Implications of Endogenous Retroelements in the Etiopathogenesis of Systemic Lupus Erythematosus
Abstract
1. Introduction
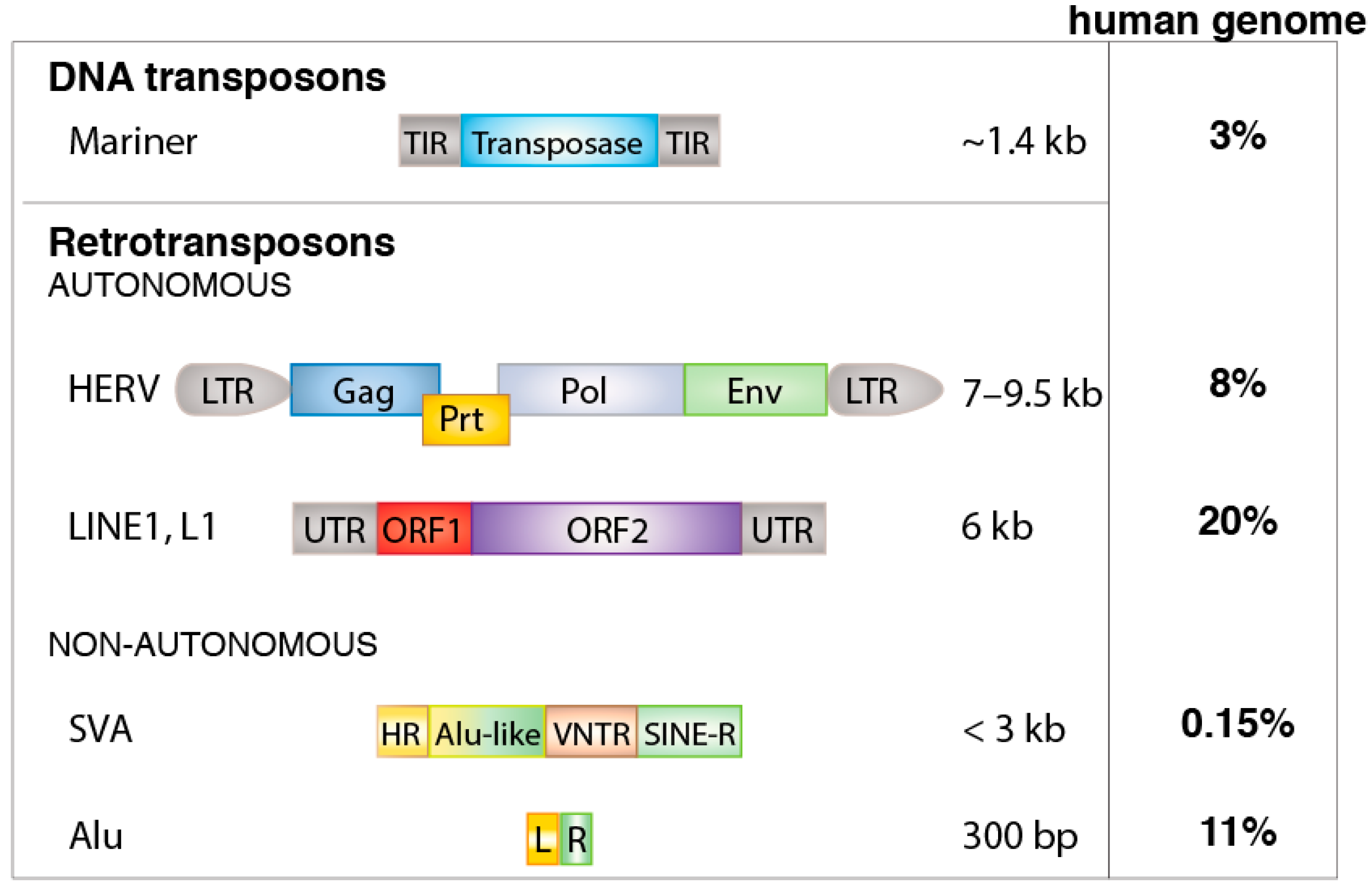
2. Transposable Elements in the Human Genome
2.1. HERVs
2.2. L1 Retrotransposons
2.3. Non-Autonomous Retroelements
3. How L1 Retrotransposons May Trigger IFN-Positive SLE
3.1. HERVs and Other Non-L1 Retrotransposons in SLE
3.2. Are Defenses Against L1 and HERVs Defective in SLE?
3.3. Subsets of SLE with Distinct Mechanisms
3.4. L1- and HERV-Related Biomarkers
3.5. Novel Therapeutic Opportunities Related to L1
4. Concluding Remarks
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Wang, L.; Wang, F.S.; Gershwin, M.E. Human autoimmune diseases: A comprehensive update. J. Intern. Med. 2015, 278, 369–395. [Google Scholar] [CrossRef] [PubMed]
- Lockshin, M.D.; Barbhaiya, M.; Izmirly, P.; Buyon, J.P.; Crow, M.K. SLE: Reconciling heterogeneity. Lupus Sci. Med. 2019, 6, e000280. [Google Scholar] [CrossRef]
- Rullo, O.J.; Tsao, B.P. Recent insights into the genetic basis of systemic lupus erythematosus. Ann. Rheum. Dis. 2013, 72 (Suppl. 2), ii56–ii61. [Google Scholar] [CrossRef]
- Hom, G.; Graham, R.R.; Modrek, B.; Taylor, K.E.; Ortmann, W.; Garnier, S.; Lee, A.T.; Chung, S.A.; Ferreira, R.C.; Pant, P.V.; et al. Association of systemic lupus erythematosus with C8orf13-BLK and ITGAM-ITGAX. N. Engl. J. Med. 2008, 358, 900–909. [Google Scholar] [CrossRef]
- Graham, R.R.; Hom, G.; Ortmann, W.; Behrens, T.W. Review of recent genome-wide association scans in lupus. J. Intern. Med. 2009, 265, 680–688. [Google Scholar] [CrossRef]
- Nath, S.K.; Han, S.; Kim-Howard, X.; Kelly, J.A.; Viswanathan, P.; Gilkeson, G.S.; Chen, W.; Zhu, C.; McEver, R.P.; Kimberly, R.P.; et al. A nonsynonymous functional variant in integrin-alpha(M) (encoded by ITGAM) is associated with systemic lupus erythematosus. Nat. Genet. 2008, 40, 152–154. [Google Scholar] [CrossRef] [PubMed]
- Boackle, S.A. Advances in lupus genetics. Curr. Opin. Rheumatol. 2013, 25, 561–568. [Google Scholar] [CrossRef] [PubMed]
- Barcellos, L.F.; May, S.L.; Ramsay, P.P.; Quach, H.L.; Lane, J.A.; Nititham, J.; Noble, J.A.; Taylor, K.E.; Quach, D.L.; Chung, S.A.; et al. High-density SNP screening of the major histocompatibility complex in systemic lupus erythematosus demonstrates strong evidence for independent susceptibility regions. PLoS Genet. 2009, 5, e1000696. [Google Scholar] [CrossRef]
- International MHC, and Autoimmunity Genetics Network; Rioux, J.D.; Goyette, P.; Vyse, T.J.; Hammarstrom, L.; Fernando, M.M.; Green, T.; De Jager, P.L.; Foisy, S.; Wang, J.; et al. Mapping of multiple susceptibility variants within the MHC region for 7 immune-mediated diseases. Proc. Natl. Acad. Sci. USA 2009, 106, 18680–18685. [Google Scholar] [CrossRef] [PubMed]
- Graham, R.R.; Ortmann, W.; Rodine, P.; Espe, K.; Langefeld, C.; Lange, E.; Williams, A.; Beck, S.; Kyogoku, C.; Moser, K.; et al. Specific combinations of HLA-DR2 and DR3 class II haplotypes contribute graded risk for disease susceptibility and autoantibodies in human SLE. Eur. J. Hum. Genet. 2007, 15, 823–830. [Google Scholar] [CrossRef]
- Bowness, P.; Davies, K.A.; Norsworthy, P.J.; Athanassiou, P.; Taylor-Weideman, J.; Borysiewicz, L.K.; Meyer, P.A.R.; Walport, M.J. Hereditary C1q deficiency and systemic lupus erythematosus. Quart. J. Med. 1994, 87, 455–464. [Google Scholar]
- Truedsson, L.; Bengtsson, A.A.; Sturfelt, G. Complement deficiencies and systemic lupus erythematosus. Autoimmunity 2007, 40, 560–566. [Google Scholar] [CrossRef] [PubMed]
- Namjou, B.; Kothari, P.H.; Kelly, J.A.; Glenn, S.B.; Ojwang, J.O.; Adler, A.; Alarcon-Riquelme, M.E.; Gallant, C.J.; Boackle, S.A.; Criswell, L.A.; et al. Evaluation of the TREX1 gene in a large multi-ancestral lupus cohort. Genes Immun. 2011, 12, 270–279. [Google Scholar] [CrossRef]
- Gaipl, U.S.; Beyer, T.D.; Heyder, P.; Kuenkele, S.; Bottcher, A.; Voll, R.E.; Kalden, J.R.; Herrmann, M. Cooperation between C1q and DNase I in the clearance of necrotic cell-derived chromatin. Arthritis Rheum. 2004, 50, 640–649. [Google Scholar] [CrossRef] [PubMed]
- Stetson, D.B.; Ko, J.S.; Heidmann, T.; Medzhitov, R. Trex1 prevents cell-intrinsic initiation of autoimmunity. Cell 2008, 134, 587–598. [Google Scholar] [CrossRef] [PubMed]
- Al-Mayouf, S.M.; Sunker, A.; Abdwani, R.; Abrawi, S.A.; Almurshedi, F.; Alhashmi, N.; Al Sonbul, A.; Sewairi, W.; Qari, A.; Abdallah, E.; et al. Loss-of-function variant in DNASE1L3 causes a familial form of systemic lupus erythematosus. Nat. Genet. 2011, 43, 1186–1188. [Google Scholar] [CrossRef] [PubMed]
- Bronson, P.G.; Chaivorapol, C.; Ortmann, W.; Behrens, T.W.; Graham, R.R. The genetics of type I interferon in systemic lupus erythematosus. Curr. Opin. Immunol. 2012, 24, 530–537. [Google Scholar] [CrossRef]
- Kariuki, S.N.; Kirou, K.A.; MacDermott, E.J.; Barillas-Arias, L.; Crow, M.K.; Niewold, T.B. Cutting edge: Autoimmune disease risk variant of STAT4 confers increased sensitivity to IFN-alpha in lupus patients in vivo. J. Immunol. 2009, 182, 34–38. [Google Scholar] [CrossRef]
- Banchereau, R.; Hong, S.; Cantarel, B.; Baldwin, N.; Baisch, J.; Edens, M.; Cepika, A.M.; Acs, P.; Turner, J.; Anguiano, E.; et al. Personalized Immunomonitoring Uncovers Molecular Networks that Stratify Lupus Patients. Cell 2016, 165, 551–565. [Google Scholar] [CrossRef]
- Niewold, T.B.; Kelly, J.A.; Flesch, M.H.; Espinoza, L.R.; Harley, J.B.; Crow, M.K. Association of the IRF5 risk haplotype with high serum interferon-alpha activity in systemic lupus erythematosus patients. Arthritis Rheum. 2008, 58, 2481–2487. [Google Scholar] [CrossRef]
