Electrospun Biodegradable Nanofibers Coated Homogenously by Cu Magnetron Sputtering Exhibit Fast Ion Release. Computational and Experimental Study
Abstract
:1. Introduction
2. Materials and Methods
2.1. Electrospinning of PCL Nanofibers
2.2. Magnetron Sputtering
2.3. Plasma COOH Coating
2.4. Chemistry and Morphology Analysis
2.5. The Ion Release Measuring
2.6. Modeling
2.7. Cell Tests
2.8. Microbiology
3. Results
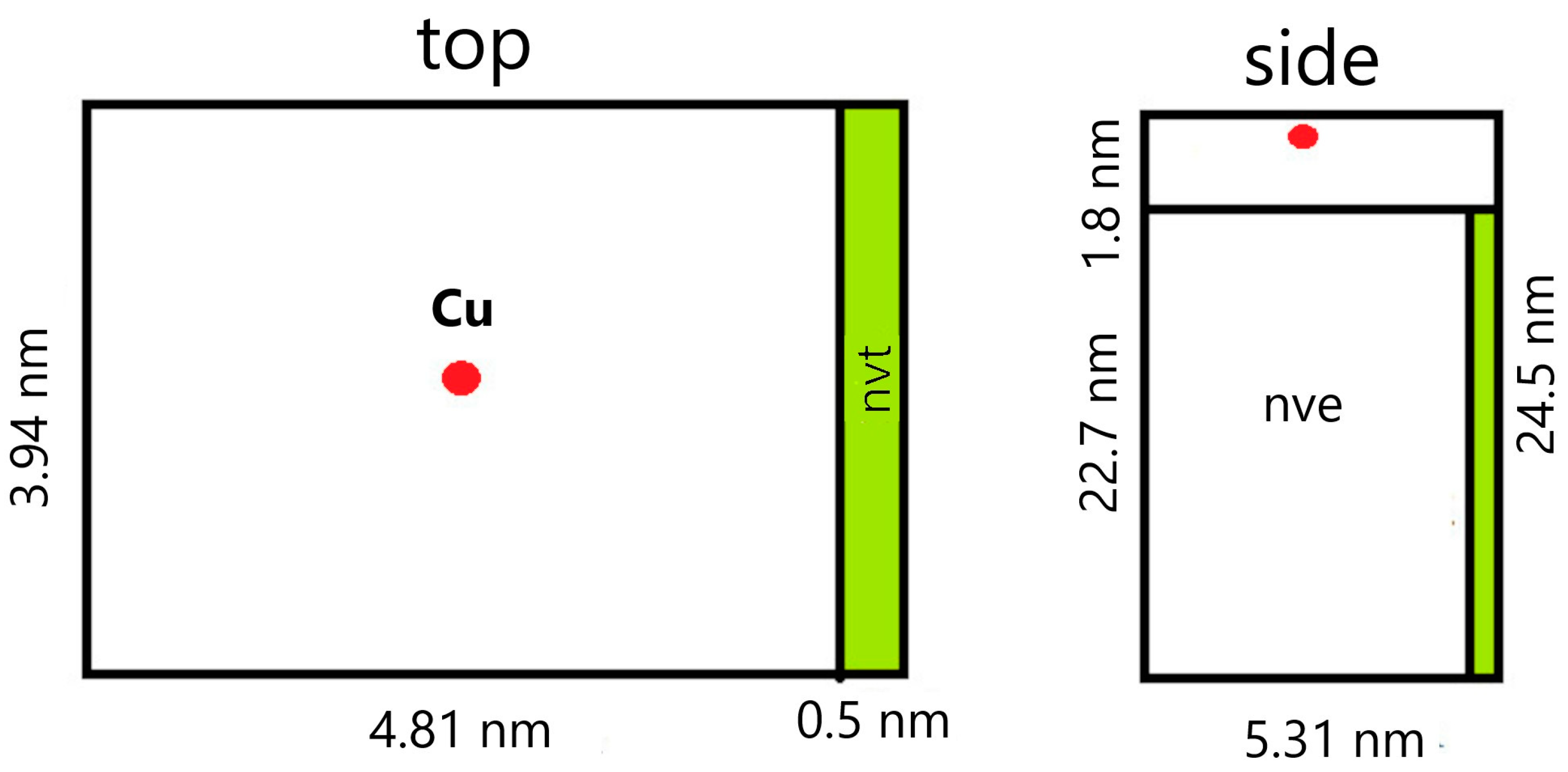
3.1. Modeling of Cu Ions Penetration Depth
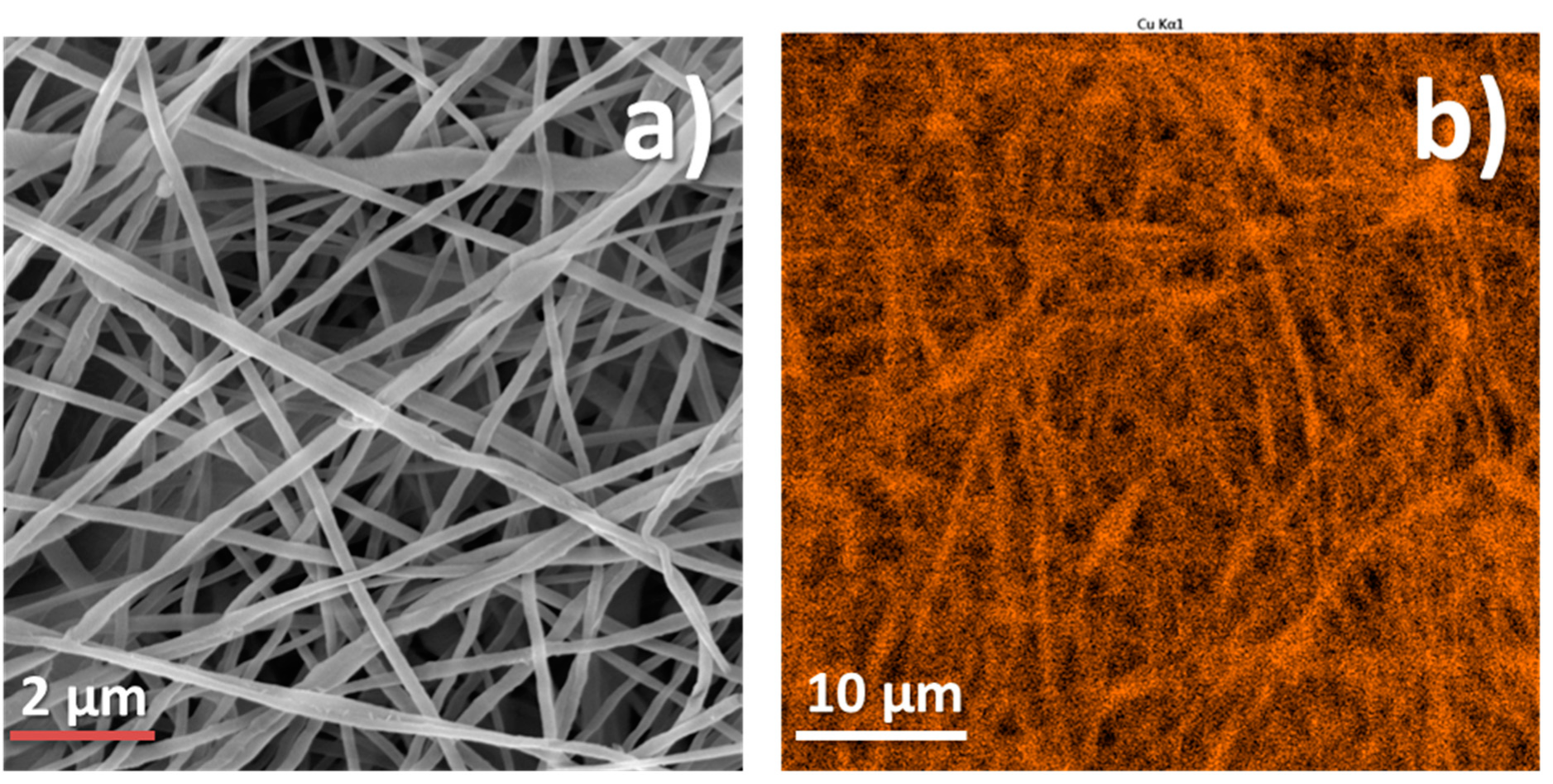
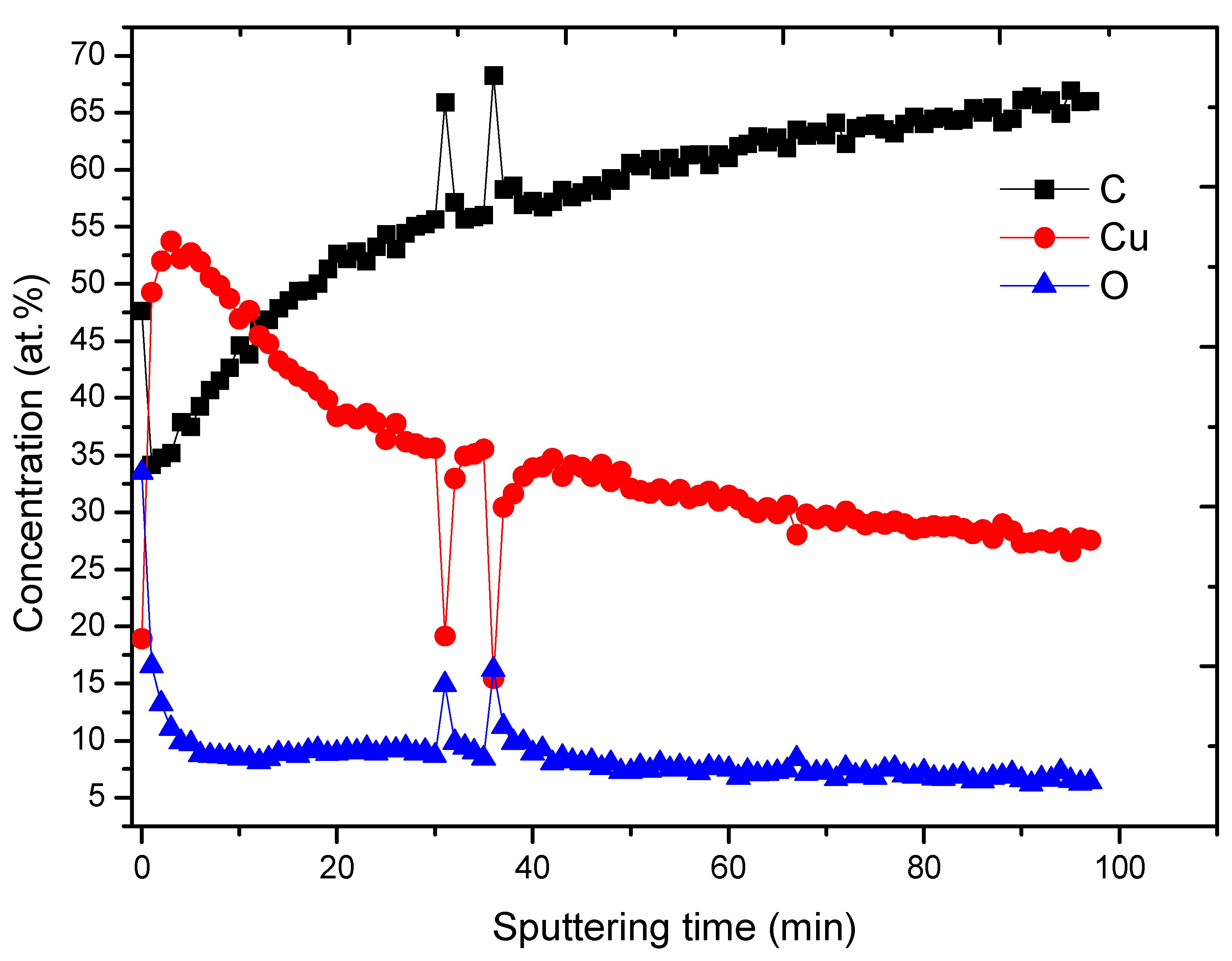
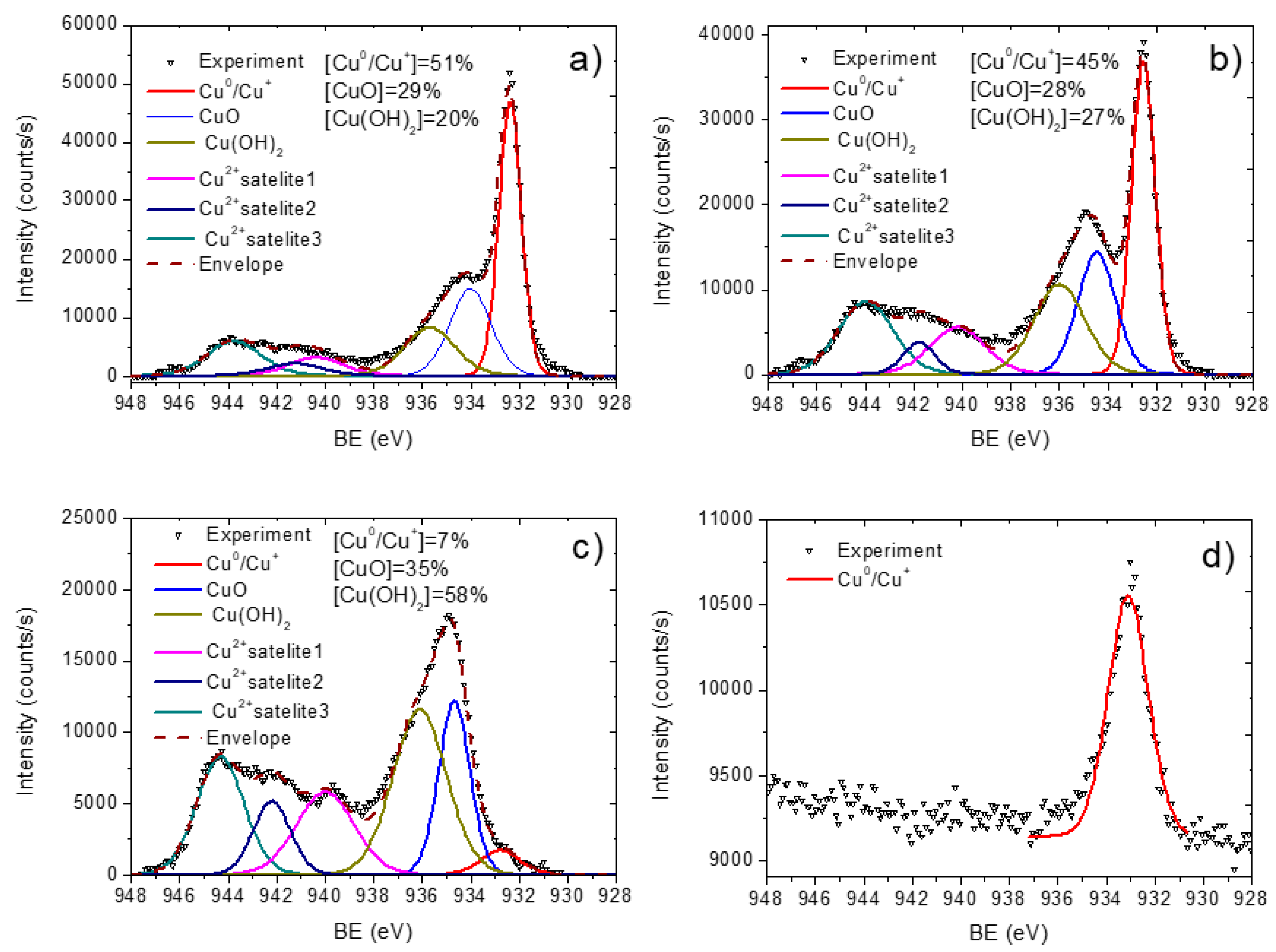
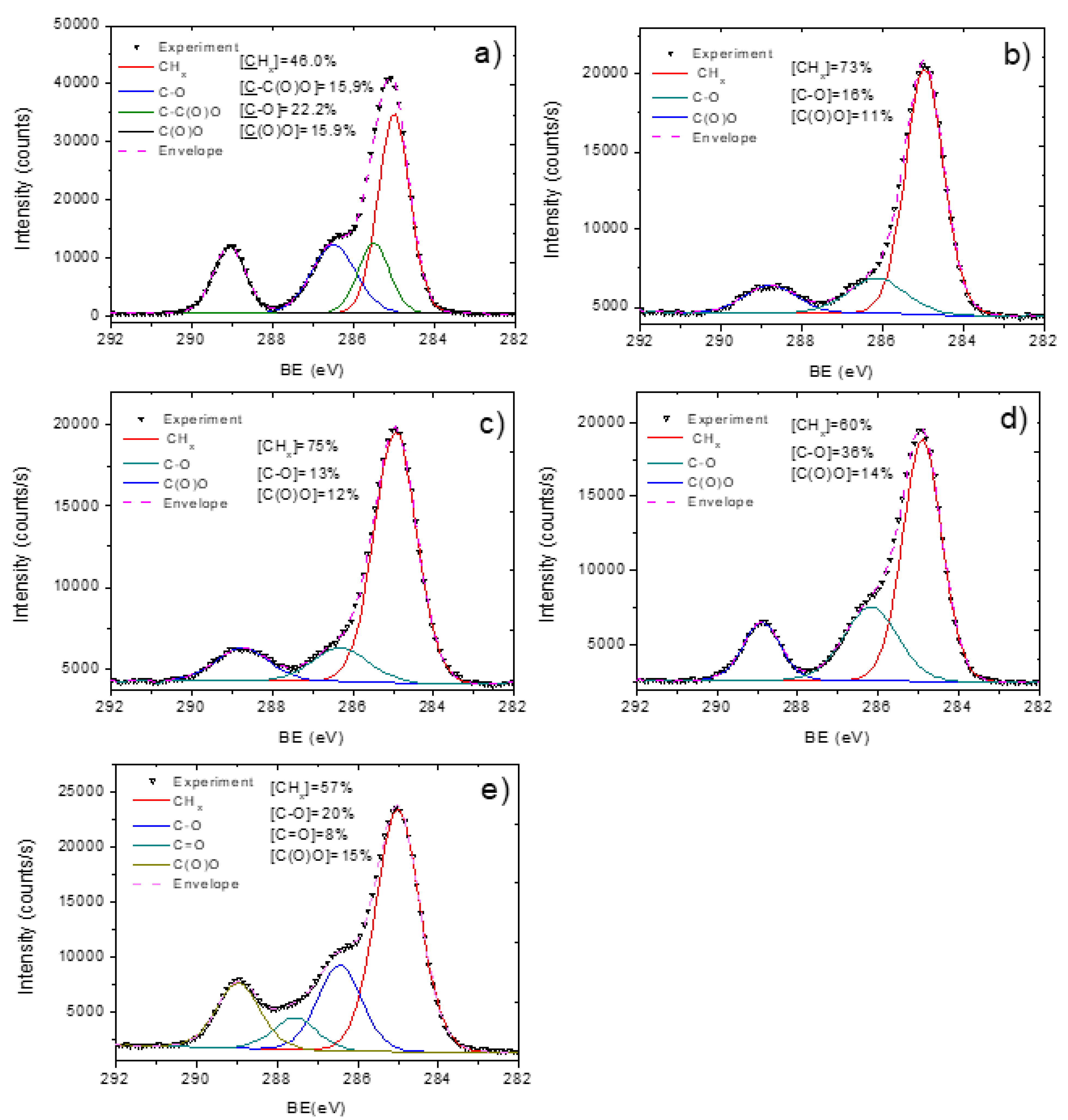
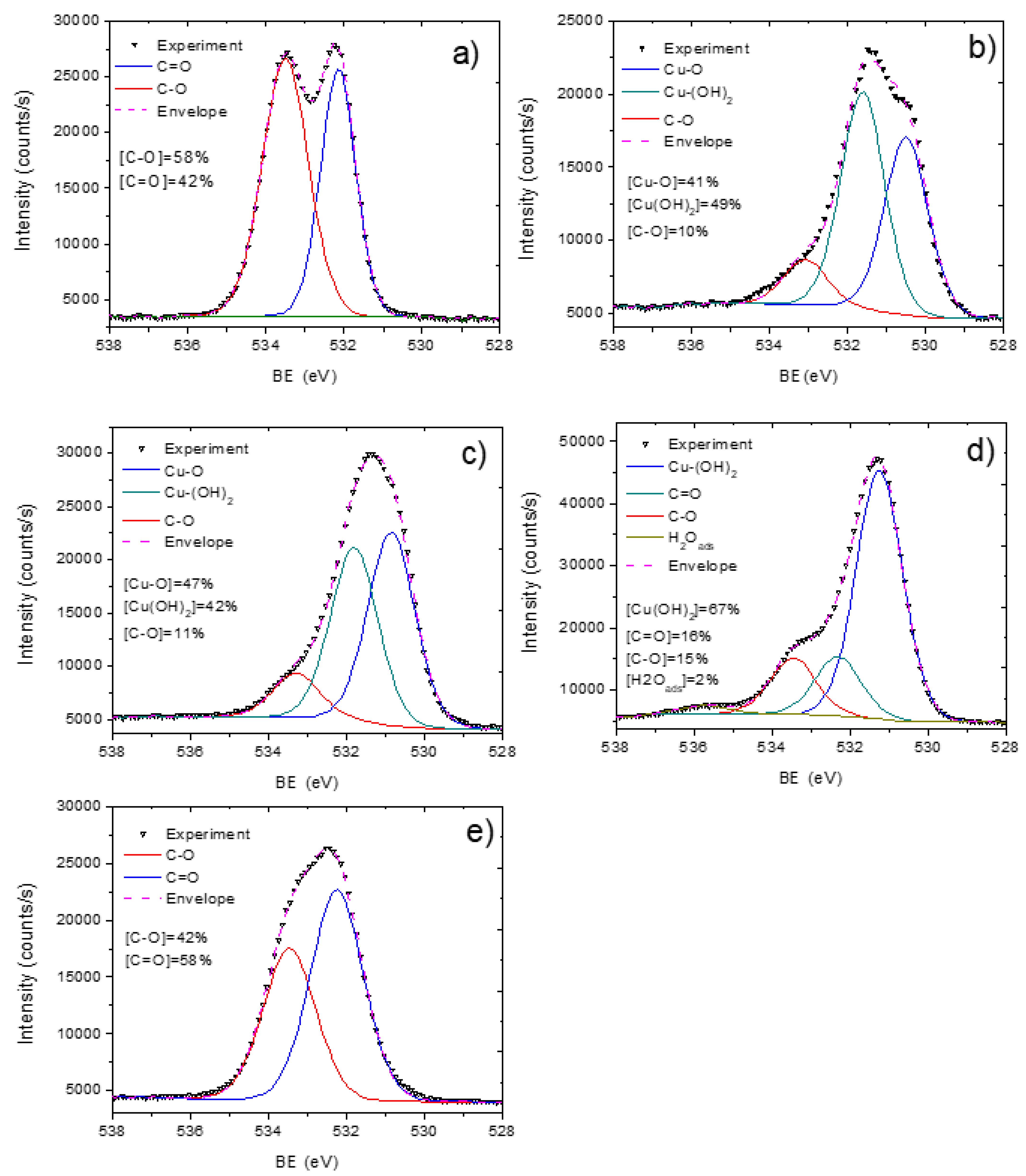
3.2. PCL-Cu Nanofibers
3.3. Cu-PCL Nanofibers Coated with Plasma Polymer
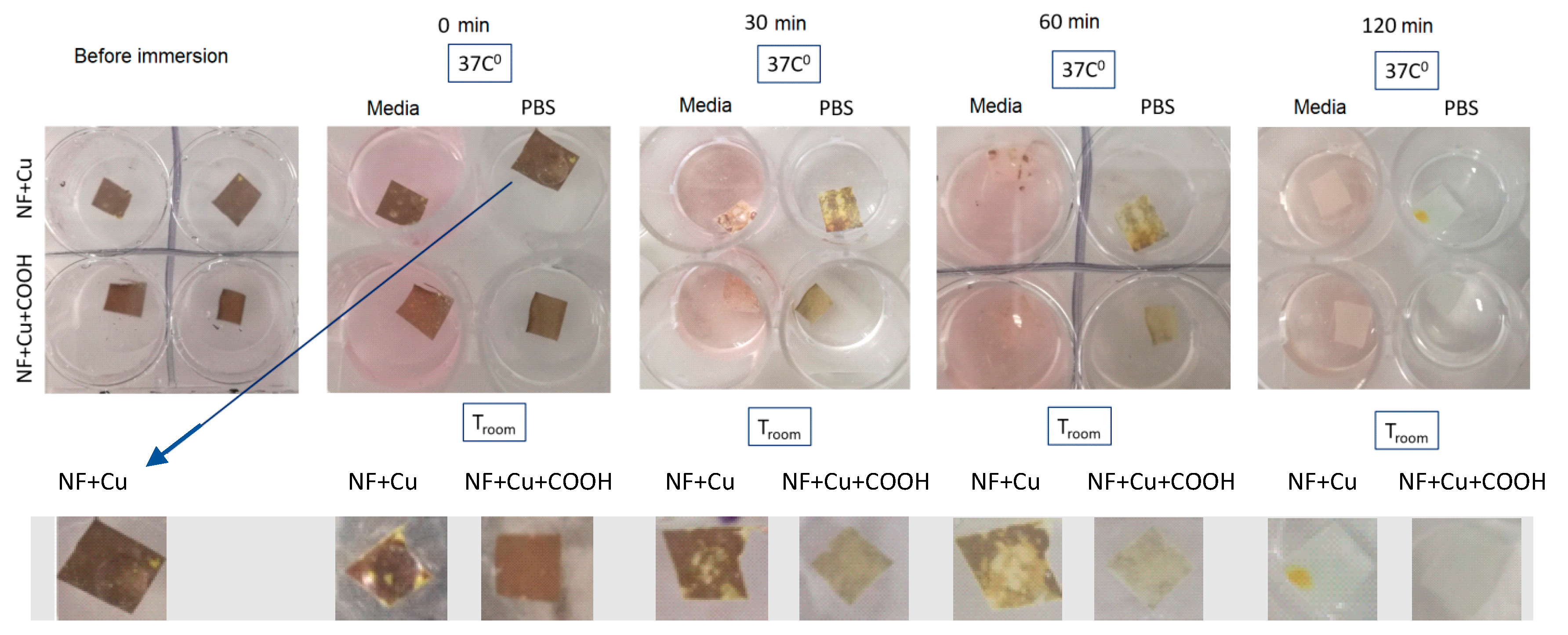
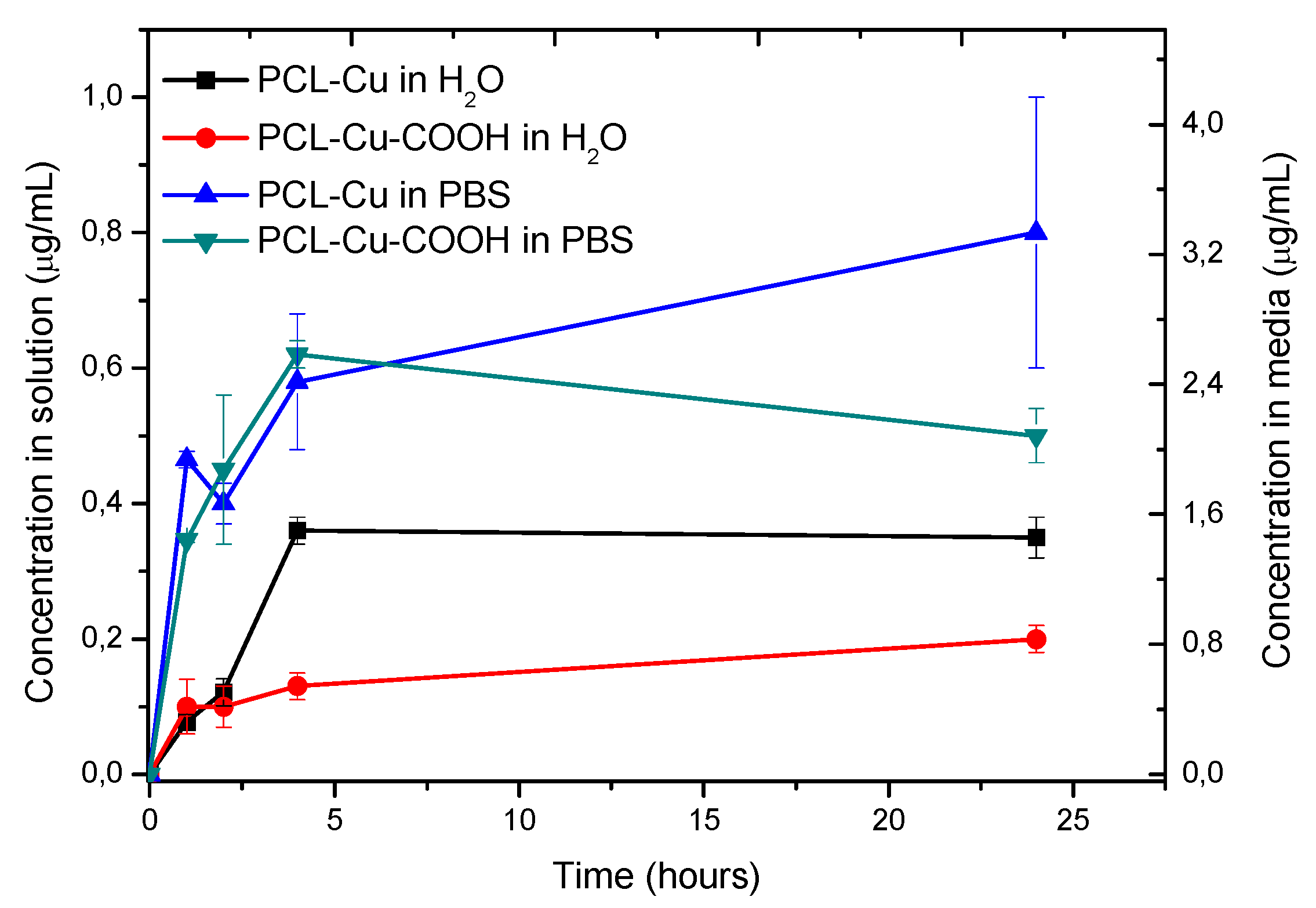
3.4. Stability in Water and Cu Ions Release
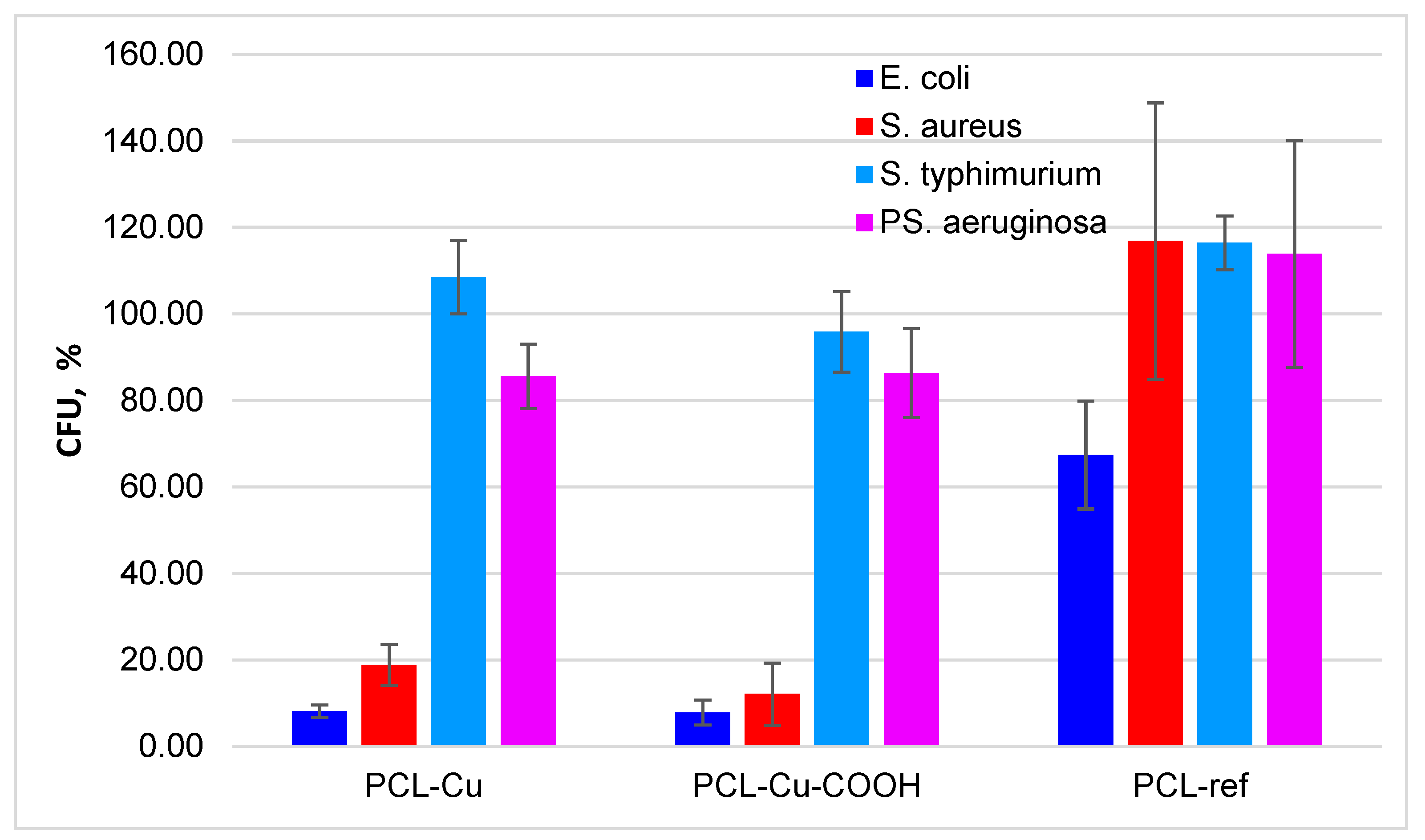
3.5. Antibacterial Properties
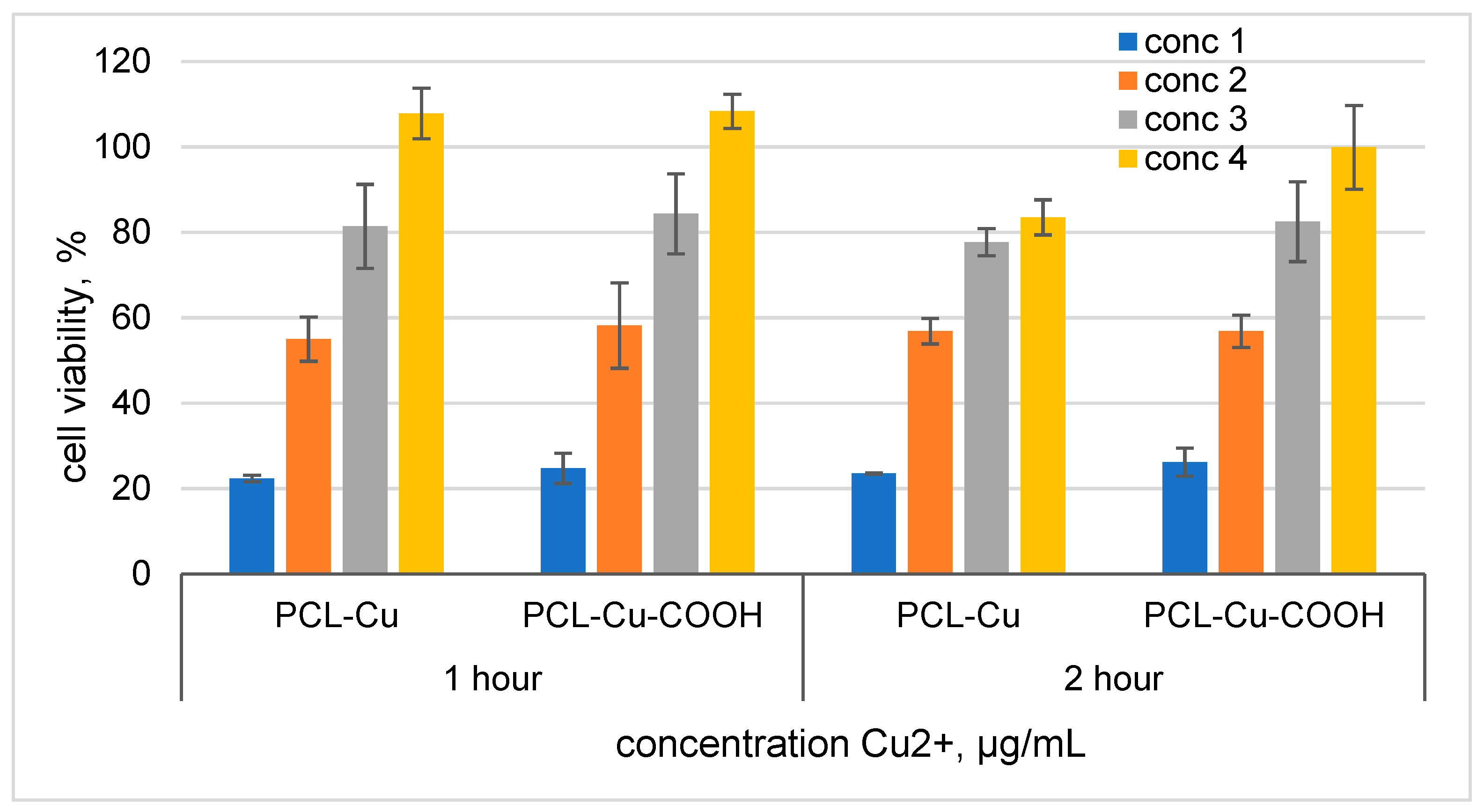
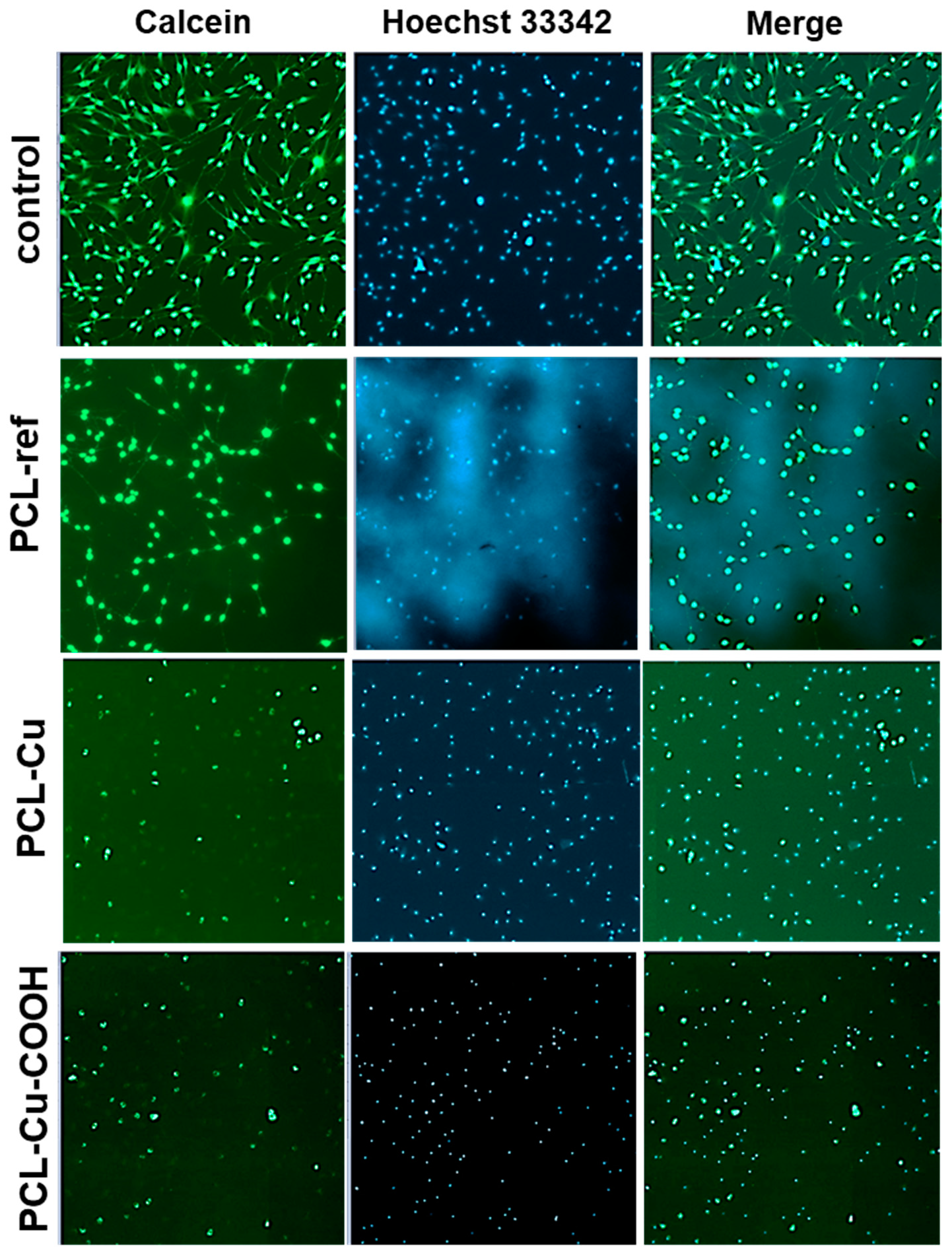
3.6. Cytotoxicity
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Xie, L.; Li, Z.; Li, X.; Wenlong, W.; Hanjiang, Y. Electrospun copper oxide nanofibers and catalysis for combustion of ammonium perchlorate. Ferroelectrics 2019, 549, 23–28. [Google Scholar] [CrossRef]
- Wang, H.; Fu, Q.; Zhang, G.; Sun, Y. The Synthesis of Ni–Cu Alloy Nanofibers via Vacuum Thermal Co-reduction Toward Hydrogen Generation from Hydrazine Decomposition. Catal. Lett. 2019, 149, 77–83. [Google Scholar] [CrossRef]
- Zheng, X.; Shen, Z.; Cheng, C.; Shi, L.; Cheng, R.; Dong, J. Electrospinning Cu–TiO2 nanofibers used for photocatalytic disinfection of bacteriophage f2: Preparation, optimization and characterization. RSC Adv. 2017, 7, 52172–52179. [Google Scholar] [CrossRef] [Green Version]
