Gas Vesicle Nanoparticles for Antigen Display
Abstract
:1. Introduction
2. Biology of GVNPs
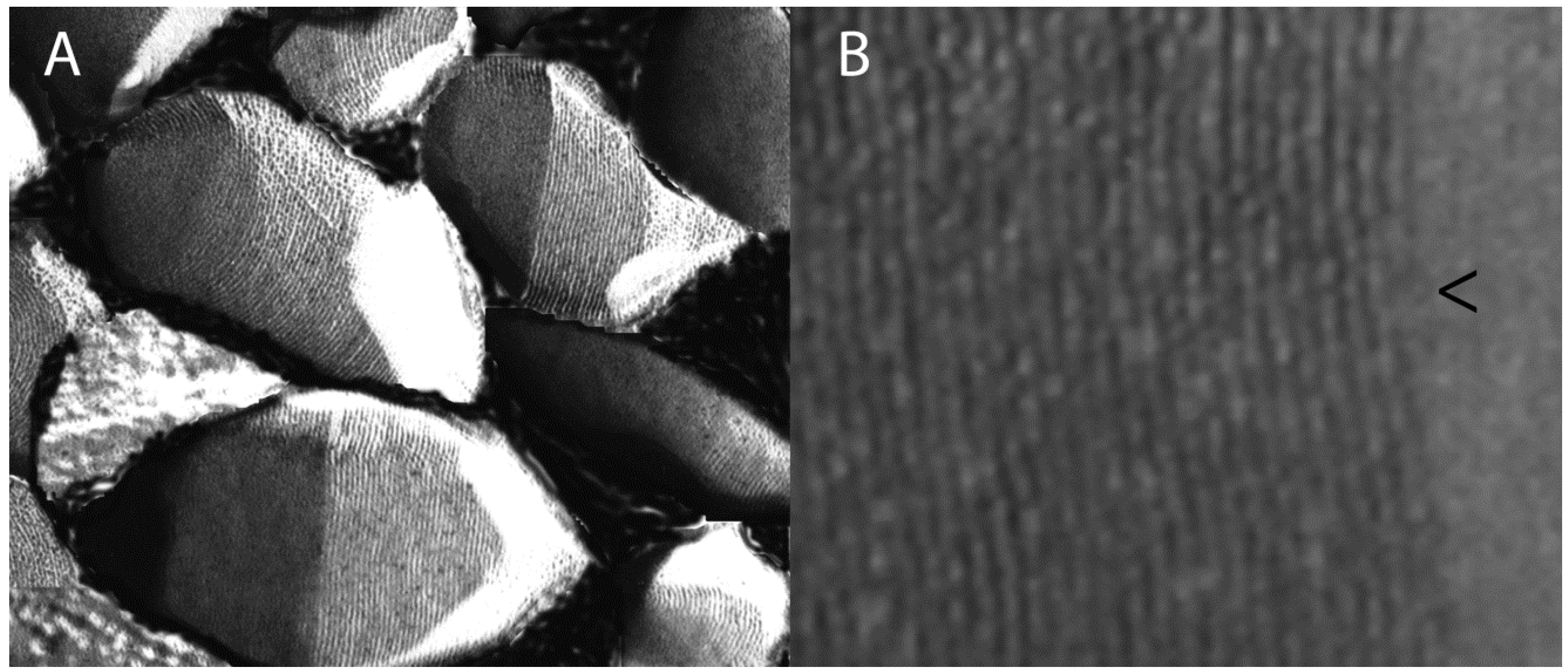

3. Gvp Gene Cluster
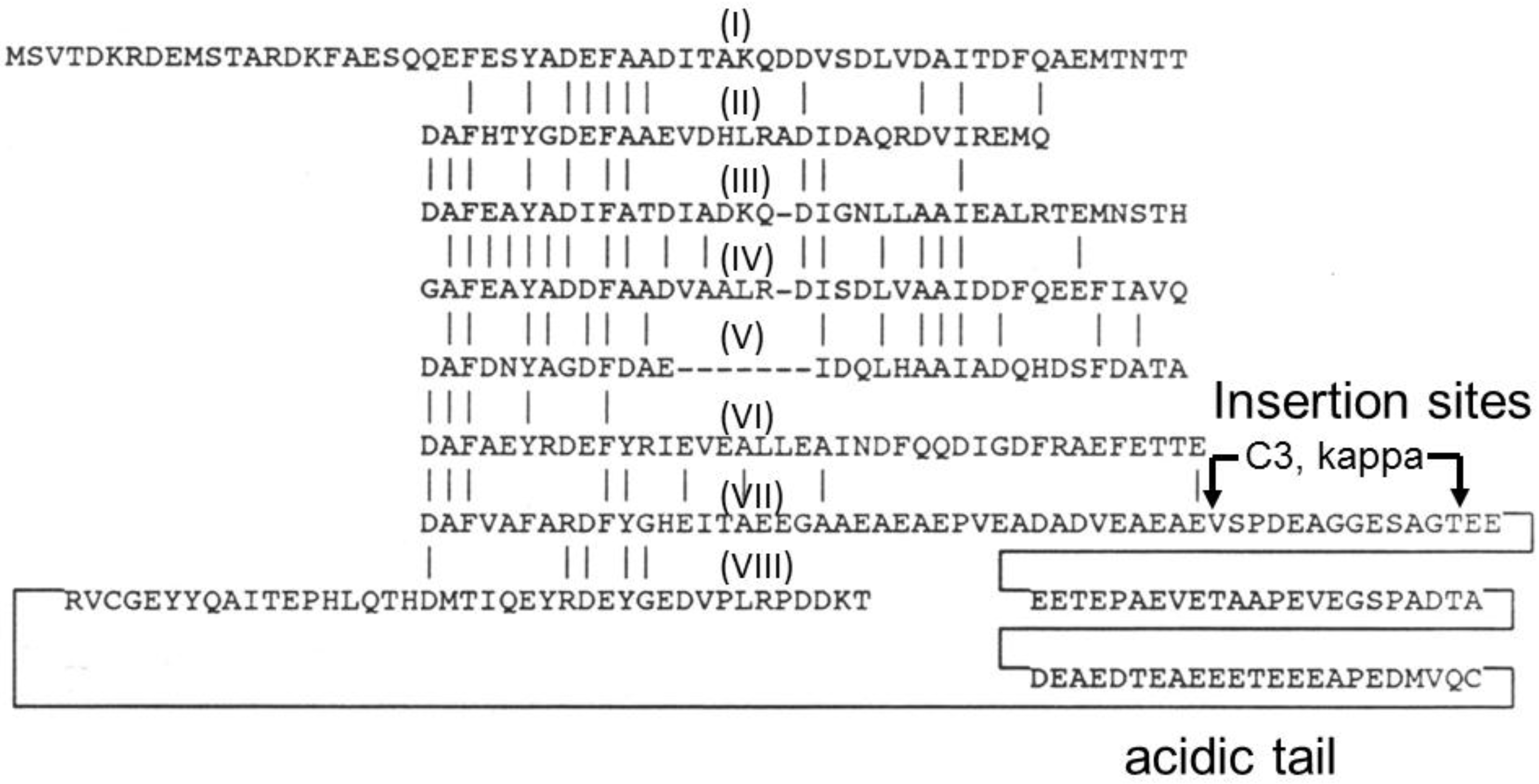
4. Gas Vesicle Proteins
| Name | MW | Functions and Characteristics |
|---|---|---|
| GvpA | 8005 | Major gas vesicle protein with predicted coil-α-β-β-α-coil fold (pfam741) |
| GvpC | 42,391 | Minor gas vesicle protein with 8 repeats and acidic tail |
| GvpD | 59,341 | Probable regulatory protein with predicted NTP binding motif |
| GvpE | 21,009 | Probable regulatory protein with possible leucine zipper domain |
| GvpF | 23,962 | Minor gas vesicle protein with coiled-coil domain (pfam 6386) |
| GvpG | 10,014 | Minor gas vesicle protein |
| GvpH | 19,883 | Predicted gene product of unknown function |
| GvpI | 16,259 | Minor gas vesicle protein of unknown function |
| GvpJ | 11,983 | Minor gas vesicle protein similar to GvpA and GvpM (pfam741) |
| GvpK | 12,695 | Predicted gene product with slight similarity to GvpC |
| GvpL | 31,994 | Minor gas vesicle protein with coiled-coil domain, laddering (pfam 6386) |
| GvpM | 9248 | Minor gas vesicle protein similar to GvpA and GvpJ (pfam741) |
| GvpN | 39,228 | Minor gas vesicle protein with NTP/AAA+ family motif |
| GvpO | 13,232 | Minor gas vesicle protein with regulatory or structural function |
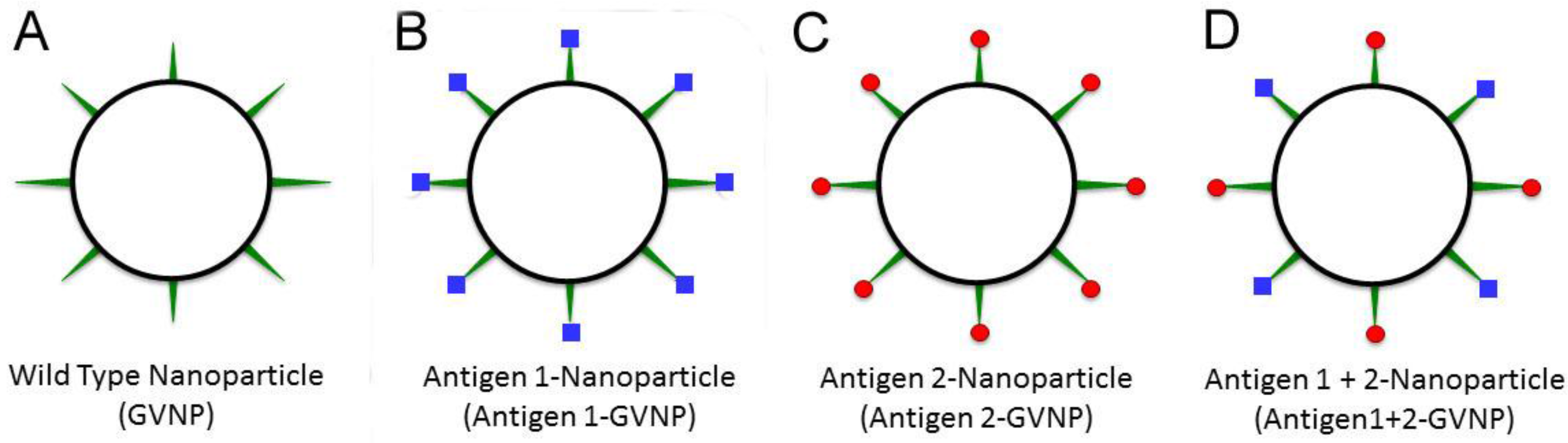
5. Antigen Display on Gas Vesicle Nanoparticles
| Antigen Displayed | Halobacterium sp. strain(s) designation | Antigen AA Length(s) | Administration Route(s) | Response(s) |
|---|---|---|---|---|
| None (Wild-type) | NRC-1 (pFM104d) or SD109 (pMS104) | N/A | IP, CC | TNF-α, IL-6, IL-12, TLR4, TLR5 |
| TNP hapten | NRC-1 | N/A | IP | IgG |
| Hexapeptide ESSGTF | SD109 (pFL2gvpC::κΔ) | 6 | IP | IgM, IgG |
| SIV Gag fragments | SD109 (pFM101d::51, 504, 705) | 17, 168, 235 | IP | IgG, mB-cell activation |