- Kariuki, S.N.; Crow, M.K.; Niewold, T.B. The PTPN22 C1858T polymorphism is associated with skewing of cytokine profiles toward high interferon-alpha activity and low tumor necrosis factor alpha levels in patients with lupus. Arthritis Rheum. 2008, 58, 2818–2823. [Google Scholar] [CrossRef] [PubMed]
- Gorman, J.A.; Hundhausen, C.; Errett, J.S.; Stone, A.E.; Allenspach, E.J.; Ge, Y.; Arkatkar, T.; Clough, C.; Dai, X.; Khim, S.; et al. The A946T variant of the RNA sensor IFIH1 mediates an interferon program that limits viral infection but increases the risk for autoimmunity. Nat. Immunol. 2017, 18, 744–752. [Google Scholar] [CrossRef] [PubMed]
- Tian, J.; Ma, Y.; Li, J.; Cen, H.; Wang, D.G.; Feng, C.C.; Li, R.J.; Leng, R.X.; Pan, H.F.; Ye, D.Q. The TLR7 7926A>G polymorphism is associated with susceptibility to systemic lupus erythematosus. Mol. Med. Rep. 2012, 6, 105–110. [Google Scholar] [CrossRef]
- Prokunina, L.; Castillejo-Lopez, C.; Oberg, F.; Gunnarsson, I.; Berg, L.; Magnusson, V.; Brookes, A.J.; Tentler, D.; Kristjansdottir, H.; Grondal, G.; et al. A regulatory polymorphism in PDCD1 is associated with susceptibility to systemic lupus erythematosus in humans. Nat. Genet. 2002, 32, 666–669. [Google Scholar] [CrossRef] [PubMed]
- Kozyrev, S.V.; Abelson, A.K.; Wojcik, J.; Zaghlool, A.; Linga Reddy, M.V.; Sanchez, E.; Gunnarsson, I.; Svenungsson, E.; Sturfelt, G.; Jonsen, A.; et al. Functional variants in the B-cell gene BANK1 are associated with systemic lupus erythematosus. Nat. Genet. 2008, 40, 211–216. [Google Scholar] [CrossRef] [PubMed]
- Namjou, B.; Kim-Howard, X.; Sun, C.; Adler, A.; Chung, S.A.; Kaufman, K.M.; Kelly, J.A.; Glenn, S.B.; Guthridge, J.M.; Scofield, R.H.; et al. PTPN22 association in systemic lupus erythematosus (SLE) with respect to individual ancestry and clinical sub-phenotypes. PLoS ONE 2013, 8, e69404. [Google Scholar] [CrossRef]
- Curran, C.S.; Gupta, S.; Sanz, I.; Sharon, E. PD-1 immunobiology in systemic lupus erythematosus. J. Autoimmun. 2019, 97, 1–9. [Google Scholar] [CrossRef]
- Jiang, S.H.; Athanasopoulos, V.; Ellyard, J.I.; Chuah, A.; Cappello, J.; Cook, A.; Prabhu, S.B.; Cardenas, J.; Gu, J.; Stanley, M.; et al. Functional rare and low frequency variants in BLK and BANK1 contribute to human lupus. Nat. Commun. 2019, 10, 2201. [Google Scholar] [CrossRef]
- Jacquemin, C.; Schmitt, N.; Contin-Bordes, C.; Liu, Y.; Narayanan, P.; Seneschal, J.; Maurouard, T.; Dougall, D.; Davizon, E.S.; Dumortier, H.; et al. OX40 Ligand Contributes to Human Lupus Pathogenesis by Promoting T Follicular Helper Response. Immunity 2015, 42, 1159–1170. [Google Scholar] [CrossRef]
- Orru, V.; Tsai, S.J.; Rueda, B.; Fiorillo, E.; Stanford, S.M.; Dasgupta, J.; Hartiala, J.; Zhao, L.; Ortego-Centeno, N.; D’Alfonso, S.; et al. A loss-of-function variant of PTPN22 is associated with reduced risk of systemic lupus erythematosus. Hum. Mol. Genet. 2009, 18, 569–579. [Google Scholar] [CrossRef]
- Hawn, T.R.; Wu, H.; Grossman, J.M.; Hahn, B.H.; Tsao, B.P.; Aderem, A. A stop codon polymorphism of Toll-like receptor 5 is associated with resistance to systemic lupus erythematosus. Proc. Natl. Acad. Sci. USA 2005, 102, 10593–10597. [Google Scholar] [CrossRef]
- Altorok, N.; Sawalha, A.H. Epigenetics in the pathogenesis of systemic lupus erythematosus. Curr. Opin. Rheumatol. 2013, 25, 569–576. [Google Scholar] [CrossRef] [PubMed]
- Costenbader, K.H.; Gay, S.; Alarcon-Riquelme, M.E.; Iaccarino, L.; Doria, A. Genes, epigenetic regulation and environmental factors: Which is the most relevant in developing autoimmune diseases? Autoimmun. Rev. 2012, 11, 604–609. [Google Scholar] [CrossRef] [PubMed]
- Yeung, K.S.; Chung, B.H.; Choufani, S.; Mok, M.Y.; Wong, W.L.; Mak, C.C.; Yang, W.; Lee, P.P.; Wong, W.H.; Chen, Y.A.; et al. Genome-Wide DNA Methylation Analysis of Chinese Patients with Systemic Lupus Erythematosus Identified Hypomethylation in Genes Related to the Type I Interferon Pathway. PLoS ONE 2017, 12, e0169553. [Google Scholar] [CrossRef] [PubMed]
- Patel, D.R.; Richardson, B.C. Dissecting complex epigenetic alterations in human lupus. Arthritis Res. Ther. 2013, 15, 201. [Google Scholar] [CrossRef]
- Shen, N.; Liang, D.; Tang, Y.; de Vries, N.; Tak, P.P. MicroRNAs--novel regulators of systemic lupus erythematosus pathogenesis. Nat. Rev. Rheumatol. 2012, 8, 701–709. [Google Scholar] [CrossRef] [PubMed]
- Stetson, D.B. Endogenous retroelements and autoimmune disease. Curr. Opin. Immunol. 2012, 24, 692–697. [Google Scholar] [CrossRef]
- Crow, M.K. Long interspersed nuclear elements (LINE-1): Potential triggers of systemic autoimmune disease. Autoimmunity 2010, 43, 7–16. [Google Scholar] [CrossRef]
- Tugnet, N.; Rylance, P.; Roden, D.; Trela, M.; Nelson, P. Human Endogenous Retroviruses (HERVs) and Autoimmune Rheumatic Disease: Is There a Link? Open Rheumatol. J. 2013, 7, 13–21. [Google Scholar] [CrossRef]
- Mustelin, T.; Ukadike, K.C. How Retroviruses and Retrotransposons in Our Genome May Contribute to Autoimmunity in Rheumatological Conditions. Front. Immunol. 2020, 11, 593891. [Google Scholar] [CrossRef]
- Belshaw, R.; Pereira, V.; Katzourakis, A.; Talbot, G.; Paces, J.; Burt, A.; Tristem, M. Long-term reinfection of the human genome by endogenous retroviruses. Proc. Natl. Acad. Sci. USA 2004, 101, 4894–4899. [Google Scholar] [CrossRef]
- Nelson, P.N.; Hooley, P.; Roden, D.; Davari Ejtehadi, H.; Rylance, P.; Warren, P.; Martin, J.; Murray, P.G.; Molecular Immunology Research, G. Human endogenous retroviruses: Transposable elements with potential? Clin. Exp. Immunol. 2004, 138, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Kano, H.; Godoy, I.; Courtney, C.; Vetter, M.R.; Gerton, G.L.; Ostertag, E.M.; Kazazian, H.H., Jr. L1 retrotransposition occurs mainly in embryogenesis and creates somatic mosaicism. Genes Dev. 2009, 23, 1303–1312. [Google Scholar] [CrossRef]
