Enhancing Disease Diagnosis: Biomedical Applications of Surface-Enhanced Raman Scattering
Abstract
1. Introduction
2. Surface Enhanced Raman Scattering: A Brief Tutorial
3. The Affinity of SERS-Active Substrates for Biochemical Molecules
4. Multivariate Chemometrics
5. Applications of SERS in Disease Diagnostics
5.1. Cancer
5.2. Microbial Infections–Pathogen Detection
5.3. SERS in Breath Analysis
6. Conclusions and Future Outlook
Author Contributions
Funding
Conflicts of Interest
References
- Trivedi, D.K.; Hollywood, K.A.; Goodacre, R. Metabolomics for the masses: The future of metabolomics in a personalized world. New Horiz. Transl. Med. 2017, 3, 294–305. [Google Scholar] [CrossRef] [PubMed]
- Ellis, D.I.; Dunn, W.B.; Griffin, J.L.; Allwood, J.W.; Goodacre, R. Metabolic fingerprinting as a diagnostic tool. Pharmacogenomics 2007, 8, 1243–1266. [Google Scholar] [CrossRef] [PubMed]
- Atkinson, A.J.; Colburn, W.A.; DeGruttola, V.G.; DeMets, D.L.; Downing, G.J.; Hoth, D.F.; Oates, J.A.; Peck, C.C.; Schooley, R.T.; Spilker, B.A.; et al. Biomarkers and surrogate endpoints: Preferred definitions and conceptual framework. Clin. Pharmacol. Ther. 2001, 69, 89–95. [Google Scholar]
- Wheelock, C.E.; Goss, V.M.; Balgoma, D.; Nicholas, B.; Brandsma, J.; Skipp, P.J.; Snowden, S.; Burg, D.; D’Amico, A.; Horvath, I.; et al. Application of ’omics technologies to biomarker discovery in inflammatory lung diseases. Eur. Respir. J. 2013, 42, 802–825. [Google Scholar] [CrossRef] [PubMed]
- Luo, P.; Yin, P.Y.; Hua, R.; Tan, Y.X.; Li, Z.F.; Qiu, G.K.; Yin, Z.Y.; Xie, X.W.; Wang, X.M.; Chen, W.B.; et al. A large-scale, multicenter serum metabolite biomarker identification study for the early detection of hepatocellular carcinoma. Hepatology 2018, 67, 662–675. [Google Scholar] [CrossRef]
- Zhang, A.H.; Sun, H.; Wang, P.; Han, Y.; Wang, X.J. Modern analytical techniques in metabolomics analysis. Analyst 2012, 137, 293–300. [Google Scholar] [CrossRef]
- Kitteringham, N.R.; Jenkins, R.E.; Lane, C.S.; Elliott, V.L.; Park, B.K. Multiple reaction monitoring for quantitative biomarker analysis in proteomics and metabolomics. J. Chromatogr. B 2009, 877, 1229–1239. [Google Scholar] [CrossRef]
- Ayalew, M.; Le-Niculescu, H.; Levey, D.F.; Jain, N.; Changala, B.; Patel, S.D.; Winiger, E.; Breier, A.; Shekhar, A.; Amdur, R.; et al. Convergent functional genomics of schizophrenia: From comprehensive understanding to genetic risk prediction. Mol. Psychiatr. 2012, 17, 887–905. [Google Scholar] [CrossRef]
- Dunn, W.B.; Broadhurst, D.I.; Atherton, H.J.; Goodacre, R.; Griffin, J.L. Systems level studies of mammalian metabolomes: The roles of mass spectrometry and nuclear magnetic resonance spectroscopy. Chem. Soc. Rev. 2011, 40, 387–426. [Google Scholar] [CrossRef]
- Goodacre, R.; Graham, D.; Faulds, K. Recent developments in quantitative SERS: Moving towards absolute quantification. TrAC Trends Anal. Chem. 2018, 102, 359–368. [Google Scholar] [CrossRef]
- Levine, R.J.; Lam, C.; Qian, C.; Yu, K.F.; Maynard, S.E.; Sachs, B.P.; Sibai, B.M.; Epstein, F.H.; Romero, R.; Thadhani, R.; et al. Soluble endoglin and other circulating antiangiogenic factors in preeclampsia. N. Engl. J. Med. 2006, 355, 992–1005. [Google Scholar] [CrossRef]
- Berger, A.G.; Restaino, S.M.; White, I.M. Vertical-flow paper SERS system for therapeutic drug monitoring of flucytosine in serum. Anal. Chim. Acta 2017, 949, 59–66. [Google Scholar] [CrossRef]
- Graham, D.; Moskovits, M.; Tian, Z.Q. SERS—Facts, figures and the future. Chem. Soc. Rev. 2017, 46, 3864–3865. [Google Scholar] [CrossRef]
- Tempero, M.A.; Malafa, M.P.; Al-Hawary, M.; Asbun, H.; Bain, A.; Behrman, S.W.; Benson, A.B.; Binder, E.; Cardin, D.B.; Cha, C.; et al. Pancreatic adenocarcinoma, version 2.2017, NCCN clinical practice guidelines in oncology. J. Natl. Compr. Cancer Netw. 2017, 15, 1028–1061. [Google Scholar] [CrossRef]
- Tang, H.B.; Meng, G.W.; Huang, Q.; Zhang, Z.; Huang, Z.L.; Zhu, C.H. Arrays of cone-shaped ZnO nanorods decorated with Ag nanoparticles as 3D surface-enhanced Raman scattering substrates for rapid detection of trace polychlorinated biphenyls. Adv. Funct. Mater. 2012, 22, 218–224. [Google Scholar] [CrossRef]
- Makam, P.; Shilpa, R.; Kandjani, A.E.; Periasamy, S.R.; Sabri, Y.M.; Madhu, C.; Bhargava, S.K.; Govindaraju, T. SERS and fluorescence-based ultrasensitive detection of mercury in water. Biosens. Bioelectron. 2018, 100, 556–564. [Google Scholar] [CrossRef]
- Zhou, H.B.; Yang, D.T.; Ivleva, N.P.; Mircescu, N.E.; Schubert, S.; Niessner, R.; Wieser, A.; Haisch, C. Label-free in situ discrimination of live and dead bacteria by surface-enhanced Raman scattering. Anal. Chem. 2015, 87, 6553–6561. [Google Scholar] [CrossRef]
- Galvan, D.D.; Yu, Q.M. Surface-enhanced Raman scattering for rapid detection and characterization of antibiotic-resistant bacteria. Adv. Healthc. Mater. 2018, 7, 1701335. [Google Scholar] [CrossRef]
- Liu, B.; Zhou, P.; Liu, X.M.; Sun, X.; Li, H.; Lin, M.S. Detection of pesticides in fruits by surface-enhanced Raman spectroscopy coupled with gold nanostructures. Food Bioprocess Technol. 2013, 6, 710–718. [Google Scholar] [CrossRef]
- Lin, M.; He, L.; Awika, J.; Yang, L.; Ledoux, D.R.; Li, H.; Mustapha, A. Detection of melamine in gluten, chicken feed, and processed foods using surface-enhanced Raman spectroscopy and HPLC. J. Food Sci. 2008, 73, T129–T134. [Google Scholar] [CrossRef]
- Subaihi, A.; Almanqur, L.; Muhamadali, H.; AlMasoud, N.; Ellis, D.I.; Trivedi, D.K.; Hollywood, K.A.; Xu, Y.; Goodacre, R. Rapid, accurate, and quantitative detection of propranolol in multiple human biofluids via surface-enhanced Raman scattering. Anal. Chem. 2016, 88, 10884–10892. [Google Scholar] [CrossRef]
- Tackman, E.C.; Trujillo, M.J.; Lockwood, T.L.E.; Merga, G.; Lieberman, M.; Camden, J.P. Identification of substandard and falsified antimalarial pharmaceuticals chloroquine, doxycycline, and primaquine using surface-enhanced Raman scattering. Anal. Methods 2018, 10, 4718–4722. [Google Scholar] [CrossRef]
- Jamieson, L.E.; Asiala, S.M.; Gracie, K.; Faulds, K.; Graham, D. Bioanalytical measurements enabled by surface-enhanced Raman scattering (SERS) probes. Annu. Rev. Anal. Chem. 2017, 10, 415–437. [Google Scholar] [CrossRef] [PubMed]
- Laing, S.; Gracie, K.; Faulds, K. Multiplex in vitro detection using SERS. Chem. Soc. Rev. 2016, 45, 1901–1918. [Google Scholar] [CrossRef] [PubMed]
