Considering Exosomal miR-21 as a Biomarker for Cancer
Abstract
:1. Introduction
2. Materials and Methods
2.1. Search Formula
2.2. Inclusion Criteria and Data Extraction
2.3. Statistics Analysis
3. Results
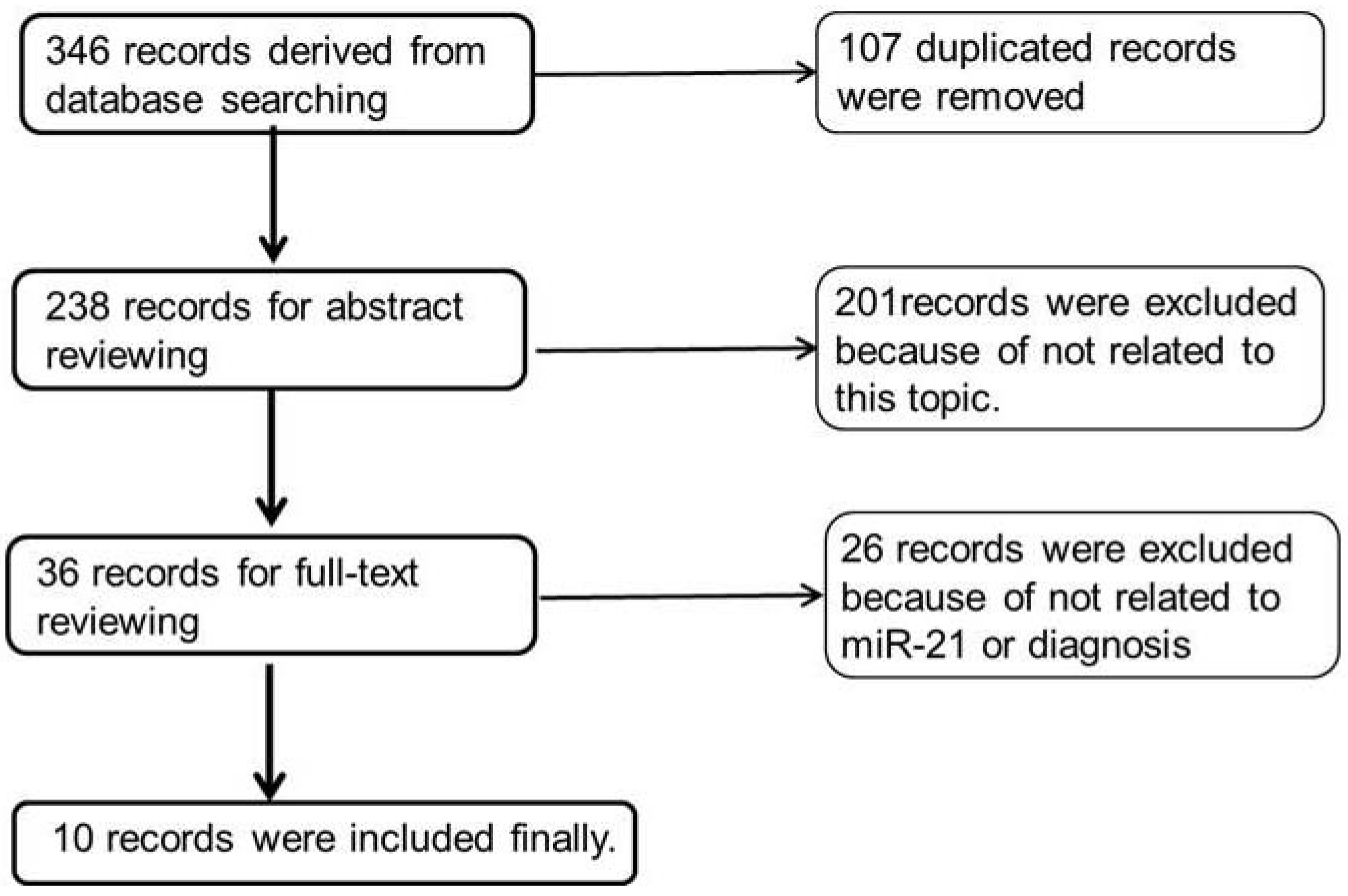
3.1. Literature Search Results and Summary of Studies
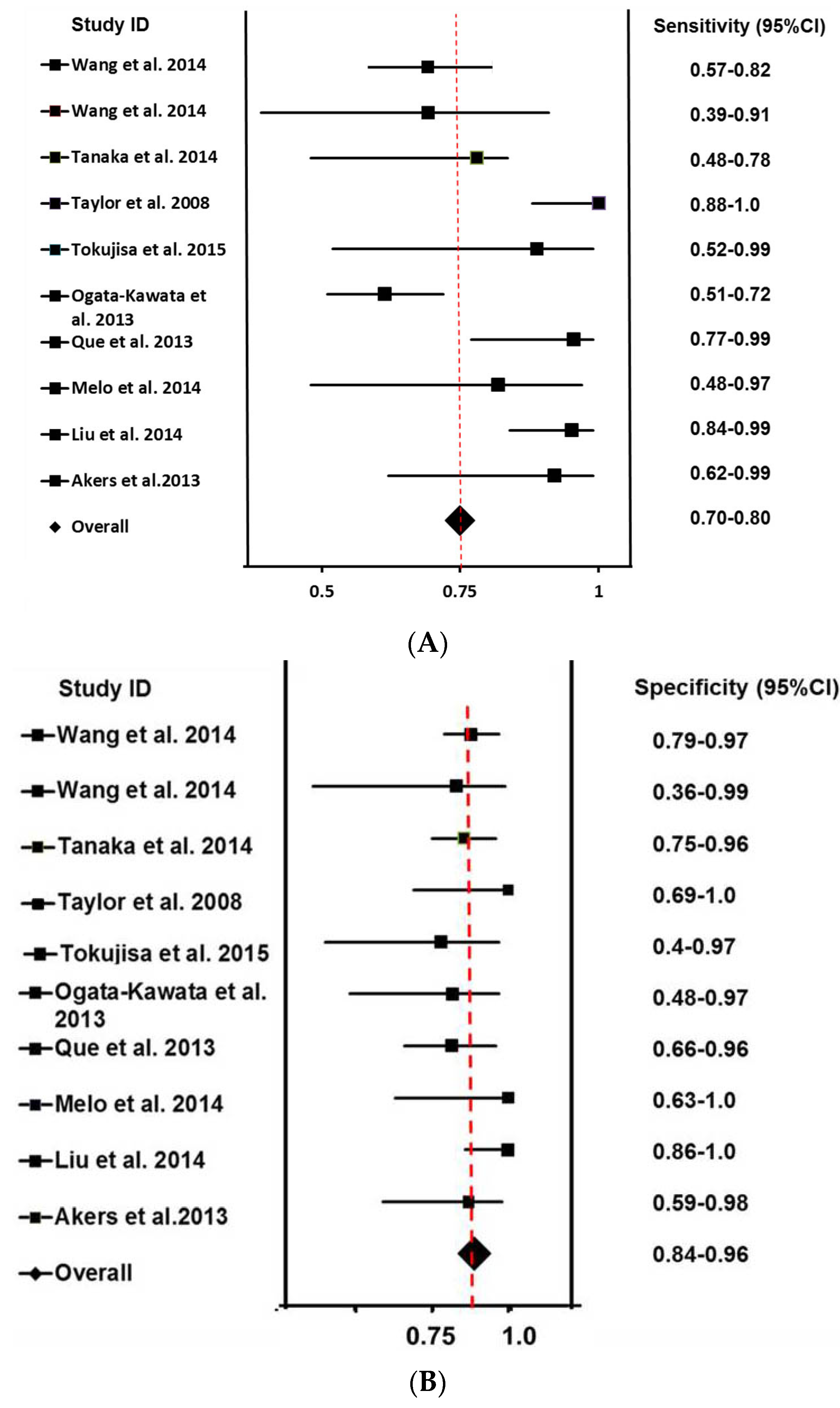
3.2. The Sensitivity and Specificity of Pooled Studies
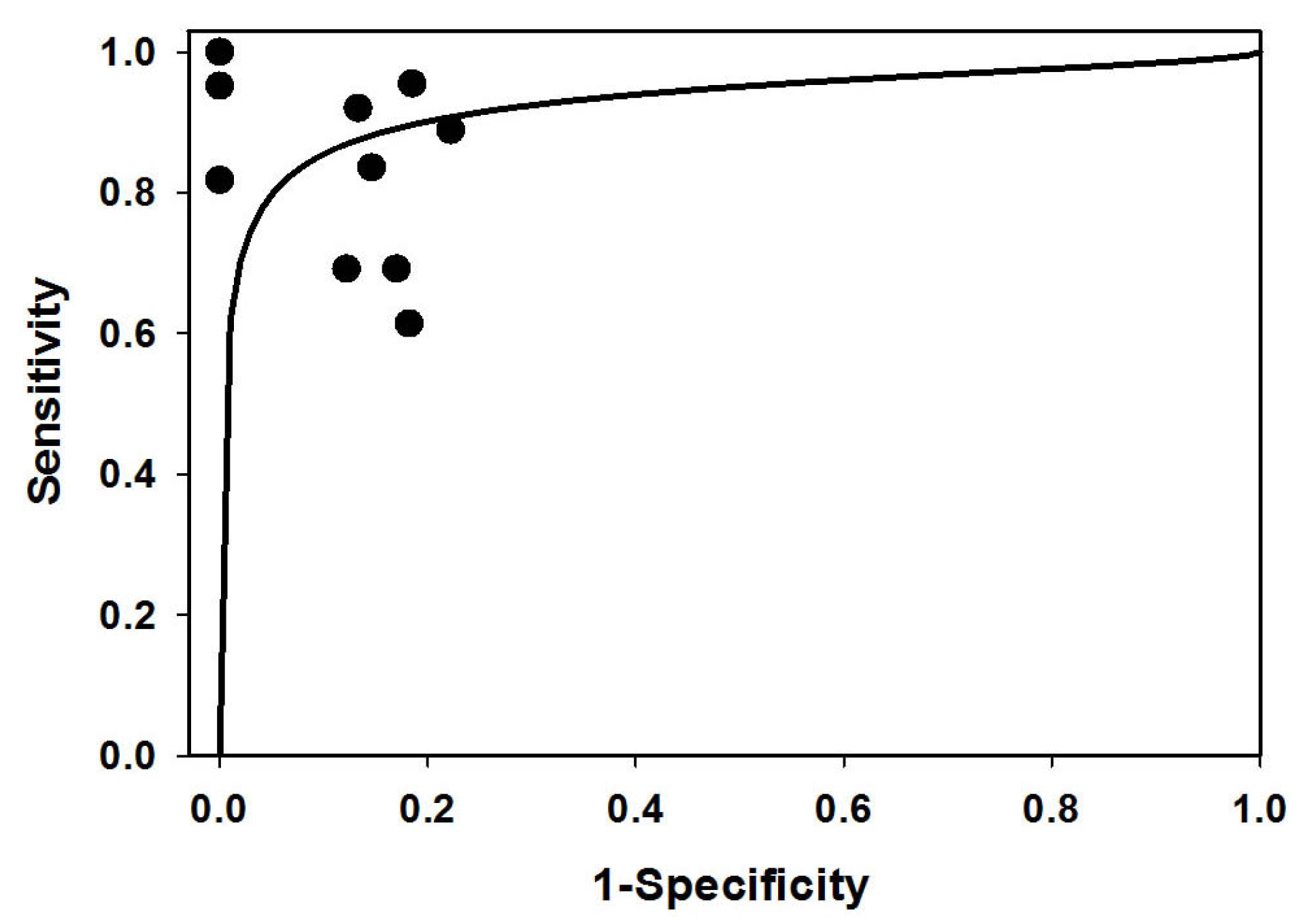
3.3. The SROC and AUC of Pooled Studies
3.4. The Cutoff Values and Endogenous Controls
4. Discussion
4.1. The Exosomal miR-21 as a Universal Biomarker for Diagnostic Cancers
4.2. The Combination of miRNA Panels and Cancer Antigens
4.3. Choosing the Right Internal Controls
4.4. Comparison of Exosomal miR-21 Expression in Other Diseases
5. Conclusions and Prospect
Acknowledgments
Conflicts of Interest
References
- Wu, R.; Jiang, Y.; Wu, Q.; Li, Q.; Cheng, D.; Xu, L.; Zhang, C.; Zhang, M.; Ye, L. Diagnostic value of microRNA-21 in the diagnosis of lung cancer: Evidence from a meta-analysis involving 11 studies. Tumour Biol. 2014, 35, 8829–8836. [Google Scholar] [CrossRef] [PubMed]
- Duffy, M.J. Role of tumor markers in patients with solid cancers: A critical review. Eur. J. Intern. Med. 2007, 18, 175–184. [Google Scholar] [CrossRef] [PubMed]
- O’Byrne, K.J.; Gatzemeier, U.; Bondarenko, I.; Barrios, C.; Eschbach, C.; Martens, U.M.; Hotko, Y.; Kortsik, C.; Paz-Ares, L.; Pereira, J.R.; et al. Molecular biomarkers in non-small-cell lung cancer: A retrospective analysis of data from the phase 3 flex study. Lancet Oncol. 2011, 12, 795–805. [Google Scholar]
- Ambros, V. The functions of animal microRNAs. Nature 2004, 431, 350–355. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef]
- Shi, J. Regulatory networks between neurotrophins and miRNAs in brain diseases and cancers. Actapharmacol. Sin. 2015, 36, 149–157. [Google Scholar] [CrossRef] [PubMed]
- Schwarzenbach, H.; Nishida, N.; Calin, G.A.; Pantel, K. Clinical relevance of circulating cell-free microRNAs in cancer. Nat. Rev. Clin. Oncol. 2014, 11, 145–156. [Google Scholar] [CrossRef] [PubMed]