- Goodier, J.L.; Kazazian, H.H., Jr. Retrotransposons revisited: The restraint and rehabilitation of parasites. Cell 2008, 135, 23–35. [Google Scholar] [CrossRef]
- Bourque, G.; Burns, K.H.; Gehring, M.; Gorbunova, V.; Seluanov, A.; Hammell, M.; Imbeault, M.; Izsvak, Z.; Levin, H.L.; Macfarlan, T.S.; et al. Ten things you should know about transposable elements. Genome Biol. 2018, 19, 199. [Google Scholar] [CrossRef] [PubMed]
- Wicker, T.; Sabot, F.; Hua-Van, A.; Bennetzen, J.L.; Capy, P.; Chalhoub, B.; Flavell, A.; Leroy, P.; Morgante, M.; Panaud, O.; et al. A unified classification system for eukaryotic transposable elements. Nat. Rev. Genet. 2007, 8, 973–982. [Google Scholar] [CrossRef] [PubMed]
- Lander, E.S.; Linton, L.M.; Birren, B.; Nusbaum, C.; Zody, M.C.; Baldwin, J.; Devon, K.; Dewar, K.; Doyle, M.; FitzHugh, W.; et al. Initial sequencing and analysis of the human genome. Nature 2001, 409, 860–921. [Google Scholar] [CrossRef] [PubMed]
- Mandal, P.K.; Kazazian, H.H., Jr. SnapShot: Vertebrate transposons. Cell 2008, 135, 192–192.e1. [Google Scholar] [CrossRef] [PubMed]
- Deininger, P. Alu elements: Know the SINEs. Genome Biol. 2011, 12, 236. [Google Scholar] [CrossRef]
- Hancks, D.C.; Kazazian, H.H., Jr. Roles for retrotransposon insertions in human disease. Mob. DNA 2016, 7, 9. [Google Scholar] [CrossRef]
- Karamitros, T.; Hurst, T.; Marchi, E.; Karamichali, E.; Georgopoulou, U.; Mentis, A.; Riepsaame, J.; Lin, A.; Paraskevis, D.; Hatzakis, A.; et al. Human Endogenous Retrovirus-K HML-2 integration within RASGRF2 is associated with intravenous drug abuse and modulates transcription in a cell-line model. Proc. Natl. Acad. Sci. USA 2018, 115, 10434–10439. [Google Scholar] [CrossRef]
- Batista, R.L.; Yamaguchi, K.; Rodrigues, A.D.S.; Nishi, M.Y.; Goodier, J.L.; Carvalho, L.R.; Domenice, S.; Costa, E.M.F.; Kazazian, H.H.; Mendonca, B.B. Mobile DNA in Endocrinology: LINE-1 Retrotransposon Causing Partial Androgen Insensitivity Syndrome. J. Clin. Endocrinol. Metab. 2019, 104, 6385–6390. [Google Scholar] [CrossRef]
- Ardeljan, D.; Taylor, M.S.; Ting, D.T.; Burns, K.H. The Human Long Interspersed Element-1 Retrotransposon: An Emerging Biomarker of Neoplasia. Clin. Chem. 2017, 63, 816–822. [Google Scholar] [CrossRef]
- Xia, Z.; Cochrane, D.R.; Anglesio, M.S.; Wang, Y.K.; Nazeran, T.; Tessier-Cloutier, B.; McConechy, M.K.; Senz, J.; Lum, A.; Bashashati, A.; et al. LINE-1 retrotransposon-mediated DNA transductions in endometriosis associated ovarian cancers. Gynecol. Oncol. 2017, 147, 642–647. [Google Scholar] [CrossRef]
- Qian, Y.; Mancini-DiNardo, D.; Judkins, T.; Cox, H.C.; Brown, K.; Elias, M.; Singh, N.; Daniels, C.; Holladay, J.; Coffee, B.; et al. Identification of pathogenic retrotransposon insertions in cancer predisposition genes. Cancer Genet. 2017, 216–217, 159–169. [Google Scholar] [CrossRef] [PubMed]
- Scott, E.C.; Gardner, E.J.; Masood, A.; Chuang, N.T.; Vertino, P.M.; Devine, S.E. A hot L1 retrotransposon evades somatic repression and initiates human colorectal cancer. Genome Res. 2016, 26, 745–755. [Google Scholar] [CrossRef]
- Xue, B.; Sechi, L.A.; Kelvin, D.J. Human Endogenous Retrovirus K (HML-2) in Health and Disease. Front. Microbiol. 2020, 11, 1690. [Google Scholar] [CrossRef] [PubMed]
- Kemp, J.R.; Longworth, M.S. Crossing the LINE Toward Genomic Instability: LINE-1 Retrotransposition in Cancer. Front. Chem. 2015, 3, 68. [Google Scholar] [CrossRef]
- Rodić, N.; Sharma, R.; Sharma, R.; Zampella, J.; Dai, L.; Taylor, M.S.; Hruban, R.H.; Iacobuzio-Donahue, C.A.; Maitra, A.; Torbenson, M.S.; et al. Long Interspersed Element-1 Protein Expression Is a Hallmark of Many Human Cancers. Am. J. Pathol. 2014, 184, 1280–1286. [Google Scholar] [CrossRef] [PubMed]
- Ma, K.; Qiu, L.; Mrasek, K.; Zhang, J.; Liehr, T.; Quintana, L.G.; Li, Z. Common fragile sites: Genomic hotspots of DNA damage and carcinogenesis. Int. J. Mol. Sci. 2012, 13, 11974–11999. [Google Scholar] [CrossRef]
- Fungtammasan, A.; Walsh, E.; Chiaromonte, F.; Eckert, K.A.; Makova, K.D. A genome-wide analysis of common fragile sites: What features determine chromosomal instability in the human genome? Genome Res. 2012, 22, 993–1005. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, R.P.; Wildschutte, J.H.; Russo, C.; Coffin, J.M. Identification, characterization, and comparative genomic distribution of the HERV-K (HML-2) group of human endogenous retroviruses. Retrovirology 2011, 8, 90. [Google Scholar] [CrossRef]
- Dewannieux, M.; Harper, F.; Richaud, A.; Letzelter, C.; Ribet, D.; Pierron, G.; Heidmann, T. Identification of an infectious progenitor for the multiple-copy HERV-K human endogenous retroelements. Genome Res. 2006, 16, 1548–1556. [Google Scholar] [CrossRef]
- Hohn, O.; Hanke, K.; Bannert, N. HERV-K(HML-2), the Best Preserved Family of HERVs: Endogenization, Expression, and Implications in Health and Disease. Front. Oncol. 2013, 3, 246. [Google Scholar] [CrossRef]
- Garcia-Montojo, M.; Doucet-O’Hare, T.; Henderson, L.; Nath, A. Human endogenous retrovirus-K (HML-2): A comprehensive review. Crit. Rev. Microbiol. 2018, 44, 715–738. [Google Scholar] [CrossRef]
- Jha, A.R.; Nixon, D.F.; Rosenberg, M.G.; Martin, J.N.; Deeks, S.G.; Hudson, R.R.; Garrison, K.E.; Pillai, S.K. Human endogenous retrovirus K106 (HERV-K106) was infectious after the emergence of anatomically modern humans. PLoS ONE 2011, 6, e20234. [Google Scholar] [CrossRef]
- Ehlhardt, S.; Seifert, M.; Schneider, J.; Ojak, A.; Zang, K.D.; Mehraein, Y. Human endogenous retrovirus HERV-K(HML-2) Rec expression and transcriptional activities in normal and rheumatoid arthritis synovia. J. Rheumatol. 2006, 33, 16–23. [Google Scholar]