| SIV Tat, Rev, Nef1 | SD109 (pMS104d::tat, rev, nef1) | 50, 81, 214 | SC | IgG1, mB-cell activation, IL-10, IL-12, IL-18 (except Nef1) |
| C. trachomatis MOMP fragments VD3, VD4 | SD109 (pMS104::VD3 *, VD4) | 48, 69 | CC | TNF-α, IL1-β *, IL-6, IL-12, TLR4, TLR5 * |
| C. trachomatis OmcB fragments 23, 420 | SD109 (pMS104::OmcB23 †, B420) | 162, 144 | CC | TNF-α, IL-6, IL-12, TLR4, TLR5 † |
| C. trachomatis PompD fragments M1, N2 | SD109 (pMS104::PompDM1, DN2 ‡) | 173, 222 | CC | TNF-α, IL-6 ‡, IL-12 ‡, TLR4 |
| S. enterica SopB fragments 4, 5 | SD109 (pSD104::SopB4, B5) | 101, 167 | IP (B5 only) | CD4+ T-cells, IFNγ, IL-2, IL-9 |
| P. falciparum CSP | SD109 (SDcsp20), Δura3ΔgvpC (pDRKcsp6) | 398 | N/T | N/T |
| P. falciparum Enolase | SD109 (SD104::Eno) | 15 | IP | IgG |
| G. princeps Luciferase | NRC-1, Δura3ΔgvpC (pDRKC3-luc) | 185 | N/T | N/T |
6. GVNP Bioengineering System
7. Future Opportunities and Challenges
8. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Storni, T.; Kqndig, T.M.; Senti, G.; Johansen, P. Immunity in response to particulate antigen-delivery systems. Adv. Drug Deliv. Rev. 2005, 57, 333–355. [Google Scholar] [CrossRef] [PubMed]
- Perrie, Y.; Mohammed, A.R.; Kirby, D.J.; McNeil, S.E.; Bramwell, V.W. Vaccine adjuvant systems: Enhancing the efficacy of sub-unit protein antigens. Int. J. Pharm. 2008, 364, 272–280. [Google Scholar] [CrossRef] [PubMed]
- Foged, C. Subunit vaccines of the future: The need for safe, customized and optimized particulate delivery systems. Ther. Deliv. 2001, 2, 1057–1077. [Google Scholar] [CrossRef]
- Zhao, L.; Seth, A.; Wibowo, N.; Zhao, C.X.; Mitter, N.; Yu, C.; Middelberg, A.P. Nanoparticle vaccines. Vaccine 2014, 32, 327–337. [Google Scholar] [CrossRef] [PubMed]
- Stuart, E.S.; Morshed, F.; Sremac, M.; DasSarma, S. Antigen presentation using novel particulate organelles from halophilic archaea. J. Biotechnol. 2001, 88, 119–128. [Google Scholar] [CrossRef]
