Paraoxonases (PON) 1, 2, and 3 Polymorphisms and PON-1 Activities in Patients with Sickle Cell Disease
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Design and Participants
2.2. Sample Preparation
2.3. Biochemical Analyses
2.4. PON-1, PON-2, and PON-3 Genotypes
2.5. PON-1 Activities
2.6. Statistical Data Analysis
3. Results
3.1. Gender and Age of Subjects from Sickle Cell Disease and Healthy Control Groups
3.2. Cholesterol and Fractions, Apo-A1 and B in Patients with Sickle Cell Disease and Healthy Controls
3.3. PON-1 Activities in Sickle Cell Disease Patients and Healthy Controls
3.4. Transferrin and Ferritin as Predictors for PON-1 Arylesterase and Paraoxonase Activities in Patients with Sickle Cell Disease
3.5. Other Biochemical Determinations
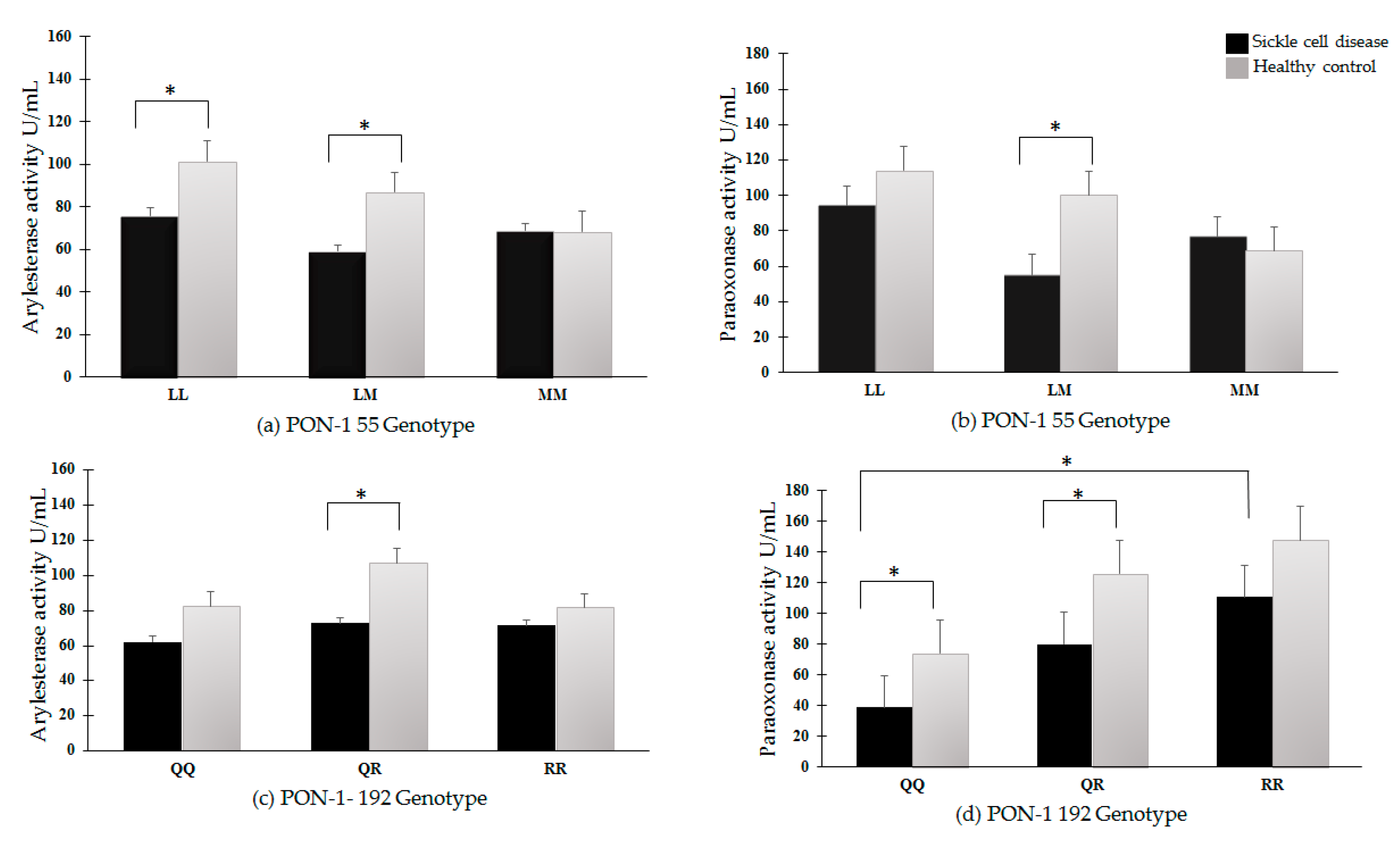
3.6. PON-1 Arylesterase and Paraoxonase Activities
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Chirico, E.N.; Pialoux, V. Role of oxidative stress in the pathogenesis of sickle cell disease. IUBMB Life 2012, 64, 72–80. [Google Scholar] [CrossRef] [PubMed]
- Kato, G.J.; Nouraie, S.M.; Gladwin, M.T. Lactate dehydrogenase and hemolysis in sickle cell disease. Blood 2013, 122, 1091–1092. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gladwin, M.T. Cardiovascular complications and risk of death in sickle-cell disease. Lancet 2016, 387, 2565–2574. [Google Scholar] [CrossRef]
- Samir, K.; Gupta, K.; Adams-Graves, P.E. Sickle Cell Pain: A Critical Reappraisal. Blood 2018, 120, 3647–3657. [Google Scholar]
- Voskou, S.; Aslan, M.; Fanis, P.; Phylactides, M.; Kleanthous, M. Oxidative stress in β-thalassaemia and sickle cell disease. Redox Biol. 2015, 6, 226–239. [Google Scholar] [CrossRef] [PubMed]
- da Guarda, C.C.; Santiago, R.P.; Fiuza, L.M.; Aleluia, M.M.; Ferreira, J.R.D.; Figueiredo, C.V.B.; Yahouedehou, S.C.M.A.; de Oliveira, R.M.; Lyra, I.M.; Gonçalves, M.D.S. Heme-mediated cell activation: The inflammatory puzzle of sickle cell anemia. Expert Rev. Hematol. 2017, 10, 533–541. [Google Scholar] [CrossRef] [PubMed]
- van Beers, E.J.; van Wijk, R. Oxidative stress in sickle cell disease; more than a DAMP squib. Clin. Hemorheol. Microcirc. 2018, 68, 239–250. [Google Scholar] [CrossRef] [PubMed]
- Valko, M.; Jomova, K.; Rhodes, C.J.; Kuča, K.; Musílek, K. Redox- and non-redox-metal-induced formation of free radicals and their role in human disease. Arch. Toxicol. 2016, 90, 1–37. [Google Scholar] [CrossRef] [PubMed]
- Mackness, M.I.; Durrington, P.N.; Mackness, B. How high-density lipoprotein protects against the effects of lipid peroxidation. Curr. Opin. Lipidol. 2000, 11, 383–388. [Google Scholar] [CrossRef]
