Concentrations of Multiple Phytoplankton Pigments in the Global Oceans Obtained from Satellite Ocean Color Measurements with MERIS
Abstract
1. Introduction
2. Data and Methods
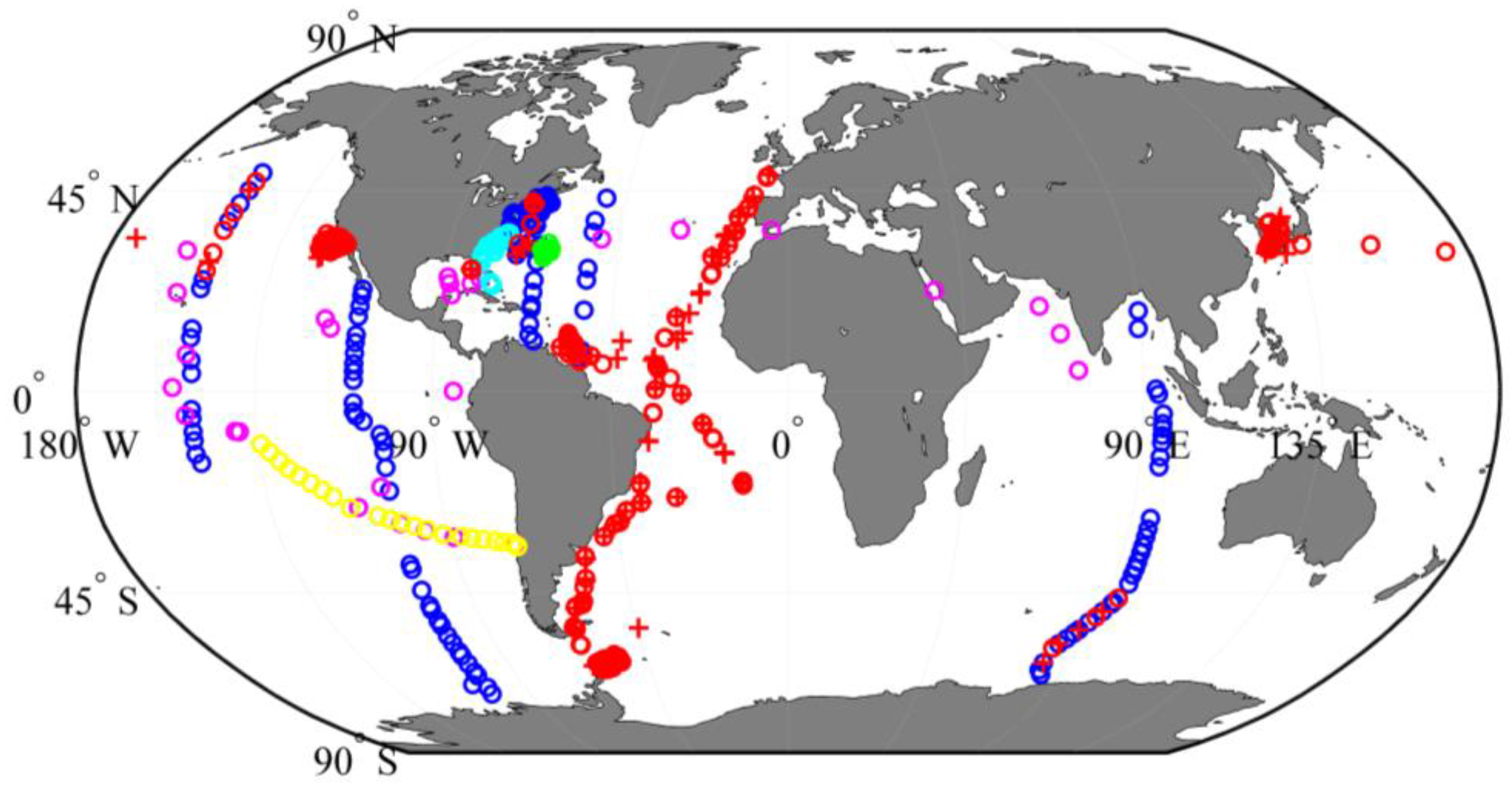
2.1. Datasets and Study Sites
2.2. Radiometric Measurements
2.3. Absorption Measurements
2.4. Pigment Concentrations
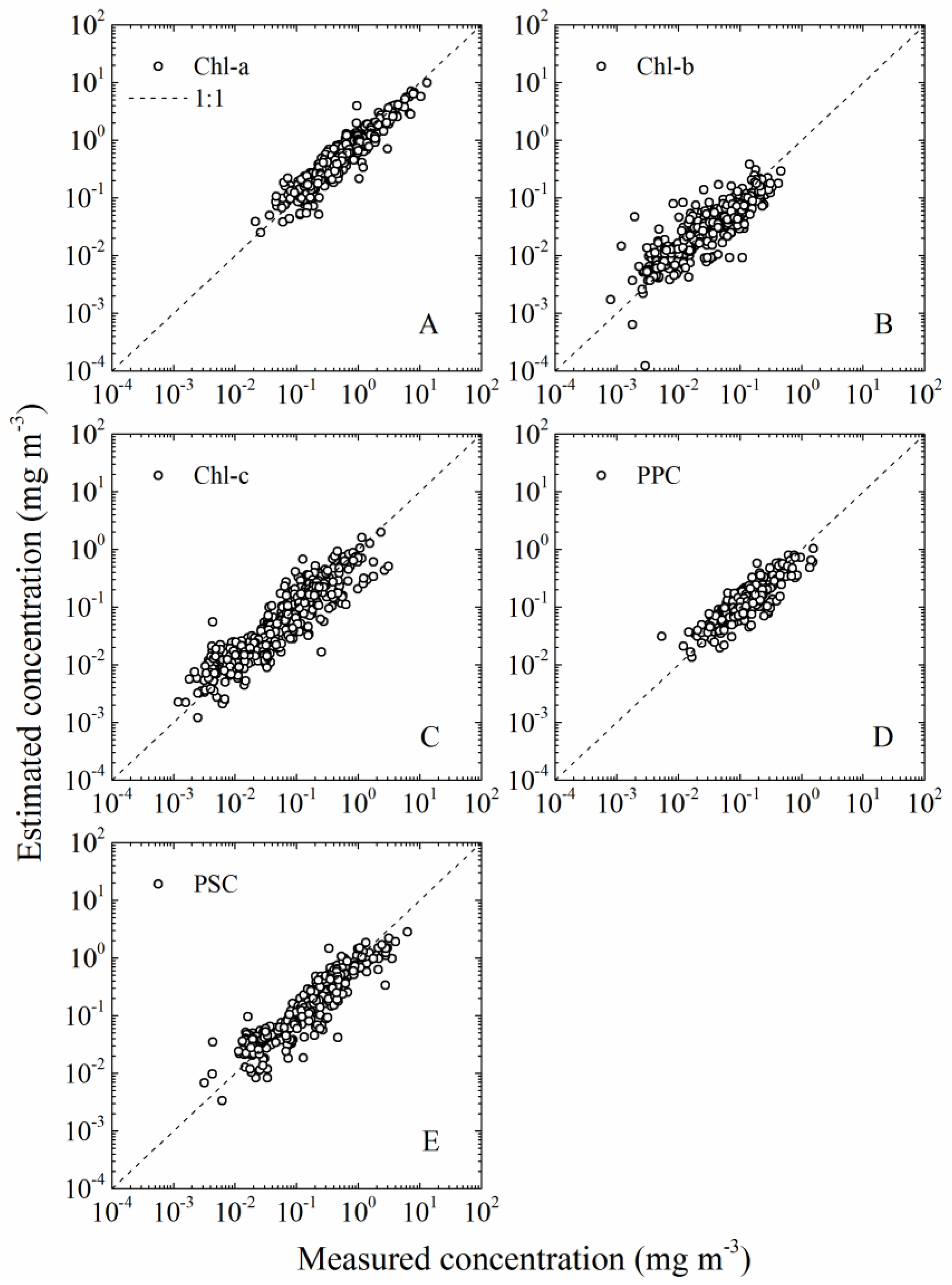
- (A)
- Total chlorophyll a (Chl-a) = chlorophyll a + divinyl chlorophyll a + chlorophyllide a;
- (B)
- Total chlorophyll b (Chl-b) = chlorophyll b + divinyl chlorophyll b;
- (C)
- chlorophyll c (Chl-c) = chlorophyll c1 + chlorophyll c2;
- (D)
- PPC = α-carotene + β-carotene + zeaxanthin + alloxanthin + diadinoxanthin;
- (E)
- PSC = 19′-hexanoyloxyfucoxanthin + fucoxanthin + 19′-butanoyloxyfucoxanthin + peridinin.