- Stuart, E.S.; Morshed, F.; Sremac, M.; DasSarma, S. Cassette-based presentation of SIV epitopes with recombinant gas vesicles from halophilic archaea. J. Biotechnol. 2004, 114, 225–237. [Google Scholar] [CrossRef] [PubMed]
- Sremac, M.; Stuart, E.S. Recombinant gas vesicles from Halobacterium sp. displaying SIV peptides demonstrate biotechnology potential as a pathogen peptide delivery vehicle. BMC Biotechnol. 2008. [Google Scholar] [CrossRef] [PubMed]
- Sremac, M.; Stuart, E.S. SIVsm Tat, Rev, and Nef1: Functional characteristics of r-GV internalization on isotypes, cytokines, and intracellular degradation. BMC Biotechnol. 2010. [Google Scholar] [CrossRef] [PubMed]
- Childs, T.S.; Webley, W.C. In vitro assessment of halobacterial gas vesicles as a Chlamydia vaccine display and delivery system. Vaccine 2012, 30, 5942–5948. [Google Scholar] [CrossRef] [PubMed]
- DasSarma, P.; Negi, V.D.; Balakrishnan, A.; Karan, R.; Barnes, S.; Ekulona, F.; Chakravortty, D.; DasSarma, S. Haloarchaeal gas vesicle nanoparticles displaying Salmonella SopB antigen reduce bacterial burden when administered with live attenuated bacteria. Vaccine 2014, 32, 4543–4549. [Google Scholar] [CrossRef] [PubMed]
- DasSarma, P.; Negi, V.D.; Balakrishnan, A.; Kim, J.-M.; Karan, R.; Chakravortty, D.; DasSarma, S. Haloarchaeal gas vesicle nanoparticles displaying Salmonella antigens as a novel approach to vaccine development. Procedia Vaccinol. 2015, 9, 16–23. [Google Scholar] [CrossRef]
- Pecher, W.T.; Kim, J.-M.; DasSarma, P.; Karan, R.; Sinnis, P.; DasSarma, S. Halobacterium Expression System for Production of Full-Length Plasmodium Falciparum Circumsporozoite Protein. In Biotechnology of Extremophiles: Advances and Challenges; Springer-Verlag Berlin: Heidelberg, Germany, in press.