- Altenhöfer, S.; Witte, I.; Teiber, J.F.; Wilgenbus, P.; Pautz, A.; Li, H.; Daiber, A.; Witan, H.; Clement, A.M.; Förstermann, U.; et al. One enzyme, two functions: PON2 prevents mitochondrial superoxide formation and apoptosis independent from its lactonase activity. J. Biol. Chem. 2010, 285, 24398–24403. [Google Scholar] [CrossRef]
- Furlong, C.E.; Marsillach, J.; Jarvik, G.P.; Costa, L.G. Paraoxonases-1, -2 and -3: What are their functions? Chem.-Biol. Interact. 2016, 259, 51–62. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Camps, J.; García-Heredia, A.; Hernández-Aguilera, A.; Joven, J. Paraoxonases, mitochondrial dysfunction and non-communicable diseases. Chem.-Biol. Interact. 2016, 259, 382–387. [Google Scholar] [CrossRef] [PubMed]
- Deakin, S.; Leviev, I.; Gomaraschi, M.; Calabresi, L.; Franceschini, G.; James, R.W. Enzymatically active paraoxonase-1 is located at the external membrane of producing cells and released by a high affinity, saturable, desorption mechanism. J. Biol. Chem. 2002, 277, 4301–4308. [Google Scholar] [CrossRef] [PubMed]
- Di Pietro, N.; Formoso, G.; Pandolfi, A. Physiology and pathophysiology of oxLDL uptake by vascular wall cells in atherosclerosis. Vasc. Pharmacol. 2016, 84, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Tiainen, S.; Kiviniemi, A.; Hautala, A.; Huikuri, H.; Ukkola, O.; Tokola, K.; Tulppo, M.; Vasankari, T. Effects of a two-year home-based exercise training program on oxidized LDL and HDL lipids in coronary artery disease patients with and without Type-2 Diabetes. Antioxidants 2018, 7, 144. [Google Scholar] [CrossRef]
- Calvieri, C.; Tanzilli, G.; Bartimoccia, S.; Cangemi, R.; Arrivi, A.; Dominici, M.; Cammisotto, V.; Viceconte, N.; Mangieri, E.; Frati, G.; et al. Interplay between oxidative stress and platelet activation in coronary thrombus of STEMI patients. Antioxidants 2018, 7, 83. [Google Scholar] [CrossRef] [PubMed]
- Levy, D.; Reichert, C.O.; Bydlowski, S.P. Paraoxonases Activities and Polymorphisms in Elderly and Old-Age Diseases: An Overview. Antioxidants 2019, 8, 118. [Google Scholar] [CrossRef]
- Eckerson, H.; Wyte, C.; La Du, B.N. The human serum paraoxonase/arylesterase polymorphism. Am. J. Hum. Genet. 1988, 43, 227–229. [Google Scholar]
- Menini, T.; Gugliucci, A. Paraoxonase 1 in neurological disorders. Redox Rep. 2014, 19, 49–58. [Google Scholar] [CrossRef]
- Bagchi, D.; Preuss, H.; Chen, C.; Blumberg, J. Oxidative Stress Status in Humans with Metabolic Syndrome. Obesity 2012, 6, 193–210. [Google Scholar]
- Camps, J.; Iftimie, S.; García-Heredia, A.; Castro, A.; Joven, J. Paraoxonases and infectious diseases. Clin. Biochem. 2017, 50, 804–811. [Google Scholar] [CrossRef] [PubMed]
