The Genetics of Hereditary Angioedema: A Review
Abstract
1. Introduction
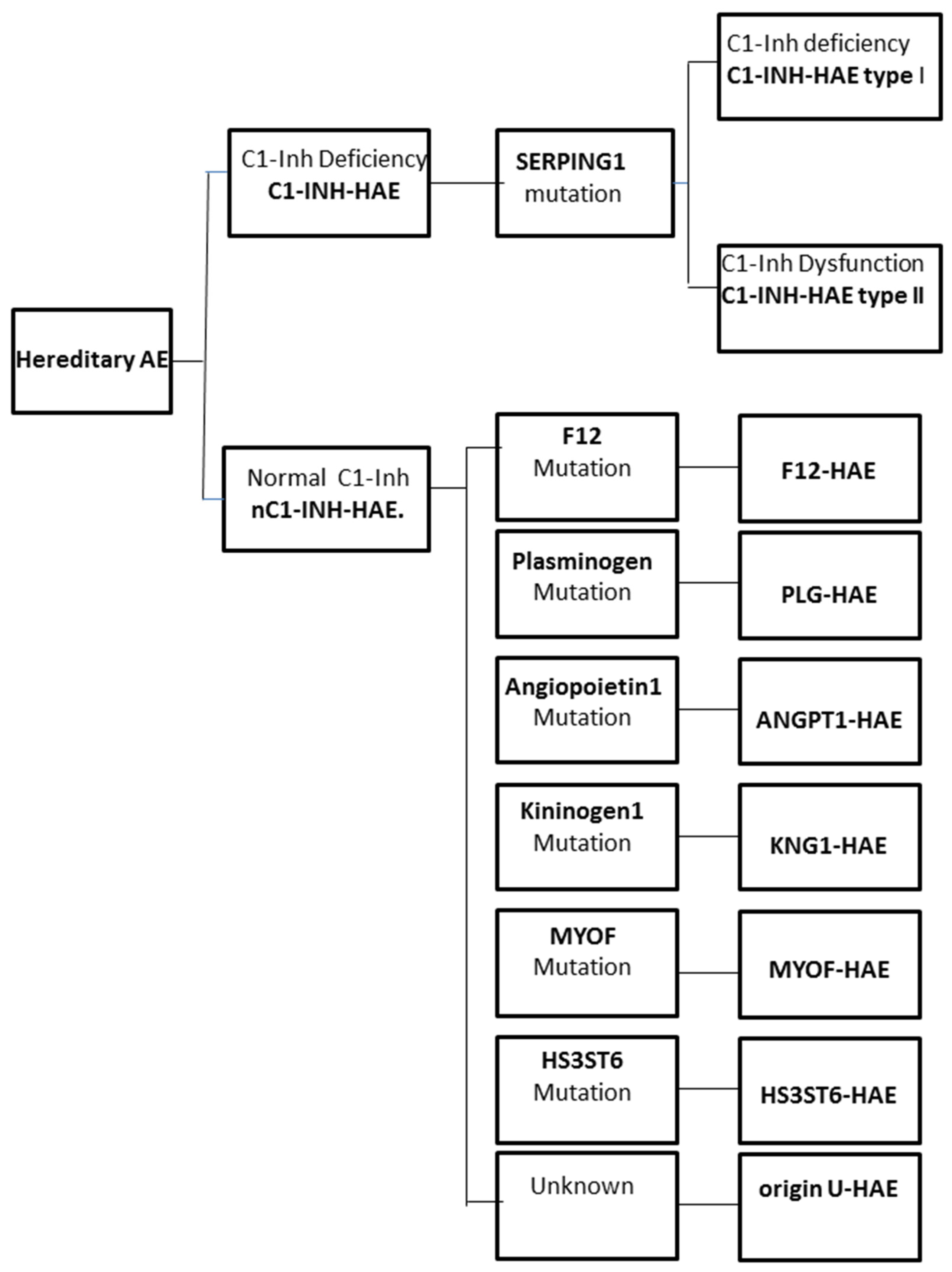
2. Hereditary Angioedema
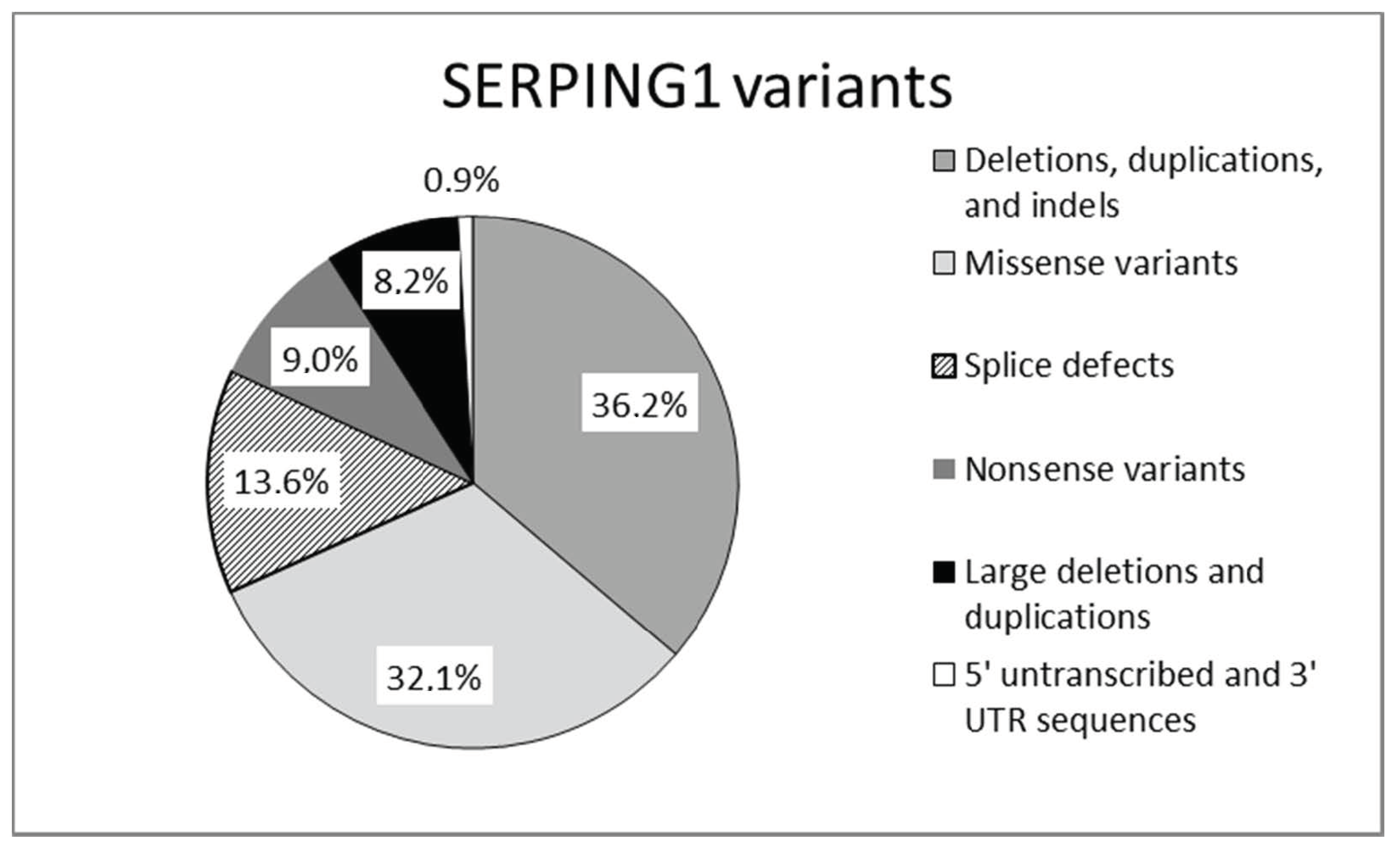
2.1. Genetics of C1-INH-HAE
2.2. The Genetics of nC1-INH-HAE
2.3. Disease-Modifying Factors
3. Diagnostic Applications of Genetic Biomarkers
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Cicardi, M.; Aberer, W.; Banerji, A.; Bas, M.; Bernstein, J.A.; Bork, K.; Caballero, T.; Farkas, H.; Grumach, A.; Kaplan, A.P.; et al. Classification, diagnosis, and approach to treatment for angioedema: Consensus report from the Hereditary Angioedema International Working Group. Allergy 2014, 69, 602–616. [Google Scholar] [CrossRef]
- Zuraw, B.L. Hereditary Angioedema. N. Engl. J. Med. 2008, 359, 1027–1036. [Google Scholar] [CrossRef] [PubMed]
- Bork, K.; Meng, G.; Staubach, P.; Hardt, J. Treatment with C1 inhibitor concentrate in abdominal pain attacks of patients with hereditary angioedema. Transfusion 2005, 45, 1774–1784. [Google Scholar] [CrossRef]
- Madsen, F.; Attermann, J.; Linneberg, A. Epidemiology of Non-hereditary Angioedema. Acta Derm. Venereol. 2012, 92, 475–479. [Google Scholar] [CrossRef]
- Kaplan, A.P. Angioedema. World Allergy Organ. J. 2008, 1, 103–113. [Google Scholar] [CrossRef] [PubMed]
- Ghazi, A.; Grant, J.A. Hereditary angioedema: Epidemiology, management, and role of icatibant. Biologics 2013, 7, 103–113. [Google Scholar] [CrossRef] [PubMed]
- Nzeako, U.C.; Frigas, E.; Tremaine, W.J. Hereditary angioedema: A broad review for clinicians. Arch. Intern. Med. 2001, 161, 2417–2429. [Google Scholar] [CrossRef] [PubMed]
- Jalaj, S.; Scolapio, J.S. Gastrointestinal manifestations, diagnosis, and management of hereditary angioedema. J. Clin. Gastro-enterol. 2013, 47, 817–823. [Google Scholar] [CrossRef]
- Xu, Y.-Y.; Zhi, Y.-X.; Liu, R.-L.; Craig, T.; Zhang, H.-Y. Upper airway edema in 43 patients with hereditary angioedema. Ann. Allergy Asthma Immunol. 2014, 112, 539–544.e1. [Google Scholar] [CrossRef]
- Zanichelli, A.; Longhurst, H.J.; Maurer, M.; Bouillet, L.; Aberer, W.; Fabien, V.; Andresen, I.; Caballero, T.; Aberer, W.; Grumach, A.; et al. Misdiagnosis trends in patients with hereditary angioedema from the real-world clinical setting. Ann Allergy Asthma Immunol. 2016, 117, 394–398. [Google Scholar] [CrossRef]