- Shukla, H.D.; DasSarma, S. Complexity of gas vesicle biogenesis in Halobacterium sp. strain NRC-1: Identification of five new proteins. J. Bacteriol. 2004, 186, 3182–3186. [Google Scholar] [CrossRef] [PubMed]
- Walsby, A.E. Gas Vesicles. Microbiol. Rev. 1994, 58, 94–144. [Google Scholar] [CrossRef] [PubMed]
- DasSarma, S.; Arora, P. Genetic analysis of gas vesicle gene cluster in haloarchaea. FEMS Microbiol. Lett. 1997, 153, 1–10. [Google Scholar] [CrossRef]
- Pfeifer, F. Distribution, formation and regulation of gas vesicles. Nat. Rev. Microbiol. 2012, 10, 705–715. [Google Scholar] [CrossRef] [PubMed]
- Zhilina, T.N.; Zavarzin, T.A. Comparative cytology of methanosarcinae and description of Methanosarcina vacuolata sp. nov. Mikrobiologiia 1979, 48, 279–285. [Google Scholar] [PubMed]
- Zinder, S.H.; Lobo, A.L. Isolation and characterization of thermophilic acetotrophic strain Methanothrix. Arch. Microbiol. 1987, 146, 315–322. [Google Scholar] [CrossRef]
- Li, N.; Cannon, M.C. Gas vesicle genes identified in Bacillus megaterium and functional expression in Escherichia coli. J. Bacteriol. 1998, 180, 2450–2458. [Google Scholar] [PubMed]
- Bentley, S.D.; Chater, K.F.; Cerdeno-Tarraga, A.M.; Challis, G.L.; Thomson, N.R.; James, K.D.; Harris, D.E.; Quail, M.A.; Kieser, H.; Harper, D.; et al. Complete genome sequence of the model actinomycete Streptomyces coelicolor A3(2). Nature 2002, 417, 141–147. [Google Scholar] [CrossRef] [PubMed]
- Shively, J.M.; Cannon, G.C.; Heinhorst, S.; Bryant, D.A.; DasSarma, S.; Bazylinski, D.; Preiss, J.; Steinbuchel, A.; Docampo, R.; Dahl, C. Bacterial and Archaeal Inclusions. In eLS (Encyclopedia of Life Sciences); John Wiley & Sons Ltd.: Chichester, UK, 2011. [Google Scholar]
- Pfeifer, F. Haloarchaea and the formation of gas vesicles. Life (Basel) 2015, 5, 385–402. [Google Scholar] [CrossRef] [PubMed]
- Walsby, A.E.; Bleything, A. The dimensions of cyanobacterial gas vesicles in relation to their efficiency in providing buoyancy and withstanding pressure. J. Gen. Microbiol. 1988, 134, 2635–2645. [Google Scholar] [CrossRef]
- Cohen-Bazire, G.R.; Kunisawa, R.; Pfennig, N. Comparative study of the structure of gas vacuoles. J. Bacteriol. 1969, 100, 1049–1061. [Google Scholar] [PubMed]
- Simon, R.D. Morphology and protein composition of gas vesicles from wild type and gas vacuole defective strains of Halobacterium salinarium strain 5. J. Gen. Microbiol. 1981, 125, 103–111. [Google Scholar] [CrossRef]
- Surek, B.U.; Pillay, B.; Rdest, U.; Beyreuther, K.; Goebel, W. Evidence for two different gas vesicle proteins and genes in Halobacterium halobium. J. Bacteriol. 1988, 70, 1746–1751. [Google Scholar]
- Stoeckenius, W.; Kunau, W.H. Further characterization of particulate fractions from lysed cell envelopes of Halobacterium halobium and isolation of gas vacuole membranes. J. Cell Biol. 1968, 38, 337–357. [Google Scholar] [CrossRef]
- Blaurock, A.E.; Wober, W. Structure of the wall of Halobacterium halobium. J. Mol. Biol. 1976, 106, 871–888. [Google Scholar] [CrossRef]
- Krantz, M.J.; Ballou, C.E. Analysis of Halobacterium halobium gas vesicles. J. Bacteriol. 1973, 114, 1058–1067. [Google Scholar] [PubMed]
- McMaster, T.J.; Miles, M.J.; Walsby, A.E. Direct observation of proteins secondary structure in gas vesicles by atomic force microscopy. Biophys. J. 1996, 70, 2432–2436. [Google Scholar] [CrossRef]
- Walsby, A.E. The pressure relationship of gas vacuoles. Proc. R. Soc. Lond. Ser. B 1971, 178, 301–326. [Google Scholar] [CrossRef]
- Sundararajan, A.; Ju, L.K. Evaluation of oxygen permeability of gas vesicles from cyanobacterium Anabaena flos-aquae. J. Biotechnol. 2000, 77, 151–156. [Google Scholar] [CrossRef]
