Application of Artificial Intelligence Techniques to Predict Survival in Kidney Transplantation: A Review
Abstract
1. Introduction
- They help to extract the factors considered by experts in their field of study when evaluating a situation or making decisions.
- They make it possible to find unknown functional relationships or properties among the entry data.
- They quickly adapt to changing environments with no need to redesign the system if the data are updated or replaced by other data.
- They can handle missing and noisy data.
- They make it possible to find relations and correlations among large amounts of data, and to generate solutions with a high degree of accuracy.
2. Results
2.1. Machine Learning Techniques
2.1.1. Decision Trees
2.1.2. Ensemble Methods
2.1.3. Artificial Neural Networks
2.1.4. Support Vector Machines
2.2. Application of Machine Learning Algorithms
2.2.1. Classification of Patient Survival
2.2.2. Modelling the Patient Survival Function
3. Discussion
4. Future Directions
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
Appendix A
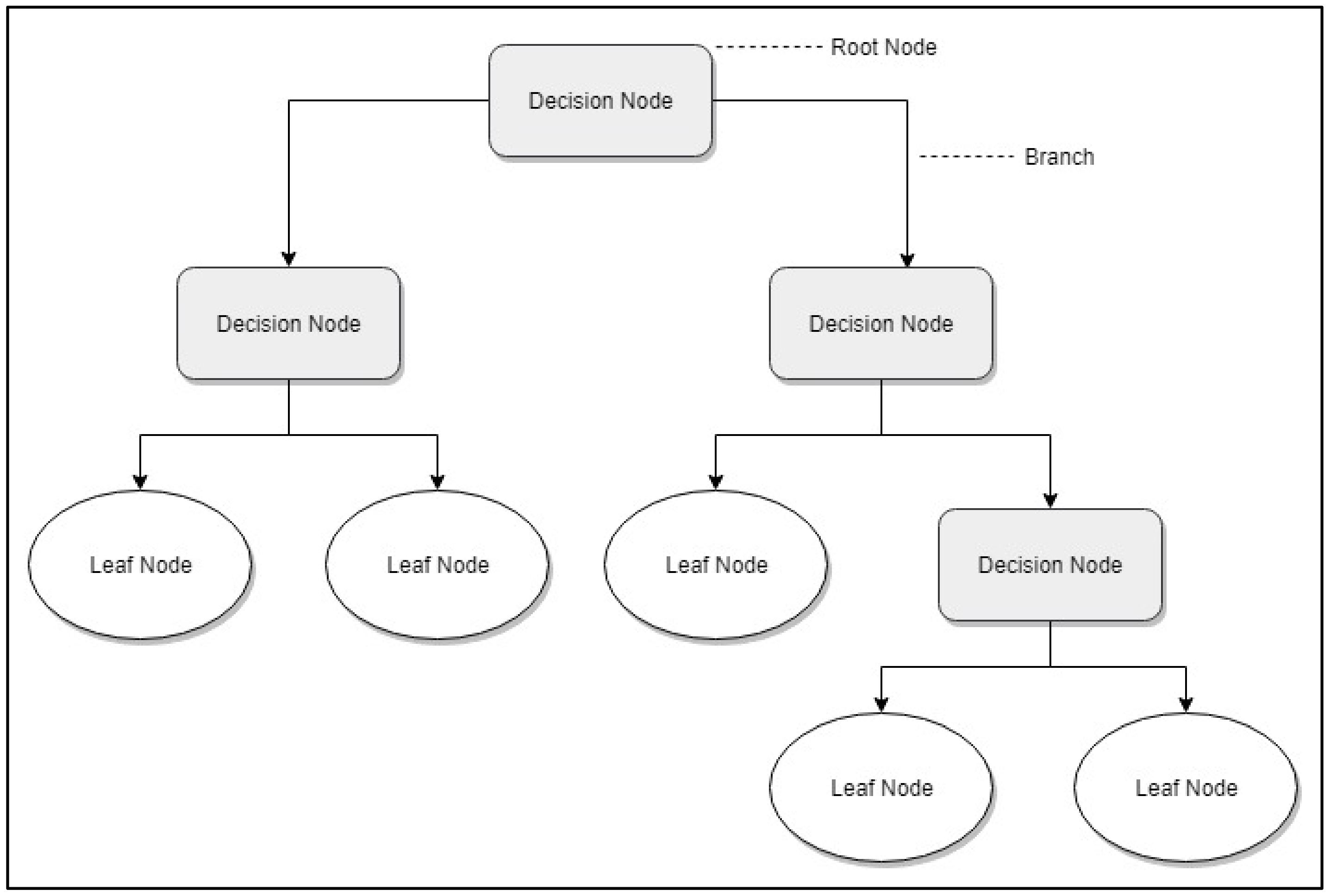
Decision Tree
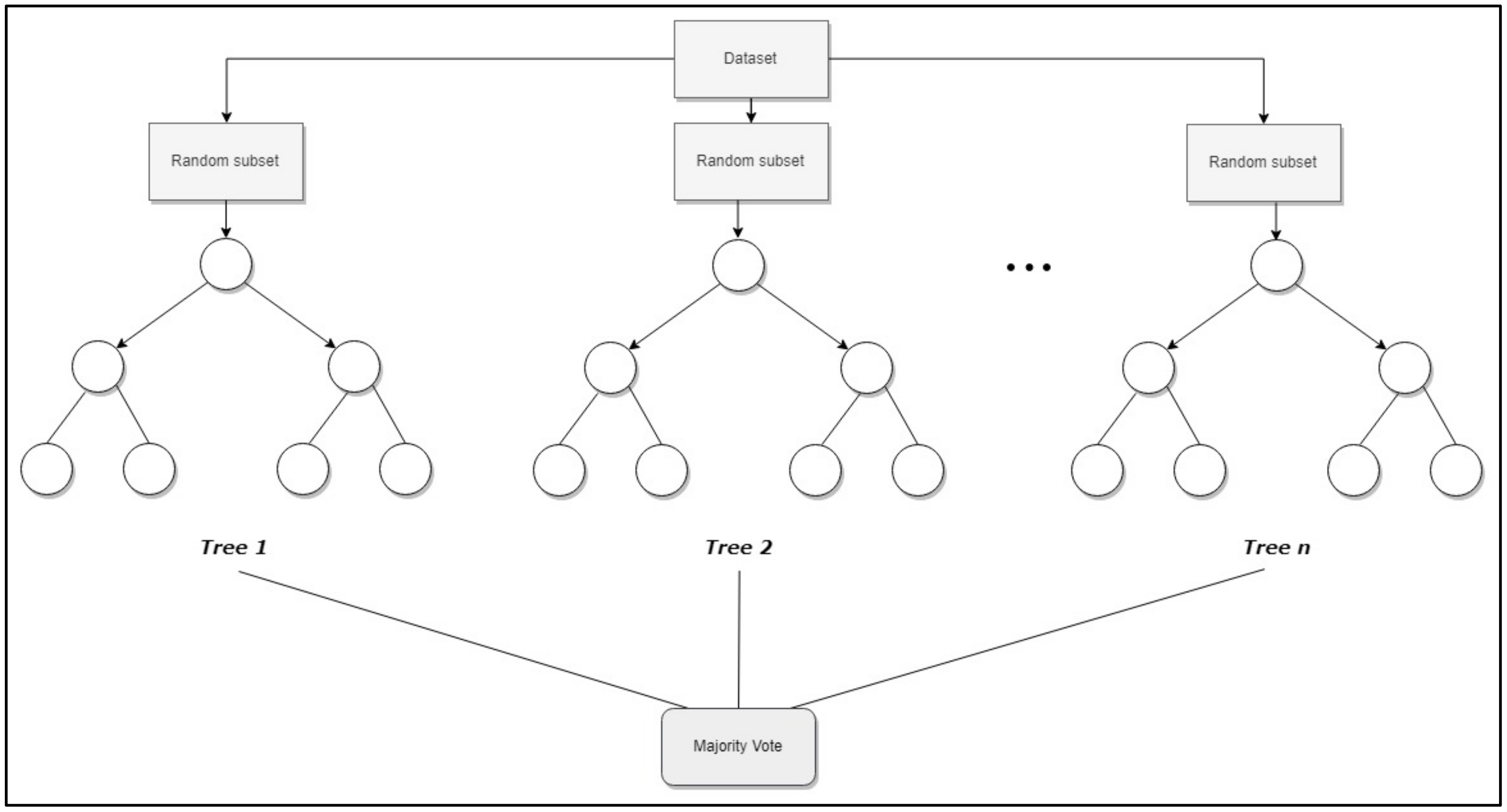
Random Forest
Artificial Neural Networks
- Input layer: receive data or signals from the environment.