- Donaldson, V.H.; Evans, R.R. A biochemical abnormality in hereditary angioneurotic edema: Absence of serum inhibitor of C’1-esterase. Am. J. Med. 1963, 35, 37–44. [Google Scholar] [CrossRef]
- Kaplan, A.P.; Joseph, K. Pathogenic mechanisms of bradykinin mediated diseases: Dysregulation of an innate inflammatory pathway. Adv. Immunol. 2014, 121, 41–89. [Google Scholar] [PubMed]
- Maurer, M.; Bader, M.; Bas, M.; Bossi, F.; Cicardi, M.; Cugno, M.; Howarth, P.; Kaplan, A.; Kojda, G.; Leeb-Lundberg, F.; et al. New topics in bradykinin research. Allergy 2011, 66, 1397–1406. [Google Scholar] [CrossRef] [PubMed]
- Banday, A.Z.; Kaur, A.; Jindal, A.K.; Rawat, A. Singh S An update on the genetics and pathogenesis of hereditary angioedema. Genes Dis. 2020, 7, 75–83. [Google Scholar] [CrossRef] [PubMed]
- Zuraw, B.L. Hereditary angioedema with normal C1 inhibitor: Four types and counting. J. Allergy Clin. Immunol. 2018, 141, 884–885. [Google Scholar] [CrossRef] [PubMed]
- Caccia, S.; Suffritti, C.; Carzaniga, T.; Berardelli, R.; Berra, S.; Martorana, V.; Fra, A.; Drouet, C.; Cicardi, M. Intermittent C1-inhibitordeficiency associated with recessive inheritance: Functional and structural insight. Sci Rep. 2018, 8, 977. [Google Scholar] [CrossRef]
- Ponard, D.; Gaboriaud, C.; Charignon, D.; Ghannam, A.; Wagenaar-Bos, I.G.A.; Roem, D.; López-Lera, A.; López-Trascasa, M.; Tosi, M.; Drouet, C. SERPING1 mutation update: Mutation spectrum and C1 Inhibitor phenotypes. Human Mutat. 2020, 41, 38–57. [Google Scholar] [CrossRef]
- Stenson, P.D.; Mort, M.; Ball, E.V.; Shaw, K.; Phillips, A.D.; Cooper, D.N. The Human Gene Mutation Database: Building a comprehensive mutation repository for clinical and molecular genetics, diagnostic testing and personalized genomic medicine. Qual. Life Res. 2014, 133, 1–9. [Google Scholar] [CrossRef]
- Kalmár, L.; Hegedüs, T.; Farkas, H.; Nagy, M.; Tordai, A. HAEdb: A novel interactive, locus-specific mutation database for the C1 inhibitor gene. Human Mutat. 2005, 25, 1–5. [Google Scholar] [CrossRef]
- Bors, A.; Csuka, D.; Varga, L.; Farkas, H.; Tordai, A.; Fust, G.; Szilagyi, A. Less severe clinical manifestations in patients with hereditary angioedema with missense C1INH gene mutations. J. Allergy Clin. Immunol. 2003, 131, 1708–1711. [Google Scholar] [CrossRef] [PubMed]