- Waaland, J.R.; Branton, D. Gas vesicle development in blue-green alga. Science 1969, 163, 1339–1341. [Google Scholar] [CrossRef] [PubMed]
- Offner, S.; Ziese, U.; Wanner, G.; Typke, D.; Pfeifer, F. Structural characteristics of halobacterial gas vesicles. Microbiology 1998, 144, 1331–1342. [Google Scholar] [CrossRef] [PubMed]
- Walsby, A.E.; Buckland, B. Isolation and purification of intact gas vesicles from blue-green algae. Nature 1969, 224, 716–717. [Google Scholar] [CrossRef]
- Belenky, M.; Meyers, R.; Herzfeld, J. Subunit structure of gas vesicles: A MALDI-TOF mass spectrometry study. Biophys. J. 2004, 86, 499–505. [Google Scholar] [CrossRef]
- Sivertsen, A.C.; Bayro, M.J.; Belenky, M.; Griffin, R.G.; Herzfeld, J. Solid-state NMR characterization of gas vesicle structure. Biophys. J. 2010, 99, 1932–1939. [Google Scholar] [CrossRef] [PubMed]
- Bayro, M.J.; Daviso, E.; Belenky, M.; Griffin, R.G.; Herzfeld, J. An amyloid organelle, solid-state NMR evidence for cross-β assembly of gas vesicles. J. Biol. Chem. 2012, 287, 3479–3484. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Walsby, A.E.; Hayes, P.K. The minor cyanobacterial gas vesicle protein, GVPc, is attached to the outer surface of the gas vesicle. J. Gen. Microbiol. 1988, 134, 2647–2657. [Google Scholar] [CrossRef]
- Halladay, J.T.; Jones, J.G.; Lin, F.; MacDonald, A.B.; DasSarma, S. The rightward gas vesicle operon in Halobacterium halobium plasmid pNRC-100: Identification of the gvpA and gvpC gene products by use of antibody probes and genetic analysis of the region downstream of gvpC. J. Bacteriol. 1993, 175, 684–692. [Google Scholar] [PubMed]
- Jones, J.G.; Young, D.C.; DasSarma, S. Structure and organization of the gas vesicle gene cluster on the Halobacterium halobium plasmid pNRC-100. Gene 1991, 102, 1017–1022. [Google Scholar] [CrossRef]
- DasSarma, S.; Arora, P.; Lin, F.; Molinari, E.; Yin, L.R.-S. Wild-type gas vesicle formation requires at least ten genes in the gvp gene cluster of Halobacterium halobium plasmid pNRC100. J. Bacteriol. 1994, 176, 7646–7652. [Google Scholar] [PubMed]
- DasSarma, S.; Damerval, T.; Jones, J.G.; Tandeau de Marsac, N. A plasmid encoded gas vesicle protein gene in a halophilic archaebacterium. Mol. Microbiol. 1987, 1, 365–370. [Google Scholar] [CrossRef] [PubMed]
- Tandeau de Marsac, M.; Mazel, D.; Bryant, D.A.; Houmard, J. Molecular cloning and nucleotide sequence of a developmentally regulated gene from the cyanobacterium Calothrix PCC 7601: A gas vesicle gene. Nucleic Acids Res. 1985, 13, 7223–7236. [Google Scholar] [CrossRef] [PubMed]
- Damerval, T.; Houmard, J.; Guglielmi, G.; Csiszar, K.; Tandeau de Marsac, N. A developmentally regulated gvpABC operon is involved in the formation of gas vesicles in the cyanobacterium Calothrix 7601. Gene 1987, 54, 83–92. [Google Scholar]
- Horne, M.; Englert, C.; Pfeifer, F. Two genes encoding gas vacuole proteins in Halobacterium halobium. Mol. Gen. Genet. 1988, 213, 459–464. [Google Scholar] [CrossRef] [PubMed]
- Ng, W.-L.; Ciufo, S.A.; Smith, T.M.; Bumgardner, R.E.; Baskin, D.; Faust, J.; Hall, B.; Loretz, C.; Seto, J.; Slagel, J.; et al. Snapshot of a large dynamic replicon from a halophilic Archaeon: Megaplasmid or minichromosome? Genome Res. 1998, 8, 1131–1141. [Google Scholar] [PubMed]
- Ng, W.V.; Kennedy, S.P.; Mahairas, G.G.; Berquist, B.; Pan, M.; Shukla, H.D.; Lasky, S.R.; Baliga, N.; Thorsson, V.; Sbrogna, J. Genome sequence of Halobacterium species NRC-1. Proc. Natl. Acad. Sci. USA 2000, 97, 12176–12181. [Google Scholar] [CrossRef] [PubMed]
