Risk Assessment of Novel Coronavirus COVID-19 Outbreaks Outside China
Abstract
1. Introduction
2. Materials and Methods
2.1. Model Ingredients
- (i)
- We estimate the cumulative number of cases in China outside Hubei province after 23 January, using a time-dependent compartmental model of the transmission dynamics.
- (ii)
- We use that number as an input to the global transportation network to generate probability distributions of the number of infected travellers arriving at destinations outside China.
- (iii)
- In a destination country, we use a Galton–Watson branching process to model the initial spread of the virus. We calculate the extinction probability of each branch initiated by a single imported case, obtaining the probability of a major outbreak as the probability that at least one branch will not go extinct.
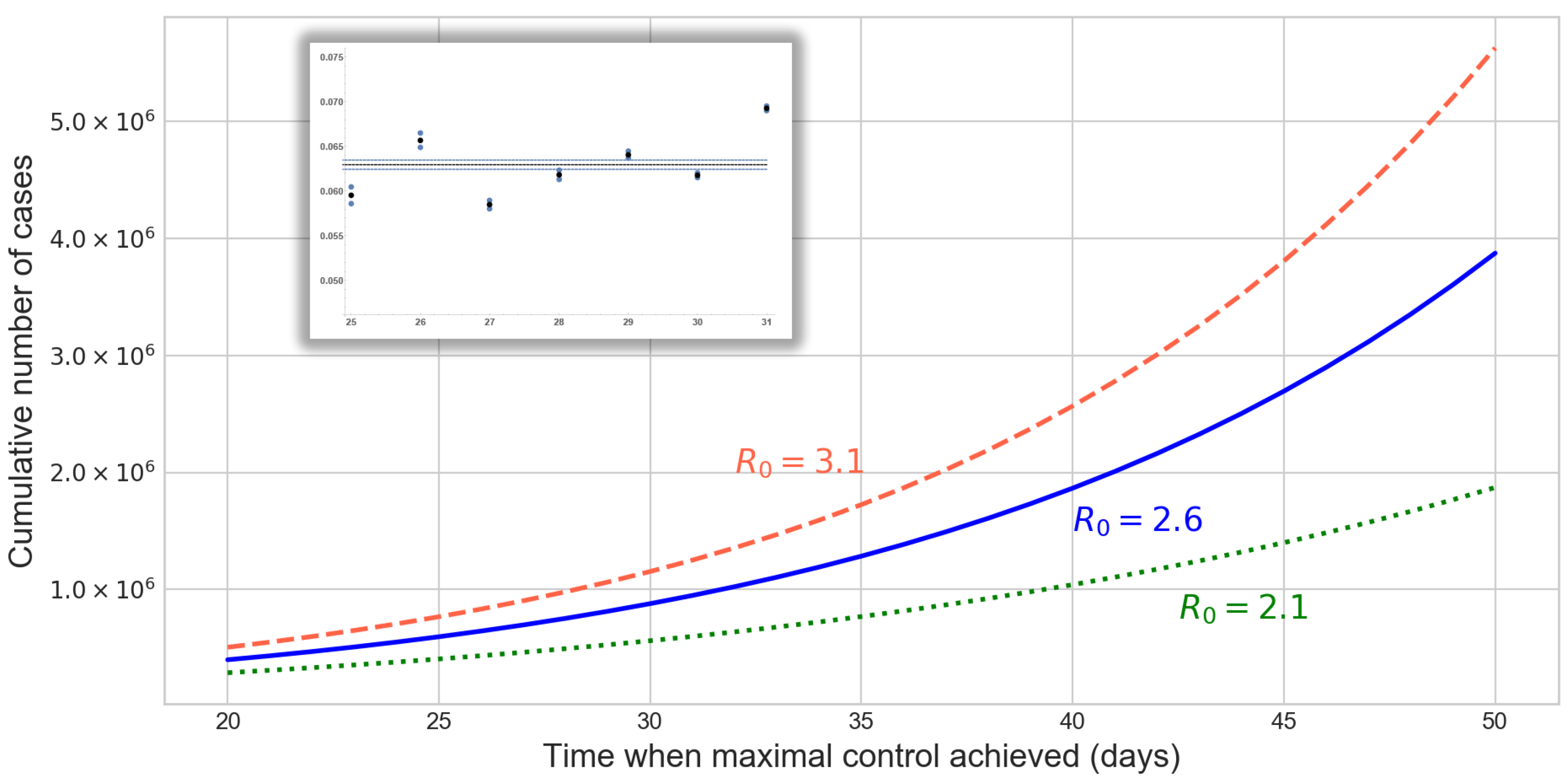
2.2. Epidemic Size in China Outside the Closed Areas of Hubei
2.3. Connectivity and Case Exportation
2.4. Probability of a Major Outbreak in a Country by Imported Cases
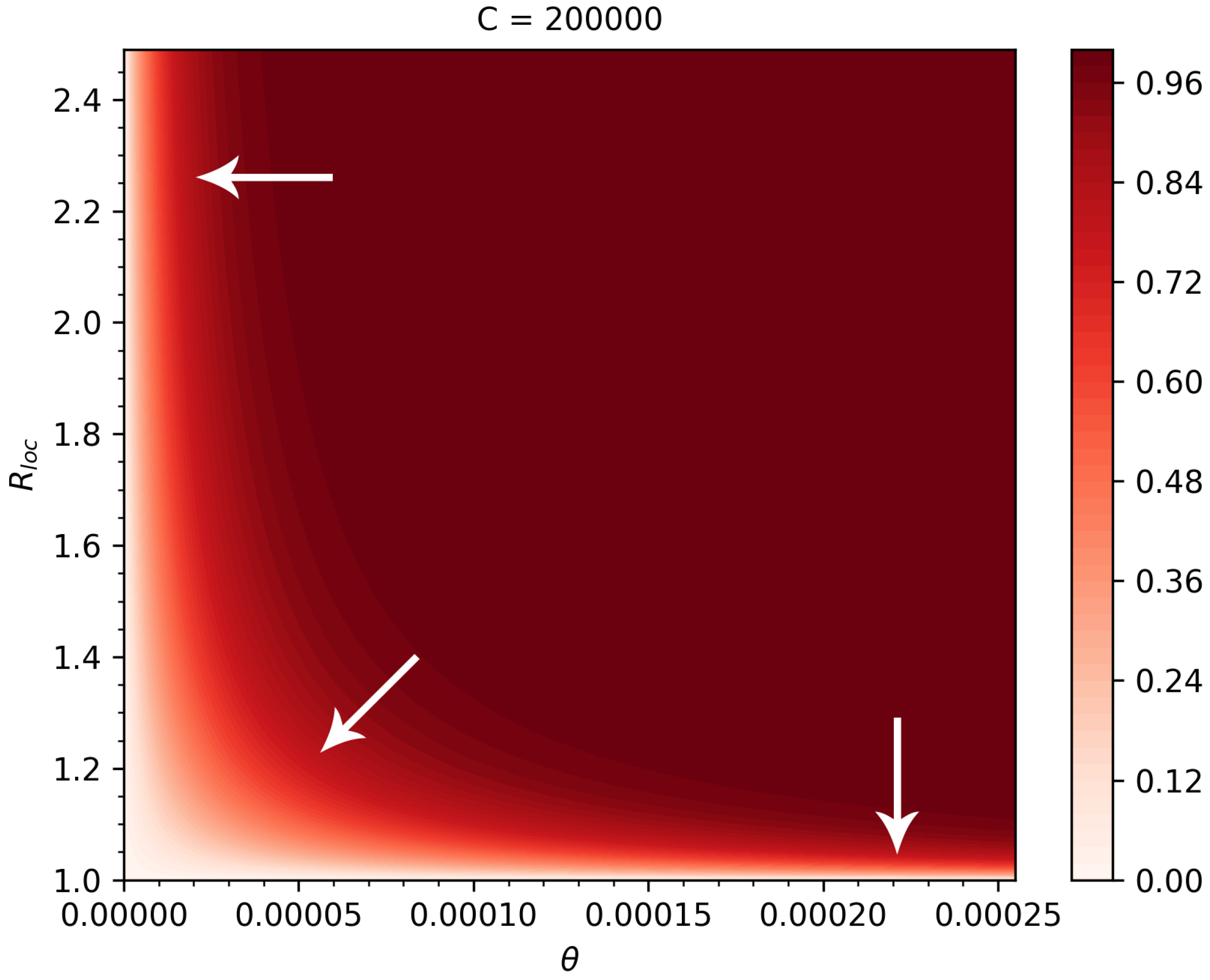
2.5. Dependence of the Risk of Major Outbreaks on Key Parameters
3. Results
3.1. Epidemic Size in China
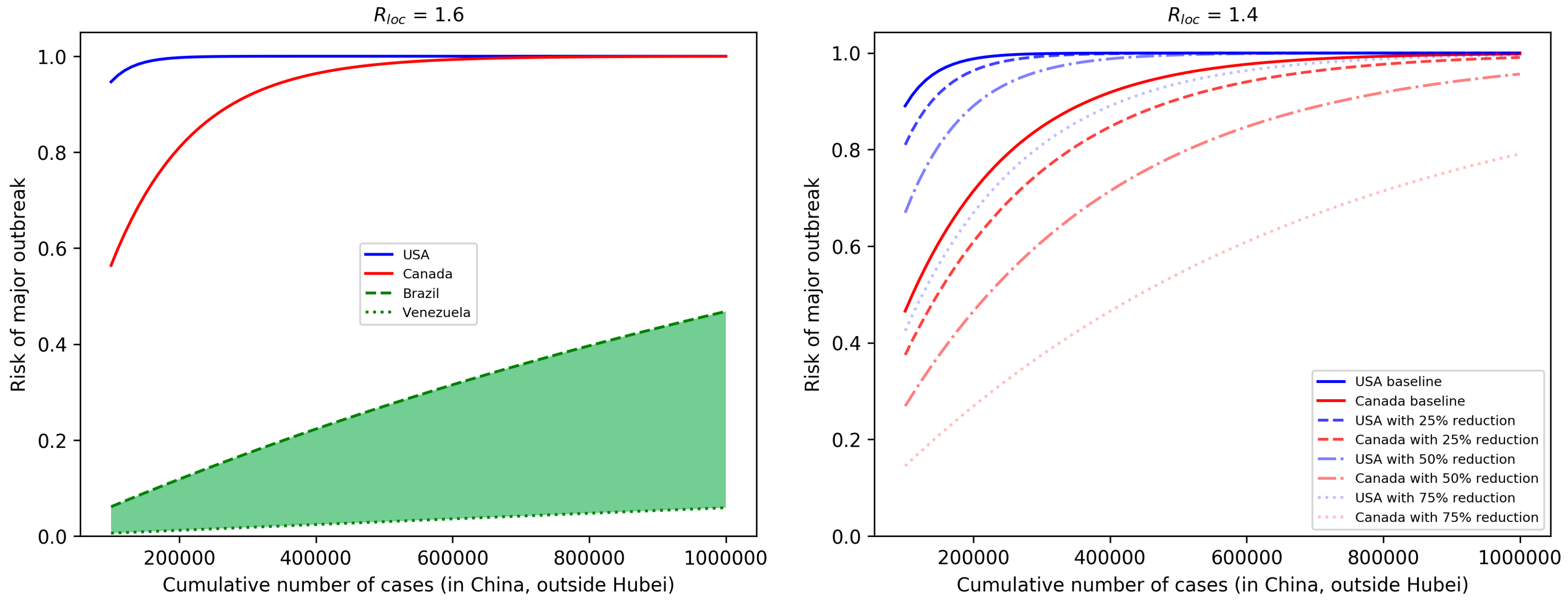
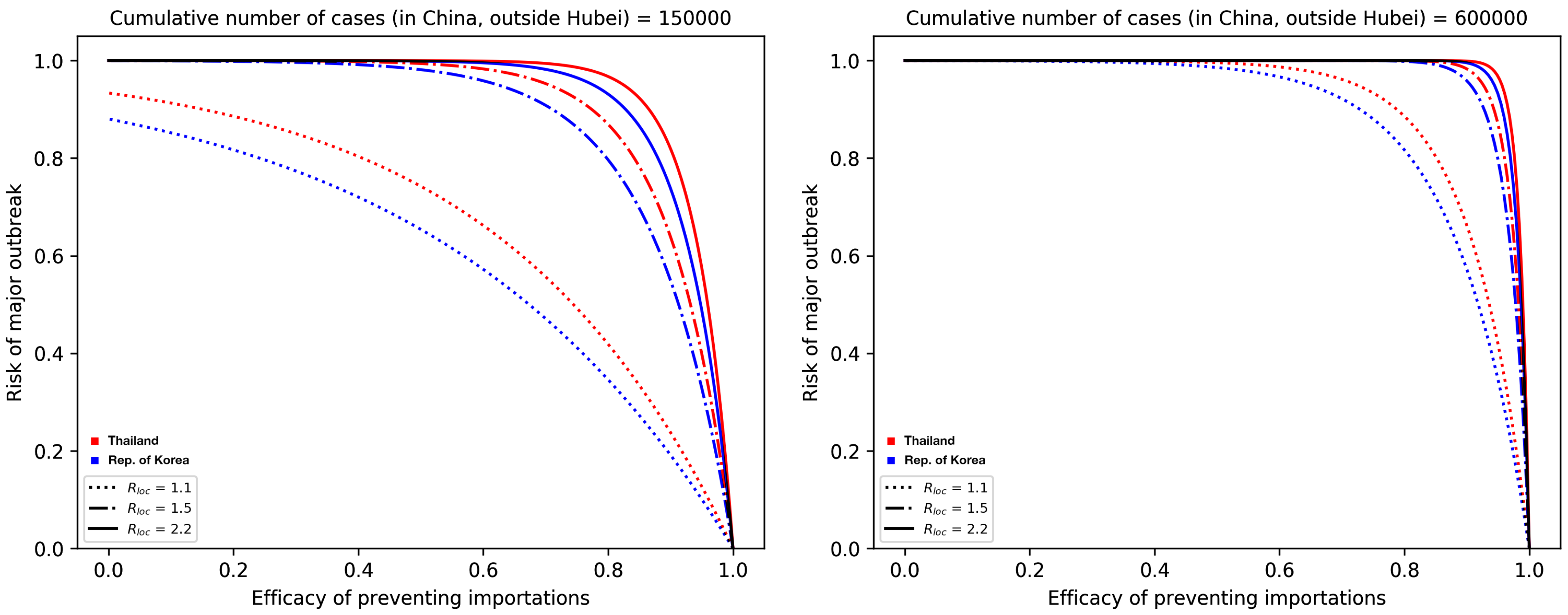
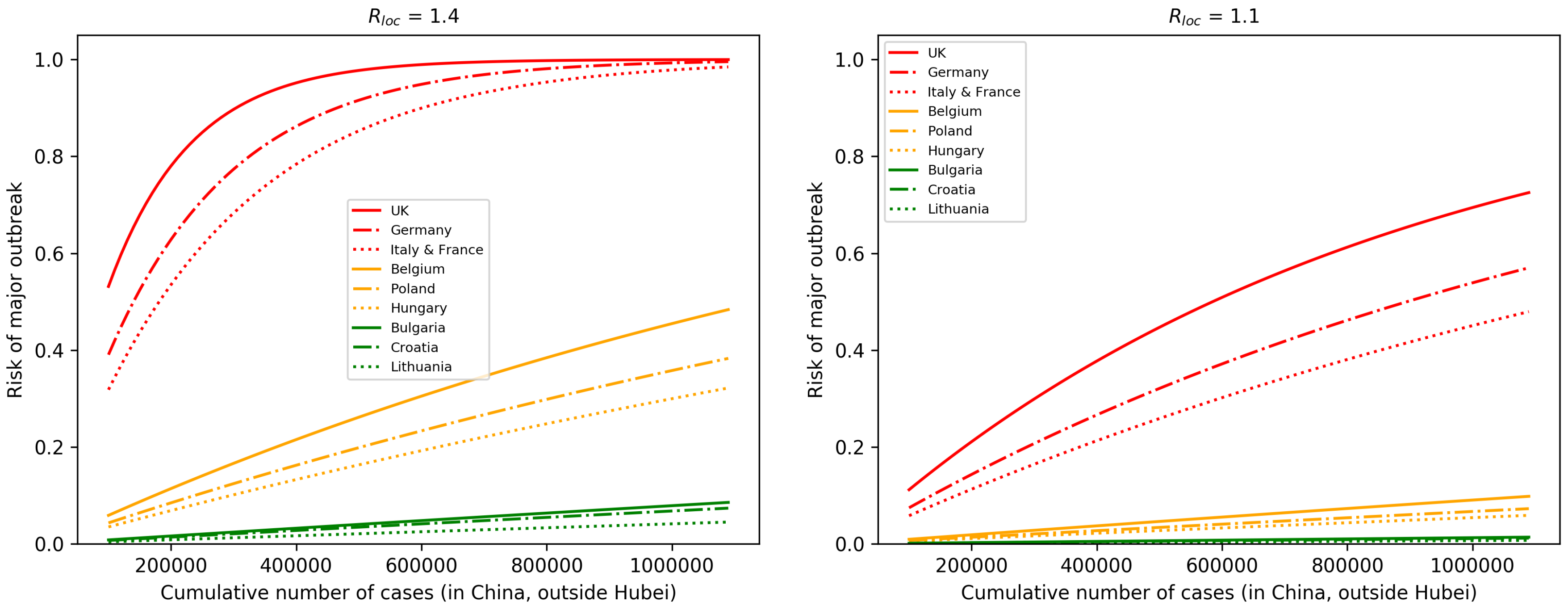
3.2. Risk of Major Outbreaks
3.3. Profile of Countries Benefiting the Most From Interventions
4. Discussion
Author Contributions
Funding
Conflicts of Interest
Appendix A. Transmission Dynamics
Appendix B. Calculating the Risk of Outbreaks by Importation
| Parameter | Interpretation | Depends on … | Typical Range |
|---|---|---|---|
