Recombinant BCG Expressing HTI Prime and Recombinant ChAdOx1 Boost Is Safe and Elicits HIV-1-Specific T-Cell Responses in BALB/c Mice
Abstract
1. Introduction
2. Materials and Methods
2.1. Construction of the BCG.HTI2auxo.int Strain Using an Antibiotic-Free Plasmid Selection System
2.2. Bacterial Cultures and Transformation
2.3. Sodium Dodecyl Sulfate–Polyacrylamide Gel Electrophoresis and Western Blot Analysis
2.4. In Vitro Stability of the BCG.HTIint Strain
2.5. Mycobacterial Genomic DNA Preparation for the Multiplex PCR Assay and for attR and attL DNA Regions PCR
2.6. Multiplex PCR Assay for M. bovis BCG Substrain Pasteur Identification
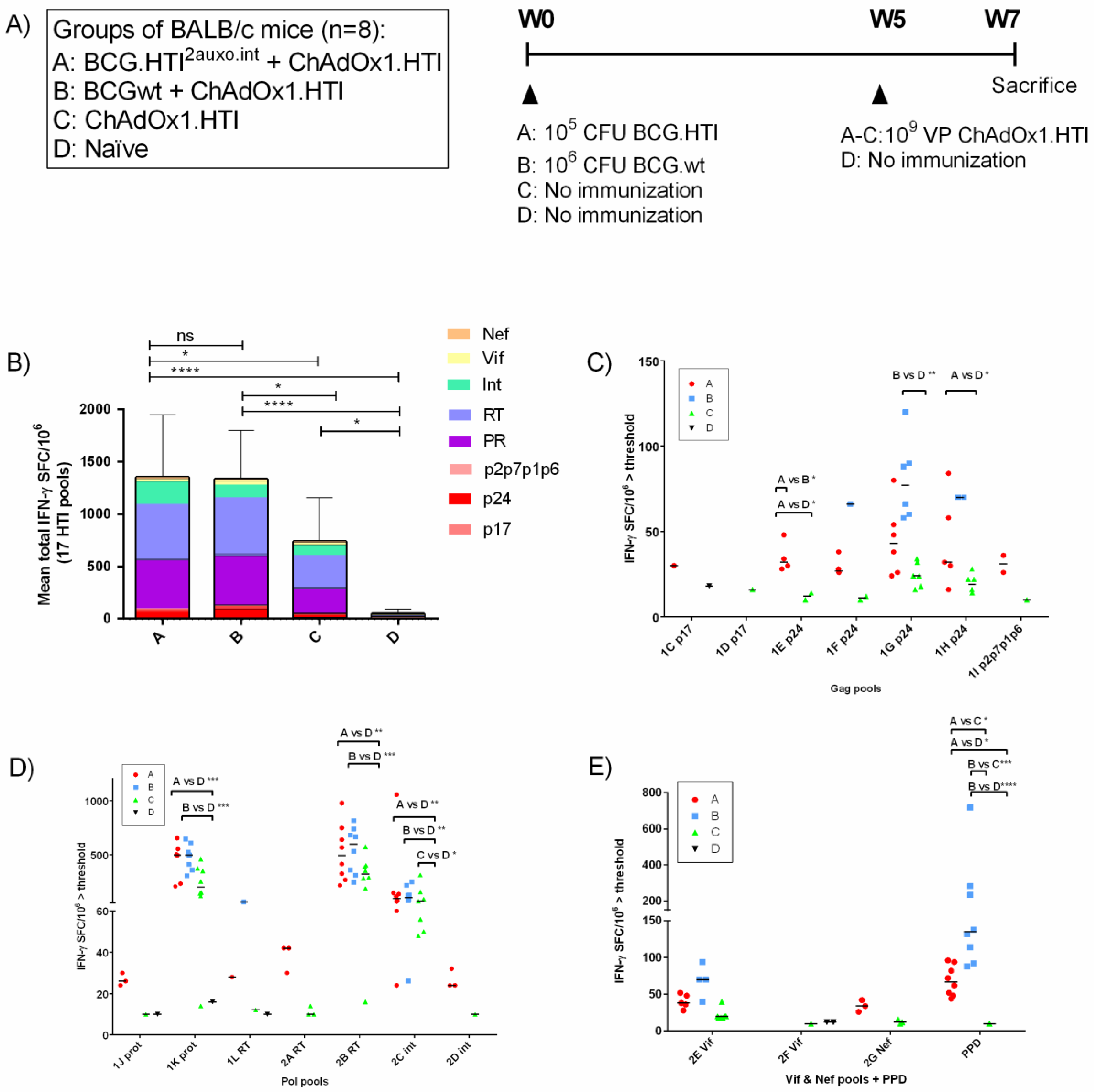
2.7. Immunization of Mice and Isolation of Splenocytes
2.8. IFN-γ ELISpot Analysis
2.9. Statistical Analysis
2.10. Ethics Statement