- DasSarma, S.; Halladay, J.T.; Jones, J.G.; Donovan, J.W.; Giannasca, P.J.; Tandeau de Marsac, N. High-frequency mutations in plasmid-encoded gas vesicle gene in Halobacterium halobium. Proc. Natl. Acad. Sci. USA 1988, 85, 6861–6865. [Google Scholar] [CrossRef] [PubMed]
- Jones, J.G.; Hackett, N.R.; Halladay, J.T.; Scothorn, D.J.; Yang, C.-F.; Ng, W.-L.; DasSarma, S. Analysis of insertion mutants reveals two new genes in the pNRC100 gas vesicle gene cluster of Halobacterium halobium. Nucleic Acids Res. 1989, 17, 7785–7793. [Google Scholar] [CrossRef] [PubMed]
- Horne, M.; Englert, C.; Wimmer, C.; Pfeifer, F. A DNA region of 9 bp contains all genes necessary for gas vesicle synthesis in halophilic archaebacteria. Mol. Microbiol. 1991, 5, 1159–1174. [Google Scholar] [CrossRef] [PubMed]
- Englert, C.; Wanner, G.; Pfeifer, F. Functional analysis of the gas vesicle gene cluster of the halophilic archaeon Haloferax mediterranei defines the vac region boundary and suggests a regulatory role for the gvpD gene or its product. Mol. Microbiol. 1992, 6, 3543–3550. [Google Scholar] [CrossRef] [PubMed]
- Pfeifer, F.; Kruger, K.; Roeder, R.; Mayr, A.; Ziesche, S.; Offner, S. Gas vesicle formation in halophilic archaea. Arch. Microbiol. 1987, 167, 259–268. [Google Scholar] [CrossRef]
- Albouy, D.; Castets, A.M.; Tandeau de Marsac, N. The gas vesicle gene cluster of the cyanobacterium Pseudanabaena sp. strain PCC 6901. DNA Seq. 2001, 12, 337–44. [Google Scholar] [CrossRef] [PubMed]
- Mlouka, A.; Comte, K.; Castets, A.M.; Bouchier, C.; Tandeau de Marsac, N. The gas vesicle gene cluster from Microcystis aeruginosa and DNA rearrangements that lead to loss of cell buoyancy. J. Bacteriol. 2004, 186, 2355–2365. [Google Scholar] [CrossRef] [PubMed]
- Kinsman, R.; Hayes, P.K. Genes encoding proteins homologous to halobacterial Gvps N, J, K, F and L are located downstream of gvpC in the cyanobacterium Anabaena flos-aquae. DNA Seq. 1997, 7, 97–106. [Google Scholar] [PubMed]
- Maeder, D.L.; Anderson, I.; Brettin, T.S.; Bruce, D.C.; Gilna, P.; Han, C.S.; Lapidus, A.; Metcalf, W.W.; Saunders, E.; Tapia, R.; et al. The Methanosarcina barkeri genome: comparative analysis with Methanosarcina acetivorans and Methanosarcina mazei reveals extensive rearrangement within methanosarcinal genomes. J. Bacteriol. 2006, 188, 7922–7931, Erratum in: J. Bacteriol. 2007, 189, 1488. [Google Scholar] [CrossRef]
- Halladay, J.T.; Ng, W.-L.; DasSarma, S. Genetic transformation of a halophilic archaebacterium with a gas vesicle gene cluster restores its ability to float. Gene 1992, 119, 131–136. [Google Scholar] [CrossRef]
- Ng, W.L.; DasSarma, S. Minimal replication origin of the 200-kilobase Halobacterium plasmid pNRC100. J. Bacteriol. 1993, 175, 4584–4596. [Google Scholar] [PubMed]
- Blaseio, U.; Pfeifer, F. Transformation of Halobacterium halobium: development of vectors and investigation of gas vesicle synthesis. Proc. Natl. Acad. Sci. USA 1990, 87, 6772–6776. [Google Scholar] [CrossRef] [PubMed]
- Offner, S.; Pfeifer, F. Complementation studies with the gas vesicle encoding p-vac region of Halobacterium salinarium PHH1 reveal a regulatory role for p-gvpDE genes. Mol. Microbiol. 1995, 170, 9–19. [Google Scholar] [CrossRef]
- Offner, S.; Gerhard, W.; Pfeifer, F. Functional studies of gvpACNO operon of Halobacterium salinarium reveal that GvpC protein shapes gas vesicles. J. Bacteriol. 1996, 178, 2071–2078. [Google Scholar] [PubMed]
- Kruger, K.; Pfeifer, F. Transcript analysis of the c-vac region and differential synthesis of the two regulatory gas vesicle proteins GvpD and GvpE in Halobacterium salinarium. J. Bacteriol. 1996, 178, 4012–4019. [Google Scholar] [PubMed]