| C | Cumulative case number in China, outside the closed areas | properties of nCoV-2019, efficacy of Chinese control | |
| Local reproduction number in destination country | destination country | ||
| Probability of a importation chance that a case from the origin travelling to and mixing into the local population of the destination country | China and destination country |
Appendix C.
References
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020, 395, 497–506. [Google Scholar] [CrossRef]
- WHO. Statement Regarding Cluster of Pneumonia Cases in Wuhan, China; World Health Organization: Geneva, Switzerland, 2020. Available online: https://www.who.int/china/news/detail/09-01-2020-who-statementregarding-cluster-of-pneumonia-cases-in-wuhan-china (accessed on 17 February 2020).
- WHO. Novel Coronavirus—Thailand (ex-China); World Health Organization: Geneva, Switzerland, 2020. Available online: https://www.who.int/csr/don/14-january-2020-novel-coronavirus-thailand-ex-china/en (accessed on 17 February 2020).
- WHO. Novel Coronavirus (2019-nCoV) Situation Report—1; World Health Organization: Geneva, Switzerland, 2020. Available online: https://www.who.int/docs/default-source/coronaviruse/situation-reports/20200121-sitrep-1-2019-ncov.pdf (accessed on 17 February 2020).
- JHU IDD Team. 2019-nCoV Global Cases by Center for Systems Science and Engineering. 2020. Available online: https://docs.google.com/spreadsheets/d/1wQVypefm946ch4XDp37uZ-wartW4V7ILdg-qYiDXUHM/edit?usp=sharing (accessed on 17 February 2020).
- CDC. 2019 Novel Coronavirus. Prevention & Treatment. Cent. Disease Control Prev. 2020. Available online: https://www.cdc.gov/coronavirus/2019-ncov/about/prevention-treatment.html (accessed on 17 February 2020).
- NPR. Chinese Authorities Begin Quarantine Of Wuhan City As Coronavirus Cases Multiply. 2020. Available online: https://www.npr.org/2020/01/23/798789671/chinese-authorities-begin-quarantine-of-wuhan-city-as-coronavirus-cases-multiply (accessed on 17 February 2020).
- Cheng, W.C.C.; Wong, S.-C.; To, K.K.W.; Ho, P.L.; Yuen, K.-Y. Preparedness and proactive infection control measures against the emerging Wuhan coronavirus pneumonia in China. J. Hosp. Infect. 2020. [Google Scholar] [CrossRef]
- Arnot, M.; Mzezewa, T. The Coronavirus: What Travelers Need to Know; The New York Times: New York, NY, USA, 2020; Available online: https://www.nytimes.com/2020/01/26/travel/Coronavirus-travel.html (accessed on 17 February 2020).