3. Results
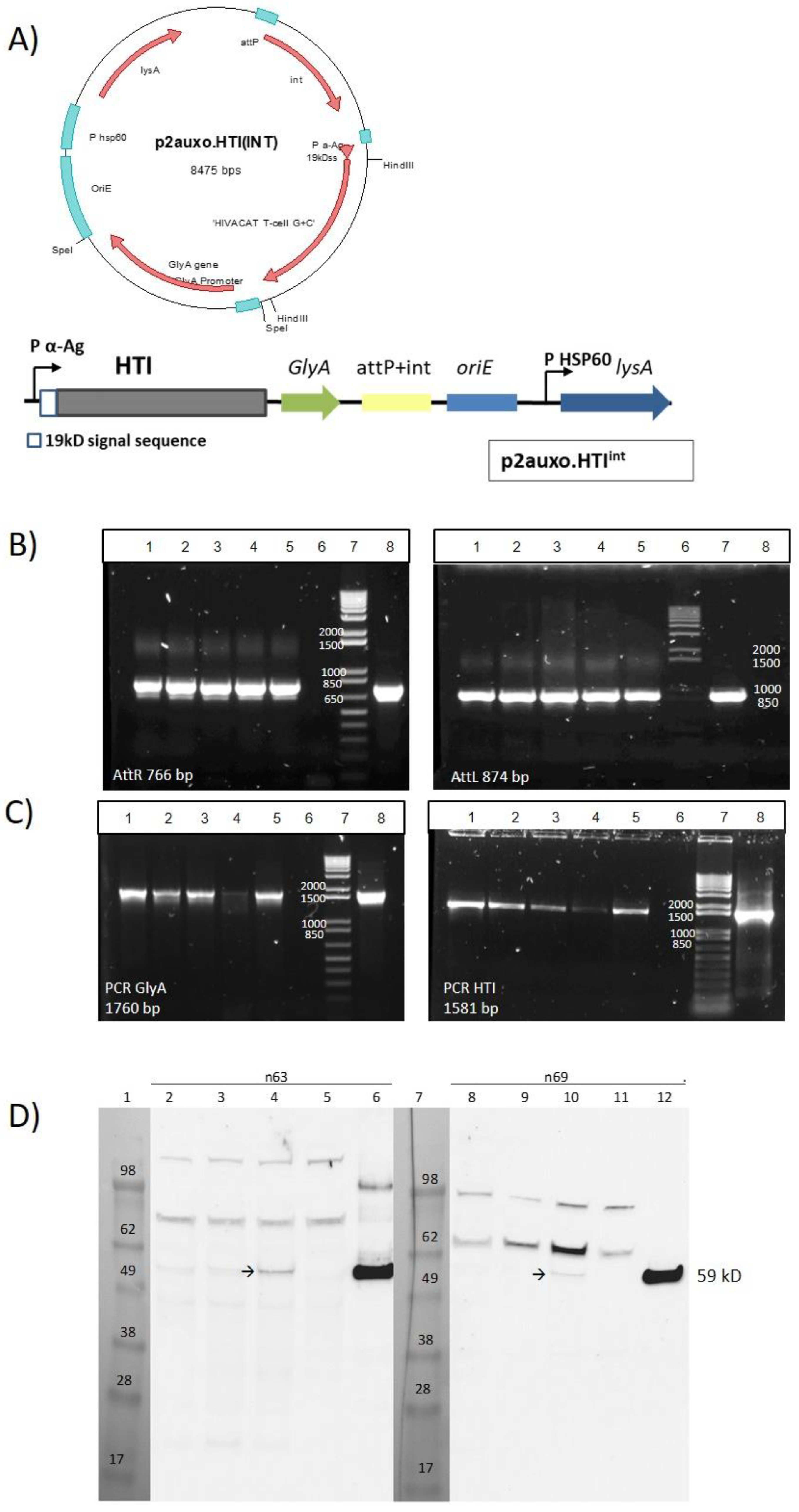
3.1. Construction of the BCG.HTI2auxo.int Vaccine Strain
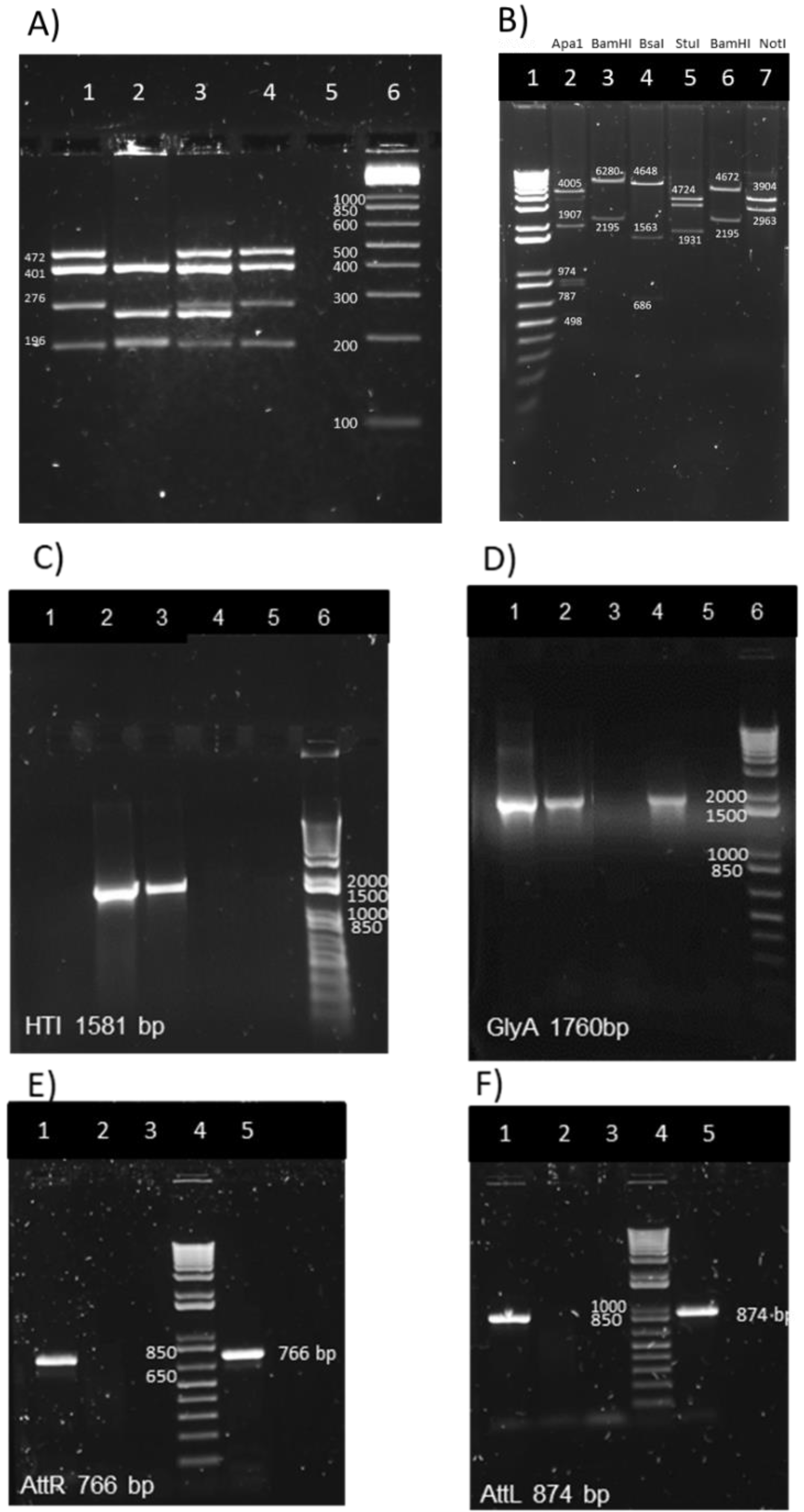
3.2. Genetic Identification and Characterization of BCG.HTI2auxo.int
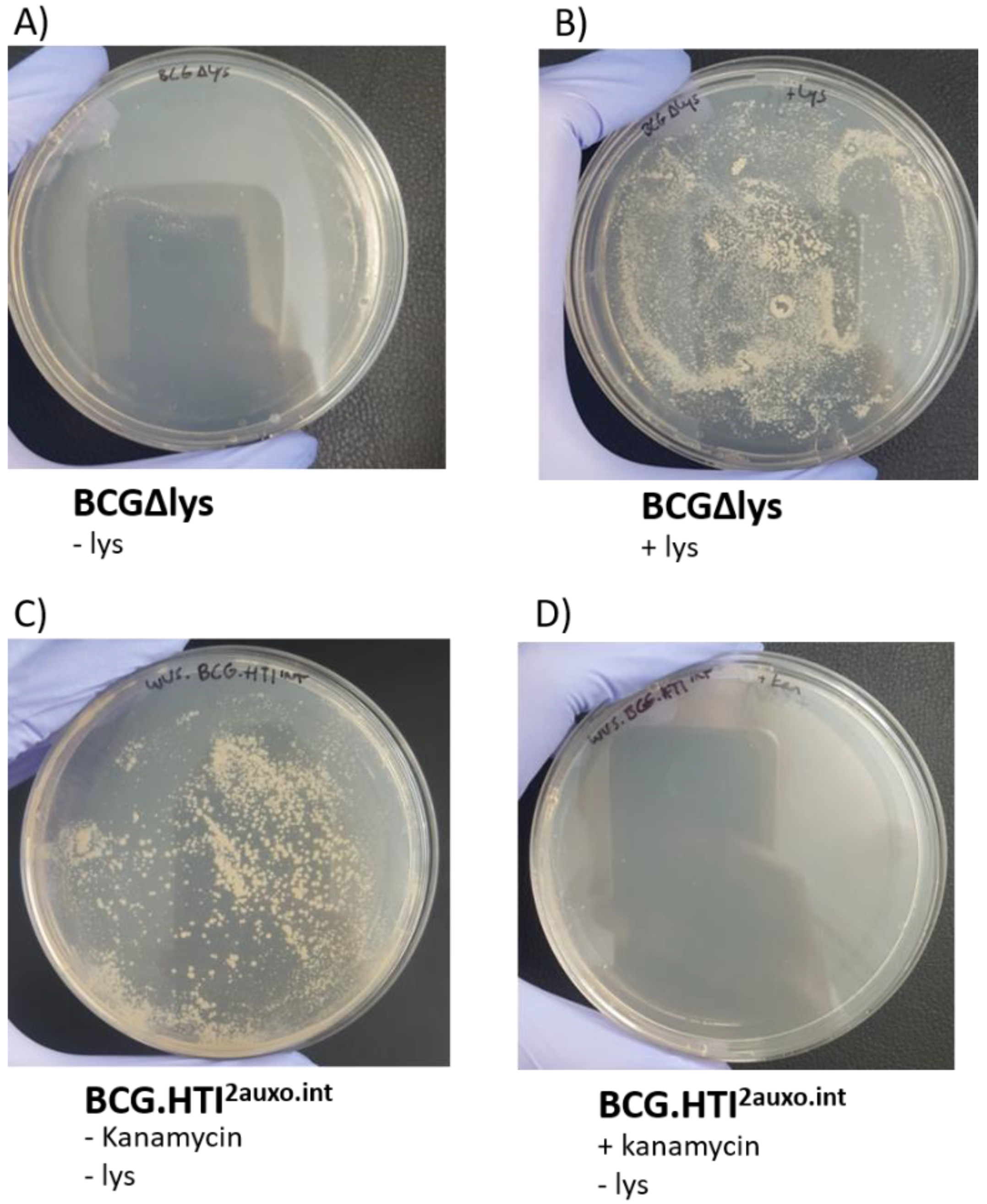
3.3. Phenotypic Characterization of BCG.HTI2auxo.int
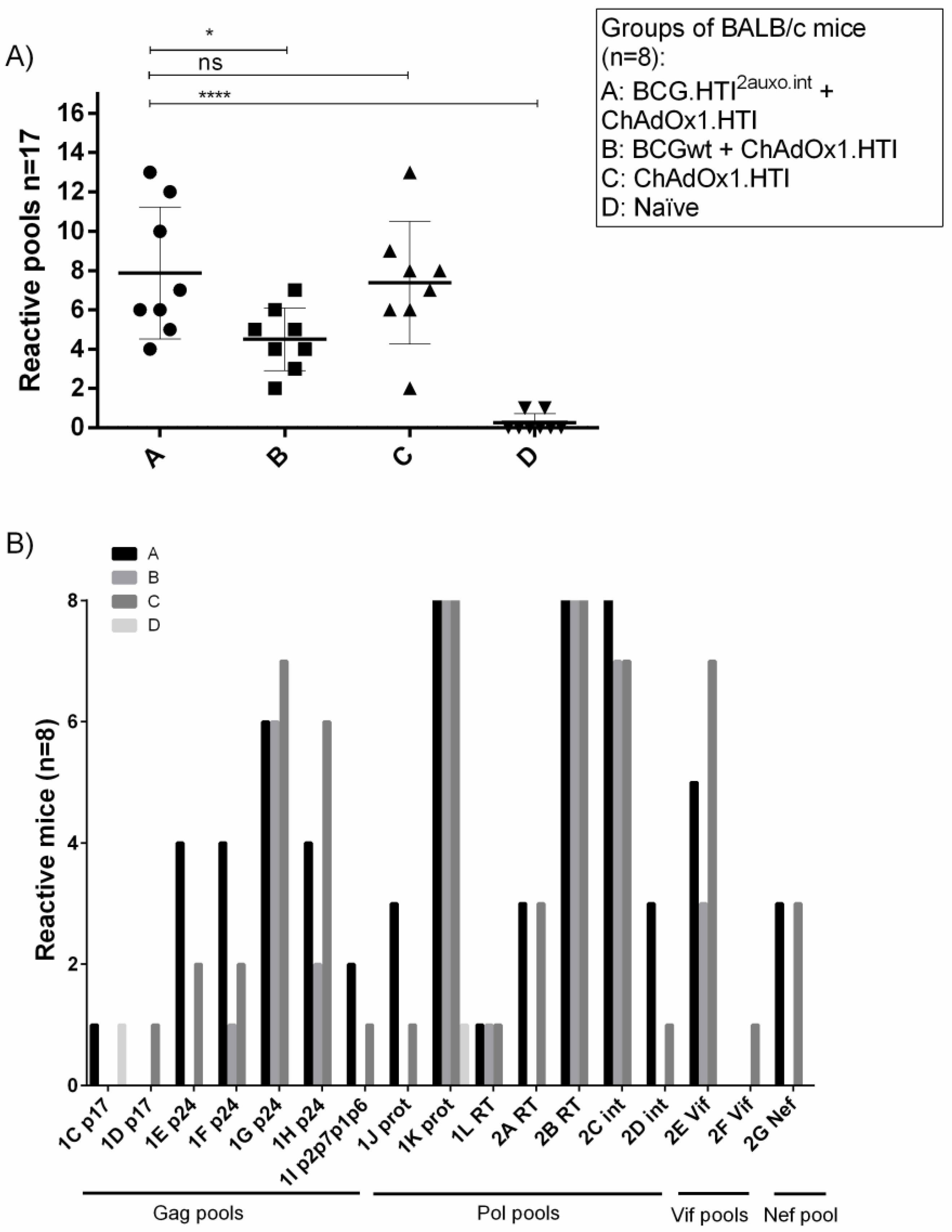