- Kruger, K.; Hermann, T.; Armbruster, V.; Pfeifer, F. The transcriptional activator GvpE for the halobacterial gas vesicle genes resembles a basic region leucine-zipper regulatory protein. J. Mol. Biol. 1998, 279, 761–771. [Google Scholar] [CrossRef] [PubMed]
- Pfeifer, F.; Zotzel, J.; Kurenbach, B.; Roder, R.; Zimmermann, P. A p-loop motif and two basic regions in the regulatory protein GvpD are important for the repression of gas vesicle formation in the archaeon Haloferax mediterranei. Microbiology 2001, 147, 63–73. [Google Scholar] [CrossRef] [PubMed]
- Plosser, P.; Pfeifer, F. A bZIP protein from halophilic archaea: structural features and dimer formation of cGvpE from Halobacterium salinarum. Mol. Microbiol. 2002, 45, 511–520. [Google Scholar] [CrossRef] [PubMed]
- Zimmermann, P.; Pfeifer, F. Regulation of the expression of gas vesicle genes in Haloferax mediterranei: Interaction of the two regulatory proteins GvpD and GvpE. Mol. Microbiol. 2003, 49, 783–794. [Google Scholar] [CrossRef] [PubMed]
- Hofacker, A.; Schmitz, K.M.; Cichonczyk, A.; Sartorius-Neef, S.; Pfeifer, F. GvpE- and GvpD-mediated transcription regulation of the p-gvp genes encoding gas vesicles in Halobacterium salinarum. Microbiology 2004, 150, 1829–1838. [Google Scholar] [CrossRef] [PubMed]
- Tavlaridou, S.; Winter, K.; Pfeifer, F. The accessory gas vesicle protein GvpM of haloarchaea and its interaction partners during gas vesicle formation. Extremophiles 2014, 18, 693–706. [Google Scholar] [CrossRef] [PubMed]
- Tavlaridou, S.; Faist, K.; Weitzel, K.; Pfeifer, F. Effect of an overproduction of accessory Gvp proteins on gas vesicle formation in Haloferax volcanii. Extremophiles 2013, 17, 277–287. [Google Scholar] [CrossRef] [PubMed]
- Walker, J.E.; Hayes, P.K.; Walsby, A.E. Homology of gas vesicle proteins in cyanobacteria and halobacteria. J. Gen. Microbiol. 1984, 130, 2709–2717. [Google Scholar] [CrossRef]
- DasSarma, S. Mechanisms of genetic variability in Halobacterium halobium: the purple membrane and gas vesicle mutations. Can. J. Microbiol. 1989, 35, 65–72. [Google Scholar] [CrossRef] [PubMed]
- Chu, L.J.; Chen, M.C.; Setter, J.; Tsai, Y.S.; Yang, H.; Fang, X.; Ting, Y.S.; Shaffer, S.A.; Taylor, G.K.; von Haller, P.D.; et al. New structural proteins of Halobacterium salinarum gas vesicle revealed by comparative proteomics analysis. J. Proteome Res. 2011, 10, 1170–1178. [Google Scholar] [CrossRef] [PubMed]
- Simon, R.D. Acrylamide gel electrophoresis of hydrophobic proteins: Gas vacuole protein. Electrophoresis 1980, 11, 172–176. [Google Scholar] [CrossRef]
- Strunk, T.; Hamacher, K.; Hoffgaard, F.; Engelhardt, H.; Zillig, M.D.; Faist, K.; Wenzel, W.; Pfeifer, F. Structural model of the gas vesicle protein GvpA and analysis of GvpA mutants in vivo. Mol. Microbiol. 2011, 81, 56–68. [Google Scholar] [CrossRef] [PubMed]
- Ezzeldin, H.M.; Klauda, J.B.; Solares, S.D. Modeling of the major gas vesicle protein, GvpA: From protein sequence to vesicle wall structure. J. Struct. Biol. 2012, 179, 18–28. [Google Scholar] [CrossRef] [PubMed]
- Daviso, E.; Belenky, M.; Griffin, R.G.; Herzfeld, J. Gas vesicles across kingdoms: A comparative solid state NMR study. J. Mol. Microbiol. Biotechnol. 2013, 23, 281–289. [Google Scholar] [CrossRef] [PubMed]
- Kinsman, R.; Walsby, A.E.; Hayes, P.K. GvpCs with reduced numbers of repeating sequence elements bind to and strengthen cyanobacterial gas vesicles. Mol. Microbiol. 1995, 17, 147–154. [Google Scholar] [CrossRef] [PubMed]
- Shukla, H.D.; DasSarma, S.; University of Maryland, Baltimore, MD, USA. Unpublished data. 2015.