- Cui, W.Y.; Yoo, H.J.; Li, Y.G.; Baek, C.; Min, J. Electrospun Nanofibers Embedded with Copper Oxide Nanoparticles to Improve Antiviral Function. J. Nanosci. Nanotechnol. 2021, 21, 4174–4178. [Google Scholar] [CrossRef]
- Abedalwafa, M.A.; Zhang, H.; Mei, Q.; Li, Y.; Wang, F. A novel method to construct antimicrobial surface by decorating polyacrylonitrile nanofibrous membrane with nanoparticles. J. Ind. Text. 2020. [Google Scholar] [CrossRef]
- Ghasemian Lemraski, E.; Jahangirian, H.; Dashti, M.; Khajehali, E.; Sharafinia, M.S.; Rafiee-Moghaddam, R.; Webster, T.J. Antimicrobial Double-Layer Wound Dressing Based on Chitosan/Polyvinyl Alcohol/Copper: In vitro and in vivo Assessment. Int. J. Nanomed. 2021, 16, 223–235. [Google Scholar] [CrossRef]
- Choi, S.-W.; Katoch, A.; Zhang, J.; Kim, S.S. Electrospun nanofibers of CuOSnO2 nanocomposite as semiconductor gas sensors for H2S detection. Sens. Actuators B Chem. 2013, 176, 585–591. [Google Scholar] [CrossRef]
- Ju, W.; Jiang, F.; Ma, H.; Pan, Z.; Zhao, Y.; Pagani, F.; Rentsch, D.; Wang, J.; Battaglia, C. Electrocatalytic Reduction of Gaseous CO2 to CO on Sn/Cu-Nanofiber-Based Gas Diffusion Electrodes. Adv. Energy Mater. 2019, 9, 1901514. [Google Scholar] [CrossRef]
- Nami-Ana, S.F.; Nasresfahani, S.; Tashkhourian, J.; Shamsipur, M.; Zargarpour, Z.; Sheikhi, M.H. Nanofibers of Polyaniline and Cu(II)–L-Aspartic Acid for a Room-Temperature Carbon Monoxide Gas Sensor. ACS Appl. Mater. Interfaces 2021, 13, 39791–39805. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.Q.; Kharaghani, D.; Nishat, N.; Sanaullah; Shahzad, A.; Hussain, T.; Kim, K.O.; Kim, I.S. The fabrications and characterizations of antibacterial PVA/Cu nanofibers composite membranes by synthesis of Cu nanoparticles from solution reduction, nanofibers reduction and immersion methods. Mater. Res. Express 2019, 6, 075051. [Google Scholar] [CrossRef]