- Boller, K.; Schonfeld, K.; Lischer, S.; Fischer, N.; Hoffmann, A.; Kurth, R.; Tonjes, R.R. Human endogenous retrovirus HERV-K113 is capable of producing intact viral particles. J. Gen. Virol. 2008, 89, 567–572. [Google Scholar] [CrossRef] [PubMed]
- Nelson, P.N.; Carnegie, P.R.; Martin, J.; Davari Ejtehadi, H.; Hooley, P.; Roden, D.; Rowland-Jones, S.; Warren, P.; Astley, J.; Murray, P.G. Demystified. Human endogenous retroviruses. Mol. Pathol. 2003, 56, 11–18. [Google Scholar] [CrossRef] [PubMed]
- Perl, A.; Fernandez, D.; Telarico, T.; Phillips, P.E. Endogenous retroviral pathogenesis in lupus. Curr. Opin. Rheumatol. 2010, 22, 483–492. [Google Scholar] [CrossRef] [PubMed]
- Mameli, G.; Erre, G.L.; Caggiu, E.; Mura, S.; Cossu, D.; Bo, M.; Cadoni, M.L.; Piras, A.; Mundula, N.; Colombo, E.; et al. Identification of a HERV-K env surface peptide highly recognized in Rheumatoid Arthritis (RA) patients: A cross-sectional case-control study. Clin. Exp. Immunol. 2017, 189, 127–131. [Google Scholar] [CrossRef]
- Talotta, R.; Atzeni, F.; Laska, M.J. The contribution of HERV-E clone 4-1 and other HERV-E members to the pathogenesis of rheumatic autoimmune diseases. APMIS 2020, 128, 367–377. [Google Scholar] [CrossRef] [PubMed]
- Karimi, A.; Esmaili, N.; Ranjkesh, M.; Zolfaghari, M.A. Expression of human endogenous retroviruses in pemphigus vulgaris patients. Mol. Biol. Rep. 2019, 46, 6181–6186. [Google Scholar] [CrossRef]
- Bengtsson, A.; Blomberg, J.; Nived, O.; Pipkorn, R.; Toth, L.; Sturfelt, G. Selective antibody reactivity with peptides from human endogenous retroviruses and nonviral poly(amino acids) in patients with systemic lupus erythematosus. Arthritis Rheum. 1996, 39, 1654–1663. [Google Scholar] [CrossRef] [PubMed]
- Blomberg, J.; Nived, O.; Pipkorn, R.; Bengtsson, A.; Erlinge, D.; Sturfelt, G. Increased antiretroviral antibody reactivity in sera from a defined population of patients with systemic lupus erythematosus. Correlation with autoantibodies and clinical manifestations. Arthritis Rheum. 1994, 37, 57–66. [Google Scholar] [CrossRef] [PubMed]
- Talal, N.; Garry, R.F.; Schur, P.H.; Alexander, S.; Dauphinee, M.J.; Livas, I.H.; Ballester, A.; Takei, M.; Dang, H. A conserved idiotype and antibodies to retroviral proteins in systemic lupus erythematosus. J. Clin. Investig. 1990, 85, 1866–1871. [Google Scholar] [CrossRef] [PubMed]
- Pelton, B.K.; North, M.; Palmer, R.G.; Hylton, W.; Smith-Burchnell, C.; Sinclair, A.L.; Malkovsky, M.; Dalgleish, A.G.; Denman, A.M. A search for retrovirus infection in systemic lupus erythematosus and rheumatoid arthritis. Ann. Rheum. Dis. 1988, 47, 206–209. [Google Scholar] [CrossRef]
- Perl, A.; Nagy, G.; Koncz, A.; Gergely, P.; Fernandez, D.; Doherty, E.; Telarico, T.; Bonilla, E.; Phillips, P.E. Molecular mimicry and immunomodulation by the HRES-1 endogenous retrovirus in SLE. Autoimmunity 2009, 41, 287–297. [Google Scholar] [CrossRef]
- Montesion, M.; Williams, Z.H.; Subramanian, R.P.; Kuperwasser, C.; Coffin, J.M. Promoter expression of HERV-K (HML-2) provirus-derived sequences is related to LTR sequence variation and polymorphic transcription factor binding sites. Retrovirology 2018, 15, 57. [Google Scholar] [CrossRef]
- Zhou, F.; Li, M.; Wei, Y.; Lin, K.; Lu, Y.; Shen, J.; Johanning, G.L.; Wang-Johanning, F. Activation of HERV-K Env protein is essential for tumorigenesis and metastasis of breast cancer cells. Oncotarget 2016, 7, 84093–84117. [Google Scholar] [CrossRef]
- Goering, W.; Schmitt, K.; Dostert, M.; Schaal, H.; Deenen, R.; Mayer, J.; Schulz, W.A. Human endogenous retrovirus HERV-K(HML-2) activity in prostate cancer is dominated by a few loci. Prostate 2015, 75, 1958–1971. [Google Scholar] [CrossRef] [PubMed]
- Kelly, M.; Lihua, S.; Zhe, Z.; Li, S.; Yoselin, P.; Michelle, P.; Sullivan Kathleen, E. Transposable element dysregulation in systemic lupus erythematosus and regulation by histone conformation and Hsp90. Clin. Immunol. 2018, 197, 6–18. [Google Scholar] [CrossRef] [PubMed]
- Elbarbary, R.A.; Lucas, B.A.; Maquat, L.E. Retrotransposons as regulators of gene expression. Science 2016, 351, aac7247. [Google Scholar] [CrossRef] [PubMed]
- Pullmann, R., Jr.; Bonilla, E.; Phillips, P.E.; Middleton, F.A.; Perl, A. Haplotypes of the HRES-1 endogenous retrovirus are associated with development and disease manifestations of systemic lupus erythematosus. Arthritis Rheum. 2008, 58, 532–540. [Google Scholar] [CrossRef]
- Ostertag, E.M.; Kazazian, H.H., Jr. Biology of mammalian L1 retrotransposons. Annu. Rev. Genet. 2001, 35, 501–538. [Google Scholar] [CrossRef]
- Brouha, B.; Schustak, J.; Badge, R.M.; Lutz-Prigge, S.; Farley, A.H.; Moran, J.V.; Kazazian, H.H., Jr. Hot L1s account for the bulk of retrotransposition in the human population. Proc. Natl. Acad. Sci. USA 2003, 100, 5280–5285. [Google Scholar] [CrossRef]
- Richardson, S.R.; Doucet, A.J.; Kopera, H.C.; Moldovan, J.B.; Garcia-Perez, J.L.; Moran, J.V. The Influence of LINE-1 and SINE Retrotransposons on Mammalian Genomes. Microbiol. Spectr. 2015, 3, MDNA3-0061-2014. [Google Scholar] [CrossRef]
- Petri, R.; Brattas, P.L.; Sharma, Y.; Jonsson, M.E.; Pircs, K.; Bengzon, J.; Jakobsson, J. LINE-2 transposable elements are a source of functional human microRNAs and target sites. PLoS Genet. 2019, 15, e1008036. [Google Scholar] [CrossRef] [PubMed]
- Feng, Q.; Moran, J.V.; Kazazian, H.H., Jr.; Boeke, J.D. Human L1 retrotransposon encodes a conserved endonuclease required for retrotransposition. Cell 1996, 87, 905–916. [Google Scholar] [CrossRef]
- Mathias, S.L.; Scott, A.F.; Kazazian, H.H., Jr.; Boeke, J.D.; Gabriel, A. Reverse transcriptase encoded by a human transposable element. Science 1991, 254, 1808–1810. [Google Scholar] [CrossRef]