2.5. Pigment Retrieval from Rrs(λ)
2.5.1. aGau(λ) from Rrs(λ)
2.5.2. aGau(λ) Versus Cpigs
3. Results
3.1. Retrievals from Rrs(λ)
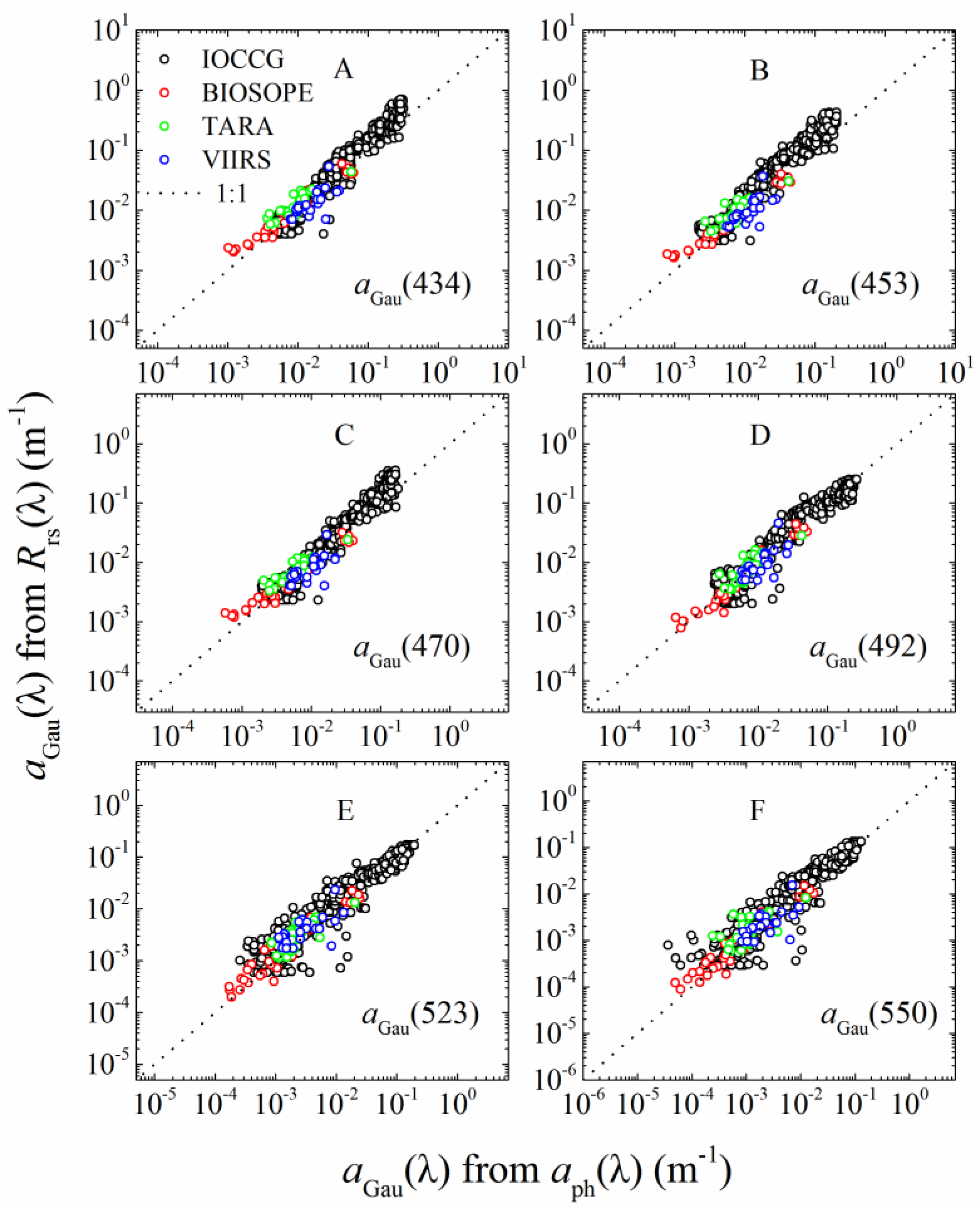
3.1.1. aGau(λ) Validation