- Liu, X.; Xu, H.; Zhang, M.; Yu, D.-G. Electrospun Medicated Nanofibers for Wound Healing: Review. Membranes 2021, 11, 770. [Google Scholar] [CrossRef] [PubMed]
- Mousavi, S.; Nejad, Z.M.; Hashemi, S.A.; Salari, M.; Gholami, A.; Ramakrishna, S.; Chiang, W.; Lai, C.W. Bioactive Agent-Loaded Electrospun Nanofiber Membranes for Accelerating Healing Process: A Review. Membranes 2021, 11, 702. [Google Scholar] [CrossRef] [PubMed]
- Essa, W.; Yasin, S.; Saeed, I.; Ali, G. Nanofiber-Based Face Masks and Respirators as COVID-19 Protection: A Review. Membrranes 2021, 11, 250. [Google Scholar] [CrossRef]
- Ullah, S.; Ullah, A.; Lee, J.; Jeong, Y.; Hashmi, M.; Zhu, C.; Joo, K., II; Cha, H.J.; Kim, I.S. Reusability Comparison of Melt-Blown vs Nanofiber Face Mask Filters for Use in the Coronavirus Pandemic. ACS Appl. Nano Mater. 2020, 3, 7231–7241. [Google Scholar] [CrossRef]
- Mozaffari, A.; Gashti, M.P.; Mirjalili, M.; Parsania, M. Argon and Argon–Oxygen Plasma Surface Modification of Gelatin Nanofibers for Tissue Engineering Applications. Membranes 2021, 11, 31. [Google Scholar] [CrossRef] [PubMed]
- Ramírez-Cedillo, E.; Ortega-Lara, W.; Rocha-Pizaña, M.; Gutierrez-Uribe, J.; Elías-Zúñiga, A.; Rodríguez, C. Electrospun Polycaprolactone Fibrous Membranes Containing Ag, TiO2 and Na2Ti6O13 Particles for Potential Use in Bone Regeneration. Membranes 2019, 9, 12. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Haider, M.K.; Ullah, A.; Sarwar, M.N.; Yamaguchi, T.; Wang, Q.; Ullah, S.; Park, S.; Kim, I.S. Fabricating Antibacterial and Antioxidant Electrospun Hydrophilic Polyacrylonitrile Nanofibers Loaded with AgNPs by Lignin-Induced In-Situ Method. Polymers 2021, 13, 748. [Google Scholar] [CrossRef] [PubMed]
- Haider, M.K.; Sun, L.; Ullah, A.; Ullah, S.; Suzuki, Y.; Park, S.; Kato, Y.; Tamada, Y.; Kim, I.S. Polyacrylonitrile/Carbon Black nanoparticle/Nano-Hydroxyapatite (PAN/nCB/HA) composite nanofibrous matrix as a potential biomaterial scaffold for bone regenerative applications. Mater. Today Commun. 2021, 27, 102259. [Google Scholar] [CrossRef]