- Maselli, L.M.F.; da Cunha, J.; Gutierrez, E.B.; Maranhão, R.C.; Spada, C.; Bydlowski, S.P. Human Paraoxonase-1 activity is related to the number of CD4+ T-Cells and is restored by antiretroviral therapy in HIV-1-infected individuals. Dis. Mark. 2014, 2014, 480201. [Google Scholar] [CrossRef] [PubMed]
- Miller, S.A.; Dykes, D.D.; Polesky, H.F. A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res. 1988, 16, 1215. [Google Scholar] [CrossRef] [PubMed]
- Miesbach, W.; Schenk, J.; Alesci, S.; Lindhoff-Last, E. Comparison of the fibrinogen Clauss assay and the fibrinogen PT derived method in patients with dysfibrinogenemia. Thromb. Res. 2010, 126, 428–433. [Google Scholar] [CrossRef] [PubMed]
- Fernvik, E.C.; Ketelhuth, D.F.J.; Russo, M.; Gidlund, M. The autoantibody repertoire against copper-or macrophage-modified LDL differs in normolipidemics and hypercholesterolemic patients. J. Clin. Immunol. 2004, 24, 170–176. [Google Scholar] [CrossRef] [PubMed]
- Brandão, S.A.; Izar, M.C.; Fischer, S.M.; Santos, A.O.; Monteiro, C.M.; Póvoa, R.M.; Helfenstein, T.; Carvalho, A.C.; Monteiro, A.M.; Ramos, E.; et al. Early increase in autoantibodies against human oxidized low-density lipoprotein in hypertensive patients after blood pressure control. Am. J. Hypertens. 2010, 23, 208–214. [Google Scholar] [CrossRef] [PubMed]
- Motti, C.; Dess, M.; Gnasso, A.; Irace, C.; Indigeno, P.; Angelucci, C.B.; Bernardini, S.; Fucci, G.; Federici, G.; Cortese, C. A multiplex PCR-based DNA assay for the detection of paraoxonase gene cluster polymorphisms. Atherosclerosis 2001, 158, 35–40. [Google Scholar] [CrossRef]
- Hegele, R.A.; Connelly, P.W.; Scherer, S.W.; Hanley, A.J.G.; Harris, S.B.; Tsui, L.C.; Zinman, B. Paraoxonase-2 gene (PON2) G148 variant associated with elevated fasting plasma glucose in noninsulin-dependent diabetes mellitus. J. Clin. Endocrinol. Metab. 1997, 82, 3373–3377. [Google Scholar] [CrossRef]
- Sentí, M.; Tomás, M.; Anglada, R.; Elosua, R.; Marrugat, J.; Covas, M.I.; Fitó, M. Interrelationship of smoking, paraoxonase activity, and leisure time physical activity: A population-based study. Eur. J. Intern. Med. 2003, 14, 178–184. [Google Scholar] [CrossRef]
- Agachan, B.; Yilmaz, H.; Karaali, Z.; Isbir, T. Paraoxonase 55 and 192 polymorphism and its relationship to serum paraoxonase activity and serum lipids in Turkish patients with non-insulin dependent diabetes mellitus. Cell Biochem. Funct. 2004, 22, 163–168. [Google Scholar] [CrossRef]
- Nur, E.; Biemond, B.J.; Otten, H.M.; Brandjes, D.P.; Schnog, J.J.B. Oxidative stress in sickle cell disease; pathophysiology and potential implications for disease management. Am. J. Hematol. 2011, 86, 484–489. [Google Scholar] [CrossRef] [PubMed]