- Feng, S.Y.; Chen, R.; Lin, J.Q.; Pan, J.J.; Chen, G.N.; Li, Y.Z.; Cheng, M.; Huang, Z.F.; Chen, J.; Zeng, H.S. Nasopharyngeal cancer detection based on blood plasma surface-enhanced Raman spectroscopy and multivariate analysis. Biosens. Bioelectron. 2010, 25, 2414–2419. [Google Scholar] [CrossRef] [PubMed]
- Shao, L.T.; Zhang, A.Y.; Rong, Z.; Wang, C.W.; Jia, X.F.; Zhang, K.H.; Xiao, R.; Wang, S.Q. Fast and non-invasive serum detection technology based on surface-enhanced Raman spectroscopy and multivariate statistical analysis for liver disease. Nanomed. Nanotechnol. Biol. Med. 2018, 14, 451–459. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.K.; Dai, X.M.; Stogin, B.B.; Wong, T.S. Ultrasensitive surface-enhanced Raman scattering detection in common fluids. Proc. Natl. Acad. Sci. USA 2016, 113, 268–273. [Google Scholar] [CrossRef] [PubMed]
- Subaihi, A.; Trivedi, D.K.; Hollywood, K.A.; Bluett, J.; Xu, Y.; Muhamadali, H.; Ellis, D.I.; Goodacre, R. Quantitative online liquid chromatography surface-enhanced Raman scattering (LC-SERS) of methotrexate and its major metabolites. Anal. Chem. 2017, 89, 6702–6709. [Google Scholar] [CrossRef] [PubMed]
- Laing, S.; Jamieson, L.E.; Faulds, K.; Graham, D. Surface-enhanced Raman spectroscopy for in vivo biosensing. Nat. Rev. Chem. 2017, 1, 0060. [Google Scholar] [CrossRef]
- Gracie, K.; Lindsay, D.; Graham, D.; Faulds, K. Bacterial meningitis pathogens identified in clinical samples using a SERS DNA detection assay. Anal. Methods 2015, 7, 1269–1272. [Google Scholar] [CrossRef]
- Gracie, K.; Correa, E.; Mabbott, S.; Dougan, J.A.; Graham, D.; Goodacre, R.; Faulds, K. Simultaneous detection and quantification of three bacterial meningitis pathogens by SERS. Chem. Sci. 2014, 5, 1030–1040. [Google Scholar] [CrossRef]
- Smekal, A. On the quantum theory of dispersal and dispersion. Z. Phys. 1925, 32, 241–244. [Google Scholar] [CrossRef]
- Raman, C.V.; Krishnan, K.S. A new type of secondary radiation. Nature 1928, 121, 501–502. [Google Scholar] [CrossRef]
- Ellis, D.I.; Goodacre, R. Metabolic fingerprinting in disease diagnosis: Biomedical applications of infrared and Raman spectroscopy. Analyst 2006, 131, 875–885. [Google Scholar] [CrossRef]
- Fleischmann, M.; Hendra, P.J.; McQuillan, A.J. Raman spectra of pyridine adsorbed at a silver electrode. Chem. Phys. Lett. 1974, 26, 163–166. [Google Scholar] [CrossRef]
- Jeanmaire, D.L.; Vanduyne, R.P. Surface Raman spectroelectrochemistry. 1. Heterocyclic, aromatic, and aliphatic-amines adsorbed on anodized silver electrode. J. Electroanal. Chem. 1977, 84, 1–20. [Google Scholar] [CrossRef]
- Albrecht, M.G.; Creighton, J.A. Anomalously intense Raman spectra of pyridine at a silver electrode. J. Am. Chem. Soc. 1977, 99, 5215–5217. [Google Scholar] [CrossRef]
- Sharma, B.; Frontiera, R.R.; Henry, A.I.; Ringe, E.; Van Duyne, R.P. SERS: Materials, applications, and the future. Mater. Today 2012, 15, 16–25. [Google Scholar] [CrossRef]
- Graham, D.; Goodacre, R.; Arnolds, H.; Masson, J.F.; Schatz, G.; Baumberg, J.; Kim, D.H.; Aizpurua, J.; Lum, W.; Silvestri, A.; et al. Theory of SERS enhancement: General discussion. Faraday Discuss. 2017, 205, 173–211. [Google Scholar] [CrossRef]
- Moskovits, M.; Suh, J.S. Surface selection-rules for surface-enhanced Raman spectroscopy—Calculations and application to the surface-enhanced Raman-spectrum of phthalazine on silver. J. Phys. Chem. 1984, 88, 5526–5530. [Google Scholar] [CrossRef]
- Willets, K.A.; Van Duyne, R.P. Localized surface plasmon resonance spectroscopy and sensing. Annu. Rev. Phys. Chem. 2007, 58, 267–297. [Google Scholar] [CrossRef]
- Stiles, P.L.; Dieringer, J.A.; Shah, N.C.; Van Duyne, R.R. Surface-enhanced Raman spectroscopy. Annu. Rev. Anal. Chem. 2008, 1, 601–626. [Google Scholar] [CrossRef]
- Schlucker, S. Surface-enhanced Raman spectroscopy: Concepts and chemical applications. Angew. Chem.-Int. Ed. 2014, 53, 4756–4795. [Google Scholar] [CrossRef]
- Qian, X.M.; Nie, S.M. Single-molecule and single-nanoparticle SERS: From fundamental mechanisms to biomedical applications. Chem. Soc. Rev. 2008, 37, 912–920. [Google Scholar] [CrossRef]
- Kneipp, K.; Wang, Y.; Kneipp, H.; Perelman, L.T.; Itzkan, I.; Dasari, R.; Feld, M.S. Single molecule detection using surface-enhanced Raman scattering (SERS). Phys. Rev. Lett. 1997, 78, 1667–1670. [Google Scholar] [CrossRef]
- Fisk, H.; Westley, C.; Turner, N.J.; Goodacre, R. Achieving optimal SERS through enhanced experimental design. J. Raman Spectrosc. 2016, 47, 59–66. [Google Scholar] [CrossRef]
- Zong, C.; Xu, M.X.; Xu, L.J.; Wei, T.; Ma, X.; Zheng, X.S.; Hu, R.; Ren, B. Surface-enhanced Raman spectroscopy for bioanalysis: Reliability and challenges. Chem. Rev. 2018, 118, 4946–4980. [Google Scholar] [CrossRef]
- Wang, Y.; Ruan, Q.Y.; Lei, Z.C.; Ling, S.C.; Zhu, Z.; Zhou, L.J.; Yang, C.Y. Highly sensitive and automated surface-enhanced Raman scattering-based immunoassay for H5N1 detection with digital microfluidics. Anal. Chem. 2018, 90, 5224–5231. [Google Scholar] [CrossRef]
- Cialla-May, D.; Zheng, X.S.; Weber, K.; Popp, J. Recent progress in surface-enhanced Raman spectroscopy for biological and biomedical applications: From cells to clinics. Chem. Soc. Rev. 2017, 46, 3945–3961. [Google Scholar] [CrossRef]
- Howes, P.D.; Chandrawati, R.; Stevens, M.M. Colloidal nanoparticles as advanced biological sensors. Science 2014, 346, 1247390. [Google Scholar] [CrossRef]
- Austin, L.A.; Mackey, M.A.; Dreaden, E.C.; El-Sayed, M.A. The optical, photothermal, and facile surface chemical properties of gold and silver nanoparticles in biodiagnostics, therapy, and drug delivery. Arch. Toxicol. 2014, 88, 1391–1417. [Google Scholar] [CrossRef] [PubMed]
- Lombardi, J.R.; Birke, R.L. A unified approach to surface-enhanced Raman spectroscopy. J. Phys. Chem. C 2008, 112, 5605–5617. [Google Scholar] [CrossRef]
- Xu, L.J.; Lei, Z.C.; Li, J.X.; Zong, C.; Yang, C.J.; Ren, B. Label-free surface-enhanced Raman spectroscopy detection of DNA with single-base sensitivity. J. Am. Chem. Soc. 2015, 137, 5149–5154. [Google Scholar] [CrossRef] [PubMed]
- Bonifacio, A.; Cervo, S.; Sergo, V. Label-free surface-enhanced Raman spectroscopy of biofluids: Fundamental aspects and diagnostic applications. Anal. Bioanal. Chem. 2015, 407, 8265–8277. [Google Scholar] [CrossRef] [PubMed]