- Yoshino, H.; Seki, N.; Itesako, T.; Chiyomaru, T.; Nakagawa, M.; Enokida, H. Aberrant expression of microRNAs in bladder cancer. Nat. Rev. Urol. 2013, 10, 396–404. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, R.U.; Miyazaki, H.; Ochiya, T. The roles of microRNAs in breast cancer. Cancers 2015, 7, 598–616. [Google Scholar] [CrossRef] [PubMed]
- Wu, K.; Li, L.; Li, S. Circulating microRNA-21 as a biomarker for the detection of various carcinomas: An updated meta-analysis based on 36 studies. Tumour Biol. 2015, 36, 1973–1981. [Google Scholar] [CrossRef] [PubMed]
- Zeng, Z.; Wang, J.; Zhao, L.; Hu, P.; Zhang, H.; Tang, X.; He, D.; Tang, S.; Zeng, Z. Potential role of microRNA-21 in the diagnosis of gastric cancer: A meta-analysis. PLoS ONE 2013, 8, e73278. [Google Scholar] [CrossRef] [PubMed]
- Colombo, M.; Raposo, G.; Thery, C. Biogenesis, secretion, and intercellular interactions of exosomes and other extracellular vesicles. Annu. Rev. Cell Dev. Biol. 2014, 30, 255–289. [Google Scholar] [CrossRef] [PubMed]
- Cai, X.; Hagedorn, C.H.; Cullen, B.R. Human microRNAs are processed from capped, polyadenylated transcripts that can also function as miRNAs. RNA 2004, 10, 1957–1966. [Google Scholar] [CrossRef] [PubMed]
- Loffler, D.; Brocke-Heidrich, K.; Pfeifer, G.; Stocsits, C.; Hackermuller, J.; Kretzschmar, A.K.; Burger, R.; Gramatzki, M.; Blumert, C.; Bauer, K.; et al. Interleukin-6 dependent survival of multiple myeloma cells involves the Stat3-mediated induction of microRNA-21 through a highly conserved enhancer. Blood 2007, 110, 1330–1333. [Google Scholar] [CrossRef] [PubMed]
- Fujita, S.; Ito, T.; Mizutani, T.; Minoguchi, S.; Yamamichi, N.; Sakurai, K.; Iba, H. miR-21 gene expression triggered by AP-1 is sustained through a double-negative feedback mechanism. J. Mol. Biol. 2008, 378, 492–504. [Google Scholar] [CrossRef] [PubMed]
- Shen, L.; Wan, Z.; Ma, Y.; Wu, L.; Liu, F.; Zang, H.; Xin, S. The clinical utility of microRNA-21 as novel biomarker for diagnosing human cancers. Tumour Biol. 2015, 36, 1993–2005. [Google Scholar] [CrossRef] [PubMed]