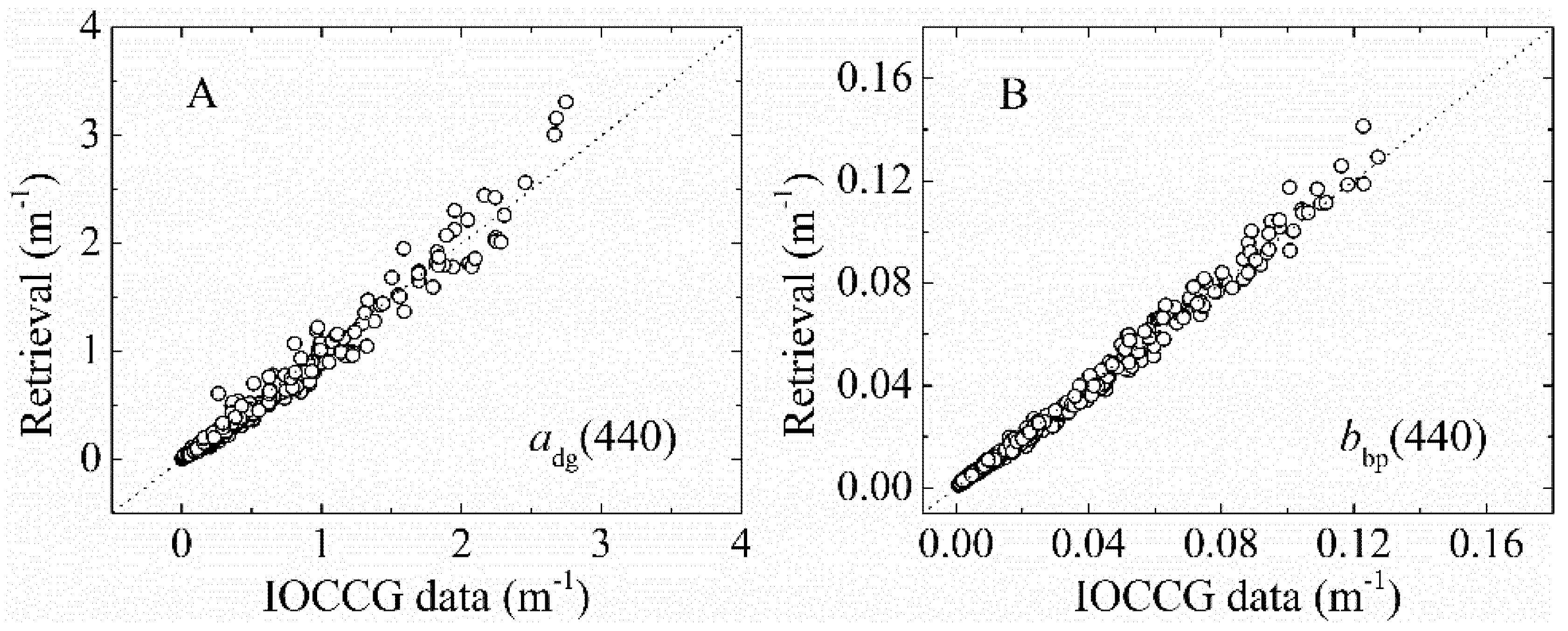
3.1.2. bbp(λ) and adg(λ) Validation
3.2. Cpigs from