3.4. The BCG.HTI2auxo.int prime-ChAdOx1.HTI Boost Regimen Elicits HIV-1-Specific T-cell Responses
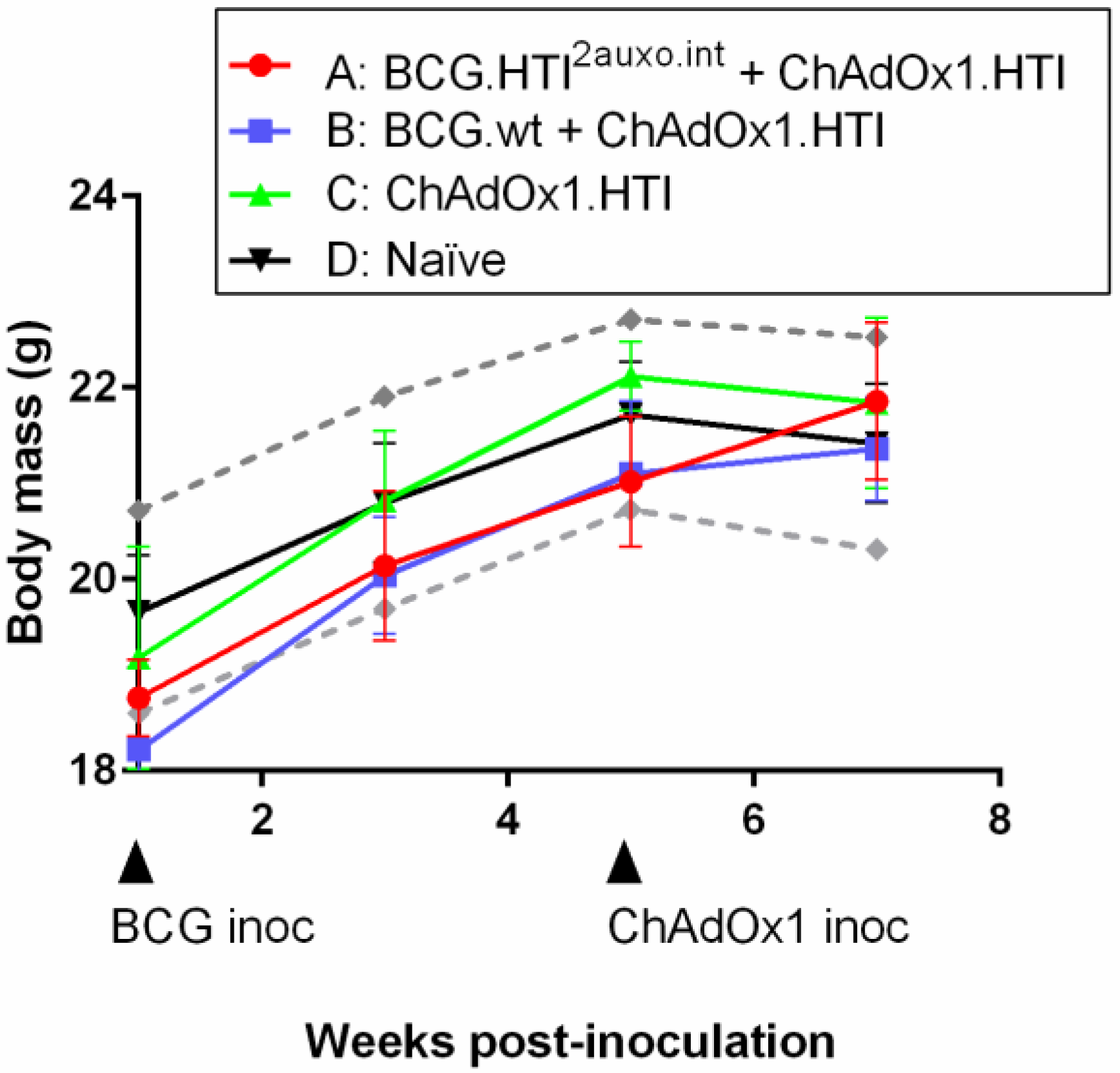
3.5. The BCG.HTI2auxo.int + ChAdOx1.HTI Prime-Boost Regimen Is Well Tolerated
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
Appendix A

References
- World Health Organization. Data and Statistics. Available online: https://www.who.int/hiv/data/2017_global_summary_web_v11.pptx (accessed on 5 February 2019).