- Output layer: provide the network response to input stimuli.
- Hidden layer: they do not receive or provide information to the environment (internal processing of the net).
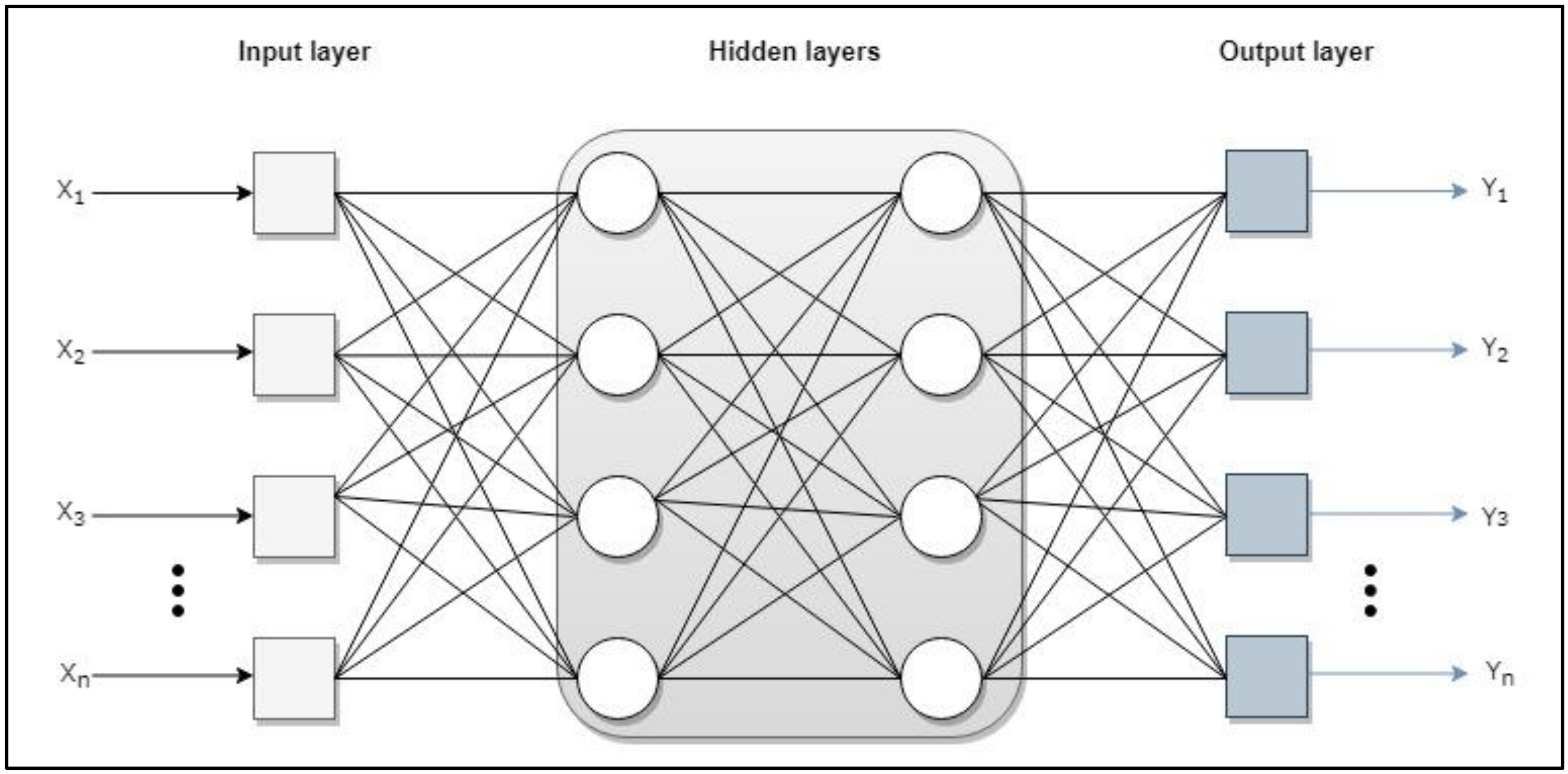
- Single-layer network. Only one input layer related to the output layer.
- Multi-layer network (MLP). One or more layers are added between the inputs and outputs, as can be seen in Figure A1. The MLP is a neural network that contains one or more layers of hidden neurons and uses the backpropagation (BP) algorithm [57] to train. This algorithm consists of two phases: feed-forward and feed-back propagation. In the feed-forward phase, the network output is calculated with all fixed weighting values while the input vector is entered from the input layer to the output one through the hidden layers. In the feed-back propagation phase, the error signal is calculated by subtracting the network output value from the expected output value. An error signal originates from an output neuron and spreads backwards layer by layer through the network [76]. It then propagates through the hidden layers to the input layer so that the weighting values are corrected. These two phases are repeated and the learning process is recycled to generate a better approach to the output. The learning process ends when the stop condition is met.