Satellite Remote Sensing
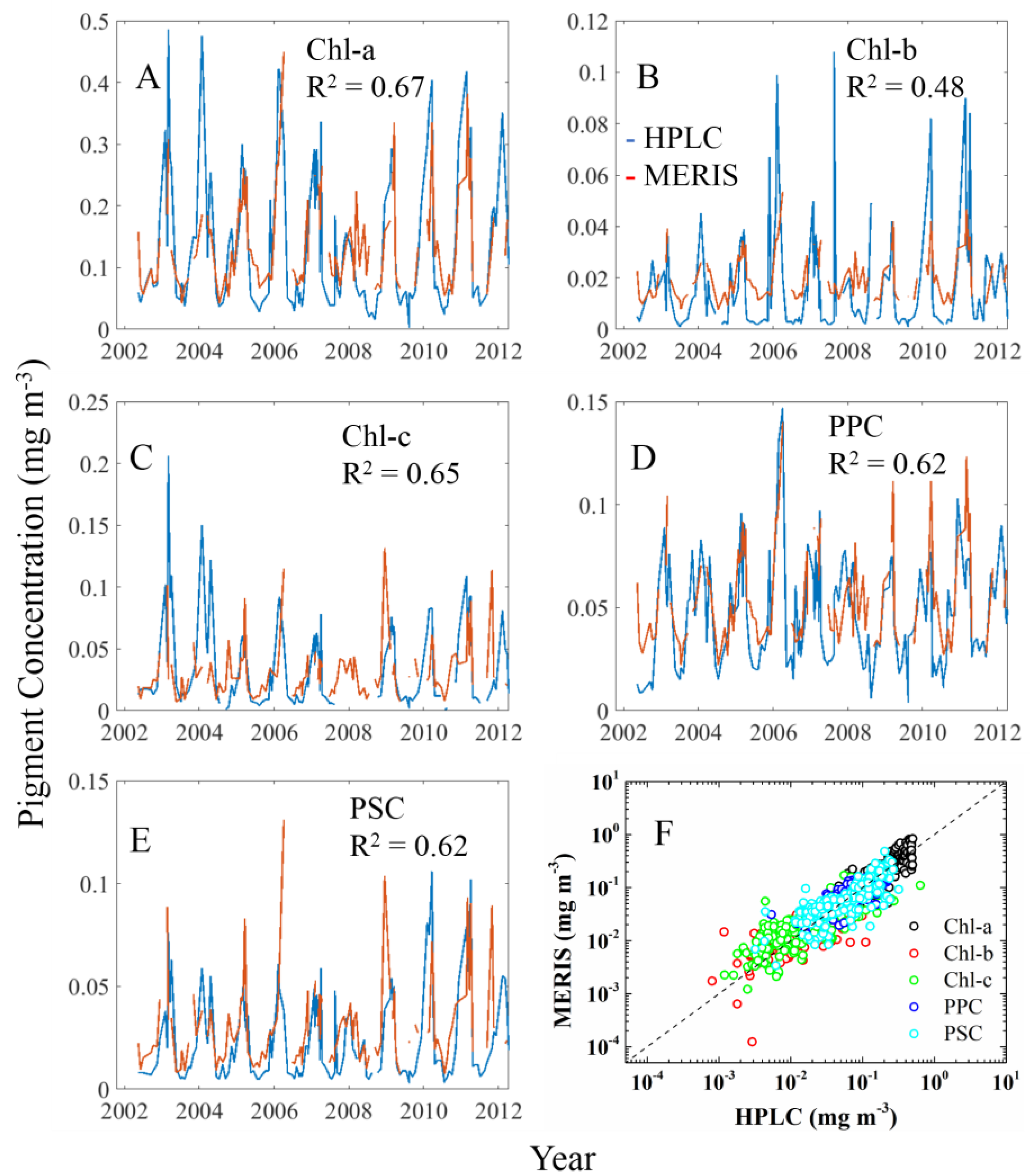
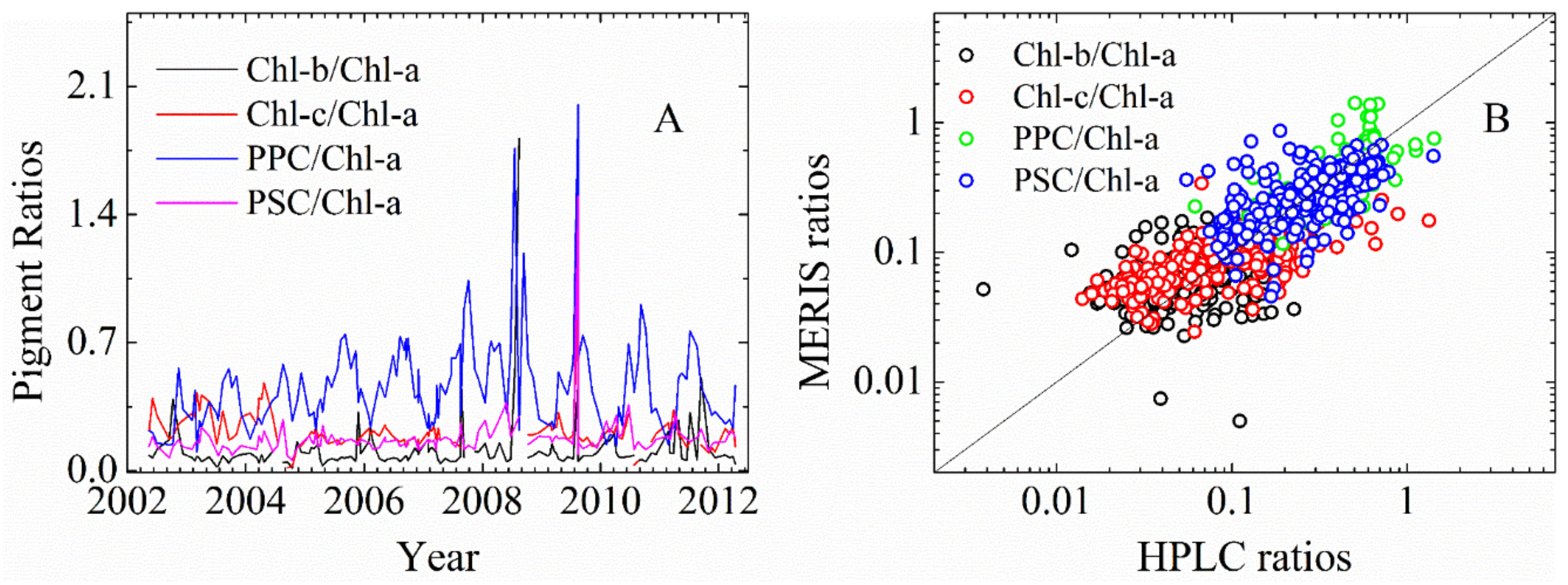