- Dos Santos, T.E.; de Sousa, G.F.; Barbosa, M.C.; Gonçalves, R.P. The role of iron overload on oxidative stress in sickle cell anemia. Biomark. Med. 2012, 6, 813–819. [Google Scholar] [CrossRef] [PubMed]
- Grdic Rajkovic, M.; Rumora, L.; Barisic, K. The paraoxonase 1, 2 and 3 in humans. Biochem. Med. 2011, 21, 122–130. [Google Scholar] [CrossRef]
- Loued, S.; Isabelle, M.; Berrougui, H.; Khalil, A. The anti-inflammatory effect of paraoxonase 1 against oxidized lipids depends on its association with high density lipoproteins. Life Sci. 2012, 90, 82–88. [Google Scholar] [CrossRef] [PubMed]
- Viktorinova, A.; Jurkovicova, I.; Fabryova, L.; Kinova, S.; Koren, M.; Stecova, A.; Svitekova, K. Abnormalities in the relationship of paraoxonase 1 with HDL and apolipoprotein A1 and their possible connection to HDL dysfunctionality in type 2 diabetes. Diabetes Res. Clin. Pract. 2018, 140, 174–182. [Google Scholar] [CrossRef] [PubMed]
- Martinelli, N.; García-Heredia, A.; Roca, H.; Aranda, N.; Arija, V.; Mackness, B.; Mackness, M.; Busti, F.; Aragonès, G.; Pedro-Botet, J.; et al. Paraoxonase-1 status in patients with hereditary hemochromatosis. J. Lipid Res. 2013, 54, 1484–1492. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Del Giudice, R.; Pesce, A.; Cozzolino, F.; Monti, M.; Relini, A.; Piccoli, R.; Arciello, A.; Monti, D.M. Effects of iron on the aggregation propensity of the N-terminal fibrillogenic polypeptide of human apolipoprotein A-I. BioMetals 2018, 31, 551–559. [Google Scholar] [CrossRef] [PubMed]
- Gradinaru, D.; Borsa, C.; Ionescu, C.; Prada, G.I. Oxidized LDL and NO synthesis-Biomarkers of endothelial dysfunction and ageing. Mech. Ageing Dev. 2015, 151, 101–113. [Google Scholar] [CrossRef]
- Reichert, C.O.; Da Cunha, J.; Levy, D.; Maselli, L.M.F.; Bydlowski, S.P.; Spada, C. Hepcidin: Homeostasis and Diseases Related to Iron Metabolism. Acta Haematol. 2017, 137, 220–236. [Google Scholar] [CrossRef]
- Reichert, C.O.; Marafon, F.; Levy, D.; Maselli, L.M.F.; Bagatini, M.D.; Blatt, S.L.; Bydlowski, S.P.; Spada, C. Influence of Hepcidin in the Development of Anemia. In Current Topics in Anemia; IntechOpen: London, UK, 2018; Volume 1, pp. 1–17. [Google Scholar]
- Ikeda, K.; Nakamura, Y.; Kiyozuka, T.; Aoyagi, J.; Hirayama, T.; Nagata, R.; Ito, H.; Iwamoto, K.; Murata, K.; Yoshii, Y.; et al. Serological profiles of urate, paraoxonase-1, ferritin and lipid in Parkinson’s disease: Changes linked to disease progression. Neurodegener. Dis. 2011, 8, 252–258. [Google Scholar] [CrossRef]
- Matusiewicz, M.; Neubauer, K.; Lewandowska, P.; Gamian, A.; Krzystek-Korpacka, M. Reduced Transferrin Levels in Active Inflammatory Bowel Disease. BioMed Res. Int. 2017, 2017, 9541370. [Google Scholar] [CrossRef] [PubMed]