- Khan, S.; Longhurst, H. Epigenetic alterations on C1-inhibitor expression may influence hereditary angioedema attack fre-quency and C4 levels. Clin. Exp. Immunol. 2020, 202, 144–145. [Google Scholar] [CrossRef] [PubMed]
- Vatsiou, S.; Zamanakou, M.; Loules, G.; Psarros, F.; Parsopoulou, F.; Csuka, D.; Valerieva, A.; Staevska, M.; Porebski, G.; Obtulowicz, K.; et al. A novel deep intronic SERPING1 variantas a cause of hereditary angioedema due to C1-inhibitor deficiency. Allergol Int. 2020, 69, 443–449. [Google Scholar] [CrossRef] [PubMed]
- Hujová, P.; Souček, P.; Grodecká, L.; Grombiříková, H.; Ravčuková, B.; Kuklínek, P.; Hakl, R.; Litzman, J.; Freiberger, T. Deep intronic mutation inSERPING1 caused hereditary angioedema through pseudoexon activation. J. Clin. Immunol 2020, 40, 435–446. [Google Scholar] [CrossRef] [PubMed]
- Loules, G.; Zamanakou, M.; Parsopoulou, F.; Vatsiou, S.; Psarros, F.; Csuka, D.; Porebski, G.; Obtulowicz, K.; Valerieva, A.; Staevska, M.; et al. Targeted next-generation sequencing for the molecular diagnosis of hereditary angioedema due to C1-inhibitor defi-ciency. Gene 2018, 667, 76–82. [Google Scholar] [CrossRef] [PubMed]
- Bork, K.; Barnstedt, S.E.; Koch, P.; Traupe, H. Hereditary angioedema with normal C1-inhibitor activity in women. Lancet 2000, 356, 213–217. [Google Scholar] [CrossRef]
- Binkley, K.E.; Davis, A. Clinical, biochemical, and genetic characterization of a novel estrogen-dependent inherited form of angioedema. J. Allergy Clin. Immunol. 2000, 106, 546–550. [Google Scholar] [CrossRef] [PubMed]
- Dewald, G.; Bork, K. Missense mutations in the coagulation factor XII (Hageman factor) gene in hereditary angioedema with normal C1 inhibitor. Biochem. Biophys. Res. Commun. 2006, 343, 1286–1289. [Google Scholar] [CrossRef] [PubMed]
- Bork, K.; Gül, D.; Hardt, J.; Dewald, G. Hereditary Angioedema with Normal C1 Inhibitor: Clinical Symptoms and Course. Am. J. Med. 2007, 120, 987–992. [Google Scholar] [CrossRef] [PubMed]
- Kiss, N.; Barabás, E.; Várnai, K.; Halász, A.; Varga, L.Á.; Prohászka, Z.; Farkas, H.; Szilágyi, Á. Novel duplication in the F12 gene in a patient with recurrent angioedema. Clin. Immunol. 2013, 149, 142–145. [Google Scholar] [CrossRef]