- Meng, F.; Henson, R.; Wehbe-Janek, H.; Ghoshal, K.; Jacob, S.T.; Patel, T. microRNA-21 regulates expression of the pten tumor suppressor gene in human hepatocellular cancer. Gastroenterology 2007, 133, 647–658. [Google Scholar] [CrossRef] [PubMed]
- Sayed, D.; Rane, S.; Lypowy, J.; He, M.; Chen, I.Y.; Vashistha, H.; Yan, L.; Malhotra, A.; Vatner, D.; Abdellatif, M. microRNA-21 targets sprouty2 and promotes cellular outgrowths. Mol. Biol. Cell 2008, 19, 3272–3282. [Google Scholar] [CrossRef] [PubMed]
- Xiong, B.; Cheng, Y.; Ma, L.; Zhang, C. miR-21 regulates biological behavior through the PTEN/PI-3 k/Akt signaling pathway in human colorectal cancer cells. Int. J. Oncol. 2013, 42, 219–228. [Google Scholar] [CrossRef] [PubMed]
- Asangani, I.A.; Rasheed, S.A.; Nikolova, D.A.; Leupold, J.H.; Colburn, N.H.; Post, S.; Allgayer, H. MicroRNA-21 (miR-21) post-transcriptionally downregulates tumor suppressor Pdcd4 and stimulates invasion, intravasation and metastasis in colorectal cancer. Oncogene 2008, 27, 2128–2136. [Google Scholar] [CrossRef] [PubMed]
- Peacock, O.; Lee, A.C.; Cameron, F.; Tarbox, R.; Vafadar-Isfahani, N.; Tufarelli, C.; Lund, J.N. Inflammation and miR-21 pathways functionally interact to downregulate PDCD4 in colorectal cancer. PLoS ONE 2014, 9, e110267. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.H.; Yue, J.; Pfeffer, S.R.; Handorf, C.R.; Pfeffer, L.M. MicroRNA miR-21 regulates the metastatic behavior of B16 melanoma cells. J. Biol. Chem. 2011, 286, 39172–39178. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Li, P.F. Foxo3a regulates apoptosis by negatively targeting miR-21. J. Biol. Chem. 2010, 285, 16958–16966. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.H.; Pfeffer, S.R.; Sims, M.; Yue, J.; Wang, Y.; Linga, V.G.; Paulus, E.; Davidoff, A.M.; Pfeffer, L.M. The oncogenic microRNA-21 inhibits the tumor suppressive activity of FBXO11 to promote tumorigenesis. J. Biol. Chem. 2015, 290, 6037–6046. [Google Scholar] [CrossRef] [PubMed]
- Martin del Campo, S.E.; Latchana, N.; Levine, K.M.; Grignol, V.P.; Fairchild, E.T.; Jaime-Ramirez, A.C.; Dao, T.V.; Karpa, V.I.; Carson, M.; Ganju, A.; et al. miR-21 enhances melanoma invasiveness via inhibition of tissue inhibitor of metalloproteinases 3 expression: In vivo effects of miR-21 inhibitor. PLoS ONE 2015, 10, e0115919. [Google Scholar]