- Phan, D.-N.; Dorjjugder, N.; Saito, Y.; Khan, M.Q.; Ullah, A.; Bie, X.; Taguchi, G.; Kim, I.-S. Antibacterial mechanisms of various copper species incorporated in polymeric nanofibers against bacteria. Mater. Today Commun. 2020, 25, 101377. [Google Scholar] [CrossRef]
- Haider, A.; Kwak, S.; Gupta, K.C.; Kang, I.-K. Antibacterial Activity and Cytocompatibility of PLGA/CuO Hybrid Nanofiber Scaffolds Prepared by Electrospinning. J. Nanomater. 2015, 2015, 1–10. [Google Scholar] [CrossRef]
- Hashmi, M.; Ullah, S.; Kim, I.S. Copper oxide (CuO) loaded polyacrylonitrile (PAN) nanofiber membranes for antimicrobial breath mask applications. Curr. Res. Biotechnol. 2019, 1, 1–10. [Google Scholar] [CrossRef]
- Permyakova, E.S.; Kiryukhantsev-Korneev, P.V.; Gudz, K.Y.; Konopatsky, A.S.; Polčak, J.; Zhitnyak, I.Y.; Gloushankova, N.A.; Shtansky, D.V.; Manakhov, A.M. Comparison of Different Approaches to Surface Functionalization of Biodegradable Polycaprolactone Scaffolds. Nanomaterials 2019, 9, 1769. [Google Scholar] [CrossRef] [Green Version]
- Plimpton, S. Fast Parallel Algorithms for Short-Range Molecular Dynamics. J. Comput. Phys. 1995, 117, 1–19. [Google Scholar] [CrossRef] [Green Version]
- Lloyd, A.; Cornil, D.; van Duin, A.C.T.; van Duin, D.; Smith, R.; Kenny, S.D.; Cornil, J.; Beljonne, D. Development of a ReaxFF potential for Ag/Zn/O and application to Ag deposition on ZnO. Surf. Sci. 2016, 645, 67–73. [Google Scholar] [CrossRef] [Green Version]
- Kohn, W.; Sham, L.J. Self-Consistent Equations Including Exchange and Correlation Effects. Phys. Rev. 1965, 140, A1133–A1138. [Google Scholar] [CrossRef] [Green Version]
- Hohenberg, P.; Kohn, W. Inhomogeneous Electron Gas. Phys. Rev. 1964, 136, B864–B871. [Google Scholar] [CrossRef] [Green Version]