- Bork, K.; Wulff, K.; Hardt, J.; Witzke, G.; Lohse, P. Characterization of a partial exon 9/intron 9 deletion in the coagulation factor XII gene (F12) detected in two Turkish families with hereditary angioedema and normal C1 inhibitor. Haemophilia 2014, 20. [Google Scholar] [CrossRef]
- Bork, K.; Wulff, K.; Meinke, P.; Wagner, N.; Hardt, J.; Witzke, G. A novel mutation in the coagulation factor 12 gene in subjects with hereditary angioedema and normal C1-inhibitor. Clin. Immunol. 2011, 141, 31–35. [Google Scholar] [CrossRef] [PubMed]
- De Maat, S.; Björkqvist, J.; Suffritti, C.; Wiesenekker, C.P.; Nagtegaal, W.; Koekman, A.; van Dooremalen, S.; Pasterkamp, G.; de Groot, P.G.; Cicardi, M.; et al. Plasmin is a natural trigger for bradykinin production in patients with hereditary angioedema with factor XII mutations. J. Allergy Clin. Immunol. 2016, 138, 1414–1423.e9. [Google Scholar] [CrossRef]
- Bork, K.; Wulff, K.; Steinmüller-Magin, L.; Braenne, I.; Staubach-Renz, P.; Witzke, G.; Hardt, J. Hereditary angioedema with a mutation in the plasminogen gene. Allergy 2018, 73, 442–450. [Google Scholar] [CrossRef] [PubMed]
- Dewald, G. A missense mutation in the plasminogen gene, within the plasminogen kringle 3 domain, in hereditary angi-oedema with normal C1 inhibitor. Biochem Biophys Res. Commun. 2018, 498, 193–198. [Google Scholar] [CrossRef] [PubMed]
- Bafunno, V.; Firinu, D.; D’Apolito, M.; Cordisco, G.; Loffredo, S.; Leccese, A.; Bova, M.; Barca, M.P.; Santacroce, R.; Cicardi, M.; et al. Mutation of the angiopoietin-1 gene (ANGPT1) associates with a new type of hereditary angioedema. J. Allergy Clin. Immunol. 2018, 141, 1009–1017. [Google Scholar] [CrossRef] [PubMed]
- Bork, K.; Wulff, K.; Rossmann, H.; Steinmüller-Magin, L.; Braenne, I.; Witzke, G.; Hardt, J. Hereditary angioedema cosegre-gating with a novel kininogen 1 gene mutation changing the N-terminal cleavage site of bradykinin. Allergy 2019, 74, 2479–2481. [Google Scholar] [CrossRef]
- Loules, G.; Parsopoulou, F.; Zamanakou, M.; Csuka, D.; Bova, M.; González-Quevedo, T.; Psarros, F.; Porebski, G.; Speletas, M.; Firinu, D.; et al. Deciphering the Genetics of Primary Angioedema with Normal Levels of C1 Inhibitor. J. Clin. Med. 2020, 9, 3402. [Google Scholar] [CrossRef]