- Wold, S.; Esbensen, K.; Geladi, P. Principal component analysis. Chemom. Intell. Lab. Syst. 1987, 2, 37–52. [Google Scholar] [CrossRef]
- Granato, D.; Santos, J.S.; Escher, G.B.; Ferreira, B.L.; Maggio, R.M. Use of principal component analysis (PCA) and hierarchical cluster analysis (HCA) for multivariate association between bioactive compounds and functional properties in foods: A critical perspective. Trends Food Sci. Technol. 2018, 72, 83–90. [Google Scholar] [CrossRef]
- Muhamadali, H.; Chisanga, M.; Subaihi, A.; Goodacre, R. Combining Raman and FT-IR spectroscopy with quantitative isotopic labeling for differentiation of E. coli cells at community and single cell levels. Anal. Chem. 2015, 87, 4578–4586. [Google Scholar] [CrossRef]
- Wold, S.; Sjostrom, M.; Eriksson, L. Pls-regression: A basic tool of chemometrics. Chemom. Intell. Lab. Syst. 2001, 58, 109–130. [Google Scholar] [CrossRef]
- Gromski, P.S.; Muhamadali, H.; Ellis, D.I.; Xu, Y.; Correa, E.; Turner, M.L.; Goodacre, R. A tutorial review: Metabolomics and partial least squares-discriminant analysis—A marriage of convenience or a shotgun wedding. Anal. Chim. Acta 2015, 879, 10–23. [Google Scholar] [CrossRef]
- Mazivila, S.J.; Olivieri, A.C. Chemometrics coupled to vibrational spectroscopy and spectroscopic imaging for the analysis of solid-phase pharmaceutical products: A brief review on non-destructive analytical methods. TrAC-Trends Anal. Chem. 2018, 108, 74–87. [Google Scholar] [CrossRef]
- Shinzawa, H.; Awa, K.; Kanematsu, W.; Ozaki, Y. Multivariate data analysis for Raman spectroscopic imaging. J. Raman Spectrosc. 2009, 40, 1720–1725. [Google Scholar] [CrossRef]
- Ellis, D.I.; Brewster, V.L.; Dunn, W.B.; Allwood, J.W.; Golovanov, A.P.; Goodacre, R. Fingerprinting food: Current technologies for the detection of food adulteration and contamination. Chem. Soc. Rev. 2012, 41, 5706–5727. [Google Scholar] [CrossRef]
- Schmidhuber, J. Deep learning in neural networks: An overview. Neural Netw. 2015, 61, 85–117. [Google Scholar] [CrossRef]
- Rumelhart, D.E.; McClelland, J.L.; Group, P.R. Parallel Distributed Processing, Experiments in the Microstructure of Cognition; MIT Press: Cambridge, MA, USA, 1986; Volumes 1–2. [Google Scholar]
- Shi, H.Y.; Wang, H.Y.; Meng, X.Y.; Chen, R.Z.; Zhang, Y.S.; Su, Y.Y.; He, Y. Setting up a surface-enhanced Raman scattering database for artificial-intelligence-based label-free discrimination of tumor suppressor genes. Anal. Chem. 2018, 90, 14216–14221. [Google Scholar] [CrossRef]
- Krauss, S.D.; Roy, R.; Yosef, H.K.; Lechtonen, T.; El-Mashtoly, S.F.; Gerwert, K.; Mosig, A. Hierarchical deep convolutional neural networks combine spectral and spatial information for highly accurate raman-microscopy-based cytopathology. J. Biophotonics 2018, 11, e201800022. [Google Scholar] [CrossRef]
- Broadhurst, D.I.; Kell, D.B. Statistical strategies for avoiding false discoveries in metabolomics and related experiments. Metabolomics 2006, 2, 171–196. [Google Scholar] [CrossRef]
- Lin, D.; Feng, S.Y.; Pan, J.J.; Chen, Y.P.; Lin, J.Q.; Chen, G.N.; Xie, S.S.; Zeng, H.S.; Chen, R. Colorectal cancer detection by gold nanoparticle based surface-enhanced Raman spectroscopy of blood serum and statistical analysis. Opt. Express 2011, 19, 13565–13577. [Google Scholar] [CrossRef]
- Chen, Y.P.; Chen, G.; Feng, S.Y.; Pan, J.J.; Zheng, X.W.; Su, Y.; Chen, Y.; Huang, Z.F.; Lin, X.Q.; Lan, F.H.; et al. Label-free serum ribonucleic acid analysis for colorectal cancer detection by surface-enhanced Raman spectroscopy and multivariate analysis. J. Biomed. Opt. 2012, 17, 067003. [Google Scholar] [CrossRef]
- Yu, Y.; Lin, Y.T.; Xu, C.X.; Lin, K.C.; Ye, Q.; Wang, X.Y.; Xie, S.S.; Chen, R.; Lin, J.Q. Label-free detection of nasopharyngeal and liver cancer using surface-enhanced Raman spectroscopy and partial ease squares combined with support vector machine. Biomed. Opt. Express 2018, 9, 6053–6066. [Google Scholar] [CrossRef]
- Carmicheal, J.; Hayashi, C.; Huang, X.; Liu, L.; Lu, Y.; Krasnoslobodtsev, A.; Lushnikov, A.; Kshirsagar, P.G.; Patel, A.; Jain, M.; et al. Label-free characterization of exosome via surface-enhanced Raman spectroscopy for the early detection of pancreatic cancer. Nanomed. Nanotechnol. Biol. Med. 2019, 16, 88–96. [Google Scholar] [CrossRef]
- Jarvis, R.M.; Goodacre, R. Discrimination of bacteria using surface-enhanced Raman spectroscopy. Anal. Chem. 2004, 76, 40–47. [Google Scholar] [CrossRef]
- Kearns, H.; Goodacre, R.; Jamieson, L.E.; Graham, D.; Faulds, K. SERS detection of multiple antimicrobial-resistant pathogens using nanosensors. Anal. Chem. 2017, 89, 12666–12673. [Google Scholar] [CrossRef]
- Lu, Y.D.; Lin, Y.S.; Zheng, Z.C.; Tang, X.Q.; Lin, J.Y.; Liu, X.J.; Liu, M.M.; Chen, G.N.; Qiu, S.F.; Zhou, T.; et al. Label free hepatitis B detection based on serum derivative surface-enhanced Raman spectroscopy combined with multivariate analysis. Biomed. Opt. Express 2018, 9, 4755–4766. [Google Scholar] [CrossRef]
- Hidi, I.J.; Jahn, M.; Weber, K.; Bocklitz, T.; Pletz, M.W.; Cialla-May, D.; Popp, J. Lab-on-a-chip-surface-enhanced Raman scattering combined with the standard addition method: Toward the quantification of nitroxoline in spiked human urine samples. Anal. Chem. 2016, 88, 9173–9180. [Google Scholar] [CrossRef]
- Alharbi, O.; Xu, Y.; Goodacre, R. Simultaneous multiplexed quantification of caffeine and its major metabolites theobromine and paraxanthine using surface-enhanced Raman scattering. Anal. Bioanal. Chem. 2015, 407, 8253–8261. [Google Scholar] [CrossRef]
- Deng, B.E.; Luo, X.; Zhang, M.; Ye, L.M.; Chen, Y. Quantitative detection of acyclovir by surface-enhanced Raman spectroscopy using a portable Raman spectrometer coupled with multivariate data analysis. Colloid Surf. B Biointerfaces 2019, 173, 286–294. [Google Scholar] [CrossRef]
- Fitzmaurice, C.; Allen, C.; Barber, R.M.; Barregard, L.; Bhutta, Z.A.; Brenner, H.; Dicker, D.J.; Chimed-Orchir, O.; Dandona, R.; Dandona, L.; et al. Global, regional, and national cancer incidence, mortality, years of life lost, years lived with disability, and disability-adjusted life-years for 32 cancer groups, 1990 to 2015 a systematic analysis for the global burden of disease study. JAMA Oncol. 2017, 3, 524–548. [Google Scholar]
- World Health Organisation. Available online: https://www.Who.Int/en/news-room/fact-sheets/detail/cancer (accessed on 12 December 2018).