- National Health Commission of the People’s Republic of China. Work begins on mobile hospital in Wuhan. 2020. Available online: http://en.nhc.gov.cn/2020-01/29/c_76034.htm (accessed on 17 February 2020).
- National Health Commission of the People’s Republic of China. Medics flood to Hubei to fight disease. 2020. Available online: http://en.nhc.gov.cn/2020-01/29/c_76031.htm (accessed on 17 February 2020).
- Parry, J. Pneumonia in China: Lack of information raises concerns among Hong Kong health workers. BMJ 2020. [Google Scholar] [CrossRef]
- Moore, M.; Gelfeld, B.; Okunogbe, A.T.; Christopher, P. Identifying Future Disease Hot Spots: Infectious Disease Vulnerability Index; RAND Corporation: Santa Monica, CA, USA, 2016; Available online: https://www.rand.org/pubs/research_reports/RR1605.html (accessed on 17 February 2020).
- Bogoch, I.I.; Watts, A.; Thomas-Bachli, A.; Huber, C.; Kraemer, M.U.G.; Khan, K. Pneumonia of Unknown Etiology in Wuhan, China: Potential for International Spread Via Commercial Air Travel. J. Travel Med. 2020. [Google Scholar] [CrossRef]
- Chinazzi, M.; Davis, J.T.; Gioannini, C.; Litvinova, M.; Pastore y Piontti, A.; Rossi, L.; Xiong, X.; Halloran, M.E.; Longini, I.M.; Vespignani, A. Preliminary assessment of the International Spreading Risk Associated with the 2019 novel Coronavirus (2019-nCoV) outbreak in Wuhan City. Lab. Model. Biol. Soc.–Techn. Syst. 2020. Available online: https://www.mobs-lab.org/uploads/6/7/8/7/6787877/wuhan_novel_coronavirus__6_.pdf (accessed on 17 February 2020).
- Imai, N.; Dorigatti, I.; Cori, A.; Donnelly, C.; Riley, S.; Ferguson, N.M. Report 2: Estimating the potential total number of novel Coronavirus cases in Wuhan City, China. Imper. Coll. London. 2020. Available online: https://www.imperial.ac.uk/media/imperial-college/medicine/sph/ide/gida-fellowships/2019-nCoV-outbreak-report-22-01-2020.pdf (accessed on 17 February 2020).
- Zhao, S.; Musa, S.S.; Lin, Q.; Ran, J.; Yang, G.; Wang, W.; Lou, Y.; Yang, L.; Gao, D.; He, D.; et al. Estimating the Unreported Number of Novel Coronavirus (2019-nCoV) Cases in China in the First Half of January 2020: A Data-Driven Modelling Analysis of the Early Outbreak. J. Clin. Med. 2020, 9, 388. [Google Scholar] [CrossRef] [PubMed]
- Nishiura, H.; Jung, S.-M.; Linton, N.M.; Kinoshita, R.; Yang, Y.; Hayashi, K.; Kobayashi, T.; Yuan, B.; Akhmetzhanov, A.R. The Extent of Transmission of Novel Coronavirus in Wuhan, China, 2020. J. Clin. Med. 2020, 9, 330. [Google Scholar] [CrossRef] [PubMed]
- Imai, N.; Cori, A.; Dorigatti, I.; Baguelin, M.; Donnelly, C.A.; Riley, S.; Ferguson, N.M. Report 3: Transmissibility of 2019-nCoV. Imper. Coll. London 2020. Available online: https://www.imperial.ac.uk/media/imperial-college/medicine/sph/ide/gida-fellowships/Imperial-2019-nCoV-transmissibility.pdf (accessed on 17 February 2020).