- Okuturlar, Y.; Akalin, N.; Kaptanogullari, O.H.; Guner, N.T.; Yilmaz, D.; Gedikbasi, A.; Soyluk, O.; Mert, M.; Serin, S.O.; Kocoglu, H.; et al. Comparison of serum paraoxonase and arylesterase activities between iron deficiency anemia patients and chronic kidney disease patients with anemia. Ren. Fail. 2016, 38, 781–786. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thalassemia, W.; Cakmak, A.; Soker, M.; Koc, A.; Erel, O. Paraoxonase and Arylesterase activity with oxidative status in children with thalassemia major. J. Pediatr. Hematol. Oncol. 2009, 31, 583–587. [Google Scholar]
- Zohaib, M.; Ansari, S.H.; Hashim, Z.; Shamsi, T.S.; Zarina, S. Serum paraoxonase activity and malondialdehyde serum concentrations remain unaffected in response to hydroxyurea therapy in β-Thalassemia patients. J. Clin. Pharmacol. 2016, 56, 869–874. [Google Scholar] [CrossRef] [PubMed]
- Kotani, K.; Tsuzaki, K.; Sakane, N. Paraoxonase-1 gene Q192R polymorphism and reactive oxygen metabolites. J. Int. Med. Res. 2012, 40, 1513–1518. [Google Scholar] [CrossRef] [PubMed]
- Atar, A.; Gedikbasi, A.; Sonmezay, E.; Kiraz, Z.K.; Abbasoglu, S.; Tasci, A.I.; Tugcu, V. Serum paraoxonase-1 gene polymorphism and enzyme activity in patients with urolithiasis. Ren. Fail. 2016, 38, 378–382. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.; Kumar, V.; Thakur, S.; Banerjee, B.D.; Rautela, R.S.; Grover, S.S.; Rawat, D.S.; Pasha, S.T.; Jain, S.K.; Ichhpujani, R.L.; et al. Paraoxonase-1 genetic polymorphisms and susceptibility to DNA damage in workers occupationally exposed to organophosphate pesticides. Toxicol. Appl. Pharmacol. 2011, 252, 130–137. [Google Scholar] [CrossRef]
- Gaidukov, L.; Rosenblat, M.; Aviram, M.; Tawfik, D.S. The 192R/Q polymorphs of serum paraoxonase PON1 differ in HDL binding, lipolactonase stimulation, and cholesterol efflux. J. Lipid Res. 2006, 47, 2492–2502. [Google Scholar] [CrossRef] [Green Version]
- Bayrak, A.; Bayrak, T.; Bodur, E.; Kılınç, K.; Demirpençe, E. The effect of HDL-bound and free PON1 on copper-induced LDL oxidation. Chem.-Biol. Interact. 2016, 257, 141–146. [Google Scholar] [CrossRef]
- Wood, J.C. Guidelines for quantifying iron overload. Hematology 2014, 2014, 210–215. [Google Scholar] [CrossRef] [Green Version]
- Brissot, P.; Ropert, M.; Le Lan, C.; Loréal, O. Non-transferrin bound iron: A key role in iron overload and iron toxicity. Biochim. Biophys. Acta Gen. Subj. 2012, 1820, 403–410. [Google Scholar] [CrossRef] [PubMed]
- Gardner, K.; Suddle, A.; Kane, P.; O’Grady, J.; Heaton, N.; Bomford, A.; Thein, S.L. How we treat sickle hepatopathy and liver transplantation in adults. Blood 2014, 123, 2302–2307. [Google Scholar] [CrossRef] [PubMed]