- Liu, L.Z.; Li, C.; Chen, Q.; Jing, Y.; Carpenter, R.; Jiang, Y.; Kung, H.F.; Lai, L.; Jiang, B.H. miR-21 induced angiogenesis through AKT and ERK activation and HIF-1α expression. PLoS ONE 2011, 6, e19139. [Google Scholar] [CrossRef] [PubMed]
- Qi, J.H.; Ebrahem, Q.; Moore, N.; Murphy, G.; Claesson-Welsh, L.; Bond, M.; Baker, A.; Anand-Apte, B. A novel function for tissue inhibitor of metalloproteinases-3 (TIMP3): Inhibition of angiogenesis by blockage of VEGF binding to VEGF receptor-2. Nat. Med. 2003, 9, 407–415. [Google Scholar] [CrossRef] [PubMed]
- Zhu, S.; Si, M.L.; Wu, H.; Mo, Y.Y. MicroRNA-21 targets the tumor suppressor gene tropomyosin 1 (TMP1). J. Biol. Chem. 2007, 282, 14328–14336. [Google Scholar] [CrossRef] [PubMed]
- Melnik, B.C. Mir-21: An environmental driver of malignant melanoma? J. Transl. Med. 2015, 13, 202. [Google Scholar] [CrossRef] [PubMed]
- Shen, J.; Todd, N.W.; Zhang, H.; Yu, L.; Lingxiao, X.; Mei, Y.; Guarnera, M.; Liao, J.; Chou, A.; Lu, C.L.; et al. Plasma microRNAs as potential biomarkers for non-small-cell lung cancer. Labo. Investig. 2011, 91, 579–587. [Google Scholar] [CrossRef] [PubMed]
- Deville, W.L.; Buntinx, F.; Bouter, L.M.; Montori, V.M.; de Vet, H.C.; van der Windt, D.A.; Bezemer, P.D. Conducting systematic reviews of diagnostic studies: Didactic guidelines. BMC Med. Res. Methodol. 2002, 2, 9. [Google Scholar] [CrossRef] [PubMed]
- Moses, L.E.; Shapiro, D.; Littenberg, B. Combining independent studies of a diagnostic test into a summary roc curve: Data-analytic approaches and some additional considerations. Stat. Med. 1993, 12, 1293–1316. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Hou, L.; Li, A.; Duan, Y.; Gao, H.; Song, X. Expression of serum exosomal microRNA-21 inhuman hepatocellular carcinoma. BioMed Res. Int. 2014, 2014, 864894. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Zhou, Y.; Lu, J.; Sun, Y.; Xiao, H.; Liu, M.; Tian, L. Combined detection of serum exosomal miR-21 and hotair as diagnostic and prognostic biomarkers for laryngeal squamous cell carcinoma. Med. Oncol. 2014, 31, 148. [Google Scholar] [CrossRef] [PubMed]