- Tighe, P.J.; Stevens, S.E.; Dempsey, S.; Le Deist, F.; Rieux-Laucat, F.; Edgar, J.D. Inactivation of the Fas gene by Alu insertion: Retrotransposition in an intron causing splicing variation and autoimmune lymphoproliferative syndrome. Genes Immun. 2002, 3 (Suppl. 1), S66–S70. [Google Scholar] [CrossRef]
- Heinrich, M.J.; Purcell, C.A.; Pruijssers, A.J.; Zhao, Y.; Spurlock, C.F., 3rd; Sriram, S.; Ogden, K.M.; Dermody, T.S.; Scholz, M.B.; Crooke, P.S., 3rd; et al. Endogenous double-stranded Alu RNA elements stimulate IFN-responses in relapsing remitting multiple sclerosis. J. Autoimmun. 2019, 100, 40–51. [Google Scholar] [CrossRef] [PubMed]
- Rice, G.I.; Kasher, P.R.; Forte, G.M.; Mannion, N.M.; Greenwood, S.M.; Szynkiewicz, M.; Dickerson, J.E.; Bhaskar, S.S.; Zampini, M.; Briggs, T.A.; et al. Mutations in ADAR1 cause Aicardi-Goutieres syndrome associated with a type I interferon signature. Nat. Genet. 2012, 44, 1243–1248. [Google Scholar] [CrossRef]
- Tossberg, J.T.; Heinrich, R.M.; Farley, V.M.; Crooke, P.S., 3rd; Aune, T.M. Adenosine-to-Inosine RNA Editing of Alu Double-Stranded (ds)RNAs Is Markedly Decreased in Multiple Sclerosis and Unedited Alu dsRNAs Are Potent Activators of Proinflammatory Transcriptional Responses. J. Immunol. 2020, 205, 2606–2617. [Google Scholar] [CrossRef] [PubMed]
- Shen, C.K.; Maniatis, T. The organization, structure, and in vitro transcription of Alu family RNA polymerase III transcription units in the human alpha-like globin gene cluster: Precipitation of in vitro transcripts by lupus anti-La antibodies. J. Mol. Appl. Genet. 1982, 1, 343–360. [Google Scholar]
- Kole, R.; Fresco, L.D.; Keene, J.D.; Cohen, P.L.; Eisenberg, R.A.; Andrews, P.G. Alu RNA-protein complexes formed in vitro react with a novel lupus autoantibody. J. Biol. Chem. 1985, 260, 11781–11786. [Google Scholar] [CrossRef]
- Hung, T.; Pratt, G.A.; Sundararaman, B.; Townsend, M.J.; Chaivorapol, C.; Bhangale, T.; Graham, R.R.; Ortmann, W.; Criswell, L.A.; Yeo, G.W.; et al. The Ro60 autoantigen binds endogenous retroelements and regulates inflammatory gene expression. Science 2015, 350, 455–459. [Google Scholar] [CrossRef] [PubMed]
- Mavragani, C.P.; Sagalovskiy, I.; Guo, Q.; Nezos, A.; Kapsogeorgou, E.K.; Lu, P.; Liang Zhou, J.; Kirou, K.A.; Seshan, S.V.; Moutsopoulos, H.M.; et al. Expression of Long Interspersed Nuclear Element 1 Retroelements and Induction of Type I Interferon in Patients With Systemic Autoimmune Disease. Arthritis Rheumatol. 2016, 68, 2686–2696. [Google Scholar] [CrossRef]
- Richardson, B.C.; Strahler, J.R.; Pivirotto, T.S.; Quddus, J.; Bayliss, G.E.; Gross, L.A.; O’Rourke, K.S.; Powers, D.; Hanash, S.M.; Johnson, M.A. Phenotypic and functional similarities between 5-azacytidine-treated T cells and a T cell subset in patients with active systemic lupus erythematosus. Arthritis Rheum. 1992, 35, 647–662. [Google Scholar] [CrossRef]
- Goldstein, N.; Leider, M.; Baer, R.L. Drug eruptions from anticonvulsant drugs. Observations on drug-induced lesions that were fixed and lupus-erythematosus-like, and on cross-reactivity between several drugs involved. Arch. Dermatol. 1963, 87, 612–617. [Google Scholar] [CrossRef]
- Zingale, S.B.; Minzer, L.; Rosenberg, B.; Lee, S.L. Drug induced lupus-like syndrome. Clinical and laboratory syndrome similar to systemic lupus erythematosus following antituberculous therapy: Report of a case. Arch. Intern. Med. 1963, 112, 63–66. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Su, G.; Wang, Z.; Shangguan, S.; Cui, X.; Zhu, J.; Kang, M.; Li, S.; Zhang, T.; Wu, F.; et al. Hypomethylation of long interspersed nucleotide element-1 in peripheral mononuclear cells of juvenile systemic lupus erythematosus patients in China. Int. J. Rheum. Dis. 2014, 17, 280–290. [Google Scholar] [CrossRef] [PubMed]
- Everett, M.A.; Olson, R.L. Response of Cutaneous Lupus Erythematosus to Ultraviolet Light. J. Invest. Dermatol. 1965, 44, 133–138. [Google Scholar] [CrossRef][Green Version]
- Bijl, M.; Kallenberg, C.G. Ultraviolet light and cutaneous lupus. Lupus 2006, 15, 724–727. [Google Scholar] [CrossRef] [PubMed]
- Del Re, B.; Giorgi, G. Long INterspersed element-1 mobility as a sensor of environmental stresses. Environ. Mol. Mutagen. 2020, 61, 465–493. [Google Scholar] [CrossRef] [PubMed]
- Leffers, H.C.B.; Lange, T.; Collins, C.; Ulff-Moller, C.J.; Jacobsen, S. The study of interactions between genome and exposome in the development of systemic lupus erythematosus. Autoimmun. Rev. 2019, 18, 382–392. [Google Scholar] [CrossRef]
- Carter, V.; LaCava, J.; Taylor, M.S.; Liang, S.Y.; Mustelin, C.; Ukadike, K.C.; Bengtsson, A.; Lood, C.; Mustelin, T. High Prevalence and Disease Correlation of Autoantibodies Against p40 Encoded by Long Interspersed Nuclear Elements in Systemic Lupus Erythematosus. Arthritis Rheumatol. 2020, 72, 89–99. [Google Scholar] [CrossRef]
- Crow, M.K. Reactivity of IgG with the p40 Protein Encoded by the Long Interspersed Nuclear Element 1 Retroelement: Comment on the Article by Carter et al. Arthritis Rheumatol. 2020, 72, 374–376. [Google Scholar] [CrossRef]
- Goodier, J.L.; Zhang, L.; Vetter, M.R.; Kazazian, H.H., Jr. LINE-1 ORF1 protein localizes in stress granules with other RNA-binding proteins, including components of RNA interference RNA-induced silencing complex. Mol. Cell. Biol. 2007, 27, 6469–6483. [Google Scholar] [CrossRef]
- Taylor, M.S.; LaCava, J.; Mita, P.; Molloy, K.R.; Huang, C.R.; Li, D.; Adney, E.M.; Jiang, H.; Burns, K.H.; Chait, B.T.; et al. Affinity proteomics reveals human host factors implicated in discrete stages of LINE-1 retrotransposition. Cell 2013, 155, 1034–1048. [Google Scholar] [CrossRef]
- De Cecco, M.; Ito, T.; Petrashen, A.P.; Elias, A.E.; Skvir, N.J.; Criscione, S.W.; Caligiana, A.; Brocculi, G.; Adney, E.M.; Boeke, J.D.; et al. L1 drives IFN in senescent cells and promotes age-associated inflammation. Nature 2019, 566, 73–78. [Google Scholar] [CrossRef]