- Rowland-Jones, S.L.; Dong, T.; Fowke, K.R.; Kimani, J.; Krausa, P.; Newell, H.; Blanchard, T.; Ariyoshi, K.; Oyugi, J.; Ngugi, E.; et al. Cytotoxic T cell responses to multiple conserved HIV epitopes in HIV-resistant prostitutes in Nairobi. J. Clin. Investig. 1998, 102, 1758–1765. [Google Scholar] [CrossRef] [PubMed]
- Koup, R.A.; Safrit, J.T.; Cao, Y.; Andrews, C.A.; McLeod, G.; Borkowsky, W.; Farthing, C.; Ho, D.D. Temporal association of cellular immune responses with the initial control of viremia in primary human immunodeficiency virus type 1 syndrome. J. Virol. 1994, 68, 4650–4655. [Google Scholar] [PubMed]
- Kuebler, P.J.; Mehrotra, M.L.; McConnell, J.J.; Holditch, S.J.; Shaw, B.I.; Tarosso, L.F.; Leadabrand, K.S.; Milush, J.M.; York, V.A.; Raposo, R.A.S.; et al. Cellular immune correlates analysis of an HIV-1 preexposure prophylaxis trial. Proc. Natl. Acad. Sci. USA 2015, 112, 8379–8384. [Google Scholar] [CrossRef] [PubMed]
- Zuñiga, R.; Lucchetti, A.; Galvan, P.; Sanchez, S.; Sanchez, C.; Hernandez, A.; Sanchez, H.; Frahm, N.; Linde, C.H.; Hewitt, H.S.; et al. Relative dominance of Gag p24-specific cytotoxic T lymphocytes is associated with human immunodeficiency virus control. J. Virol. 2006, 80, 3122–3125. [Google Scholar] [CrossRef] [PubMed]
- Janes, H.; Friedrich, D.P.; Krambrink, A.; Smith, R.J.; Kallas, E.G.; Horton, H.; Casimiro, D.R.; Carrington, M.; Geraghty, D.E.; Gilbert, P.B.; et al. Vaccine-Induced Gag-Specific T Cells Are Associated With Reduced Viremia After HIV-1 Infection. J. Infect. Dis. 2013, 208, 1231–1239. [Google Scholar] [CrossRef] [PubMed]
- Mothe, B.; Hu, X.; Llano, A.; Rosati, M.; Olvera, A.; Kulkarni, V.; Valentin, A.; Alicea, C.; Pilkington, G.R.; Sardesai, N.Y.; et al. A human immune data-informed vaccine concept elicits strong and broad T-cell specificities associated with HIV-1 control in mice and macaques. J. Transl. Med. 2015, 13, 60. [Google Scholar] [CrossRef] [PubMed]
- Iyer, S.S.; Amara, R.R. DNA/MVA Vaccines for HIV/AIDS. Vaccines 2014, 2, 160–178. [Google Scholar] [CrossRef] [PubMed]
- Donnelly, J.J.; Liu, M.A.; Ulmer, J.B. Antigen Presentation and DNA Vaccines. Am. J. Respir. Crit. Care Med. 2000, 162, S190–S193. [Google Scholar] [CrossRef] [PubMed]
- Corr, M.; Lee, D.J.; Carson, D.A.; Tighe, H. Gene vaccination with naked plasmid DNA: Mechanism of CTL priming. J. Exp. Med. 1996, 184, 1555–1560. [Google Scholar] [CrossRef]
- Van Faassen, H.; Dudani, R.; Krishnan, L.; Sad, S. Prolonged antigen presentation, APC-, and CD8+ T cell turnover during mycobacterial infection: Comparison with Listeria monocytogenes. J. Immunol. 2004, 172, 3491–3500. [Google Scholar] [CrossRef]
- Ravn, P.; Boesen, H.; Pedersen, B.K.; Andersen, P. Human T cell responses induced by vaccination with Mycobacterium bovis bacillus Calmette-Guérin. J. Immunol. 1997, 158, 1949–1955. [Google Scholar] [PubMed]
- Averill, L.E.; Cavallo, U.; Wallis, R.S.; Boom, W.H.; Bona, M.; Mincek, M.; Pascopella, L.; Jacobs, W.R.; Ellner, J.J. Screening of a cosmid library of Mycobacterium bovis BCG in Mycobacterium smegmatis for novel T-cell stimulatory antigens. Res. Microbiol. 1993, 144, 349–362. [Google Scholar] [CrossRef]
- World Health Organization. BCG Vaccine. Available online: http://www.who.int/biologicals/areas/vaccines/bcg/en/ (accessed on 6 September 2018).