- Chikkaveeraiah, B.V.; Bhirde, A.A.; Morgan, N.Y.; Eden, H.S.; Chen, X.Y. Electrochemical immunosensors for detection of cancer protein biomarkers. ACS Nano 2012, 6, 6546–6561. [Google Scholar] [CrossRef]
- Wu, L.; Qu, X.G. Cancer biomarker detection: Recent achievements and challenges. Chem. Soc. Rev. 2015, 44, 2963–2997. [Google Scholar] [CrossRef]
- Wang, W.S.; Lin, J.K.; Lin, T.C.; Chiou, T.J.; Liu, J.H.; Yen, C.C.; Chen, W.S.; Jiang, J.K.; Yang, S.H.; Wang, H.S.; et al. EIA versus RIA in detecting carcinoembryonic antigen level of patients with metastatic colorectal cancer. Hepato-Gastroenterol. 2004, 51, 136–141. [Google Scholar]
- Lim, S.; Janzer, A.; Becker, A.; Zimmer, A.; Schule, R.; Buettner, R.; Kirfel, J. Lysine-specific demethylase 1 (LSD1) is highly expressed in ER-negative breast cancers and a biomarker predicting aggressive biology. Carcinogenesis 2010, 31, 512–520. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.E.; Kim, Y.J.; Song, S.; Lee, K.N.; Seong, W.K. A simple electrochemical immunosensor platform for detection of apolipoprotein A1 (Apo-A1) as a bladder cancer biomarker in urine. Sens. Actuator B Chem. 2019, 278, 103–109. [Google Scholar] [CrossRef]
- Ambrosi, A.; Airo, F.; Merkoci, A. Enhanced gold nanoparticle based ELISA for a breast cancer biomarker. Anal. Chem. 2010, 82, 1151–1156. [Google Scholar] [CrossRef]
- Fitzgerald, S.; O’Reilly, J.A.; Wilson, E.; Joyce, A.; Farrell, R.; Kenny, D.; Kay, E.W.; Fitzgerald, J.; Byrne, B.; Kijanka, G.S.; et al. Measurement of the IgM and IgG autoantibody immune responses in human serum has high predictive value for the presence of colorectal cancer. Clin. Colorectal Cancer 2018, in press. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.D.; Peng, J.; Jiang, L.P.; Zhu, J.J. Fluorescent immunosensor based on CuS nanoparticles for sensitive detection of cancer biomarker. Analyst 2014, 139, 649–655. [Google Scholar] [CrossRef]
- Zhou, Y.F.; Huang, X.L.; Xiong, S.C.; Li, X.M.; Zhan, S.N.; Zeng, L.F.; Xiong, Y.H. Dual-mode fluorescent and colorimetric immunoassay for the ultrasensitive detection of alpha-fetoprotein in serum samples. Anal. Chim. Acta 2018, 1038, 112–119. [Google Scholar] [CrossRef] [PubMed]
- Chon, H.; Lee, S.; Son, S.W.; Oh, C.H.; Choo, J. Highly sensitive immunoassay of lung cancer marker carcinoembryonic antigen using surface-enhanced Raman scattering of hallow gold nanospheres. Anal. Chem. 2009, 81, 3029–3034. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.F.; Lipert, R.J.; Jain, M.; Kaur, S.; Chakraboty, S.; Torres, M.P.; Batra, S.K.; Brand, R.E.; Porter, M.D. Detection of the potential pancreatic cancer marker MUC4 in serum using surface-enhanced Raman scattering. Anal. Chem. 2011, 83, 2554–2561. [Google Scholar] [CrossRef]
- Li, J.R.; Wang, J.; Grewal, Y.S.; Howard, C.B.; Raftery, L.J.; Mahler, S.; Wang, Y.L.; Trau, M. Multiplexed sers detection of soluble cancer protein biomarkers with gold-silver alloy nanoboxes and nanoyeast single-chain variable fragments. Anal. Chem. 2018, 90, 10377–10384. [Google Scholar] [CrossRef]
- Nguyen, A.H.; Lee, J.; Choi, H.I.; Kwak, H.S.; Sim, S.J. Fabrication of plasmon length-based surface-enhanced Raman scattering for multiplex detection on microfluidic device. Biosens. Bioelectron. 2015, 70, 358–365. [Google Scholar] [CrossRef]
- Ma, H.; Sun, X.Y.; Chen, L.; Han, X.X.; Zhao, B.; Lu, H.; He, C.Y. Antibody-free discrimination of protein biomarkers in human serum based on surface-enhanced Raman spectroscopy. Anal. Chem. 2018, 90, 12342–12346. [Google Scholar] [CrossRef]
- Qian, X.M.; Peng, X.H.; Ansari, D.O.; Yin-Goen, Q.; Chen, G.Z.; Shin, D.M.; Yang, L.; Young, A.N.; Wang, M.D.; Nie, S.M. In vivo tumor targeting and spectroscopic detection with surface-enhanced Raman nanoparticle tags. Nat. Biotechnol. 2008, 26, 83–90. [Google Scholar] [CrossRef]
- Faulds, K.; McKenzie, F.; Smith, W.E.; Graham, D. Quantitative simultaneous multianalyte detection of DNA by dual-wavelength surface-enhanced resonance Raman scattering. Angew. Chem.-Int. Ed. 2007, 46, 1829–1831. [Google Scholar] [CrossRef]
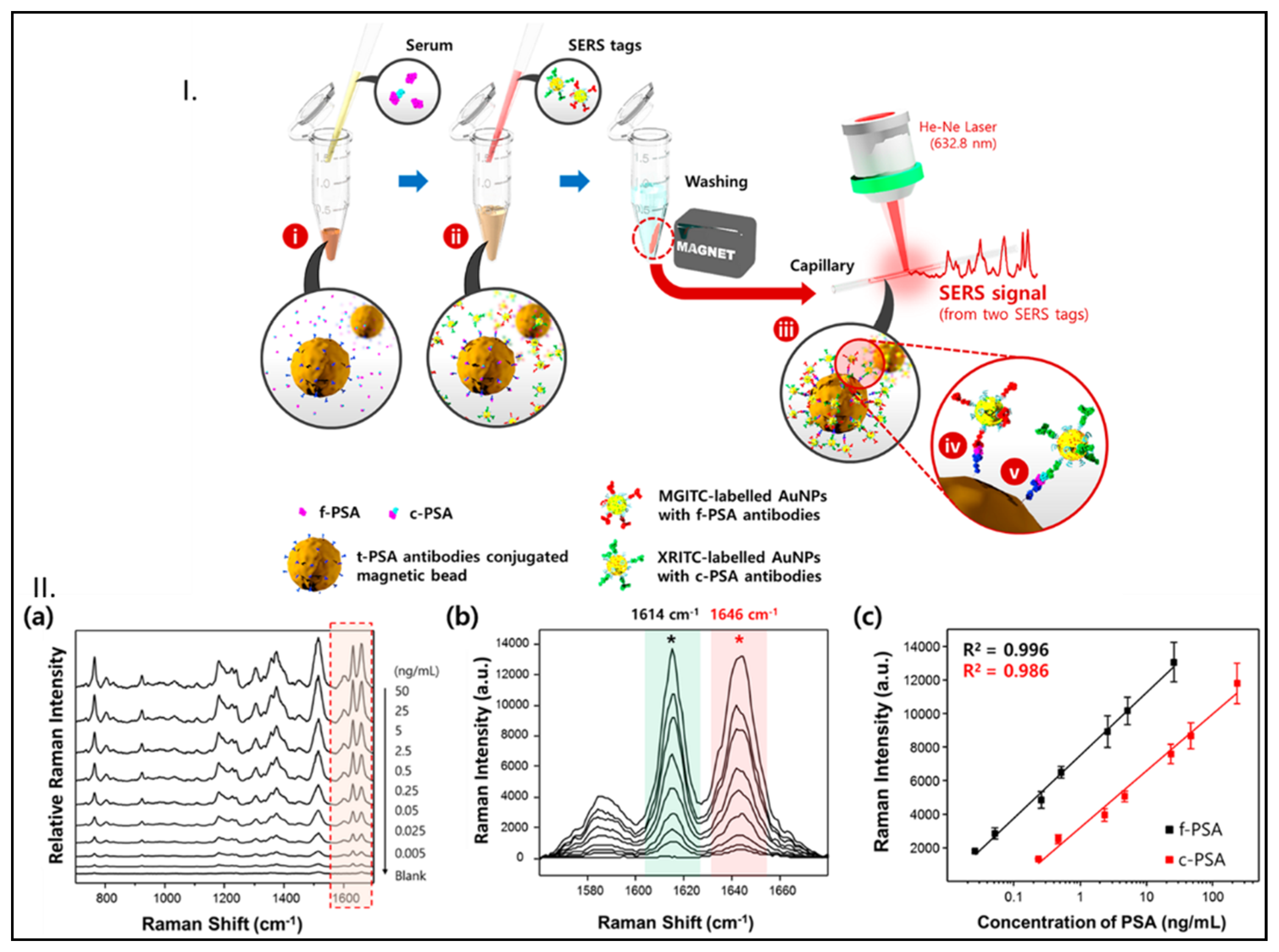
- Cheng, Z.; Choi, N.; Wang, R.; Lee, S.; Moon, K.C.; Yoon, S.Y.; Chen, L.X.; Choo, J. Simultaneous detection of dual prostate specific antigens using surface-enhanced Raman scattering-based immunoassay for accurate diagnosis of prostate cancer. ACS Nano 2017, 11, 4926–4933. [Google Scholar] [CrossRef]
- Banaei, N.; Foley, A.; Houghton, J.M.; Sun, Y.B.; Kim, B. Multiplex detection of pancreatic cancer biomarkers using a SERS-based immunoassay. Nanotechnology 2017, 28, 455101. [Google Scholar] [CrossRef]
- Hwang, H.; Chon, H.; Choo, J.; Park, J.K. Optoelectrofluidic sandwich immunoassays for detection of human tumor marker using surface-enhanced Raman scattering. Anal. Chem. 2010, 82, 7603–7610. [Google Scholar] [CrossRef]
- Neng, J.; Harpster, M.H.; Zhang, H.; Mecham, J.O.; Wilson, W.C.; Johnson, P.A. A versatile SERS-based immunoassay for immunoglobulin detection using antigen-coated gold nanoparticles and malachite green-conjugated protein A/G. Biosens. Bioelectron. 2010, 26, 1009–1015. [Google Scholar] [CrossRef]
- Gao, R.K.; Cheng, Z.Y.; Demello, A.J.; Choo, J. Wash-free magnetic immunoassay of the PSA cancer marker using SERS and droplet microfluidics. Lab Chip 2016, 16, 1022–1029. [Google Scholar] [CrossRef]
- Perozziello, G.; Candeloro, P.; Gentile, F.; Nicastri, A.; Perri, A.; Coluccio, M.L.; Adamo, A.; Pardeo, F.; Catalano, R.; Parrotta, E.; et al. Microfluidics & nanotechnology: Towards fully integrated analytical devices for the detection of cancer biomarkers. RSC Adv. 2014, 4, 55590–55598. [Google Scholar]
- Wee, E.J.H.; Wang, Y.L.; Tsao, S.C.H.; Trau, M. Simple, sensitive and accurate multiplex detection of clinically important melanoma DNA mutations in circulating tumour DNA with SERS nanotags. Theranostics 2016, 6, 1506–1513. [Google Scholar] [CrossRef]
- Goodacre, R. Metabolomics of a superorganism. J. Nutr. 2007, 137, 259S–266S. [Google Scholar] [CrossRef]
- World Health Organisation Global Statistics. p. 30. Available online: http://apps.Who.Int/iris/bitstream/handle/10665/255336/9789241565486-eng.Pdf?Sequence=1 (accessed on 3 January 2018).