- Kresse, G.; Hafner, J. Ab initio molecular-dynamics simulation of the liquid-metal–amorphous-semiconductor transition in germanium. Phys. Rev. B 1994, 49, 14251–14269. [Google Scholar] [CrossRef] [PubMed]
- Kresse, G.; Furthmüller, J. Efficiency of ab-initio total energy calculations for metals and semiconductors using a plane-wave basis set. Comput. Mater. Sci. 1996, 6, 15–50. [Google Scholar] [CrossRef]
- Chatani, Y.; Okita, Y.; Tadokoro, H.; Yamashita, Y. Structural Studies of Polyesters. III. Crystal Structure of Poly-ε-caprolactone. Polym. J. 1970, 1, 555–562. [Google Scholar] [CrossRef] [Green Version]
- Claros, M.; Setka, M.; Jimenez, Y.P.; Vallejos, S. AACVD Synthesis and Characterization of Iron and Copper Oxides Modified ZnO Structured Films. Nanomaterials 2020, 10, 471. [Google Scholar] [CrossRef] [Green Version]
- Parmigiani, F.; Sangaletti, L. The Cu2p X-ray photoelectron core-lines in copper oxide based high temperature superconductors. J. Electron Spectros. Relat. Phenom. 1994, 66, 223–239. [Google Scholar] [CrossRef]
- Biesinger, M.C. Advanced analysis of copper X-ray photoelectron spectra. Surf. Interface Anal. 2017, 1325–1334. [Google Scholar] [CrossRef]
- Nakamura, T.; Tomizuka, H.; Takahashi, M.; Hoshi, T. Methods of Powder Sample Mounting and Their Evaluations in XPS Analysis. Hyomen. Kagaku 1995, 16, 515–520. [Google Scholar] [CrossRef] [Green Version]
- Manakhov, A.; Kiryukhantsev-Korneev, P.; Michlíček, M.; Permyakova, E.; Dvořáková, E.; Polčák, J.; Popov, Z.; Visotin, M.; Shtansky, D.V. Grafting of carboxyl groups using CO2/C2H4/Ar pulsed plasma: Theoretical modeling and XPS derivatization. Appl. Surf. Sci. 2018, 435, 1220–1227. [Google Scholar] [CrossRef]
- Yang, Z.; Peng, H.; Wang, W.; Liu, T. Copper (II) Ions and Copper Nanoparticles-Loaded Chemically Modified Cotton Cellulose Fibers with Fair Antibacterial Properties. J. Appl. Polym. Sci. 2009, 113, 757–766. [Google Scholar]