- Roth, A.; Gustafson, P.; Nhaga, A.; Djana, Q.; Poulsen, A.; Garly, M.-L.; Jensen, H.; Sodemann, M.; Rodriques, A.; Aaby, P. BCG vaccination scar associated with better childhood survival in Guinea-Bissau. Int. J. Epidemiol. 2005, 34, 540–547. [Google Scholar] [CrossRef] [PubMed]
- Post, C.L.; Victora, C.G.; Valente, J.G.; Leal, M.d.C.; Niobey, F.M.; Sabroza, P.C. Prognostic factors of hospital mortality from diarrhea or pneumonia in infants younger than 1 year old. A case-control study. Rev. Saude Publica 1992, 26, 369–378. [Google Scholar] [CrossRef] [PubMed]
- Kleinnijenhuis, J.; Quintin, J.; Preijers, F.; Joosten, L.A.B.; Ifrim, D.C.; Saeed, S.; Jacobs, C.; van Loenhout, J.; de Jong, D.; Stunnenberg, H.G.; et al. Bacille Calmette-Guerin induces NOD2-dependent nonspecific protection from reinfection via epigenetic reprogramming of monocytes. Proc. Natl. Acad. Sci. USA 2012, 109, 17537–17542. [Google Scholar] [CrossRef] [PubMed]
- Kilpeläinen, A.; Maya-Hoyos, M.; Saubí, N.; Soto, C.Y.; Joseph Munne, J. Advances and challenges in recombinant Mycobacterium bovis BCG-based HIV vaccine development: Lessons learned. Expert Rev. Vaccines 2018, 17, 1005–1020. [Google Scholar] [CrossRef]
- Melancon-Kaplan, J.; Hunter, S.W.; McNeil, M.; Stewart, C.; Modlin, R.L.; Rea, T.H.; Convit, J.; Salgame, P.; Mehra, V.; Bloom, B.R. Immunological significance of Mycobacterium leprae cell walls. Proc. Natl. Acad. Sci. USA 1988, 85, 1917–1921. [Google Scholar] [CrossRef] [PubMed]
- Oiso, R.; Fujiwara, N.; Yamagami, H.; Maeda, S.; Matsumoto, S.; Nakamura, S.; Oshitani, N.; Matsumoto, T.; Arakawa, T.; Kobayashi, K. Mycobacterial trehalose 6,6’-dimycolate preferentially induces type 1 helper T cell responses through signal transducer and activator of transcription 4 protein. Microb. Pathog. 2005, 39, 35–43. [Google Scholar] [CrossRef]
- Fujita, Y.; Naka, T.; Doi, T.; Yano, I. Direct molecular mass determination of trehalose monomycolate from 11 species of mycobacteria by MALDI-TOF mass spectrometry. Microbiology 2005, 151, 1443–1452. [Google Scholar] [CrossRef]
- Brightbill, H.D.; Libraty, D.H.; Krutzik, S.R.; Yang, R.B.; Belisle, J.T.; Bleharski, J.R.; Maitland, M.; Norgard, M.V.; Plevy, S.E.; Smale, S.T.; et al. Host defense mechanisms triggered by microbial lipoproteins through toll-like receptors. Science 1999, 285, 732–736. [Google Scholar] [CrossRef]
- Gheorghiu, M.; Lagrange, P.H.; Fillastre, C. The stability and immunogenicity of a dispersed-grown freeze-dried Pasteur BCG vaccine. J. Biol. Stand. 1988, 16, 15–26. [Google Scholar] [CrossRef]
- Matsuo, K.; Yasutomi, Y. Mycobacterium bovis Bacille Calmette-Guérin as a Vaccine Vector for Global Infectious Disease Control. Tuberc. Res. Treat. 2011, 2011, 9. [Google Scholar]
- Joseph, J.; Saubi, N.; Pezzat, E.; Gatell, J.M. Progress towards an HIV vaccine based on recombinant Bacillus Calmette–Guérin: Failures and challenges. Expert Rev. Vaccines 2006, 5, 827–838. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization BCG vaccine. WHO position paper. Relev. Epidemiol. Hebd. 2004, 79, 27–38. [Google Scholar]
- Hopkins, R.; Bridgeman, A.; Joseph, J.; Gilbert, S.C.; McShane, H.; Hanke, T. Dual neonate vaccine platform against HIV-1 and M. tuberculosis. PLoS ONE 2011, 6, e20067. [Google Scholar] [CrossRef] [PubMed]
- Hopkins, R.; Bridgeman, A.; Bourne, C.; Mbewe-Mvula, A.; Sadoff, J.C.; Both, G.W.; Joseph, J.; Fulkerson, J.; Hanke, T. Optimizing HIV-1-specific CD8+ T-cell induction by recombinant BCG in prime-boost regimens with heterologous viral vectors. Eur. J. Immunol. 2011, 41, 3542–3552. [Google Scholar] [CrossRef]
- Chapman, R.; Stutz, H.; Jacobs, W.; Shephard, E.; Williamson, A.L.L. Priming with Recombinant Auxotrophic BCG Expressing HIV-1 Gag, RT and Gp120 and Boosting with Recombinant MVA Induces a Robust T Cell Response in Mice. PLoS ONE 2013, 8, 8. [Google Scholar] [CrossRef]
- Hart, B.E.; Asrican, R.; Lim, S.Y.; Sixsmith, J.D.; Lukose, R.; Souther, S.J.R.; Rayasam, S.D.G.; Saelens, J.W.; Chen, C.J.; Seay, S.A.; et al. Stable Expression of Lentiviral Antigens by Quality-Controlled Recombinant Mycobacterium bovis BCG Vectors. Clin. Vaccine Immunol. 2015, 22, 726–741. [Google Scholar] [CrossRef]
- Yu, J.S.; Peacock, J.W.; Jacobs, W.R., Jr.; Frothingham, R.; Letvin, N.L.; Liao, H.X.; Haynes, B.F. Recombinant Mycobacterium bovis Bacillus Calmette-Guérin Elicits Human Immunodeficiency Virus Type 1 Envelope-Specific T Lymphocytes at Mucosal Sites. Clin. Vaccine Immunol. 2007, 14, 886. [Google Scholar] [CrossRef]