- Jiao, H.; Wachsmuth, L.; Kumari, S.; Schwarzer, R.; Lin, J.; Eren, R.O.; Fisher, A.; Lane, R.; Young, G.R.; Kassiotis, G.; et al. Z-nucleic-acid sensing triggers ZBP1-dependent necroptosis and inflammation. Nature 2020, 580, 391–395. [Google Scholar] [CrossRef]
- Wang, R.; Li, H.; Wu, J.; Cai, Z.Y.; Li, B.; Ni, H.; Qiu, X.; Chen, H.; Liu, W.; Yang, Z.H.; et al. Gut stem cell necroptosis by genome instability triggers bowel inflammation. Nature 2020, 580, 386–390. [Google Scholar] [CrossRef]
- Bengtsson, A.A.; Sturfelt, G.; Truedsson, L.; Blomberg, J.; Alm, G.; Vallin, H.; Ronnblom, L. Activation of type I interferon system in systemic lupus erythematosus correlates with disease activity but not with antiretroviral antibodies. Lupus 2000, 9, 664–671. [Google Scholar] [CrossRef] [PubMed]
- Baechler, E.C.; Batliwalla, F.M.; Karypis, G.; Gaffney, P.M.; Ortmann, W.A.; Espe, K.J.; Shark, K.B.; Grande, W.J.; Hughes, K.M.; Kapur, V.; et al. Interferon-inducible gene expression signature in peripheral blood cells of patients with severe lupus. Proc. Natl. Acad. Sci. USA 2003, 100, 2610–2615. [Google Scholar] [CrossRef] [PubMed]
- Bennett, L.; Palucka, A.K.; Arce, E.; Cantrell, V.; Borvak, J.; Banchereau, J.; Pascual, V. Interferon and granulopoiesis signatures in systemic lupus erythematosus blood. J. Exp. Med. 2003, 197, 711–723. [Google Scholar] [CrossRef] [PubMed]
- Crow, M.K.; Kirou, K.A.; Wohlgemuth, J. Microarray analysis of interferon-regulated genes in SLE. Autoimmunity 2003, 36, 481–490. [Google Scholar] [CrossRef] [PubMed]
- Ronnblom, L.; Alm, G.V.; Eloranta, M.L. Type I interferon and lupus. Curr. Opin. Rheumatol. 2009, 21, 471–477. [Google Scholar] [CrossRef]
- Zhao, K.; Du, J.; Peng, Y.; Li, P.; Wang, S.; Wang, Y.; Hou, J.; Kang, J.; Zheng, W.; Hua, S.; et al. LINE1 contributes to autoimmunity through both RIG-I- and MDA5-mediated RNA sensing pathways. J. Autoimmun. 2018, 90, 105–115. [Google Scholar] [CrossRef]
- Melki, I.; Rose, Y.; Uggenti, C.; Van Eyck, L.; Fremond, M.L.; Kitabayashi, N.; Rice, G.I.; Jenkinson, E.M.; Boulai, A.; Jeremiah, N.; et al. Disease-associated mutations identify a novel region in human STING necessary for the control of type I interferon signaling. J. Allergy Clin. Immunol. 2017, 140, 543–552.e5. [Google Scholar] [CrossRef]
- Bodewes, I.L.A.; Huijser, E.; van Helden-Meeuwsen, C.G.; Tas, L.; Huizinga, R.; Dalm, V.; van Hagen, P.M.; Groot, N.; Kamphuis, S.; van Daele, P.L.A.; et al. TBK1: A key regulator and potential treatment target for interferon positive Sjogren’s syndrome, systemic lupus erythematosus and systemic sclerosis. J. Autoimmun. 2018, 91, 97–102. [Google Scholar] [CrossRef]
- An, J.; Durcan, L.; Karr, R.M.; Briggs, T.A.; Rice, G.I.; Teal, T.H.; Woodward, J.J.; Elkon, K.B. Expression of Cyclic GMP-AMP Synthase in Patients with Systemic Lupus Erythematosus. Arthritis Rheumatol. 2017, 69, 800–807. [Google Scholar] [CrossRef] [PubMed]
- Ronnblom, L.; Eloranta, M.L.; Alm, G.V. The type I interferon system in systemic lupus erythematosus. Arthritis Rheum. 2006, 54, 408–420. [Google Scholar] [CrossRef] [PubMed]
- Ronnblom, L. The importance of the type I interferon system in autoimmunity. Clin. Exp. Rheumatol. 2016, 34, 21–24. [Google Scholar] [PubMed]
- Arbuckle, M.R.; McClain, M.T.; Rubertone, M.V.; Scofield, R.H.; Dennis, G.J.; James, J.A.; Harley, J.B. Development of autoantibodies before the clinical onset of systemic lupus erythematosus. N. Engl. J. Med. 2003, 349, 1526–1533. [Google Scholar] [CrossRef]
- Reynier, F.; Verjat, T.; Turrel, F.; Imbert, P.E.; Marotte, H.; Mougin, B.; Miossec, P. Increase in human endogenous retrovirus HERV-K (HML-2) viral load in active rheumatoid arthritis. Scand. J. Immunol. 2009, 70, 295–299. [Google Scholar] [CrossRef]
- Talal, N.; Flescher, E.; Dang, H. Are endogenous retroviruses involved in human autoimmune disease? J. Autoimmun. 1992, 5 (Suppl. A), 61–66. [Google Scholar] [CrossRef]
- Barat, C.; Schatz, O.; Le Grice, S.; Darlix, J.L. Analysis of the interactions of HIV1 replication primer tRNA(Lys,3) with nucleocapsid protein and reverse transcriptase. J. Mol. Biol. 1993, 231, 185–190. [Google Scholar] [CrossRef]
- Ji, X.; Klarmann, G.J.; Preston, B.D. Effect of human immunodeficiency virus type 1 (HIV-1) nucleocapsid protein on HIV-1 reverse transcriptase activity in vitro. Biochemistry 1996, 35, 132–143. [Google Scholar] [CrossRef]
- Ariumi, Y. Guardian of the Human Genome: Host Defense Mechanisms against LINE-1 Retrotransposition. Front. Chem. 2016, 4, 28. [Google Scholar] [CrossRef]
- Yang, F.; Wang, P.J. Multiple LINEs of retrotransposon silencing mechanisms in the mammalian germline. Semin. Cell Dev. Biol. 2016, 59, 118–125. [Google Scholar] [CrossRef] [PubMed]
- Tsutsumi, Y. Hypomethylation of the retrotransposon LINE-1 in malignancy. Jpn. J. Clin. Oncol. 2000, 30, 289–290. [Google Scholar] [CrossRef] [PubMed]
- Nawrocki, M.J.; Majewski, D.; Puszczewicz, M.; Jagodzinski, P.P. Decreased mRNA expression levels of DNA methyltransferases type 1 and 3A in systemic lupus erythematosus. Rheumatol. Int. 2017, 37, 775–783. [Google Scholar] [CrossRef] [PubMed]
- Tunbak, H.; Enriquez-Gasca, R.; Tie, C.H.C.; Gould, P.A.; Mlcochova, P.; Gupta, R.K.; Fernandes, L.; Holt, J.; van der Veen, A.G.; Giampazolias, E.; et al. The HUSH complex is a gatekeeper of type I interferon through epigenetic regulation of LINE-1s. Nat. Commun. 2020, 11, 5387. [Google Scholar] [CrossRef]
- Hunter, R.G.; Murakami, G.; Dewell, S.; Seligsohn, M.; Baker, M.E.; Datson, N.A.; McEwen, B.S.; Pfaff, D.W. Acute stress and hippocampal histone H3 lysine 9 trimethylation, a retrotransposon silencing response. Proc. Natl. Acad. Sci. USA 2012, 109, 17657–17662. [Google Scholar] [CrossRef]