- Riou, J.; Althaus, C.L. Pattern of early human-to-human transmission of Wuhan 2019-nCoV. bioRχiv 2020. [Google Scholar] [CrossRef]
- Liu, T.; Hu, J.; Kang, M.; Lin, L.; Zhong, H.; Xiao, J.; He, G.; Song, T.; Huang, Q.; Rong, Z.; et al. Transmission dynamics of 2019 novel coronavirus (2019-nCoV). bioRχiv 2020. [Google Scholar] [CrossRef]
- Read, J.M.; Bridgen, J.R.E.; Cummings, D.A.T.; Ho, A.; Jewell, C.P. Novel coronavirus 2019-nCoV: Early estimation of epidemiological parameters and epidemic predictions. medRχiv 2020. [Google Scholar] [CrossRef]
- Majumder, M.; Mandl, K.D. Early Transmissibility Assessment of a Novel Coronavirus in Wuhan, China. SSRN 2020. [Google Scholar] [CrossRef]
- Li, Q.; Guan, X.; Wu, P.; Wang, X.; Zhou, L.; Tong, Y.; Ren, R.; Leung, K.S.M.; Lau, E.H.Y.; Wong, J.Y.; et al. Early Transmission Dynamics in Wuhan, China, of Novel Coronavirus–Infected Pneumonia. N. Engl. J. Med. 2020. [Google Scholar] [CrossRef] [PubMed]
- Kucharski, A.; Russell, T.; Diamond, C.; CMMID nCoV Working Group; Funk, S.; Eggo, R.M. Analysis of early transmission dynamics of nCoV in Wuhan. 2020. Available online: https://cmmid.github.io/ncov/wuhan_early_dynamics (accessed on 17 February 2020).
- Shen, M.; Peng, Z.; Xiao, Y.; Zhang, L. Modelling the epidemic trend of the 2019 novel coronavirus outbreak in China. bioRχiv 2020. Available online: https://www.biorxiv.org/content/10.1101/2020.01.23.916726v1 (accessed on 17 February 2020).
- Leung, K.; Wu, J.T.; Leung, G.M. Nowcasting and forecasting the potential domestic and international spread of the 2019-nCoV outbreak originating in Wuhan, China: A modelling study. Lancet 2020. to appear. [Google Scholar] [CrossRef]
- Backer, J.A.; Klinkenberg, D.; Wallinga, J. The incubation period of 2019-nCoV infections among travellers from Wuhan, China. medRχiv 2020. [Google Scholar] [CrossRef]
- Linton, N.M.; Kobayashi, T.; Yang, Y.; Hayashi, K.; Akhmetzhanov, A.R.; Jung, S.-M.; Yuan, B.; Kinoshita, R.; Nishiura, H. Epidemiological characteristics of novel coronavirus infection: A statistical analysis of publicly available case data. medRχiv 2020. [Google Scholar] [CrossRef]
- Zheng, Q.; Meredith, H.; Grantz, K.; Bi, Q.; Jones, F.; Lauer, S.; JHU IDD Team. Real-time estimation of the novel coronavirus incubation time. 2020. Available online: https://github.com/HopkinsIDD/ncov_incubation (accessed on 17 February 2020).
- Fisman, D.N.; Hauck, T.S.; Tuite, A.R.; Greer, A.L. An IDEA for Short Term Outbreak Projection: Nearcasting Using the Basic Reproduction Number. PLoS ONE 2013, 8, 12. [Google Scholar] [CrossRef] [PubMed]
- EpiRisk. Available online: http://epirisk.net (accessed on 17 February 2020).
- Quilty, B.; Clifford, S.; CMMID nCoV Working Group; Flasche, S.; Eggo, R.M. Effectiveness of airport screening at detecting travellers infected with 2019-nCoV. 2020. Available online: https://cmmid.github.io/ncov/airport_screening_report/airport_screening_preprint_2020_01_28.pdf (accessed on 17 February 2020).
- Britton, T. Stochastic epidemic models: A survey. Math. Biosci. 2020, 225, 24–35. [Google Scholar] [CrossRef] [PubMed]
- Riley, S.; Fraser, C.; Donnelly, C.A.; Ghani, A.C.; Abu-Raddad, L.J.; Hedley, A.J.; Leung, G.M.; Ho, L.-M.; Lam, T.-H.; Thach, T.Q.; et al. Transmission Dynamics of the Etiological Agent of SARS in Hong Kong: Impact of Public Health Interventions. Science 2003, 300, 1961–1966. [Google Scholar] [CrossRef] [PubMed]
- Park, S.W.; Champredon, D.; Earn, D.J.; Li, M.; Weitz, J.S.; Grenfell, B.T.; Dushoff, J. Reconciling early-outbreak estimates of the basic reproductive number and its uncertainty: A new framework and applications to the novel coronavirus (2019-nCoV) outbreak. medRχiv 2020. [Google Scholar] [CrossRef]
- Wearing, H.J.; Rohani, P.; Keeling, M.J. Appropriate Models for the Management of Infectious Diseases. PLoS Med. 2005, 2. [Google Scholar] [CrossRef] [PubMed]
- Lauer, S.; Zlojutro, A.; Rey, D.; Dong, E.; JHU IDD Team; UNSW Sydney rCITI Team. Update January 31: Modeling the Spreading Risk of 2019-nCoV. 2020. Available online: https://systems.jhu.edu/research/public-health/ncov-model-2 (accessed on 17 February 2020).