- Ariano, A.; D’Apolito, M.; Bova, M.; Bellanti, F.; Loffredo, S.; D’Andrea, G.; Intrieri, M.; Petraroli, A.; Maffione, A.B.; Spadaro, G.; et al. A myoferlin gain-of-function variant associates with a new type of hereditary angioedema. Allergy 2020, 75, 2989–2992. [Google Scholar] [CrossRef]
- Bork, K.; Wulff, K.; Möhl, B.S.; Steinmüller-Magin, L.; Witzke, G.; Hardt, J.; Meinke, P. Novel hereditary angioedema linked with a heparan sulfate 3-O-sulfotransferase 6 gene mutation. J. Allergy Clin. Immunol. 2021, 0091. [Google Scholar] [CrossRef]
- Recke, A.; Massalme, E.G.; Jappe, U.; Steinmüller-Magin, L.; Schmidt, J.; Hellenbroich, Y.; Hüning, I.; Gillessen-Kaesbach, G.; Zillikens, D.; Hartmann, K. Identification of the recently described plasminogen gene mutation p.Lys330Glu in a family from Northern Germany with hereditary angioedema. Clin. Transl. Allergy 2019, 9, 1–4. [Google Scholar] [CrossRef]
- D’Apolito, M.; Santacroce, R.; Colia, A.L.; Cordisco, G.; Maffione, A.B.; Margaglione, M. Angiopoietin-1 haploinsufficiency affects the endothelial barrier and causes hereditary angioedema. Clin. Exp. Allergy 2019, 49, 626–635. [Google Scholar] [CrossRef]
- Cagini, N.; Veronez, C.L.; Azevedo, B.F.; Constantino-Silva, R.N.; Martin, R.P.; Da Silva, J.; Grumach, A.S.; Pesquero, J.B. In silico Analysis Of Alterations In ANGPT1 Gene Supports A New Pathway Responsible To Mediate Hereditary Angioedema In Brazilian Patients with No Mutations in SERPING1 and F12 Genes. J. Allergy Clin. Immunol. 2018, 141, AB46. [Google Scholar] [CrossRef]
- Veronez, C.L.; Aabom, A.; Martin, R.P.; Filippelli-Silva, R.; Gonçalves, R.F.; Nicolicht, P.; Mendes, A.R.; Da Silva, J.; Guilarte, M.; Grumach, A.S.; et al. Genetic Variation of Kallikrein-Kinin System and Related Genes in Patients with Hereditary Angioedema. Front. Med. 2019, 6, 6–28. [Google Scholar] [CrossRef] [PubMed]
- Speletas, M.; Szilágyi, Á.; Csuka, D.; Koutsostathis, N.; Psarros, F.; Moldovan, D.; Magerl, M.; Kompoti, M.; Varga, L.; Maurer, M.; et al. F12-46C/T polymorphism as modifier of the clinical phenotype of hereditary angioedema. Allergy 2015, 70, 1661–1664. [Google Scholar] [CrossRef] [PubMed]