- Ogata-Kawata, H.; Izumiya, M.; Kurioka, D.; Honma, Y.; Yamada, Y.; Furuta, K.; Gunji, T.; Ohta, H.; Okamoto, H.; Sonoda, H.; et al. Circulating exosomal microRNAs as biomarkers of colon cancer. PLoS ONE 2014, 9, e92921. [Google Scholar] [CrossRef] [PubMed]
- Que, R.; Ding, G.; Chen, J.; Cao, L. Analysis of serum exosomal microRNAs and clinicopathologic features of patients with pancreatic adenocarcinoma. World J. Surg. Oncol. 2013, 11, 219. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, Y.; Kamohara, H.; Kinoshita, K.; Kurashige, J.; Ishimoto, T.; Iwatsuki, M.; Watanabe, M.; Baba, H. Clinical impact of serum exosomal microRNA-21 as a clinical biomarker in human esophageal squamous cell carcinoma. Cancer 2013, 119, 1159–1167. [Google Scholar] [CrossRef] [PubMed]
- Taylor, D.D.; Gercel-Taylor, C. MicroRNA signatures of tumor-derived exosomes as diagnostic biomarkers of ovarian cancer. Gynecol. Oncol. 2008, 110, 13–21. [Google Scholar] [CrossRef] [PubMed]
- Melo, S.A.; Sugimoto, H.; O’Connell, J.T.; Kato, N.; Villanueva, A.; Vidal, A.; Qiu, L.; Vitkin, E.; Perelman, L.T.; Melo, C.A.; et al. Cancer exosomes perform cell-independent microRNA biogenesis and promote tumorigenesis. Cancer Cell 2014, 26, 707–721. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Sun, H.; Wang, X.; Yu, Q.; Li, S.; Yu, X.; Gong, W. Increased exosomal microRNA-21 and microRNA-146a levels in the cervicovaginal lavage specimens of patients with cervical cancer. Int. J. Mol. Sci. 2014, 15, 758–773. [Google Scholar] [CrossRef] [PubMed]
- Akers, J.C.; Ramakrishnan, V.; Kim, R.; Skog, J.; Nakano, I.; Pingle, S.; Kalinina, J.; Hua, W.; Kesari, S.; Mao, Y.; et al. MiR-21 in the extracellular vesicles (EVs) of cerebrospinal fluid (CSF): A platform for glioblastoma biomarker development. PLoS ONE 2013, 8, e78115. [Google Scholar]
- Tokuhisa, M.; Ichikawa, Y.; Kosaka, N.; Ochiya, T.; Yashiro, M.; Hirakawa, K.; Kosaka, T.; Makino, H.; Akiyama, H.; Kunisaki, C.; et al. Exosomal miRNAs from peritoneum lavage fluid as potential prognostic biomarkers of peritoneal metastasis in gastric cancer. PLoS ONE 2015, 10, e0130472. [Google Scholar] [CrossRef] [PubMed]