- Cui, H.; Liu, M.; Yu, W.; Cao, Y.; Zhou, H.; Yin, J.; Liu, H.; Que, S.; Wang, J.; Huang, C.; et al. Copper Peroxide-Loaded Gelatin Sponges for Wound Dressings with Antimicrobial and Accelerating Healing Properties. ACS Appl. Mater. Interfaces 2021, 13, 26800–26807. [Google Scholar] [CrossRef]
- Fowler, L.; Engqvist, H.; Öhman-Mägi, C. Effect of Copper Ion Concentration on Bacteria and Cells. Materials 2019, 12, 3798. [Google Scholar] [CrossRef] [Green Version]
- Lei, J.; Yao, G.; Sun, Z.; Wang, B.; Yu, C.; Zheng, S. Fabrication of a novel antibacterial TPU nanofiber membrane containing Cu-loaded zeolite and its antibacterial activity toward Escherichia coli. J. Mater. Sci. 2019, 54, 11682–11693. [Google Scholar] [CrossRef]
- Chaturvedi, K.S.; Henderson, J.P. Pathogenic adaptations to host-derived antibacterial copper. Front. Cell. Infect. Microbiol. 2014, 5, 1–12. [Google Scholar] [CrossRef] [Green Version]
- Achard, M.E.S.; Tree, J.J.; Holden, J.A.; Simpfendorfer, K.R.; Wijburg, O.L.C.; Strugnell, R.A.; Schembri, M.A.; Sweet, M.J.; Jennings, M.P.; McEwan, A.G. The multi-copper-ion oxidase CueO of Salmonella enterica serovar typhimurium is required for systemic virulence. Infect. Immun. 2010, 78, 2312–2319. [Google Scholar] [CrossRef] [Green Version]
- Figueiredo, R.; Card, R.M.; Nunez-Garcia, J.; Mendonça, N.; Da Silva, G.J.; Anjum, M.F. Multidrug-Resistant Salmonella enterica Isolated from Food Animal and Foodstuff May Also Be Less Susceptible to Heavy Metals. Foodborne Pathog. Dis. 2019, 16, 166–172. [Google Scholar] [CrossRef]
- Molteni, C.; Abicht, H.K.; Solioz, M. Killing of bacteria by copper surfaces involves dissolved copper. Appl. Environ. Microbiol. 2010, 76, 4099–4101. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Harris, E.D.; Rayton, J.K.; Balthrop, J.E.; DiSilvestro, R.A.; Garcia-de-Quevedo, M. Copper and the synthesis of elastin and collagen. Ciba Found. Symp. 1980, 79, 163–182. [Google Scholar]