- Grivčeva-Panovska, V.; Košnik, M.; Korošec, P.; Andrejević, S.; Karadža-Lapić, L.; Rijavec, M. Hereditary angioedema due to C1-inhibitor deficiency in Macedonia: Clinical characteristics, novel SERPING1 mutations and genetic factors modifying the clinical phenotype. Ann. Med. 2018, 50, 269–276. [Google Scholar] [CrossRef] [PubMed]
- Corvillo, F.; de la Morena Barrio, M.E.; Marcos Bravo, C.; López-Trascasa, M.; Vicente, V.; Emsley, J.; Caballero, T.; Corral, J.; Lopez Lera, A. The FXII c.-4T>C Polymorphism as a Disease Modifier in Patients with Hereditary Angioedema Due to the FXII p.Thr328Lys Variant. Front. Genet. 2020, 11, 1033. [Google Scholar] [CrossRef]
- Gianni, P.; Loules, G.; Zamanakou, M.; Kompoti, M.; Csuka, D.; Psarros, F.; Magerl, M.; Moldovan, D.; Maurer, M.; Speletas, M.G.; et al. Genetic Determinants of C1 Inhibitor Deficiency Angioedema Age of Onset. Int. Arch. Allergy Immunol. 2017, 174, 200–204. [Google Scholar] [CrossRef]
- Germenis, A.E.; Margaglione, M.; Pesquero, J.B.; Farkas, H.; Cichon, S.; Csuka, D.; López Lera, A.; Rijavec, M.; Jolles, S.; Szilagyi, A.; et al. International consensus on the use of genetics in the management of hereditary angioedema. J. Allergy Clin. Immunol Pract. 2020, 8, 901–911. [Google Scholar] [CrossRef]
| F12 | T328K T328R c.971_1018 + 24del72 c.892_909dup |
| PLG | K330E |
| ANGPT1 | A119S A8V Q370H |
| KNG1 | M379K |
| MYOF | R217S |
| HS3ST6 | T144S |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Santacroce, R.; D'Andrea, G.; Maffione, A.B.; Margaglione, M.; d'Apolito, M. The Genetics of Hereditary Angioedema: A Review. J. Clin. Med. 2021, 10, 2023. https://doi.org/10.3390/jcm10092023
Santacroce R, D'Andrea G, Maffione AB, Margaglione M, d'Apolito M. The Genetics of Hereditary Angioedema: A Review. Journal of Clinical Medicine. 2021; 10(9):2023. https://doi.org/10.3390/jcm10092023
Chicago/Turabian StyleSantacroce, Rosa, Giovanna D'Andrea, Angela Bruna Maffione, Maurizio Margaglione, and Maria d'Apolito. 2021. "The Genetics of Hereditary Angioedema: A Review" Journal of Clinical Medicine 10, no. 9: 2023. https://doi.org/10.3390/jcm10092023
APA StyleSantacroce, R., D'Andrea, G., Maffione, A. B., Margaglione, M., & d'Apolito, M. (2021). The Genetics of Hereditary Angioedema: A Review. Journal of Clinical Medicine, 10(9), 2023. https://doi.org/10.3390/jcm10092023