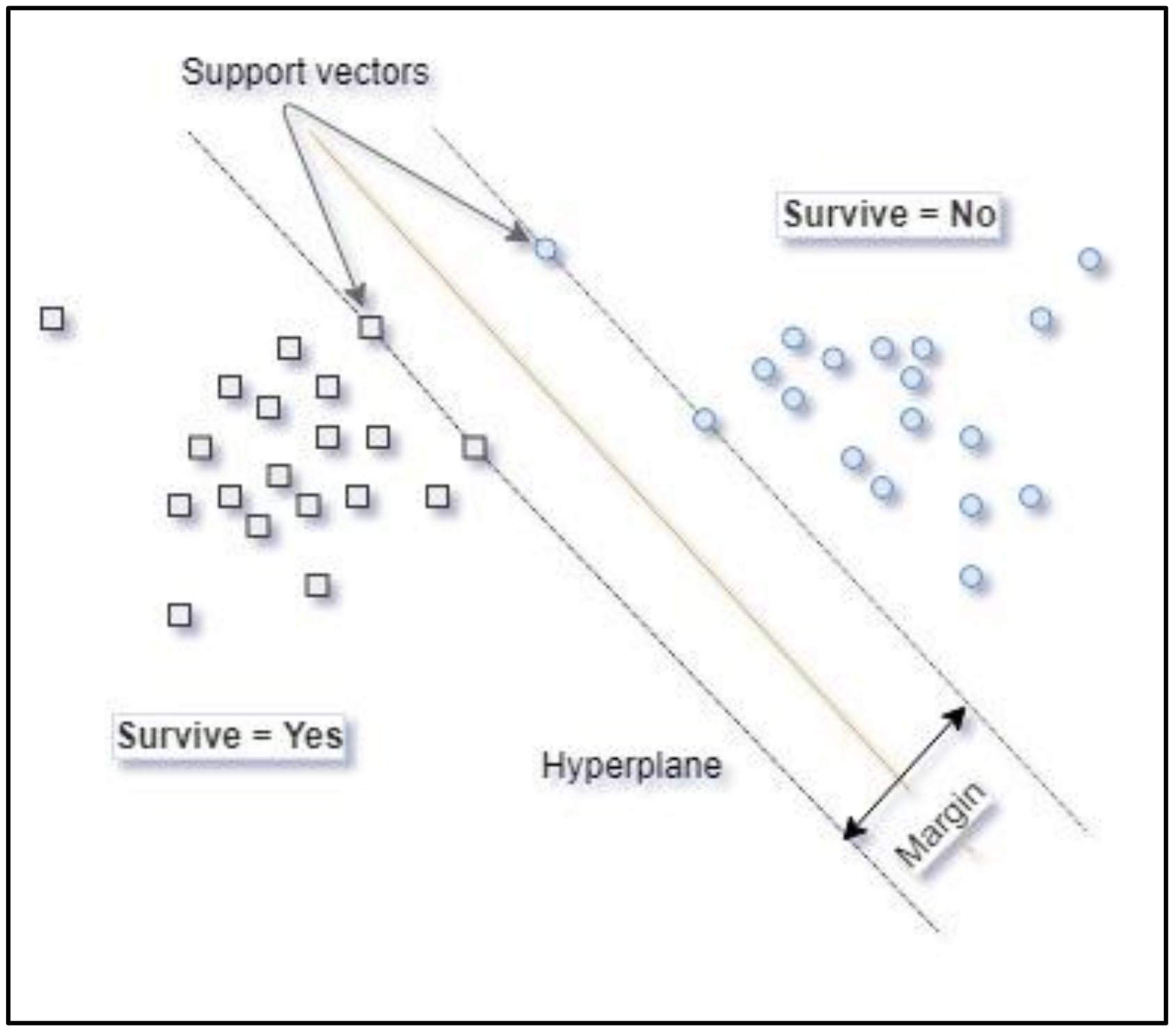
Support Vector Machine
Appendix B
Bayesian Belief Networks
Conditional Inference Trees
Estimated Post-Transplant Survival
Information Gain
K-Nearest Neighbours
Kidney Donor Risk Index
Kidney Donor Profile Index
Weibull Model
References
- Levey, A.S.; Coresh, J.; Bolton, K.; Culleton, B.; Harvey, K.S.; Ikizler, T.A.; Johnson, C.A.; Kausz, A.; Kimmel, P.L.; Kusek, J.; et al. K/DOQI clinical practice guidelines for chronic kidney disease: Evaluation, classification, and stratification. Am. J. Kidney Dis. 2002, 39, S1–S266. [Google Scholar]
- Era-Edta—PRESS RELEASES. Available online: https://web.era-edta.org/press-releases (accessed on 3 June 2019).
- Informe de Diálisis y Trasplante 2017. Available online: http://www.registrorenal.es/download/documentacion/Informe_REER_2017.pdf (accessed on 17 May 2019).
- Ojo, A.O.; Hanson, J.A.; Wolfe, R.A.; Leichtman, A.B.; Agodoa, L.Y.; Port, F.K. Long-term survival in renal transplant recipients with graft function. Kidney Int. 2000, 57, 307–313. [Google Scholar] [CrossRef] [PubMed]