- Cucci, L.M.; Satriano, C.; Marzo, T.; La Mendola, D. Angiogenin and copper crossing in wound healing. Int. J. Mol. Sci. 2021, 22, 704. [Google Scholar] [CrossRef]
- Manakhov, A.; Permyakova, E.; Ershov, S.; Miroshnichenko, S.; Pykhtina, M.; Beklemishev, A.; Kovalskii, A.; Solovieva, A. XPS Modeling of Immobilized Recombinant Angiogenin and Apoliprotein A1 on Biodegradable Nanofibers. Nanomaterials 2020, 10, 879. [Google Scholar] [CrossRef]
- Solovieva, A.; Miroshnichenko, S.; Kovalskii, A.; Permyakova, E.; Popov, Z.; Dvořáková, E.; Kiryukhantsev-Korneev, P.; Obrosov, A.; Polčak, J.; Zajíčková, L.; et al. Immobilization of Platelet-Rich Plasma onto COOH Plasma-Coated PCL Nanofibers Boost Viability and Proliferation of Human Mesenchymal Stem Cells. Polymers 2017, 9, 736. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chaturvedi, K.S.; Hung, C.S.; Crowley, J.R.; Stapleton, A.E.; Henderson, J.P. The siderophore yersiniabactin binds copper to protect pathogens during infection. Nat. Chem. Biol. 2012, 8, 731–736. [Google Scholar] [CrossRef] [Green Version]

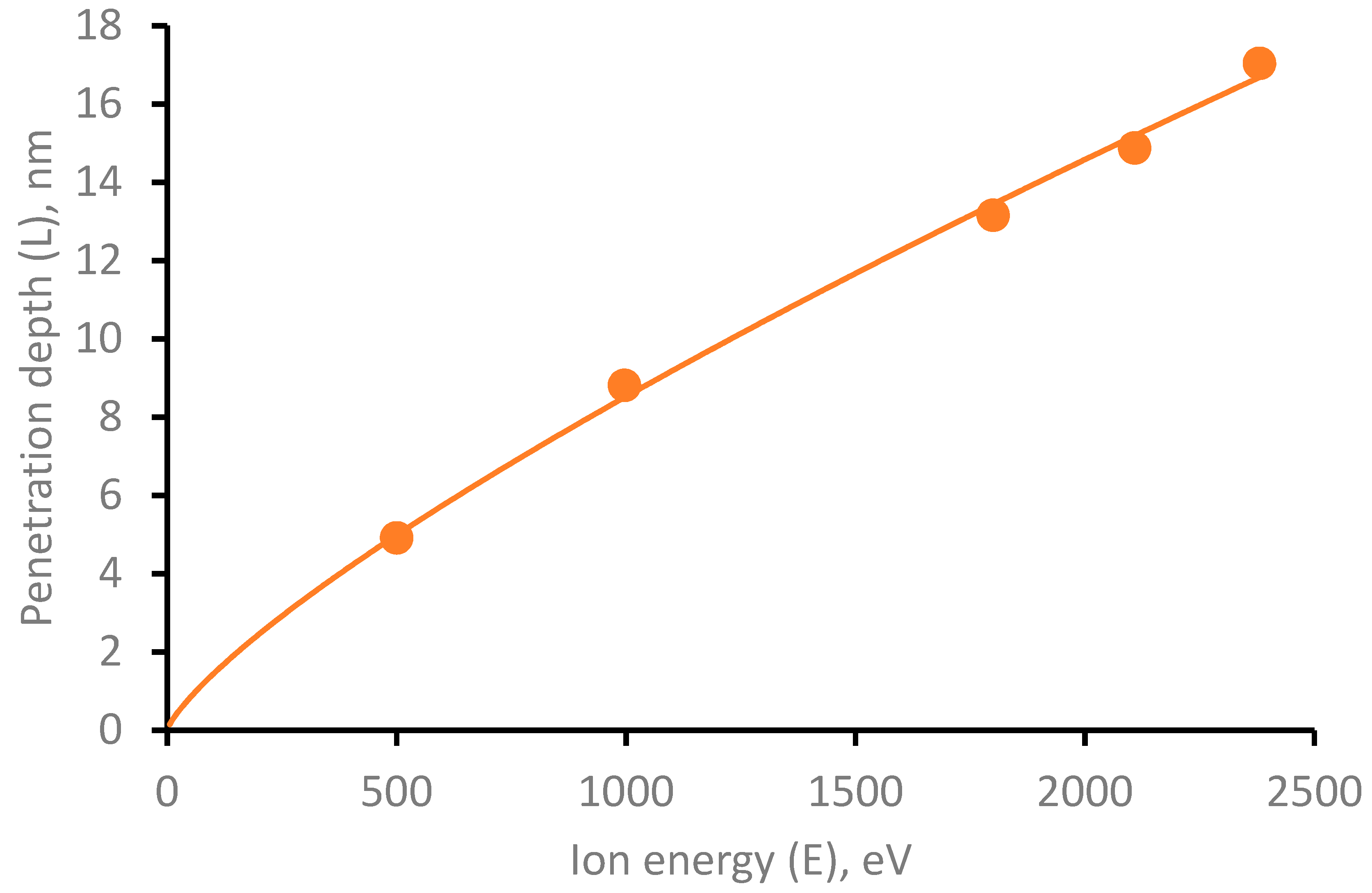


| Dimer | E, eV (PBE) | E, eV (ReaxFF) | ΔE, eV | R, Ǻ (PBE) | R, Ǻ (ReaxFF) |
|---|---|---|---|---|---|
| Cu-Cu | −2.75 | −1.38 | −1.36 | 2.22 | 2.33 |
| Cu-H | −4.27 | −2.58 | −1.69 | 1.46 | 1.53 |
| Cu-C | −4.28 | −2.65 | −1.63 | 1.76 | 1.61 |
| Cu-O | −5.25 | −4.05 | −1.21 | 1.69 | 1.75 |
| Sample Name | Cu | O | C |
|---|---|---|---|
| PCL-ref | 0.0 | 26.1 | 73.9 |
| PCL-Cu | 20.4 | 29.0 | 50.6 |
| PCL-Cu-PBS-2h | 17.2 | 35.7 | 47.1 |
| PCL-Cu-PBS-24h | 8.6 | 43.5 | 47.9 |
| PCL-COOH | 0.0 | 27.5 | 72.5 |
| PCL-Cu-COOH | 0.4 | 26.5 | 73.1 |
| Sample Name | PCL-Cu | PCL-Cu-COOH | PCL-Ref | Control | |
|---|---|---|---|---|---|
| Microorganism | |||||
| E. coli ATCC25922 | 7.9 ± 2.9 | 8.1 ± 1.4 | 67.4 ± 12.5 | 100 ± 7.9 | |