- Wang, X.; Jiang, C.; Fu, B.; Zhu, R.; Diao, F.; Xu, N.; Chen, Z.; Tao, W.; Li, C.J. MILI, a PIWI family protein, inhibits melanoma cell migration through methylation of LINE1. Biochem. Biophys. Res. Commun. 2015, 457, 514–519. [Google Scholar] [CrossRef]
- Kuramochi-Miyagawa, S.; Watanabe, T.; Gotoh, K.; Totoki, Y.; Toyoda, A.; Ikawa, M.; Asada, N.; Kojima, K.; Yamaguchi, Y.; Ijiri, T.W.; et al. DNA methylation of retrotransposon genes is regulated by Piwi family members MILI and MIWI2 in murine fetal testes. Genes Dev. 2008, 22, 908–917. [Google Scholar] [CrossRef]
- Russell, S.J.; Stalker, L.; LaMarre, J. PIWIs, piRNAs and Retrotransposons: Complex battles during reprogramming in gametes and early embryos. Reprod. Domest. Anim. 2017, 52 (Suppl. 4), 28–38. [Google Scholar] [CrossRef]
- Imgenberg-Kreuz, J.; Sandling, J.K.; Almlof, J.C.; Nordlund, J.; Signer, L.; Norheim, K.B.; Omdal, R.; Ronnblom, L.; Eloranta, M.L.; Syvanen, A.C.; et al. Genome-wide DNA methylation analysis in multiple tissues in primary Sjogren’s syndrome reveals regulatory effects at interferon-induced genes. Ann. Rheum. Dis. 2016, 75, 2029–2036. [Google Scholar] [CrossRef]
- Sukapan, P.; Promnarate, P.; Avihingsanon, Y.; Mutirangura, A.; Hirankarn, N. Types of DNA methylation status of the interspersed repetitive sequences for LINE-1, Alu, HERV-E and HERV-K in the neutrophils from systemic lupus erythematosus patients and healthy controls. J. Hum. Genet. 2014, 59, 178–188. [Google Scholar] [CrossRef]
- Mavragani, C.P.; Nezos, A.; Sagalovskiy, I.; Seshan, S.; Kirou, K.A.; Crow, M.K. Defective regulation of L1 endogenous retroelements in primary Sjogren’s syndrome and systemic lupus erythematosus: Role of methylating enzymes. J. Autoimmun. 2018, 88, 75–82. [Google Scholar] [CrossRef]
- Lee, J.; Kalia, V.; Perera, F.; Herbstman, J.; Li, T.; Nie, J.; Qu, L.R.; Yu, J.; Tang, D. Prenatal airborne polycyclic aromatic hydrocarbon exposure, LINE1 methylation and child development in a Chinese cohort. Environ. Int. 2017, 99, 315–320. [Google Scholar] [CrossRef]
- Breton, C.V.; Yao, J.; Millstein, J.; Gao, L.; Siegmund, K.D.; Mack, W.; Whitfield-Maxwell, L.; Lurmann, F.; Hodis, H.; Avol, E.; et al. Prenatal Air Pollution Exposures, DNA Methyl Transferase Genotypes, and Associations with Newborn LINE1 and Alu Methylation and Childhood Blood Pressure and Carotid Intima-Media Thickness in the Children’s Health Study. Environ. Health Perspect. 2016, 124, 1905–1912. [Google Scholar] [CrossRef]
- Cornacchia, E.; Golbus, J.; Maybaum, J.; Strahler, J.; Hanash, S.; Richardson, B. Hydralazine and procainamide inhibit T cell DNA methylation and induce autoreactivity. J. Immunol. 1988, 140, 2197–2200. [Google Scholar]
- Yung, R.L.; Quddus, J.; Chrisp, C.E.; Johnson, K.J.; Richardson, B.C. Mechanism of drug-induced lupus. I. Cloned Th2 cells modified with DNA methylation inhibitors in vitro cause autoimmunity in vivo. J. Immunol. 1995, 154, 3025–3035. [Google Scholar] [PubMed]
- Lieberman, M.W.; Beach, L.R.; Palmiter, R.D. Ultraviolet radiation-induced metallothionein-I gene activation is associated with extensive DNA demethylation. Cell 1983, 35, 207–214. [Google Scholar] [CrossRef]
- Yang, Y.G.; Lindahl, T.; Barnes, D.E. Trex1 exonuclease degrades ssDNA to prevent chronic checkpoint activation and autoimmune disease. Cell 2007, 131, 873–886. [Google Scholar] [CrossRef] [PubMed]
- Pokatayev, V.; Hasin, N.; Chon, H.; Cerritelli, S.M.; Sakhuja, K.; Ward, J.M.; Morris, H.D.; Yan, N.; Crouch, R.J. RNase H2 catalytic core Aicardi-Goutieres syndrome-related mutant invokes cGAS-STING innate immune-sensing pathway in mice. J. Exp. Med. 2016, 213, 329–336. [Google Scholar] [CrossRef] [PubMed]
- Thomas, C.A.; Tejwani, L.; Trujillo, C.A.; Negraes, P.D.; Herai, R.H.; Mesci, P.; Macia, A.; Crow, Y.J.; Muotri, A.R. Modeling of TREX1-Dependent Autoimmune Disease using Human Stem Cells Highlights L1 Accumulation as a Source of Neuroinflammation. Cell Stem Cell 2017, 21, 319–331.e8. [Google Scholar] [CrossRef]
- Lim, Y.W.; Sanz, L.A.; Xu, X.; Hartono, S.R.; Chedin, F. Genome-wide DNA hypomethylation and RNA:DNA hybrid accumulation in Aicardi-Goutieres syndrome. Elife 2015, 4. [Google Scholar] [CrossRef]
- Crow, Y.J.; Chase, D.S.; Lowenstein Schmidt, J.; Szynkiewicz, M.; Forte, G.M.; Gornall, H.L.; Oojageer, A.; Anderson, B.; Pizzino, A.; Helman, G.; et al. Characterization of human disease phenotypes associated with mutations in TREX1, RNASEH2A, RNASEH2B, RNASEH2C, SAMHD1, ADAR, and IFIH1. Am. J. Med. Genet. A 2015, 167A, 296–312. [Google Scholar] [CrossRef] [PubMed]
- Crow, Y.J.; Manel, N. Aicardi-Goutieres syndrome and the type I interferonopathies. Nat. Rev. Immunol. 2015, 15, 429–440. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Du, J.; Goodier, J.L.; Hou, J.; Kang, J.; Kazazian, H.H., Jr.; Zhao, K.; Yu, X.F. Aicardi-Goutieres syndrome protein TREX1 suppresses L1 and maintains genome integrity through exonuclease-independent ORF1p depletion. Nucleic Acids Res. 2017, 45, 4619–4631. [Google Scholar] [CrossRef] [PubMed]
- Rice, G.I.; Meyzer, C.; Bouazza, N.; Hully, M.; Boddaert, N.; Semeraro, M.; Zeef, L.A.H.; Rozenberg, F.; Bondet, V.; Duffy, D.; et al. Reverse-Transcriptase Inhibitors in the Aicardi-Goutieres Syndrome. N. Engl. J. Med. 2018, 379, 2275–2277. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Zhang, J.; Jia, R.; Cheng, V.; Xu, X.; Qiao, W.; Guo, F.; Liang, C.; Cen, S. The MOV10 helicase inhibits LINE-1 mobility. J. Biol. Chem. 2013, 288, 21148–21160. [Google Scholar] [CrossRef]