- Ellis, D.I.; Muhamadali, H.; Chisanga, M.; Goodacre, R. Omics methods for the detection of foodborne pathogens. Encycl. Food Chem. 2019, 1, 364–370. [Google Scholar]
- World Health Organisation. Available online: https://www.Who.Int/sustainable-development/housing/health-risks/waterborne-disease/en/ (accessed on 29 December 2018).
- Food Consulting Strategically. Available online: https://www.Focos-food.Com/campylobacter-on-the-rise-in-germany-every-second-chicken-in-germany-is-contaminated/ (accessed on 12 January 2019).
- Lazcka, O.; Del Campo, F.J.; Munoz, F.X. Pathogen detection: A perspective of traditional methods and biosensors. Biosens. Bioelectron. 2007, 22, 1205–1217. [Google Scholar] [CrossRef] [PubMed]
- Frieden, T.R.; Sterling, T.R.; Munsiff, S.S.; Watt, C.J.; Dye, C. Tuberculosis. Lancet 2003, 362, 887–899. [Google Scholar] [CrossRef]
- Muhamadali, H.; Weaver, D.; Subaihi, A.; AlMasoud, N.; Trivedi, D.K.; Ellis, D.I.; Linton, D.; Goodacre, R. Chicken, beams, and Campylobacter: Rapid differentiation of foodborne bacteria via vibrational spectroscopy and MALDI-mass spectrometry. Analyst 2016, 141, 111–122. [Google Scholar] [CrossRef] [PubMed]
- Santiago-Felipe, S.; Tortajada-Genaro, L.A.; Puchades, R.; Maquieira, A. Recombinase polymerase and enzyme-linked immunosorbent assay as a DNA amplification-detection strategy for food analysis. Anal. Chim. Acta 2014, 811, 81–87. [Google Scholar] [CrossRef]
- Pang, B.; Zhao, C.; Li, L.; Song, X.L.; Xu, K.; Wang, J.; Liu, Y.S.; Fu, K.Y.; Bao, H.; Song, D.D.; et al. Development of a low-cost paper-based ELISA method for rapid Escherichia coli O157:H7 detection. Anal. Biochem. 2018, 542, 58–62. [Google Scholar] [CrossRef]
- Catala, C.; Mir-Simon, B.; Feng, X.T.; Cardozo, C.; Pazos-Perez, N.; Pazos, E.; Gomez-de Pedro, S.; Guerrini, L.; Soriano, A.; Vila, J.; et al. Online SERS quantification of Staphylococcus aureus and the application to diagnostics in human fluids. Adv. Mater. Technol. 2016, 1, 1600163. [Google Scholar] [CrossRef]
- Muhamadali, H.; Subaihi, A.; Mohammadtaheri, M.; Xu, Y.; Ellis, D.I.; Ramanathan, R.; Bansal, V.; Goodacre, R. Rapid, accurate, and comparative differentiation of clinically and industrially relevant microorganisms via multiple vibrational spectroscopic fingerprinting. Analyst 2016, 141, 5127–5136. [Google Scholar] [CrossRef]
- Tien, N.; Lin, T.H.; Hung, Z.C.; Lin, H.S.; Wang, I.K.; Chen, H.C.; Chang, C.T. Diagnosis of bacterial pathogens in the urine of urinary-tract-infection patients using surface-enhanced Raman spectroscopy. Molecules 2018, 23, 3374. [Google Scholar] [CrossRef]
- Shanmukh, S.; Jones, L.; Zhao, Y.P.; Driskell, J.D.; Tripp, R.A.; Dluhy, R.A. Identification and classification of respiratory syncytial virus (RSV) strains by surface-enhanced Raman spectroscopy and multivariate statistical techniques. Anal. Bioanal. Chem. 2008, 390, 1551–1555. [Google Scholar] [CrossRef]
- Chen, Y.; Premasiri, W.R.; Ziegler, L.D. Surface-enhanced Raman spectroscopy of Chlamydia trachomatis and Neisseria gonorrhoeae for diagnostics, and extra-cellular metabolomics and biochemical monitoring. Sci. Rep. 2018, 8, 5163. [Google Scholar] [CrossRef]
- Witkowska, E.; Jagielski, T.; Kaminska, A. Genus- and species-level identification of dermatophyte fungi by surface-enhanced Raman spectroscopy. Spectroc. Acta Part A Mol. Biomol. Spectr. 2018, 192, 285–290. [Google Scholar] [CrossRef] [PubMed]
- Chisanga, M.; Muhamadali, H.; Kimber, R.; Goodacre, R. Quantitative detection of isotopically enriched E. coli cells by SERS. Faraday Discuss. 2017, 205, 331–343. [Google Scholar] [CrossRef]
- Efrima, S.; Zeiri, L. Understanding SERS of bacteria. J. Raman Spectrosc. 2009, 40, 277–288. [Google Scholar] [CrossRef]
- Jarvis, R.M.; Brooker, A.; Goodacre, R. Surface-enhanced Raman spectroscopy for bacterial discrimination utilizing a scanning electron microscope with a Raman spectroscopy interface. Anal. Chem. 2004, 76, 5198–5202. [Google Scholar] [CrossRef] [PubMed]
- Dina, N.E.; Gherman, A.M.R.; Chis, V.; Sarbu, C.; Wieser, A.; Bauer, D.; Haisch, C. Characterization of clinically relevant fungi via SERS fingerprinting assisted by novel chemometric models. Anal. Chem. 2018, 90, 2484–2492. [Google Scholar] [CrossRef]
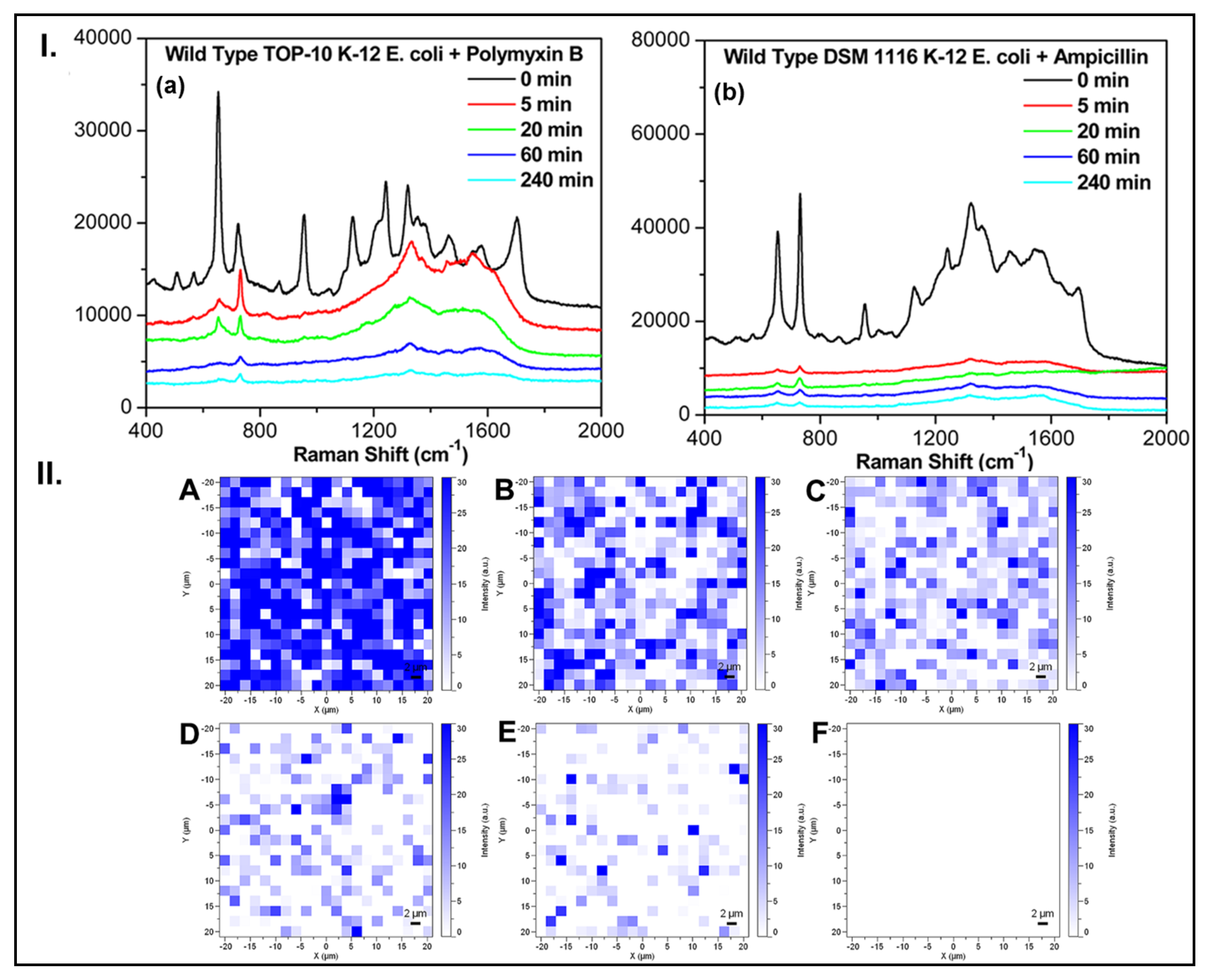
- Liu, C.Y.; Han, Y.Y.; Shih, P.H.; Lian, W.N.; Wang, H.H.; Lin, C.H.; Hsueh, P.R.; Wang, J.K.; Wang, Y.L. Rapid bacterial antibiotic susceptibility test based on simple surface-enhanced Raman spectroscopic biomarkers. Sci. Rep. 2016, 6, 23375. [Google Scholar] [CrossRef]
- Berry, D.; Mader, E.; Lee, T.K.; Woebken, D.; Wang, Y.; Zhu, D.; Palatinszky, M.; Schintimeister, A.; Schmid, M.C.; Hanson, B.T.; et al. Tracking heavy water (D2O) incorporation for identifying and sorting active microbial cells. Proc. Natl. Acad. Sci. USA 2015, 112, E194–E203. [Google Scholar] [CrossRef]
- Wang, Y.; Song, Y.Z.; Tao, Y.F.; Muhamadali, H.; Goodacre, R.; Zhou, N.Y.; Preston, G.M.; Xu, J.; Huang, W.E. Reverse and multiple stable isotope probing to study bacterial metabolism and interactions at the single cell level. Anal. Chem. 2016, 88, 9443–9450. [Google Scholar] [CrossRef]