- Hariharan, S.; Mcbride, M.A.; Cherikh, W.S.; Tolleris, C.B.; Bresnahan, B.A.; Johnson, C.P. Post-transplant renal function in the first year predicts long-term kidney transplant survival. Kidney Int. 2002, 62, 311–318. [Google Scholar] [CrossRef] [PubMed]
- Lamb, K.E.; Lodhi, S.; Meier-Kriesche, H.-U. Long-Term Renal Allograft Survival in the United States: A Critical Reappraisal. Am. J. Transplant. 2011, 11, 450–462. [Google Scholar] [CrossRef] [PubMed]
- Yoo, K.D.; Noh, J.; Lee, H.; Kim, D.K.; Lim, C.S.; Kim, Y.H.; Lee, J.P.; Kim, G.; Kim, Y.S. A Machine Learning Approach Using Survival Statistics to Predict Graft Survival in Kidney Transplant Recipients: A Multicenter Cohort Study. Sci. Rep. 2017, 7, 8904. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.; Mukhopadhyay, K. Survival analysis in clinical trials: Basics and must know areas. Perspect. Clin. Res. 2011, 2, 145–148. [Google Scholar] [CrossRef] [PubMed]
- Schluchter, M.D. Methods for the analysis of informatively censored longitudinal data. Stat. Med. 1992, 11, 1861–1870. [Google Scholar] [CrossRef]
- Kleinbaum, D.G. (Ed.) Survival Analysis—A Self-Learning Text, 3rd ed.; Springer: New York, NY, USA, 2012; Available online: https://www.springer.com/gp/book/9781441966452 (accessed on 3 June 2019).