- Hammoudi, N.; Lionnet, F.; Redheuil, A.; Montalescot, G. Cardiovascular manifestations of sickle cell disease. Eur. Heart J. 2019, 40, 217. [Google Scholar] [CrossRef] [PubMed]
- Ballas, S.K. Iron overload is a determinant of morbidity and mortality in adult patients with sickle cell disease. Semin. Hematol. 2001, 38, 30–36. [Google Scholar] [CrossRef]
| PON-1 Primers | Sequences 5′→3′ |
| PON-1 55F | GAG TGA TGT ATA GCC CCA GTT TC |
| PON-1 55R | AGT CCA TTA GGC AGT ATC TCCg |
| PON-1 192F | TTG AAT GAT GTT GCT GTG GGA CCT GAG |
| PON-1 192R | CGA CCA CGC TAA ACC CAA ATA CAT CTC CCA GaA |
| PON-2 Primers | Sequences 5′→3′ |
| PON-2 148F | AGT GGA AAT TTT TAA ATT TGA AGC AG |
| PON-2 148R | TTG TTT GCA AAT GCT GGG GAT |
| PON-2 311F | GGT TCT CCG CAT CCA GAA CAT TgaA |
| PON-2 311R | TGT TAA GaT ATC GCA TCA TGC C |
| Gender | Groups | ||
|---|---|---|---|
| Sickle Cell Disease (n = 43) | Healthy Control (n = 43) | P-Value (1) | |
| Female | 30 (69.77%) | 30 (69.77%) | 1.0 |
| Male | 13 (30.23%) | 13 (30.23%) | 1.0 |
| Age (years) | 38.1 ± 11.72 | 37.8 ± 11.32 | 0.904 |
| Parameters | Groups | ||||||
|---|---|---|---|---|---|---|---|
| Sickle Cell Disease (n = 43) | Healthy Controls (n = 43) | P Value (1) | R Value (2) | R Value (3) | |||
| Mean ± SD | Min−Max | Mean ± SD | Min−Max | ||||
| Total cholesterol (mg/dL) | 132.51 ± 27.93 | 78−197 | 181.44 ± 35.99 | 104−277 | 0.01 | 0.407 | 0.098 |
| HDL-C (mg/dL) | 36.16 ± 11.38 | 14−66 | 50.65 ± 13.17 | 27−82 | 0.01 | 0.334 | 0.296 |
| LDL-C (mg/dL) | 71.18 ± 23.78 | 29−126 | 104.93 ± 32.23 | 44−208 | 0.01 | 0.270 | 0.060 |
| VLDL-C (mg/dL) | 25.27 ± 9.44 | 11−56 | 25.4 ± 10.06 | 11−56 | >0.05 | 0.131 | −0.215 |
| Triglycerides (mg/dL) | 125.81 ± 47.14 | 57−278 | 129.07 ± 65.04 | 56−373 | >0.05 | 0.111 | −0.227 |
| Apolipoprotein A1 | 107.78 ± 21.21 | 75.5−164.2 | 162.0 ± 26.6 | 113.3−225 | 0.01 | 0.427 | 0.259 |
| Apolipoprotein B | 65.13 ± 20.3 | 33.8−107.9 | 89.03 ± 22.32 | 53.6−161.5 | 0.01 | 0.170 | −0.102 |
| ApoB:ApoAI ratio | 0.62 ± 0.20 | 0.21−1.07 | 0.56 ± 0.16 | 0.31−1.14 | 0.08 | −0.092 | 0.330 |
| TG:HDL-C ratio | 3.99 ± 2.54 | 0.9−15.1 | 2.85 ± 1.93 | 0.8−10.4 | 0.006 | −0.120 | 0.034 |
| LDL-C:HDL-C ratio | 2.10 ± 0.80 | 0.5−4.8 | 2.17 ± 0.82 | 0.8−4.7 | >0.05 | −0.144 | −0.340 |
| C−reactive protein(mg/L) | 7.7 ± 9.3 | 0.8−59.4 | 2.69 ± 3.32 | 0.2−14.7 | 0.001 | −0.179 | −0.229 |
| Anti−oxLDL (U/mL) | 2.84 ± 1.52 | 0.91−10.33 | 1.74 ± 0.74 | 0.61−4.5 | 0.001 | −0.107 | −0.086 |
| Paraoxonase−1 activities | |||||||
| Paraoxonase (U/mL) | 80.3 ± 45.8 | 12−184 | 100.1 ± 55.2 | 24−232 | 0.078 | - | - |
| Arylesterase (U/mL) | 69.9 ± 20.3 | 24.5−122 | 89.7 ± 27.3 | 1−161 | 0.001 | - | - |