- Leask, F. Renishaw diagnostics Announces CE Marking of RenDx Multiplex Assay System and Fungiplex Assay. SelectScience. 2015. Available online: http://www.selectscience.net/industry-news/renishaw-diagnostics-announces-cemarking-of-rendx-multiplex-assay-system-andfungiplex-assay/?artID=38483 (accessed on 30 December 2018).
- Neng, J.; Li, Y.N.; Driscoll, A.J.; Wilson, W.C.; Johnson, P.A. Detection of multiple pathogens in serum using silica-encapsulated nanotags in a surface-enhanced Raman scattering-based immunoassay. J. Agric. Food Chem. 2018, 66, 5707–5712. [Google Scholar] [CrossRef] [PubMed]
- Sebba, D.; Lastovich, A.G.; Kuroda, M.; Fallows, E.; Johnson, J.; Ahouidi, A.; Honko, A.N.; Fu, H.; Nielson, R.; Carruthers, E.; et al. A point-of-care diagnostic for differentiating Ebola from endemic febrile diseases. Sci. Transl. Med. 2018, 10, eaat0944. [Google Scholar] [CrossRef] [PubMed]
- Muhlig, A.; Bocklitz, T.; Labugger, I.; Dees, S.; Henk, S.; Richter, E.; Andres, S.; Merker, M.; Stockel, S.; Weber, K.; et al. LOC-SERS: A promising closed system for the identification of mycobacteria. Anal. Chem. 2016, 88, 7998–8004. [Google Scholar] [CrossRef]
- Willner, M.R.; McMillan, K.S.; Graham, D.; Vikesland, P.J.; Zagnoni, M. Surface-enhanced Raman scattering based microfluidics for single-cell analysis. Anal. Chem. 2018, 90, 12004–12010. [Google Scholar] [CrossRef] [PubMed]
- Patel, I.S.; Premasiri, W.R.; Moir, D.T.; Ziegler, L.D. Barcoding bacterial cells: A SERS-based methodology for pathogen identification. J. Raman Spectrosc. 2008, 39, 1660–1672. [Google Scholar] [CrossRef]
- Shikha, S.; Salafi, T.; Cheng, J.T.; Zhang, Y. Versatile design and synthesis of nano-barcodes. Chem. Soc. Rev. 2017, 46, 7054–7093. [Google Scholar] [CrossRef]
- Bos, L.D.; Sterk, P.J.; Fowler, S.J. Breathomics in the setting of asthma and chronic obstructive pulmonary disease. J. Allergy Clin. Immunol. 2016, 138, 970–976. [Google Scholar] [CrossRef] [PubMed]
- Rattray, N.J.W.; Hamrang, Z.; Trivedi, D.K.; Goodacre, R.; Fowler, S.J. Taking your breath away: Metabolomics breathes life in to personalized medicine. Trends Biotechnol. 2014, 32, 538–548. [Google Scholar] [CrossRef] [PubMed]
- Alonso, M.; Sanchez, J.M. Analytical challenges in breath analysis and its application to exposure monitoring. TrAC-Trends Anal. Chem. 2013, 44, 78–89. [Google Scholar] [CrossRef]
- de Vries, R.; Dagelet, Y.W.F.; Spoor, P.; Snoey, E.; Jak, P.M.C.; Brinkman, P.; Dijkers, E.; Bootsma, S.K.; Elskamp, F.; de Jongh, F.H.C.; et al. Clinical and inflammatory phenotyping by breathomics in chronic airway diseases irrespective of the diagnostic label. Eur. Resp. J. 2018, 51, 1701817. [Google Scholar] [CrossRef]
- Neerincx, A.H.; Vijverberg, S.J.H.; Bos, L.D.J.; Brinkman, P.; van der Schee, M.P.; de Vries, R.; Sterk, P.J.; Maitland-van der Zee, A.H. Breathomics from exhaled volatile organic compounds in pediatric asthma. Pediatr. Pulmonol. 2017, 52, 1616–1627. [Google Scholar] [CrossRef] [PubMed]
- Aizpurua, J.; Arnolds, H.; Baumberg, J.; Bruzas, I.; Chikkaraddy, R.; Chisanga, M.; Dawson, P.; Deckert, V.; Delfino, I.; de Nijs, B.; et al. Ultrasensitive and towards single molecule SERS: General discussion. Faraday Discuss. 2017, 205, 291–330. [Google Scholar] [CrossRef] [PubMed]
- Wong, C.L.; Dinish, U.S.; Buddharaju, K.D.; Schmidt, M.S.; Olivo, M. Surface-enhanced Raman scattering (SERS)-based volatile organic compounds (VOCs) detection using plasmonic bimetallic nanogap substrate. Appl. Phys. A Mater. Sci. Process. 2014, 117, 687–692. [Google Scholar] [CrossRef]
- Wong, C.L.; Dinish, U.S.; Schmidt, M.S.; Olivo, M. Non-labeling multiplex surface-enhanced Raman scattering (SERS) detection of volatile organic compounds (VOCs). Anal. Chim. Acta 2014, 844, 54–60. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.S.; Zhang, Y.X.; Pan, F.; Liu, J.; Wang, K.; Zhang, C.L.; Cheng, S.L.; Lu, L.G.; Zhang, W.; Zhang, Z.; et al. Breath analysis based on surface-enhanced Raman scattering sensors distinguishes early and advanced gastric cancer patients from healthy persons. ACS Nano 2016, 10, 8169–8179. [Google Scholar] [CrossRef]
- Mason, E.A.; Kronstadt, B. Graham’s laws of diffusion and effusion. J. Chem. Educ. 1967, 44, 740. [Google Scholar] [CrossRef]
- Kim, S.; Kim, D.H.; Park, S.G. Highly sensitive and on-site NO2 SERS sensors operated under ambient conditions. Analyst 2018, 143, 3006–3010. [Google Scholar] [CrossRef]
- Zhang, Z.; Yu, W.; Wang, J.; Luo, D.; Qiao, X.Z.; Qin, X.Y.; Wang, T. Ultrasensitive surface-enhanced Raman scattering sensor of gaseous aldehydes as biomarkers of lung cancer on dendritic Ag nanocrystals. Anal. Chem. 2017, 89, 1416–1420. [Google Scholar] [CrossRef]
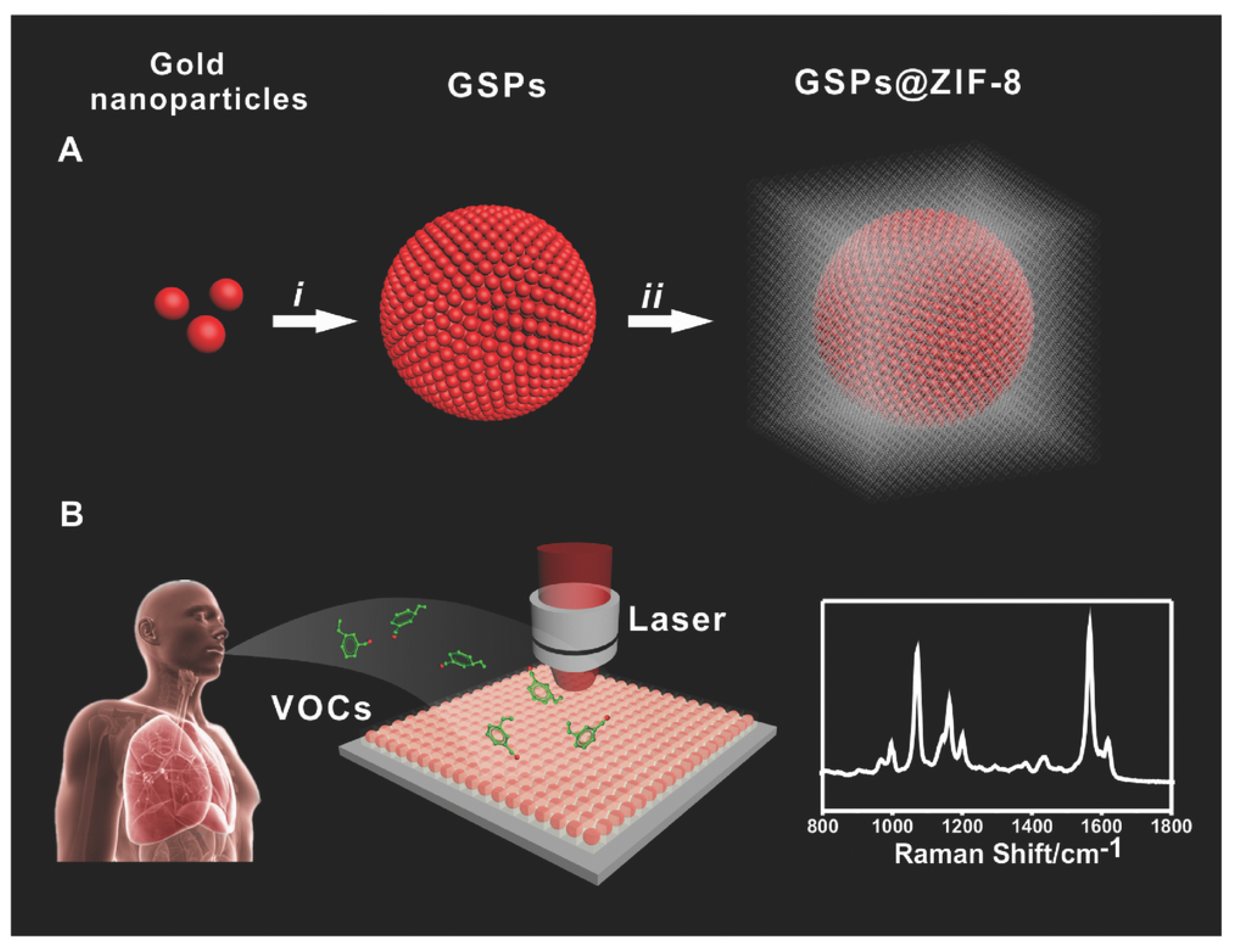
- Qiao, X.Z.; Su, B.S.; Liu, C.; Song, Q.; Luo, D.; Mo, G.; Wang, T. Selective surface-enhanced Raman scattering for quantitative detection of lung cancer biomarkers in superparticle@MOF structure. Adv. Mater. 2018, 30, 1702275. [Google Scholar] [CrossRef]
- Lawal, O.; Knobel, H.; Weda, H.; Bos, L.D.; Nijsen, T.M.E.; Goodacre, R.; Fowler, S.J. Volatile organic compound signature from co-culture of lung epithelial cell line with Pseudomonas aeruginosa. Analyst 2018, 143, 3148–3155. [Google Scholar] [CrossRef]
- Lemfack, M.C.; Gohlke, B.O.; Toguem, S.M.T.; Preissner, S.; Piechulla, B.; Preissner, R. mVOC 2.0: A database of microbial volatiles. Nucleic Acids Res. 2018, 46, D1261–D1265. [Google Scholar] [CrossRef] [PubMed]