- Klein, J.P.; Moeschberger, M.L. Survival Analysis: Techniques for Censored and Truncated Data; Springer Science & Business Media: New York, NY, USA, 2006; ISBN 0-387-21645-6. [Google Scholar]
- Coemans, M.; Süsal, C.; Döhler, B.; Anglicheau, D.; Giral, M.; Bestard, O.; Legendre, C.; Emonds, M.-P.; Kuypers, D.; Molenberghs, G.; et al. Analyses of the short- and long-term graft survival after kidney transplantation in Europe between 1986 and 2015. Kidney Int. 2018, 94, 964–973. [Google Scholar] [CrossRef]
- Huang, S.-T.; Yu, T.-M.; Chuang, Y.-W.; Chung, M.-C.; Wang, C.-Y.; Fu, P.-K.; Ke, T.-Y.; Li, C.-Y.; Lin, C.-L.; Wu, M.-J.; et al. The Risk of Stroke in Kidney Transplant Recipients with End-Stage Kidney Disease. Int. J. Environ. Res. Public Health 2019, 16, 326. [Google Scholar] [CrossRef]
- Kaplan, E.L.; Meier, P. Nonparametric Estimation from Incomplete Observations. J. Am. Stat. Assoc. 1958, 53, 457–481. [Google Scholar] [CrossRef]
- Cox, D.R. Regression Models and Life-Tables. J. R. Stat. Soc. Ser. B (Methodol.) 1972, 34, 187–202. [Google Scholar] [CrossRef]
- Sagiroglu, S.; Sinanc, D. Big data: A review. In Proceedings of the 2013 International Conference on Collaboration Technologies and Systems (CTS), San Diego, CA, USA, 20–24 May 2013; pp. 42–47. [Google Scholar]
- Gandomi, A.; Haider, M. Beyond the hype: Big data concepts, methods, and analytics. Int. J. Inf. Manag. 2015, 35, 137–144. [Google Scholar] [CrossRef]
- Ferroni, P.; Zanzotto, F.M.; Riondino, S.; Scarpato, N.; Guadagni, F.; Roselli, M. Breast Cancer Prognosis Using a Machine Learning Approach. Cancers 2019, 11, 328. [Google Scholar] [CrossRef] [PubMed]
- Lim, E.-C.; Park, J.H.; Jeon, H.J.; Kim, H.-J.; Lee, H.-J.; Song, C.-G.; Hong, S.K. Developing a Diagnostic Decision Support System for Benign Paroxysmal Positional Vertigo Using a Deep-Learning Model. J. Clin. Med. 2019, 8, 633. [Google Scholar] [CrossRef] [PubMed]
- Calvert, J.; Saber, N.; Hoffman, J.; Das, R. Machine-Learning-Based Laboratory Developed Test for the Diagnosis of Sepsis in High-Risk Patients. Diagnostics 2019, 9, 20. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Shen, L. Skin Lesion Analysis towards Melanoma Detection Using Deep Learning Network. Sensors 2018, 18, 556. [Google Scholar] [CrossRef]
- Heo, S.-J.; Kim, Y.; Yun, S.; Lim, S.-S.; Kim, J.; Nam, C.-M.; Park, E.-C.; Jung, I.; Yoon, J.-H. Deep Learning Algorithms with Demographic Information Help to Detect Tuberculosis in Chest Radiographs in Annual Workers’ Health Examination Data. Int. J. Environ. Res. Public. Health 2019, 16, 250. [Google Scholar] [CrossRef]
- Khalifa, F.; Shehata, M.; Soliman, A.; El-Ghar, M.A.; El-Diasty, T.; Dwyer, A.C.; El-Melegy, M.; Gimel’farb, G.; Keynton, R.; El-Baz, A. A generalized MRI-based CAD system for functional assessment of renal transplant. In Proceedings of the 2017 IEEE 14th International Symposium on Biomedical Imaging (ISBI 2017), Melbourne, VIC, Australia, 18–21 April 2017; pp. 758–761. [Google Scholar]
- Bishop, C.M. Pattern Recognition and Machine Learning; Springer: New York, NY, USA, 2006; ISBN 0-387-31073-8. [Google Scholar]
- Barr, A.; Feigenbaum, E.A. The Handbook of Artificial Intelligence; Kaufmann, W., Ed.; HeurisTech Press: Stanford/Los Altos, CA, USA, 1981; Volume 1. [Google Scholar]
- Prompramote, S.; Chen, Y.; Chen, Y.-P.P. Machine Learning in Bioinformatics. In Bioinformatics Technologies; Chen, Y.-P.P., Ed.; Springer: Berlin/Heidelberg, Germany, 2005; pp. 117–153. ISBN 978-3-540-26888-8. [Google Scholar]
- Pubmeddev Home-PubMed-NCBI. Available online: https://www.ncbi.nlm.nih.gov/pubmed/ (accessed on 11 July 2019).
- ScienceDirect.com—Science, Health and Medical Journals, Full Text Articles and Books. Available online: https://www.sciencedirect.com/ (accessed on 11 July 2019).
- Dblp: Computer Science Bibliography. Available online: https://dblp.uni-trier.de/ (accessed on 10 October 2019).
- Luck, M.; Sylvain, T.; Cardinal, H.; Lodi, A.; Bengio, Y. Deep Learning for Patient-Specific Kidney Graft Survival Analysis. Available online: https://arxiv.org/abs/1705.10245 (accessed on 19 February 2020).