- Dahabreh, I.J.; Trikalinos, T.A.; Lau, J.; Schmid, C. An empirical Assessment of Bivariate Methods for Meta-Analysis of Test Accuracy; Agency for Healthcare Research and Quality: Rockville, MD, USA, 2012. [Google Scholar]
- Wei, J.; Gao, W.; Zhu, C.; Liu, Y.; Mei, Z.; Cheng, T.; Shu, Y. Identification of plasma microRNA-21 as a biomarker for early detection and chemosensitivity of non-small lung cancer. Chin. J. Caner 2011, 30, 407–414. [Google Scholar] [CrossRef]
- He, X.Y.; Liao, Y.D.; Guo, X.Q.; Wang, R.; Xiao, Z.Y.; Wang, Y.G. Prognostic role of microRNA-21 expression in brain tumors: A meta-analysis. Mol. Neurobiol. 2016, 53, 1856–1861. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Zheng, Z.; Guo, J.; Ding, X. Correlation and quantitation of microRNA aberrant expression in tissues and sera from patients with breast tumor. Gynecol. Oncol. 2010, 119, 586–593. [Google Scholar] [CrossRef] [PubMed]
- Shen, J.; Liu, Z.; Todd, N.W.; Zhang, H.; Liao, J.; Yu, L.; Guarnera, M.A.; Li, R.; Cai, L.; Zhan, M.; et al. Diagnosis of lung cancer in individuals with solitary pulmonary nodules by plasma microRNA biomarkers. BMC Cancer 2011, 11, 374. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Yu, L.; Gao, X.; Hu, J.; Wang, J.; Dai, Z.; Wang, J.F.; Zhang, Z.; Lu, S.; Huang, X.; et al. Plasma microRNA panel to diagnose hepatitis b virus-related hepatocellular carcinoma. J. Clin. Oncol. 2011, 29, 4781–4788. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Cai, F.; Zhang, B.; Barekati, Z.; Zhong, X.Y. The level of circulating miRNA-10b and miRNA-373 in detecting lymph node metastasis of breast cancer: Potential biomarkers. Tumour Biol. 2013, 34, 455–462. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Gao, J.; Du, Y.; Li, Z.; Ren, Y.; Gu, J.; Wang, X.; Gong, Y.; Wang, W.; Kong, X. Combination of plasma microRNAs with serum CA19-9 for early detection of pancreatic cancer. Int. J. Cancer 2012, 131, 683–691. [Google Scholar] [CrossRef] [PubMed]