| S. aureus ATCC25923 | 12.1 ± 7.2 | 18.8 ± 4.8 | 146.9 ± 92.0 | 100 ± 34.6 | |
| S. typhimurium ATCC14028 | 95.9 ± 9.3 | 108.5 ± 8.5 | 116.4 ± 6.2 | 100 ± 8.1 | |
| PS. aeruginosa ATCC27853 | 86.3 ± 10.3 | 85.6 ± 7.5 | 143.9 ± 76.2 | 100 ± 40.7 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Manakhov, A.M.; Sitnikova, N.A.; Tsygankova, A.R.; Alekseev, A.Y.; Adamenko, L.S.; Permyakova, E.; Baidyshev, V.S.; Popov, Z.I.; Blahová, L.; Eliáš, M.; et al. Electrospun Biodegradable Nanofibers Coated Homogenously by Cu Magnetron Sputtering Exhibit Fast Ion Release. Computational and Experimental Study. Membranes 2021, 11, 965. https://doi.org/10.3390/membranes11120965
Manakhov AM, Sitnikova NA, Tsygankova AR, Alekseev AY, Adamenko LS, Permyakova E, Baidyshev VS, Popov ZI, Blahová L, Eliáš M, et al. Electrospun Biodegradable Nanofibers Coated Homogenously by Cu Magnetron Sputtering Exhibit Fast Ion Release. Computational and Experimental Study. Membranes. 2021; 11(12):965. https://doi.org/10.3390/membranes11120965
Chicago/Turabian StyleManakhov, Anton M., Natalya A. Sitnikova, Alphiya R. Tsygankova, Alexander Yu. Alekseev, Lyubov S. Adamenko, Elizaveta Permyakova, Victor S. Baidyshev, Zakhar I. Popov, Lucie Blahová, Marek Eliáš, and et al. 2021. "Electrospun Biodegradable Nanofibers Coated Homogenously by Cu Magnetron Sputtering Exhibit Fast Ion Release. Computational and Experimental Study" Membranes 11, no. 12: 965. https://doi.org/10.3390/membranes11120965
APA StyleManakhov, A. M., Sitnikova, N. A., Tsygankova, A. R., Alekseev, A. Y., Adamenko, L. S., Permyakova, E., Baidyshev, V. S., Popov, Z. I., Blahová, L., Eliáš, M., Zajíčková, L., & Solovieva, A. O. (2021). Electrospun Biodegradable Nanofibers Coated Homogenously by Cu Magnetron Sputtering Exhibit Fast Ion Release. Computational and Experimental Study. Membranes, 11(12), 965. https://doi.org/10.3390/membranes11120965