- Lauridsen, R.K.; Sommer, L.M.; Johansen, H.K.; Rindzevicius, T.; Molin, S.; Jelsbak, L.; Engelsen, S.B.; Boisen, A. SERS detection of the biomarker hydrogen cyanide from Pseudomonas aeruginosa cultures isolated from cystic fibrosis patients. Sci. Rep. 2017, 7, 45264. [Google Scholar] [CrossRef] [PubMed]
- Bos, L.D.J.; Sterk, P.J.; Schultz, M.J. Volatile metabolites of pathogens: A systematic review. PLoS Pathog. 2013, 9, e1003311. [Google Scholar] [CrossRef] [PubMed]
- Park, K.J.; Wu, C.; Mercer-Smith, A.R.; Dodson, R.A.; Moersch, T.L.; Koonath, P.; Pipino, A.C.R.; Lu, H.W.; Yang, Y.W.; Sapirstein, V.S.; et al. Raman system for sensitive and selective identification of volatile organic compounds. Sens. Actuator B Chem. 2015, 220, 491–499. [Google Scholar] [CrossRef]
- Junger, M.; Vautz, W.; Kuhns, M.; Hofmann, L.; Ulbricht, S.; Baumbach, J.I.; Quintel, M.; Perl, T. Ion mobility spectrometry for microbial volatile organic compounds: A new identification tool for human pathogenic bacteria. Appl. Microbiol. Biotechnol. 2012, 93, 2603–2614. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.J.; Bean, H.D.; Wargo, M.J.; Leclair, L.W.; Hill, J.E. Detecting bacterial lung infections: In vivo evaluation of in vitro volatile fingerprints. J. Breath Res. 2013, 7, 016003. [Google Scholar] [CrossRef] [PubMed]
- Kelly, J.; Patrick, R.; Patrick, S.; Bell, S.E.J. Surface-enhanced Raman spectroscopy for the detection of a metabolic product in the headspace above live bacterial cultures. Angew. Chem.-Int. Ed. 2018, 57, 15686–15690. [Google Scholar] [CrossRef] [PubMed]
- Subaihi, A.; Muhamadali, H.; Mutter, S.T.; Blanch, E.; Ellis, D.I.; Goodacre, R. Quantitative detection of codeine in human plasma using surface-enhanced Raman scattering via adaptation of the isotopic labelling principle. Analyst 2017, 142, 1099–1105. [Google Scholar] [CrossRef]
- Itoh, N.; Bell, S.E.J. High dilution surface-enhanced Raman spectroscopy for rapid determination of nicotine in e-liquids for electronic cigarettes. Analyst 2017, 142, 994–998. [Google Scholar] [CrossRef]
- Westley, C.; Xu, Y.; Thilaganathan, B.; Carnell, A.J.; Turner, N.J.; Goodacre, R. Absolute quantification of uric acid in human urine using surface-enhanced Raman scattering with the standard addition method. Anal. Chem. 2017, 89, 2472–2477. [Google Scholar] [CrossRef]
- Premasiri, W.R.; Lee, J.C.; Sauer-Budge, A.; Theberge, R.; Costello, C.E.; Ziegler, L.D. The biochemical origins of the surface-enhanced Raman spectra of bacteria: A metabolomics profiling by SERS. Anal. Bioanal. Chem. 2016, 408, 4631–4647. [Google Scholar] [CrossRef] [PubMed]
- Kubryk, P.; Niessner, R.; Ivleva, N.P. The origin of the band at around 730 cm−1 in the SERS spectra of bacteria: A stable isotope approach. Analyst 2016, 141, 2874–2878. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.Y.; Wei, Q.L.; Ma, F.; Li, X.; Liu, F.Y.; Zhou, M. Surface-enhanced Raman nanoparticles for tumor theranostics applications. Acta Pharm. Sin. B 2018, 8, 349–359. [Google Scholar] [CrossRef] [PubMed]
- Baker, M.J.; Byrne, H.J.; Chalmers, J.; Gardner, P.; Goodacre, R.; Henderson, A.; Kazarian, S.G.; Martin, F.L.; Moger, J.; Stone, N.; et al. Clinical applications of infrared and Raman spectroscopy: state of play and future challenges. Analyst 2018, 143, 1735–1757. [Google Scholar] [CrossRef] [PubMed]
- Neugebauer, U.; Schmid, U.; Baumann, K.; Ziebuhr, W.; Kozitskaya, S.; Deckert, V.; Schmitt, M.; Popp, J. Towards a detailed understanding of bacterial metabolism—Spectroscopic characterization of Staphylococcus epidermidis. ChemPhysChem 2007, 8, 124–137. [Google Scholar] [CrossRef] [PubMed]
- Ashton, L.; Hogwood, C.E.M.; Tait, A.S.; Kuligowski, J.; Smales, C.M.; Bracewell, D.G.; Dickson, A.J.; Goodacre, R. UV resonance Raman spectroscopy: A process analytical tool for host cell DNA and RNA dynamics in mammalian cell lines. J. Chem. Technol. Biotechnol. 2015, 90, 237–243. [Google Scholar] [CrossRef]
- Matousek, P.; Stone, N. Development of deep subsurface Raman spectroscopy for medical diagnosis and disease monitoring. Chem. Soc. Rev. 2016, 45, 1794–1802. [Google Scholar] [CrossRef]
- Stone, N.; Kerssens, M.; Lloyd, G.R.; Faulds, K.; Graham, D.; Matousek, P. Surface-enhanced spatially offset Raman spectroscopic (SESORS) imaging—The next dimension. Chem. Sci. 2011, 2, 776–780. [Google Scholar] [CrossRef]
- Matousek, P.; Stone, N. Recent advances in the development of Raman spectroscopy for deep non-invasive medical diagnosis. J. Biophotonics 2013, 6, 7–19. [Google Scholar] [CrossRef]
- Nicolson, F.; Jamieson, L.E.; Mabbott, S.; Plakas, K.; Shand, N.C.; Detty, M.R.; Graham, D.; Faulds, K. Through tissue imaging of a live breast cancer tumour model using handheld surface-enhanced spatially offset resonance Raman spectroscopy (SESORRS). Chem. Sci. 2018, 9, 3788–3792. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, R.; Jiang, S.; Zhang, Y.; Dong, Z.C. Probing adsorption configurations of small molecules on surfaces by single-molecule tip-enhanced Raman spectroscopy. ChemPhysChem 2019, 20, 37–41. [Google Scholar] [CrossRef]