- Tapak, L.; Hamidi, O.; Amini, P.; Poorolajal, J. Prediction of Kidney Graft Rejection Using Artificial Neural Network. Healthc. Inform. Res. 2017, 23, 277–284. [Google Scholar] [CrossRef]
- Mark, E.; Goldsman, D.; Gurbaxani, B.; Keskinocak, P.; Sokol, J. Using machine learning and an ensemble of methods to predict kidney transplant survival. PLoS ONE 2019, 14, e0209068. [Google Scholar] [CrossRef]
- Bae, S.; Massie, A.B.; Thomas, A.G.; Bahn, G.; Luo, X.; Jackson, K.R.; Ottmann, S.E.; Brennan, D.C.; Desai, N.M.; Coresh, J.; et al. Who can tolerate a marginal kidney? Predicting survival after deceased donor kidney transplant by donor–recipient combination. Am. J. Transplant. 2019, 19, 425–433. [Google Scholar] [CrossRef]
- Atallah, D.M.; Badawy, M.; El-Sayed, A.; Ghoneim, M.A. Predicting kidney transplantation outcome based on hybrid feature selection and KNN classifier. Multimed. Tools Appl. 2019, 78, 20383–20407. [Google Scholar] [CrossRef]
- Nematollahi, M.; Akbari, R.; Nikeghbalian, S.; Salehnasab, C. Classification Models to Predict Survival of Kidney Transplant Recipients Using Two Intelligent Techniques of Data Mining and Logistic Regression. Int. J. Organ Transplant. Med. 2017, 8, 119–122. [Google Scholar] [PubMed]
- Shahmoradi, L.; Langarizadeh, M.; Pourmand, G.; Fard, Z.A.; Borhani, A. Comparing Three Data Mining Methods to Predict Kidney Transplant Survival. Acta Inform. Med. 2016, 24, 322–327. [Google Scholar] [CrossRef] [PubMed]
- Topuz, K.; Zengul, F.D.; Dag, A.; Almehmi, A.; Yildirim, M.B. Predicting graft survival among kidney transplant recipients: A Bayesian decision support model. Decis. Support Syst. 2018, 106, 97–109. [Google Scholar] [CrossRef]
- Kamiński, B.; Jakubczyk, M.; Szufel, P. A framework for sensitivity analysis of decision trees. Cent. Eur. J. Oper. Res. 2018, 26, 135–159. [Google Scholar] [CrossRef] [PubMed]
- Marqués, M.P. Minería de Datos a Través de Ejemplos; RC Libros: Madrid, Spain, 2014; ISBN 978-84-941801-4-9. [Google Scholar]
- Chen, X.; Wang, M.; Zhang, H. The use of classification trees for bioinformatics. Wiley Interdiscip. Rev. Data Min. Knowl. Discov. 2011, 1, 55–63. [Google Scholar] [CrossRef]
- Van Arendonk, K.J.; Chow, E.K.H.; James, N.T.; Orandi, B.J.; Ellison, T.A.; Smith, J.M.; Colombani, P.M.; Segev, D.L. Choosing the Order of Deceased Donor and Living Donor Kidney Transplantation in Pediatric Recipients: A Markov Decision Process Model. Transplantation 2015, 99, 360–366. [Google Scholar] [CrossRef]
- Song, Y.; Lu, Y. Decision tree methods: Applications for classification and prediction. Shanghai Arch. Psychiatry 2015, 27, 130–135. [Google Scholar]
- Valera, V.A.; Walter, B.A.; Yokoyama, N.; Koyama, Y.; Iiai, T.; Okamoto, H.; Hatakeyama, K. Prognostic Groups in Colorectal Carcinoma Patients Based on Tumor Cell Proliferation and Classification and Regression Tree (CART) Survival Analysis. Ann. Surg. Oncol. 2007, 14, 34–40. [Google Scholar] [CrossRef]
- Schmidt, G.W.; Broman, A.T.; Hindman, H.B.; Grant, M.P. Vision Survival after Open Globe Injury Predicted by Classification and Regression Tree Analysis. Ophthalmology 2008, 115, 202–209. [Google Scholar] [CrossRef]
- Lewis, R.J. An Introduction to Classification and Regression Tree (CART) Analysis. Available online: https://www.researchgate.net/profile/Roger_Lewis6/publication/240719582_An_Introduction_to_Classification_and_Regression_Tree_CART_Analysis/links/0046352d3fb18f1740000000/An-Introduction-to-Classification-and-Regression-Tree-CART-Analysis.pdf (accessed on 19 February 2020).
- Information on See5/C5.0. Available online: https://www.rulequest.com/see5-info.html (accessed on 11 October 2019).
- Bhargava, N.; Sharma, G.; Bhargava, R.; Mathuria, M. Decision tree analysis on j48 algorithm for data mining. Proc. Int. J. Adv. Res. Comput. Sci. Softw. Eng. 2013, 3, 1114–1119. [Google Scholar]
- Brijain, M.; Patel, R.; Kushik, M.; Rana, K. A Survey on Decision Tree Algorithm for Classification. Available online: http://citeseerx.ist.psu.edu/viewdoc/download;jsessionid=3BE1C9D0B289A615AC53E2737280C589?doi=10.1.1.673.2797&rep=rep1&type=pdf (accessed on 19 February 2020).
- Murthy, S.K. Automatic construction of decision trees from data: A multi-disciplinary survey. Data Min. Knowl. Discov. 1998, 2, 345–389. [Google Scholar] [CrossRef]
- LeBlanc, M.; Crowley, J. Relative risk trees for censored survival data. Biometrics 1992, 48, 411–425. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Li, Y.; Reddy, C.K. Machine learning for survival analysis: A survey. ACM Comput. Surv. (CSUR) 2019, 51, 110. [Google Scholar] [CrossRef]
- MacKay, D.J.; Mac Kay, D.J. Information Theory, Inference and Learning Algorithms; Cambridge University Press: Padstow, UK, 2003; ISBN 0-521-64298-1. [Google Scholar]
- Hothorn, T.; Lausen, B.; Benner, A.; Radespiel-Tröger, M. Bagging survival trees. Stat. Med. 2004, 23, 77–91. [Google Scholar] [CrossRef]
- Hothorn, T.; Bühlmann, P.; Dudoit, S.; Molinaro, A.; Van Der Laan, M.J. Survival ensembles. Biostatistics 2006, 7, 355–373. [Google Scholar] [CrossRef]
- Breiman, L. Random forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Ishwaran, H.; Lu, M. Random survival forests. Ann. Appl. Stat. 2008, 2, 841–860. [Google Scholar] [CrossRef]
- McCulloch, W.S.; Pitts, W. A logical calculus of the ideas immanent in nervous activity. Bull. Math. Biophys. 1943, 5, 115–133. [Google Scholar] [CrossRef]
- Rosenblatt, F. Principles of Neurodynamics. Perceptrons and the Theory of Brain Mechanisms; Cornell Aeronautical Lab Inc.: Buffalo, NY, USA, 1961. [Google Scholar]
- Shmueli, G.; Bruce, P.C.; Stephens, M.L.; Patel, N.R. Data Mining for Business Analytics: Concepts, Techniques, and Applications with JMP Pro; John Wiley & Sons: Hoboken, NJ, USA, 2016; ISBN 978-1-118-87752-4. [Google Scholar]
- Cortes, C.; Vapnik, V. Support-Vector Networks. Mach. Learn. 1995, 20, 273–297. [Google Scholar] [CrossRef]
- Bootstrap Forest. Available online: https://www.jmp.com/support/help/14-2/bootstrap-forest.shtml (accessed on 11 October 2019).