- Ami, Y.; Izumi, Y.; Matsuo, K.; Someya, K.; Kanekiyo, M.; Horibata, S.; Yoshino, N.; Sakai, K.; Shinohara, K.; Matsumoto, S.; et al. Priming-boosting vaccination with recombinant Mycobacterium bovis bacillus Calmette-Guérin and a nonreplicating vaccinia virus recombinant leads to long-lasting and effective immunity. J. Virol. 2005, 79, 12871–12879. [Google Scholar] [CrossRef]
- Mahant, A.; Saubi, N.; Eto, Y.; Guitart, N.; Gatell, J.M.; Hanke, T.; Joseph, J. Preclinical development of BCG.HIVA2auxo.int, harboring an integrative expression vector, for a HIV-TB Pediatric vaccine. Enhancement of stability and specific HIV-1 T-cell immunity. Hum. Vaccines Immunother. 2017, 13, 1798–1810. [Google Scholar] [CrossRef]
- Kanekiyo, M.; Matsuo, K.; Hamatake, M.; Hamano, T.; Ohsu, T.; Matsumoto, S.; Yamada, T.; Yamazaki, S.; Hasegawa, A.; Yamamoto, N.; et al. Mycobacterial Codon Optimization Enhances Antigen Expression and Virus-Specific Immune Responses in Recombinant Mycobacterium bovis Bacille Calmette-Guerin Expressing Human Immunodeficiency Virus Type 1 Gag. J. Virol. 2005, 79, 8716–8723. [Google Scholar] [CrossRef]
- Bedwell, J.; Kairo, S.K.; Behr, M.A.; Bygraves, J.A. Identification of substrains of BCG vaccine using multiplex PCR. Vaccine 2001, 19, 2146–2151. [Google Scholar] [CrossRef]
- Ondondo, B.; Murakoshi, H.; Clutton, G.; Abdul-Jawad, S.; Wee, E.G.T.; Gatanaga, H.; Oka, S.; McMichael, A.J.; Takiguchi, M.; Korber, B.; et al. Novel Conserved-region T-cell Mosaic Vaccine With High Global HIV-1 Coverage Is Recognized by Protective Responses in Untreated Infection. Mol. Ther. 2016, 24, 832–842. [Google Scholar] [CrossRef]
- Joseph, J.; Fernández-Lloris, R.; Pezzat, E.; Saubi, N.; Cardona, P.J.; Mothe, B.; Gatell, J.M. Molecular characterization of heterologous HIV-1gp120 gene expression disruption in mycobacterium bovis BCG host strain: A critical issue for engineering Mycobacterial based-vaccine vectors. J. Biomed. Biotechnol. 2010, 2010, 1–10. [Google Scholar] [CrossRef]
- Chapman, R.; Chege, G.; Shephard, E.; Stutz, H.; Williamson, A.L. Recombinant Mycobacterium bovis BCG as an HIV vaccine vector. Curr. HIV Res. 2010, 8, 282–298. [Google Scholar] [CrossRef]
- Im, E.J.; Saubi, N.; Virgili, G.; Sander, C.; Teoh, D.; Gatell, J.M.; McShane, H.; Joseph, J.; Hanke, T. Vaccine platform for prevention of tuberculosis and mother-to-child transmission of human immunodeficiency virus type 1 through breastfeeding. J. Virol. 2007, 81, 9408–9418. [Google Scholar] [CrossRef]
- Saubi, N.; Gea-Mallorquí, E.; Ferrer, P.; Hurtado, C.; Sánchez-Úbeda, S.; Eto, Y.; Gatell, J.M.; Hanke, T.; Joseph, J. Engineering new mycobacterial vaccine design for HIV-TB pediatric vaccine vectored by lysine auxotroph of BCG. Mol. Ther. Methods Clin. Dev. 2014, 1, 14017. [Google Scholar] [CrossRef]
- Pavelka, M.S.; Jacobs, W.R. Comparison of the construction of unmarked deletion mutations in Mycobacterium smegmatis, Mycobacterium bovis bacillus Calmette-Guerin, and Mycobacterium tuberculosis H37RV by allelic exchange. J. Bacteriol. 1999, 181, 4780–4789. [Google Scholar]
- Lee, M.H.; Pascopella, L.; Jacobs, W.R.; Hatfull, G.F. Site-specific integration of mycobacteriophage L5: Integration-proficient vectors for Mycobacterium smegmatis, Mycobacterium tuberculosis, and bacille Calmette-Guérin. Proc. Natl. Acad. Sci. USA 1991, 88, 3111–3115. [Google Scholar] [CrossRef]
- Kilpeläinen, A.; Saubi, N.; Guitart, N.; Moyo, N.; Wee, E.G.; Hanke, T.H.; Joseph, J. Priming with recombinant BCG expressing novel HIV-1 mosaic immunogens and boosting with recombinant ChAdOx1 is safe, stable and elicits HIV-1-specific T-cell responses in BALB/c mice. Front. Immunol. 2019, 10, 923. [Google Scholar] [CrossRef]
- World Health Organization. Global HIV & AIDS Statistics—2018 Fact Sheet UNAIDS. Available online: http://www.unaids.org/en/resources/fact-sheet (accessed on 8 October 2018).
- Segueni, N.; Tritto, E.; Bourigault, M.L.; Rose, S.; Erard, F.; Le Bert, M.; Jacobs, M.; Di Padova, F.; Stiehl, D.P.; Moulin, P.; et al. Controlled Mycobacterium tuberculosis infection in mice under treatment with anti-IL-17A or IL-17F antibodies, in contrast to TNFα neutralization. Sci. Rep. 2016, 6, 36923. [Google Scholar] [CrossRef]
- Cohen, G.B.; Gandhi, R.T.; Davis, D.M.; Mandelboim, O.; Chen, B.K.; Strominger, J.L.; Baltimore, D. The selective downregulation of class I major histocompatibility complex proteins by HIV-1 protects HIV-infected cells from NK cells. Immunity 1999, 10, 661–671. [Google Scholar] [CrossRef]
- Institute, A.N.R. HLA Class I and Class II Sequence Alignments. Updates. Available online: http://www.ebi.ac.uk/ipd/imgt/hla/ (accessed on 10 July 2019).