- Xiang, M.; Zeng, Y.; Yang, R.; Xu, H.; Chen, Z.; Zhong, J.; Xie, H.; Xu, Y.; Zeng, X. U6 is not a suitable endogenous control for the quantification of circulating microRNAs. Biochem. Biophys. Res. Commun. 2014, 454, 210–214. [Google Scholar] [CrossRef] [PubMed]
- Popov, A.; Szabo, A.; Mandys, V. Small nucleolar RNA U91 is a new internal control for accurate microRNAs quantification in pancreatic cancer. BMC Cancer 2015, 15, 774. [Google Scholar] [CrossRef] [PubMed]
- Lou, G.; Ma, N.; Xu, Y.; Jiang, L.; Yang, J.; Wang, C.; Jiao, Y.; Gao, X. Differential distribution of U6 (RNU6–1) expression in human carcinoma tissues demonstrates the requirement for caution in the internal control gene selection for microRNA quantification. Int. J. Mol. Med. 2015, 36, 1400–1408. [Google Scholar] [CrossRef] [PubMed]
- Kroh, E.M.; Parkin, R.K.; Mitchell, P.S.; Tewari, M. Analysis of circulating microRNA biomarkers in plasma and serum using quantitative reverse transcription-PCR (qRT-PCR). Methods 2010, 50, 298–301. [Google Scholar] [CrossRef] [PubMed]
- Ell, B.; Mercatali, L.; Ibrahim, T.; Campbell, N.; Schwarzenbach, H.; Pantel, K.; Amadori, D.; Kang, Y. Tumor-induced osteoclast miRNA changes as regulators and biomarkers of osteolytic bone metastasis. Cancer Cell 2013, 24, 542–556. [Google Scholar] [CrossRef] [PubMed]
- McDermott, A.M.; Kerin, M.J.; Miller, N. Identification and validation of miRNAs as endogenous controls for RQ-PCR in blood specimens for breast cancer studies. PLoS ONE 2013, 8, e83718. [Google Scholar]
- Rice, J.; Roberts, H.; Rai, S.N.; Galandiuk, S. Housekeeping genes for studies of plasma microRNA: A need for more precise standardization. Surgery 2015, 158, 1345–1351. [Google Scholar] [CrossRef] [PubMed]
- Goswami, S.; Banerjee, A.; Kumari, B.; Bandopadhyay, B.; Bhattacharya, N.; Basu, N.; Vrati, S.; Banerjee, A. Differential expression and significance of circulating microRNAs in cerebrospinal fluid of acute encephalitis patients infected with Japanese Encephalitis Virus. Mol. Neurobiol. 2016. [Google Scholar] [CrossRef] [PubMed]
| Study ID | Patients | Controls | Cancer | Specimen | 2 × 2 Table | |||
|---|---|---|---|---|---|---|---|---|
| TP | FN | FP | TN | |||||