- Nishiura, H.; Kobayashi, T.; Yang, Y.; Hayashi, K.; Miyama, T.; Kinoshita, R.; Linton, N.M.; Jung, S.-M.; Yuan, B.; Suzuki, A.; et al. The Rate of Underascertainment of Novel Coronavirus (2019-nCoV) Infection: Estimation Using Japanese Passengers Data on Evacuation Flights. J. Clin. Med. 2020, 9, 419. [Google Scholar] [CrossRef] [PubMed]
- Bolyai Institute, University of Szeged. Risk assessment of novel coronavirus 2019-nCoV outbreaks outside China. Github 2020. Available online: https://github.com/zsvizi/corona-virus-2020 (accessed on 17 February 2020).
| Incubation Period | Method of Estimation | Reference | |
|---|---|---|---|
| 2.6 (1.5–3.5) | - | Epidemic Simulations | [19] |
| 2.2 (1.4–3.8) | - | Stochastic Simulations | [20] |
| 2.9 (2.3–3.6) | 4.8 days | Exp. Growth, Max. Likelihood Est. | [21] |
| 2.56 (2.49–2.63) | - | Exp. Growth, Max. Likelihood Est. | [17] |
| 3.11 (2.3–4.1) | - | SEIR | [22] |
| 2.5 (2.0–3.1) | - | Incidence Decay and Exponential Adjustment model | [23] |
| 2.2 (1.4–3.9) | 5.2 days (4.1–7.0) | Renewal Equations | [24] |
| −(1.4–4.0) | - | SEIR | [25] |
| 4.71 (4.5–4.9) | days (–) | Dec. 2019, SEIJR, MCMC | [26] |
| 2.08 (1.9–2.2) | - | Jan. 2020, SEIJR, MCMC | [26] |
| 2.68 (2.4–2.9) | - | SEIR, MCMC | [27] |
| - | 5.8 days 4.6–7.9) | Weibull | [28] |
| - | 4.6 days (3.3–5.8) | Weibull incl. Wuhan | [29] |
| - | 5.0 days (4.0–5.8) | Weibull excl. Wuhan | [29] |
| - | 5.1 days (4.4–6.1) | LogNormal | [30] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Boldog, P.; Tekeli, T.; Vizi, Z.; Dénes, A.; Bartha, F.A.; Röst, G. Risk Assessment of Novel Coronavirus COVID-19 Outbreaks Outside China. J. Clin. Med. 2020, 9, 571. https://doi.org/10.3390/jcm9020571
Boldog P, Tekeli T, Vizi Z, Dénes A, Bartha FA, Röst G. Risk Assessment of Novel Coronavirus COVID-19 Outbreaks Outside China. Journal of Clinical Medicine. 2020; 9(2):571. https://doi.org/10.3390/jcm9020571
Chicago/Turabian StyleBoldog, Péter, Tamás Tekeli, Zsolt Vizi, Attila Dénes, Ferenc A. Bartha, and Gergely Röst. 2020. "Risk Assessment of Novel Coronavirus COVID-19 Outbreaks Outside China" Journal of Clinical Medicine 9, no. 2: 571. https://doi.org/10.3390/jcm9020571
APA StyleBoldog, P., Tekeli, T., Vizi, Z., Dénes, A., Bartha, F. A., & Röst, G. (2020). Risk Assessment of Novel Coronavirus COVID-19 Outbreaks Outside China. Journal of Clinical Medicine, 9(2), 571. https://doi.org/10.3390/jcm9020571