- Zou, H.; Hastie, T. Regularization and variable selection via the elastic net. J. R. Stat. Soc. Ser. B (Stat. Methodol.) 2005, 67, 301–320. [Google Scholar] [CrossRef]
- Hothorn, T.; Hornik, K.; Zeileis, A. Unbiased Recursive Partitioning: A Conditional Inference Framework. J. Comput. Graph. Stat. 2006, 15, 651–674. [Google Scholar] [CrossRef]
- Cutler, A.; Breiman, L. Archetypal analysis. Technometrics 1994, 36, 338–347. [Google Scholar] [CrossRef]
- OPTN Estimated Post Transplant Score (EPTS) Guide-Updated. 2019. Available online: https://optn.transplant.hrsa.gov/media/1511/guide_to_calculating_interpreting_epts.pdf (accessed on 3 June 2019).
- Efron, B. Bootstrap methods: Another look at the jackknife. In Breakthroughs in Statistics; Springer: New York, NY, USA, 1992; pp. 569–593. [Google Scholar]
- Friedman, J.; Hastie, T.; Tibshirani, R. The Elements of Statistical Learning; Springer: New York, NY, USA, 2001; Volume 1. [Google Scholar]
- Leeaphorn, N.; Thongprayoon, C.; Chon, W.J.; Cummings, L.S.; Mao, M.A.; Cheungpasitporn, W. Outcomes of kidney retransplantation after graft loss as a result of BK virus nephropathy in the era of newer immunosuppressant agents. Am. J. Transplant. 2019. [Google Scholar] [CrossRef]
- Redfield, R.R.; Gupta, M.; Rodriguez, E.; Wood, A.; Abt, P.L.; Levine, M.H. Graft and patient survival outcomes of a third kidney transplant. Transplantation 2015, 99, 416–423. [Google Scholar] [CrossRef]
- Risk Factors for Second Renal Allografts Immunosuppressed with Cyclosporine—Abstract—Europe PMC. Available online: https://europepmc.org/article/med/1871798 (accessed on 15 February 2020).
- Mukras, R.; Wiratunga, N.; Lothian, R.; Chakraborti, S.; Harper, D.J. Information Gain Feature Selection for Ordinal Text Classification using Probability Re-distribution. Available online: https://pdfs.semanticscholar.org/08cd/c505cfa896185e600f27671d515244e2e41a.pdf?_ga=2.22802107.10072257.1582093750-839507928.1572830582 (accessed on 19 February 2020).
- Decision Trees | SpringerLink. Available online: https://link.springer.com/chapter/10.1007/0-387-25465-X_9 (accessed on 29 December 2019).
- Patel, N.; Upadhyay, S. Study of various decision tree pruning methods with their empirical comparison in WEKA. Int. J. Comput. Appl. 2012, 60, 20–25. [Google Scholar] [CrossRef]
- Quinlan, J.R. C4. 5: Programs for Machine Learning; Elsevier: San Francisco, CA, USA, 2014; ISBN 0-08-050058-7. [Google Scholar]
- Breiman, L. Bagging predictors. Mach. Learn. 1996, 24, 123–140. [Google Scholar] [CrossRef]
- Haykin, S. Neural Networks: A Comprehensive Foundation; Prentice Hall PTR: Upper Saddle River, NJ, USA, 1994; ISBN 0-02-352761-7. [Google Scholar]
- Hearst, M.A.; Dumais, S.T.; Osuna, E.; Platt, J.; Scholkopf, B. Support vector machines. IEEE Intell. Syst. Their Appl. 1998, 13, 18–28. [Google Scholar] [CrossRef]
- Cheng, J.; Greiner, R. Learning Bayesian Belief Network Classifiers: Algorithms and System; Springer: Heidelberg, Germany, 2001; pp. 141–151. [Google Scholar]
- Weinberger, K.Q.; Saul, L.K. Distance metric learning for large margin nearest neighbor classification. J. Mach. Learn. Res. 2009, 10, 207–244. [Google Scholar]
- OPTN A Guide to Calculating and Interpreting the Kidney Donor Profle Index (KDPI). Available online: https://optn.transplant.hrsa.gov/media/1512/guide_to_calculating_interpreting_kdpi.pdf (accessed on 3 June 2019).