- Berezin, S.; Aviv, Y.; Aviv, H.; Goldberg, E.; Tischler, Y.R. Replacing a century old technique—Modern spectroscopy can supplant Gram staining. Sci. Rep. 2017, 7, 3810. [Google Scholar] [CrossRef]
- Ji, M.B.; Arbel, M.; Zhang, L.L.; Freudiger, C.W.; Hou, S.S.; Lin, D.D.; Yang, X.J.; Bacskai, B.J.; Xie, X.S. Label-free imaging of amyloid plaques in Alzheimer’s disease with stimulated Raman scattering microscopy. Sci. Adv. 2018, 4, eaat7715. [Google Scholar] [CrossRef]
- Liao, C.S.; Wang, P.; Huang, C.Y.; Lin, P.; Eakins, G.; Bentley, R.T.; Liang, R.G.; Cheng, J.X. In vivo and in situ spectroscopic imaging by a handheld stimulated Raman scattering microscope. ACS Photonics 2018, 5, 947–954. [Google Scholar] [CrossRef]
- Evans, C.L.; Xie, X.S. Coherent anti-Stokes Raman scattering microscopy: Chemical imaging for biology and medicine. Annu. Rev. Anal. Chem. 2008, 1, 883–909. [Google Scholar] [CrossRef]
- Weng, S.; Xu, X.Y.; Li, J.S.; Wong, S.T.C. Combining deep learning and coherent anti-Stokes Raman scattering imaging for automated differential diagnosis of lung cancer. J. Biomed. Opt. 2017, 22, 106017. [Google Scholar] [CrossRef]
| Enhanced Raman Techniques | Brief Description |
|---|---|
| Resonance Raman (RR) scattering | RR occurs when the incident frequency excites electronic energy (EE) states of specific molecules; e.g., aromatic chromophores, and this aids the detection of pathogens [161]. If the EE level is excited by laser frequency in the ultraviolet (UV) region, the technique is UVRR, and can be susceptible to photodissociation [162]. |
| Spatially offset Raman spectroscopy (SORS) | SORS collects distinctive chemical information and images from deep subsurfaces (including analytes in opaque containers) of a sample. SORS spectral signals are recorded when backscattered Raman photons are collected at points spatially offset (Δx) from the point of illumination (x). Negligible photodissociation, allowing for noninvasive deep medical diagnostics [163]. |
| Surface-enhanced spatially offset Raman spectroscopy (SESORS) | SORS combined with NPs that enable SERS: that is to say subsurface information is measured from molecules in the vicinity of or chemically bonded to SERS substrates. Negligible fluorescence and excellent background contrast, specificity and sensitivity with improved detection limit for various disease markers [164,165]. |
| Surface-enhanced spatially offset resonance Raman spectroscopy (SESORRS) | A variant of SESORS where incident frequency matches the EE of molecules near SERS-active substrates. SESORRS increases spectral signals further by orders of magnitude to provide extra biochemical selectivity and sensitivity, theoretically better than SESORS, as demonstrated by Fay et al. for breast cancer detection [166]. |
| Tip-enhanced Raman spectroscopy (TERS) | Similar to the SERS phenomenon, but here a single SERS-active AFM probe whose sharp pointed apex (tip) is covered in NPs and scans through biomolecules on a sample surface, resulting in highly confined plasmonic enhancement (electrostatic lightning rod and SPR effects). Improves lateral spatial resolution to as low as 10 nm, about the diameter of the tip probe. Achieves single molecule detection and discrimination of bacterial pathogens [167,168]. |
| Stimulated Raman scattering (SRS) | Two lasers provide a pump (ωp) (similar to conventional Raman) and Stokes (ωs) beam frequencies that intersect at the sample surface. The energy difference (Δω = ωp − ωs) between the beams matches the frequency (Ωvib) of molecular bond vibrations, leading to larger scattering cross-section as a consequence of stimulated Raman excitation. No nonresonance background [169], making SRS ideal for in vivo medical imaging to improve disease diagnostics [170]. |
| Coherent anti-Stokes Raman scattering (CARS) | Like SRS, CARS employs two lasers of frequencies ωp and ωs. When molecular bonds whose Ωvib coincide with Δω (as in SRS), anti-Stokes (as) lines are produced at frequency ωas = 2ωp − ωs. Thus, analytes are excited twice, from the ground to first and second excited states before relaxing back to the ground state. Though prone to nonresonance background effects which may limit the quantification of target analytes [171], CARS is effectively applied for disease detection including differential diagnostics of cancers [172]. |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chisanga, M.; Muhamadali, H.; Ellis, D.I.; Goodacre, R. Enhancing Disease Diagnosis: Biomedical Applications of Surface-Enhanced Raman Scattering. Appl. Sci. 2019, 9, 1163. https://doi.org/10.3390/app9061163
Chisanga M, Muhamadali H, Ellis DI, Goodacre R. Enhancing Disease Diagnosis: Biomedical Applications of Surface-Enhanced Raman Scattering. Applied Sciences. 2019; 9(6):1163. https://doi.org/10.3390/app9061163
Chicago/Turabian StyleChisanga, Malama, Howbeer Muhamadali, David I. Ellis, and Royston Goodacre. 2019. "Enhancing Disease Diagnosis: Biomedical Applications of Surface-Enhanced Raman Scattering" Applied Sciences 9, no. 6: 1163. https://doi.org/10.3390/app9061163
APA StyleChisanga, M., Muhamadali, H., Ellis, D. I., & Goodacre, R. (2019). Enhancing Disease Diagnosis: Biomedical Applications of Surface-Enhanced Raman Scattering. Applied Sciences, 9(6), 1163. https://doi.org/10.3390/app9061163