| Sickle Cell Disease (n = 43) | Arylesterase Activity | Paraoxonase Activity | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Parameters | Mean ± SD | Min−Max | RV (1) | β Value | Standard Error | P Value | Model (1) | β value | Standard Error | P Value | Model (2) |
| Lactate dehydrogenase (U/L) | 1162 ± 426.78 | 589−2326 | 240−480 | −0.237 | 0.009 | −0.229 | R2 = 0.011; P = 0.513 | −0.146 | −0.02 | −0.449 | R2 = 0.020; P = 0.381 |
| Total bilirubin (mg/dL) | 4.23 ± 2.58 | 1.13−13.8 | 0.20−1.00 | 6.230 | 50.53 | 0.343 | R2 = 0.015; P = 0.748 | −7.057 | 110.04 | 0.277 | R2 = 0.022; P = 0.652 |
| Indirect bilirubin (mg/dL) | 2.98 ± 1.68 | 0.66−7.92 | 0.10−0.60 | −3.904 | 50.32 | 0.359 | R2 = 0.048; P = 0.603 | 4.738 | 109.58 | 0.261 | R2 = 0.071; P = 0.432 |
| Direct bilirubin (mg/dL) | 1.24 ± 1.63 | 0.3−9.67 | <0.30 | −4.134 | 50.71 | 0.320 | R2 = 0.074; P = 0.582 | 4.260 | 110.44 | 0.300 | R2 = 0.098; P = 0.429 |
| Hemoglobin (g/dL) | 8.27 ± 1.03 | 6.2−10.6 | 12.0−16.0 | 0.052 | 10.91 | 0.925 | R2 = 0.001; P = 0.878 | −0.902 | 23.71 | 0.109 | R2 = 0.002; P = 0.772 |
| Hematocrit (%) | 23.53 ± 3.02 | 17.6−31.25 | 35−47 | −0.187 | 3.58 | 0.725 | R2 = 0.014; P = 0.758 | 0.728 | 7.78 | 0.176 | R2 = 0.013; P = 0.779 |
| Reticulocyte (%) | 12.08 ± 6.03 | 4.06−35.02 | 0.5−2.7 | −0.031 | 0.655 | 0.870 | R2 = 0.049; P = 0.599 | −0.295 | 1.42 | 0.129 | R2 = 0.135; P = 0.144 |
| Leucocytes (×103/mm3) | 10.73 ± 3.45 | 6.33−21.4 | 4−11 | −0.394 | 1.03 | 0.029 | R2 = 0.168; P = 0.146 | −0.224 | 2.24 | 0.204 | R2 = 0.173; P = 0.135 |
| Ferritin (ng/dL) | 568.36 ± 917.94 | 56−4872 | 15−150 | 0.451 | 0.015 | 0.227 | R2 = 0.047; P = 0.277 | 0.252 | 0.032 | 0.166 | R2 = 0.000; P = 0.923 |
| Transferrin (mg/dL) | 231.17 ± 42.97 | 150−335 | 250−380 | 0.507 | 0.091 | 0.018 | R2 = 0.250; P = 0.032 | 0.633 | 0.273 | 0.003 | R2 = 0.316; P = 0.011 |
| Transferrin saturation (%) | 45.11 ± 18.32 | 11.09−91.9 | 20−40 | −0.415 | 1.226 | 0.739 | R2 = 0.266; P = 0.064 | 0.064 | 2.590 | 0.959 | R2 = 0.316; P = 0.030 |
| Iron (µg/dL) | 120 ± 49.54 | 41−244 | 37−145 | 0.580 | 0.454 | 0.482 | R2 = 0.332; P = 0.055 | −0.520 | 0.960 | 0.966 | R2 = 0.378; P = 0.028 |
| TIBC (µg/dL) | 274.88 ± 62.58 | 178−489 | 228−428 | −0.734 | 0.202 | 0.298 | R2 = 0.367; P = 0.069 | −0.335 | 0.427 | 0.626 | R2 = 0.385; P = 0.054 |
| Fibrinogen (mg/dL) | 350.72 ± 108.9 | 178−712 | 150−400 | −0.30 | 0.031 | 0.856 | R2 = 0.063; P = 0.272 | −0.002 | 0.991 | 0.340 | R2 = 0.053; P = 0.340 |
| Polymorphisms | Groups | ||
|---|---|---|---|
| Sickle Cell Disease (n = 43) | Healthy Control (n = 43) | P-Value (1) | |
| PON-1 Q192R | |||
| 10 (23.2) | 24 (55.8) | 0.0064 | |
| QR | 19 (44.2) | 13 (30.2) | |
| RR | 14 (30.6) | 6 (13.9) | |
| Q allele | 20 (45) | 30 (71) | |
| R allele | 24 (55) | 13 (29) | |