- Zhang, H. The optimality of naive Bayes. AA 2004, 1, 3. [Google Scholar]
- Cox, C.; Chu, H.; Schneider, M.F.; Muñoz, A. Parametric survival analysis and taxonomy of hazard functions for the generalized gamma distribution. Stat. Med. 2007, 26, 4352–4374. [Google Scholar] [CrossRef]
- Zhang, Z. Parametric regression model for survival data: Weibull regression model as an example. Ann. Transl. Med. 2016, 4, 484. [Google Scholar] [CrossRef]
| Authors | Data Source | Population | Methods Used |
|---|---|---|---|
| Yoo et al., 2017 [7] | Three centers in Korea (1997–2012) | 3117 | Survival decision tree, bagging and random forest |
| Mark et al., 2019 [32] | United Network for Organ Sharing (UNOS) (1987–2014) | 100,000 | Combines predictions from random survival forest constructed from conditional inference trees with a Cox proportional hazards model |
| Bae et al., 2019 [33] | Organ Procurement and Transplantation Network (OPTN) (2005–2016) | 120,818 kidney recipients and 376,272 candidates on the waiting list | Combinations of The Kidney Donor Profile Index and the Estimated Post Transplant Survival score using random survival forest, and waitlist survival by Estimated Post Transplant Survival score using Weibull regressions |
| Atallah et al., 2019 [34] | Urology and Nephrology Center, Mansoura, Egypt (1976–2017) | 2728 | Merges information gain with Naïve Bayes and k-nearest neighbour |
| Nematollahi et al., 2017 [35] | Nemazee Hospital, Shiraz, southern Iran (2008–2012) | 717 | Artificial neural network and support vector machines |
| Tapak et al, 2017 [31] | Ekbatan or Besaat hospitals, Iran (1994–2011) | 378 | Artificial neuronal networks |
| Shahmoradi et al., 2016 [36] | Sina Hospital Urology Research Center, Iran (2007–2013) | 513 | C5.0, artificial neural networks and classification and regression trees |
| Luck et al., 2017 [30] | Scientific Registry of Transplant Recipients (2000–2014) | 131,709 | Artificial neural network taking into account two kinds of information loss, the presence of ties and the presence of censoring. |
| Topuz et al., 2018 [37] | UNOS (2004–2015) | 31,207 | Feature selection with support vector machine, artificial neural network and bootstrap to construct a Bayesian belief network |
| Authors | Method Used | Benchmark | Best Performance | Accuracy | Sensitivity |
|---|---|---|---|---|---|
| Nematollahi et al., 2017 [35] | ANN and SVM | Logistic regression | SVM | 90.40 | 98.20 |
| Tapak et al, 2017 [31] | ANN | Logistic regression | ANN | 75 | 91 |
| Shahmoradi et al., 2016 [36] | C5.0, ANN and CART | N/A | C5.0 | 87.21 | 90.85 |
| Atallah et al., 2019 [34] | Merges information gain with naïve Bayes and K-NN | J48, Naïve Bayes, ANN, RF, SVM, KNN | Proposed method | 80.77 | 80.40 |
| Topuz et al., 2018 [37] | Feature selection with SVM, ANN and bootstrap to construct a Bayesian Belief Network | N/A | Proposed method | 68.40 | 41.00 |
| Authors | Method Used | Benchmark | Best Performance | C-Index |
|---|---|---|---|---|
| Yoo et al., 2017 [7] | Survival decision tree, bagging, and RF | Decision tree and Cox regression | Survival decision tree | 0.80 |
| Mark et al., 2019 [32] | Combines predictions from RSF constructed from conditional inference trees with a Cox proportional hazards model | EPTS, Cox model, random forest | Proposed method | 0.724 |
| Luck et al., 2017 [30] | Artificial neural network taking into account two kinds of information loss, the presence of ties and the presence of censoring. | Traditional Cox model using Efron’s method | Proposed method | 0.6550 |
| Bae et al., 2019 [33] | Combinations of KDPI and EPTS using RSF | KDRI | Proposed method | 0.637 |
| Recipient/Donor Factors |
|---|
| The 3-month serum creatinine level post-transplant |
| Recipient age |
| Kidney cold ischemic time |
| Donor age |
| Discharge time creatinine |
| Body mass index |
| Pre-transplant dialysis |
| Recipient functional status at registration |
| Recipient diabetes at registration |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Díez-Sanmartín, C.; Sarasa Cabezuelo, A. Application of Artificial Intelligence Techniques to Predict Survival in Kidney Transplantation: A Review. J. Clin. Med. 2020, 9, 572. https://doi.org/10.3390/jcm9020572
Díez-Sanmartín C, Sarasa Cabezuelo A. Application of Artificial Intelligence Techniques to Predict Survival in Kidney Transplantation: A Review. Journal of Clinical Medicine. 2020; 9(2):572. https://doi.org/10.3390/jcm9020572
Chicago/Turabian StyleDíez-Sanmartín, Covadonga, and Antonio Sarasa Cabezuelo. 2020. "Application of Artificial Intelligence Techniques to Predict Survival in Kidney Transplantation: A Review" Journal of Clinical Medicine 9, no. 2: 572. https://doi.org/10.3390/jcm9020572
APA StyleDíez-Sanmartín, C., & Sarasa Cabezuelo, A. (2020). Application of Artificial Intelligence Techniques to Predict Survival in Kidney Transplantation: A Review. Journal of Clinical Medicine, 9(2), 572. https://doi.org/10.3390/jcm9020572