- Hayes, P.K.; Lazarus, C.M.; Bees, A.; Walker, J.E.; Walsby, A.E. The protein encoded by gvpC is a minor component of gas vesicles isolated from the cyanobacteria Anabaena flos-aquae and Microcystis sp. Mol. Microbiol. 1988, 2, 545–552. [Google Scholar] [CrossRef] [PubMed]
- Becker, S.; Hayes, P.K.; Walsby, A.E. Different GvpC length variants are transcribed within single filaments of the cyanobacterium Planktothrix rubescens. Microbiology 2005, 151, 59–67. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Karan, R.; Capes, M.D.; DasSarma, S. Function and biotechnology of extremophilic enzymes in low water activity. Aquat. Biosyst. 2012. [Google Scholar] [CrossRef] [PubMed]
- DasSarma, S.; DasSarma, P. Halophiles and their enzymes: Negativity put to good use. Curr. Opin. Microbiol. 2015, 25, 120–126. [Google Scholar] [CrossRef] [PubMed]
- Lupas, A. Predicting coiled-coil regions in proteins. Curr. Opin. Struct. Biol. 1997, 7, 388–393. [Google Scholar] [CrossRef]
- Xu, B.Y.; Dai, Y.N.; Zhou, K.; Liu, Y.T.; Sun, Q.; Ren, Y.M.; Chen, Y.; Zhou, C.Z. Structure of the gas vesicle protein GvpF from the cyanobacterium Microcystis aeruginosa. Acta Crystallogr. Sect. D Biol. Crystallogr. 2014, 70, 3013–3022. [Google Scholar] [CrossRef] [PubMed]
- Neuwald, A.F.; Aravind, L.; Spouge, J.L.; Koonin, E.V. AAA+: A class of chaperone-like ATPases associated with the assembly, operation, and disassembly of protein complexes. Genome Res. 1999, 9, 27–43. [Google Scholar] [PubMed]
- DasSarma, S.; Halladay, J.; Ng, W. Recombinant Vector and Process for Cell Flotation. U.S. Patent 6,008,051, 28 December 1999. [Google Scholar]
- DasSarma, S.; Morshed, F.; Stuart, E.S.; Black, S. Recombinant Gas Vesicles and Uses Thereof. U.S. Patent 5,824,309, 20 October 1998. [Google Scholar]
- Capiralla, H.; DasSarma, S.; University of Maryland, Baltimore, MD, USA. Unpublished data. 2015.
- Negi, V.D.; Singhamahapatra, S.; Chakravortty, D. Salmonella enterica serovar Typhimurium strain lacking pmrG-HM-D provides excellent protection against salmonellosis in murine typhoid model. Vaccine 2007, 25, 5315–5323. [Google Scholar] [CrossRef] [PubMed]
- Nagarajan, A.G.; Balasundaram, S.V.; Janice, J.; Karnam, G.; Eswarappa, S.M.; Chakravortty, D. SopB of Salmonella enterica serovar Typhimurium is a potential DNA vaccine candidate in conjugation with live attenuated bacteria. Vaccine 2009, 27, 2804–2811. [Google Scholar] [CrossRef] [PubMed]
- DasSarma, S.; Karan, R.; DasSarma, P.; Barnes, S.; Ekulona, F.; Smith, B. An improved genetic system for bioengineering buoyant gas vesicle nanoparticles from Haloarchaea. BMC Biotechnol. 2013. [Google Scholar] [CrossRef] [PubMed]
- Kennedy, S.P.; Ng, W.V.; Salzberg, S.L.; Hood, L.; DasSarma, S. Understanding the adaptation of Halobacterium species NRC-1 to its extreme environment through computational analysis of its genome sequence. Genome Res. 2001, 11, 1641–1650. [Google Scholar] [CrossRef] [PubMed]
- Jarori, G.K.; Dutta, S.; DasSarma, P.; DasSarma, S.; Tata Institute of Fundamental Research, Mumbai, India and University of Maryland, Baltimore, MD, USA. Unpublished data. 2015.
- Karan, R.; Capes, M.D.; DasSarma, P.; DasSarma, S. Cloning, overexpression, purification, and characterization of a polyextremophilic β-galactosidase from the Antarctic haloarchaeon Halorubrum lacusprofundi. BMC Biotechnol. 2013. [Google Scholar] [CrossRef] [PubMed]
- Karan, R.; DasSarma, P.; Balcer-Kubiczek, E.; Weng, R.R.; Liao, C.C.; Goodlett, D.R.; Ng, W.V.; DasSarma, S. Bioengineering radioresistance by overproduction of RPA, a mammalian-type single-stranded DNA-binding protein, in a halophilic archaeon. Appl. Microbiol. Biotechnol. 2014, 98, 1737–1747. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
DasSarma, S.; DasSarma, P. Gas Vesicle Nanoparticles for Antigen Display. Vaccines 2015, 3, 686-702. https://doi.org/10.3390/vaccines3030686
DasSarma S, DasSarma P. Gas Vesicle Nanoparticles for Antigen Display. Vaccines. 2015; 3(3):686-702. https://doi.org/10.3390/vaccines3030686
Chicago/Turabian StyleDasSarma, Shiladitya, and Priya DasSarma. 2015. "Gas Vesicle Nanoparticles for Antigen Display" Vaccines 3, no. 3: 686-702. https://doi.org/10.3390/vaccines3030686
APA StyleDasSarma, S., & DasSarma, P. (2015). Gas Vesicle Nanoparticles for Antigen Display. Vaccines, 3(3), 686-702. https://doi.org/10.3390/vaccines3030686