- Joosten, S.A.; van Meijgaarden, K.E.; van Weeren, P.C.; Kazi, F.; Geluk, A.; Savage, N.D.L.; Drijfhout, J.W.; Flower, D.R.; Hanekom, W.A.; Klein, M.R.; et al. Mycobacterium tuberculosis Peptides Presented by HLA-E Molecules Are Targets for Human CD8+ T-Cells with Cytotoxic as well as Regulatory Activity. PLoS Pathog. 2010, 6, e1000782. [Google Scholar] [CrossRef]
- Hansen, S.G.; Ford, J.C.; Lewis, M.S.; Ventura, A.B.; Hughes, C.M.; Coyne-Johnson, L.; Whizin, N.; Oswald, K.; Shoemaker, R.; Swanson, T.; et al. Profound early control of highly pathogenic SIV by an effector memory T-cell vaccine. Nature 2011, 473, 523–527. [Google Scholar] [CrossRef]
- Tomasec, P.; Braud, V.M.; Rickards, C.; Powell, M.B.; McSharry, B.P.; Gadola, S.; Cerundolo, V.; Borysiewicz, L.K.; McMichael, A.J.; Wilkinson, G.W. Surface expression of HLA-E, an inhibitor of natural killer cells, enhanced by human cytomegalovirus gpUL40. Science 2000, 287, 1031. [Google Scholar] [CrossRef]
- Andrieu, J.M.; Chen, S.; Lai, C.; Guo, W.; Lu, W. Mucosal SIV Vaccines Comprising Inactivated Virus Particles and Bacterial Adjuvants Induce CD8+ T-Regulatory Cells that Suppress SIV-Positive CD4+ T-Cell Activation and Prevent SIV Infection in the Macaque Model. Front. Immunol. 2014, 5, 297. [Google Scholar] [CrossRef]
- Jongwe, T.I.; Chapman, R.; Douglass, N.; Chetty, S.; Chege, G.; Williamson, A.L. HIV-1 Subtype C Mosaic Gag Expressed by BCG and MVA Elicits Persistent Effector T Cell Responses in a Prime-Boost Regimen in Mice. PLoS ONE 2016, 11, e0159141. [Google Scholar] [CrossRef]
- Sun, J.; Hou, J.; Li, D.; Liu, Y.; Hu, N.; Hao, Y.; Fu, J.; Hu, Y.; Shao, Y. Enhancement of HIV-1 DNA vaccine immunogenicity by BCG-PSN, a novel adjuvant. Vaccine 2013, 31, 472–479. [Google Scholar] [CrossRef]
- Van der Meer, J.W.M.; Joosten, L.A.B.; Riksen, N.; Netea, M.G. Trained immunity: A smart way to enhance innate immune defence. Mol. Immunol. 2015, 68, 40–44. [Google Scholar] [CrossRef]
- Kleinnijenhuis, J.; Quintin, J.; Preijers, F.; Joosten, L.A.B.; Jacobs, C.; Xavier, R.J.; van der Meer, J.W.M.; van Crevel, R.; Netea, M.G. BCG-induced trained immunity in NK cells: Role for non-specific protection to infection. Clin. Immunol. 2014, 155, 213–219. [Google Scholar] [CrossRef]
- Haynes, B.F.; Gilbert, P.B.; McElrath, M.J.; Zolla-Pazner, S.; Tomaras, G.D.; Alam, S.M.; Evans, D.T.; Montefiori, D.C.; Karnasuta, C.; Sutthent, R.; et al. Immune-correlates analysis of an HIV-1 vaccine efficacy trial. N. Engl. J. Med. 2012, 366, 1275–1286. [Google Scholar] [CrossRef]
- Lin, L.; Finak, G.; Ushey, K.; Seshadri, C.; Hawn, T.R.; Frahm, N.; Scriba, T.J.; Mahomed, H.; Hanekom, W.; Bart, P.A.; et al. COMPASS identifies T-cell subsets correlated with clinical outcomes. Nat. Biotechnol. 2015, 33, 610–616. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kilpeläinen, A.; Saubi, N.; Guitart, N.; Olvera, A.; Hanke, T.; Brander, C.; Joseph, J. Recombinant BCG Expressing HTI Prime and Recombinant ChAdOx1 Boost Is Safe and Elicits HIV-1-Specific T-Cell Responses in BALB/c Mice. Vaccines 2019, 7, 78. https://doi.org/10.3390/vaccines7030078
Kilpeläinen A, Saubi N, Guitart N, Olvera A, Hanke T, Brander C, Joseph J. Recombinant BCG Expressing HTI Prime and Recombinant ChAdOx1 Boost Is Safe and Elicits HIV-1-Specific T-Cell Responses in BALB/c Mice. Vaccines. 2019; 7(3):78. https://doi.org/10.3390/vaccines7030078
Chicago/Turabian StyleKilpeläinen, Athina, Narcís Saubi, Núria Guitart, Alex Olvera, Tomáš Hanke, Christian Brander, and Joan Joseph. 2019. "Recombinant BCG Expressing HTI Prime and Recombinant ChAdOx1 Boost Is Safe and Elicits HIV-1-Specific T-Cell Responses in BALB/c Mice" Vaccines 7, no. 3: 78. https://doi.org/10.3390/vaccines7030078
APA StyleKilpeläinen, A., Saubi, N., Guitart, N., Olvera, A., Hanke, T., Brander, C., & Joseph, J. (2019). Recombinant BCG Expressing HTI Prime and Recombinant ChAdOx1 Boost Is Safe and Elicits HIV-1-Specific T-Cell Responses in BALB/c Mice. Vaccines, 7(3), 78. https://doi.org/10.3390/vaccines7030078