| PON-1 L55M | |||
| LL | 21 (48.8) | 20 (46.5) | >0.05 |
| LM | 10 (23.2) | 14 (34.8) | |
| MM | 12 (27.9) | 9 (18.6) | |
| L allele | 26 (60) | 28 (64) | |
| M allele | 17 (40) | 15 (36) | |
| PON-2 A148G | |||
| AA | 24 (55.8) | 30 (69.7) | 0.178 |
| AG | 12 (27.9) | 11(25.6) | |
| GG | 7 (16.3) | 2 (4.6) | |
| A allele | 30 (70) | 36 (83) | |
| G allele | 13 (30) | 7 (17) | |
| PON-2 S311C | |||
| SS | 23 (53.4) | 23 (53.4) | 0.385 |
| SC | 16 (37.2) | 12 (27.9) | |
| CC | 4 (9.3) | 8 (18.6) | |
| S allele | 31 (72) | 29 (67) | |
| C allele | 12 (28) | 14 (33) | |
| PON-3 G10340T | |||
| GG | 1 (2.3) | 2 (4.6) | 0.621 |
| GT | 14 (32.5) | 17 (39.5) | |
| TT | 28 (65.1) | 24 (55.8) | |
| G allele | 8 (19) | 10 (24) | |
| T allele | 35 (81) | 33 (76) | |
| PON-3 A2115T | |||
| AA | 0 (0) | 1 (2.3) | 0.713 |
| AT | 3 (6.9) | 4 (9.3) | |
| TT | 40 (93) | 38 (88.3) | |
| A allele | 1 (3) | 3 (7) | |
| T allele | 42 (97) | 40 (93) | |
| PON-3 A45486C | |||
| AA | 0 (0) | 0 (0) | 0.676 |
| AC | 2 (4.6) | 4 (9.3) | |
| CC | 41 (95.3) | 39 (90.6) | |
| A allele | 1 (2) | 2 (5) | |
| C allele | 42 (98) | 41 (95) | |
| PON-3 C55146T | |||
| CC | 41 (95.3) | 38 (88.4) | 0.433 |
| CT | 2 (4.6) | 5 (11.6) | |
| TT | 0 (0) | 0 (0) | |
| C allele | 42 (98) | 40 (94) | |
| T allele | 1 (2) | 3 (6) | |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Reichert, C.O.; de Macedo, C.G.; Levy, D.; Sini, B.C.; Monteiro, A.M.; Gidlund, M.; Maselli, L.M.F.; Gualandro, S.F.M.; Bydlowski, S.P. Paraoxonases (PON) 1, 2, and 3 Polymorphisms and PON-1 Activities in Patients with Sickle Cell Disease. Antioxidants 2019, 8, 252. https://doi.org/10.3390/antiox8080252
Reichert CO, de Macedo CG, Levy D, Sini BC, Monteiro AM, Gidlund M, Maselli LMF, Gualandro SFM, Bydlowski SP. Paraoxonases (PON) 1, 2, and 3 Polymorphisms and PON-1 Activities in Patients with Sickle Cell Disease. Antioxidants. 2019; 8(8):252. https://doi.org/10.3390/antiox8080252
Chicago/Turabian StyleReichert, Cadiele Oliana, Carolina Garcia de Macedo, Débora Levy, Bruno Carnevale Sini, Andréia Moreira Monteiro, Magnus Gidlund, Luciana Morganti Ferreira Maselli, Sandra Fátima Menosi Gualandro, and Sérgio Paulo Bydlowski. 2019. "Paraoxonases (PON) 1, 2, and 3 Polymorphisms and PON-1 Activities in Patients with Sickle Cell Disease" Antioxidants 8, no. 8: 252. https://doi.org/10.3390/antiox8080252
APA StyleReichert, C. O., de Macedo, C. G., Levy, D., Sini, B. C., Monteiro, A. M., Gidlund, M., Maselli, L. M. F., Gualandro, S. F. M., & Bydlowski, S. P. (2019). Paraoxonases (PON) 1, 2, and 3 Polymorphisms and PON-1 Activities in Patients with Sickle Cell Disease. Antioxidants, 8(8), 252. https://doi.org/10.3390/antiox8080252