| Wang 2014 [33] | 52 | 49 | LSCC | Serum exosome | 36 | 16 | 6 | 43 |
| Wang 2014 [34] | 13 | 30 | HCC (III-IV) | Serum exosome | 9 | 4 | 12 | 18 |
| Tanaka 2013 [37] | 44 | 41 | ESCC | Serum exosome | 28 | 16 | 6 | 35 |
| Taylor 2008 [38] | 30 | 10 | OC | Serum exosome | 30 | 0 | 0 | 10 |
| Tokuhisa 2015 [42] | 9 | 9 | GC | PLF exosome | 8 | 1 | 2 | 7 |
| Ogata-Kawata 2013 [35] | 88 | 11 | CC | Serum exosome | 54 | 34 | 2 | 9 |
| Que 2013 [36] | 22 | 27 | PC | Serum exosome | 21 | 1 | 5 | 22 |
| Liu 2014 [40] | 45 | 25 | Cervical cancer | CLF exosome | 40 | 5 | 0 | 25 |
| Melo 2014 [39] | 11 | 8 | BC | Serum exosome | 9 | 2 | 0 | 8 |
| Akers 2013 [41] | 13 | 14 | Glioblastoma | CSF-EV | 11 | 2 | 1 | 13 |
| Cancer | Cutoff Point | Internal Control | qPCR | Exosome Isolation | |
|---|---|---|---|---|---|
| Fold | 2−ΔΔCT | ||||
| HCC [33] | 5 | 0.03 ** | U6 | SYBR Green | Reagent (Life Tech.) |
| PC [36] | 4.05 | 0.06 ** | U6 | TaqMan | Ultracentrifugation |
| ESCC [37] | 5.66 * | 0.02 | miR-16 | TaqMan | ExoQuick (SBI) |
| LSCC [34] | 4.55 * | 0.043 | U6 | SYBR Green | ExoQuick (SBI) |
| CC [35] | 6.56 * | 0.0108 | miR-451 | TaqMan | Ultracentrifugation |
| GC [42] | 3.5 * | 0.088 | miR-16 | TagMan | Ultracentrifugation |
| OC [38] | 11 intensity | None | microarry | None | MACS |
| Cervical C. [40] cancer [40] | 3.0-fold | None | None | TaqMan | Ultracentrifugation |
| Glioblastoma [41] | 0.25/EV | None | None | absolute | Ultracentrifugation |
© 2016 by the author; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shi, J. Considering Exosomal miR-21 as a Biomarker for Cancer. J. Clin. Med. 2016, 5, 42. https://doi.org/10.3390/jcm5040042
Shi J. Considering Exosomal miR-21 as a Biomarker for Cancer. Journal of Clinical Medicine. 2016; 5(4):42. https://doi.org/10.3390/jcm5040042
Chicago/Turabian StyleShi, Jian. 2016. "Considering Exosomal miR-21 as a Biomarker for Cancer" Journal of Clinical Medicine 5, no. 4: 42. https://doi.org/10.3390/jcm5040042
APA StyleShi, J. (2016). Considering Exosomal miR-21 as a Biomarker for Cancer. Journal of Clinical Medicine, 5(4), 42. https://doi.org/10.3390/jcm5040042