- Arjan-Odedra, S.; Swanson, C.M.; Sherer, N.M.; Wolinsky, S.M.; Malim, M.H. Endogenous MOV10 inhibits the retrotransposition of endogenous retroelements but not the replication of exogenous retroviruses. Retrovirology 2012, 9, 53. [Google Scholar] [CrossRef]
- Choi, J.; Hwang, S.Y.; Ahn, K. Interplay between RNASEH2 and MOV10 controls LINE-1 retrotransposition. Nucleic Acids Res. 2018, 46, 1912–1926. [Google Scholar] [CrossRef]
- Taylor, M.S.; LaCava, J.; Dai, L.; Mita, P.; Burns, K.H.; Rout, M.P.; Boeke, J.D. Characterization of L1-Ribonucleoprotein Particles. Methods Mol. Biol. 2016, 1400, 311–338. [Google Scholar] [CrossRef]
- Lian, H.; Wei, J.; Zang, R.; Ye, W.; Yang, Q.; Zhang, X.N.; Chen, Y.D.; Fu, Y.Z.; Hu, M.M.; Lei, C.Q.; et al. ZCCHC3 is a co-sensor of cGAS for dsDNA recognition in innate immune response. Nat. Commun. 2018, 9, 3349. [Google Scholar] [CrossRef]
- Lian, H.; Zang, R.; Wei, J.; Ye, W.; Hu, M.M.; Chen, Y.D.; Zhang, X.N.; Guo, Y.; Lei, C.Q.; Yang, Q.; et al. The Zinc-Finger Protein ZCCHC3 Binds RNA and Facilitates Viral RNA Sensing and Activation of the RIG-I-like Receptors. Immunity 2018, 49, 438–448.e5. [Google Scholar] [CrossRef]
- Rice, G.I.; Bond, J.; Asipu, A.; Brunette, R.L.; Manfield, I.W.; Carr, I.M.; Fuller, J.C.; Jackson, R.M.; Lamb, T.; Briggs, T.A.; et al. Mutations involved in Aicardi-Goutieres syndrome implicate SAMHD1 as regulator of the innate immune response. Nat. Genet. 2009, 41, 829–832. [Google Scholar] [CrossRef]
- Bogerd, H.P.; Wiegand, H.L.; Doehle, B.P.; Lueders, K.K.; Cullen, B.R. APOBEC3A and APOBEC3B are potent inhibitors of LTR-retrotransposon function in human cells. Nucleic Acids Res. 2006, 34, 89–95. [Google Scholar] [CrossRef] [PubMed]
- Refsland, E.W.; Harris, R.S. The APOBEC3 family of retroelement restriction factors. Curr. Top. Microbiol. Immunol. 2013, 371, 1–27. [Google Scholar] [CrossRef] [PubMed]
- Mustelin, T.; Lood, C.; Giltiay, N.V. Sources of Pathogenic Nucleic Acids in Systemic Lupus Erythematosus. Front. Immunol. 2019, 10, 1028. [Google Scholar] [CrossRef]
- Barrat, F.J.; Meeker, T.; Gregorio, J.; Chan, J.H.; Uematsu, S.; Akira, S.; Chang, B.; Duramad, O.; Coffman, R.L. Nucleic acids of mammalian origin can act as endogenous ligands for Toll-like receptors and may promote systemic lupus erythematosus. J. Exp. Med. 2005, 202, 1131–1139. [Google Scholar] [CrossRef]
- Ronnblom, L.; Alm, G.V. A pivotal role for the natural interferon alpha-producing cells (plasmacytoid dendritic cells) in the pathogenesis of lupus. J. Exp. Med. 2001, 194, F59–F63. [Google Scholar] [CrossRef] [PubMed]
- Lloyd, W.; Schur, P.H. Immune complexes, complement, and anti-DNA in exacerbations of systemic lupus erythematosus. Medicine 1981, 60, 208–217. [Google Scholar] [CrossRef]
- Kalunian, K.C.; Merrill, J.T.; Maciuca, R.; McBride, J.M.; Townsend, M.J.; Wei, X.; Davis, J.C., Jr.; Kennedy, W.P. A Phase II study of the efficacy and safety of rontalizumab (rhuMAb interferon-alpha) in patients with systemic lupus erythematosus (ROSE). Ann. Rheum. Dis. 2016, 75, 196–202. [Google Scholar] [CrossRef]
- Khamashta, M.; Merrill, J.T.; Werth, V.P.; Furie, R.; Kalunian, K.; Illei, G.G.; Drappa, J.; Wang, L.; Greth, W.; on behalf of the CD1067 Study Investigators. Sifalimumab, an anti-interferon-alpha monoclonal antibody, in moderate to severe systemic lupus erythematosus: A randomised, double-blind, placebo-controlled study. Ann. Rheum. Dis. 2016, 75, 1909–1916. [Google Scholar] [CrossRef]
- Tanaka, Y.; Tummala, R. Anifrolumab, a monoclonal antibody to the type I interferon receptor subunit 1, for the treatment of systemic lupus erythematosus: An overview from clinical trials. Mod. Rheumatol. 2020, 30, 1–12. [Google Scholar] [CrossRef]
- Shao, W.H.; Shu, D.H.; Zhen, Y.; Hilliard, B.; Priest, S.O.; Cesaroni, M.; Ting, J.P.; Cohen, P.L. Prion-like Aggregation of Mitochondrial Antiviral Signaling Protein in Lupus Patients Is Associated with Increased Levels of Type I Interferon. Arthritis Rheumatol. 2016, 68, 2697–2707. [Google Scholar] [CrossRef] [PubMed]
- Rodero, M.P.; Decalf, J.; Bondet, V.; Hunt, D.; Rice, G.I.; Werneke, S.; McGlasson, S.L.; Alyanakian, M.A.; Bader-Meunier, B.; Barnerias, C.; et al. Detection of interferon alpha protein reveals differential levels and cellular sources in disease. J. Exp. Med. 2017, 214, 1547–1555. [Google Scholar] [CrossRef] [PubMed]
- An, J.; Woodward, J.J.; Minie, M.; Sasaki, T.; Elkon, K.B. Novel Anti-Malarial Drug Derivative Inhibited Type I Interferon Production and Autoimmune Inflammation through Inhibition of CGAS-Sting Pathway in Trex1-/- Mouse (abstract 2984). Arthritis Rheumatol. 2016, 68. Available online: https://acrabstracts.org/abstract/novel-anti-malarial-drug-derivative-inhibited-type-i-interferon-production-and-autoimmune-inflammation-through-inhibition-of-cgas-sting-pathway-in-trex1-mouse/ (accessed on 13 February 2021).


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ukadike, K.C.; Mustelin, T. Implications of Endogenous Retroelements in the Etiopathogenesis of Systemic Lupus Erythematosus. J. Clin. Med. 2021, 10, 856. https://doi.org/10.3390/jcm10040856
Ukadike KC, Mustelin T. Implications of Endogenous Retroelements in the Etiopathogenesis of Systemic Lupus Erythematosus. Journal of Clinical Medicine. 2021; 10(4):856. https://doi.org/10.3390/jcm10040856
Chicago/Turabian StyleUkadike, Kennedy C., and Tomas Mustelin. 2021. "Implications of Endogenous Retroelements in the Etiopathogenesis of Systemic Lupus Erythematosus" Journal of Clinical Medicine 10, no. 4: 856. https://doi.org/10.3390/jcm10040856
APA StyleUkadike, K. C., & Mustelin, T. (2021). Implications of Endogenous Retroelements in the Etiopathogenesis of Systemic Lupus Erythematosus. Journal of Clinical Medicine, 10(4), 856. https://doi.org/10.3390/jcm10040856





