rule for Hematology Patients with Invasive Aspergillosis Based on Serum Galactomannan Kinetics. Journal of Clinical Medicine. 2020; 9(2):610. https://doi.org/10.3390/jcm9020610
Chicago/Turabian StyleMercier, Toine, Joachim Wera, Louis Y. A. Chai, Katrien Lagrou, and Johan Maertens. 2020. "A Mortality Prediction Rule for Hematology Patients with Invasive Aspergillosis Based on Serum Galactomannan Kinetics" Journal of Clinical Medicine 9, no. 2: 610. https://doi.org/10.3390/jcm9020610
APA StyleMercier, T., Wera, J., Chai, L. Y. A., Lagrou, K., & Maertens, J. (2020). A Mortality Prediction Rule for Hematology Patients with Invasive Aspergillosis Based on Serum Galactomannan Kinetics. Journal of Clinical Medicine, 9(2), 610. https://doi.org/10.3390/jcm9020610