3.2.1. Cpigs Validation and Their Seasonal Variation
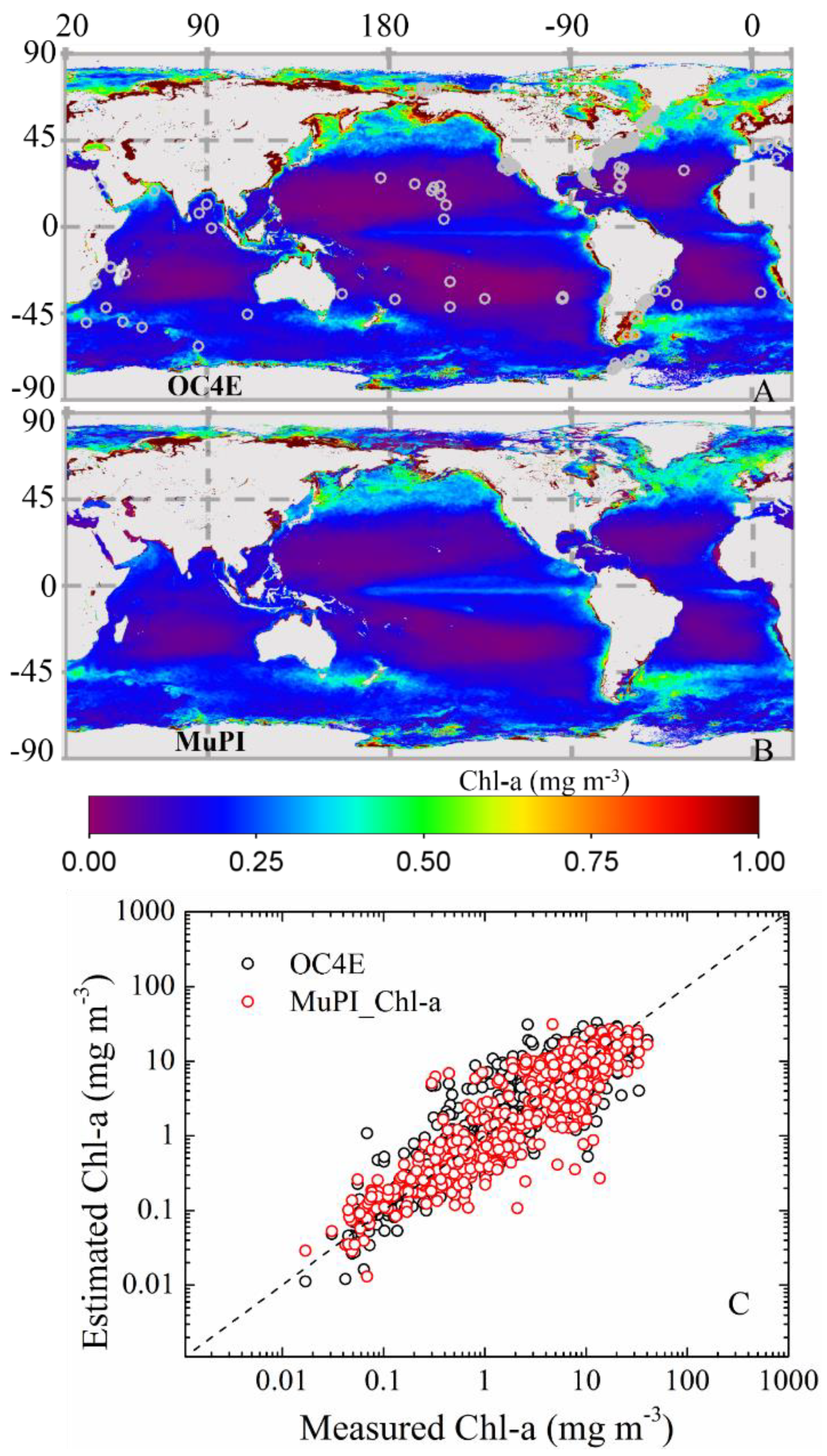
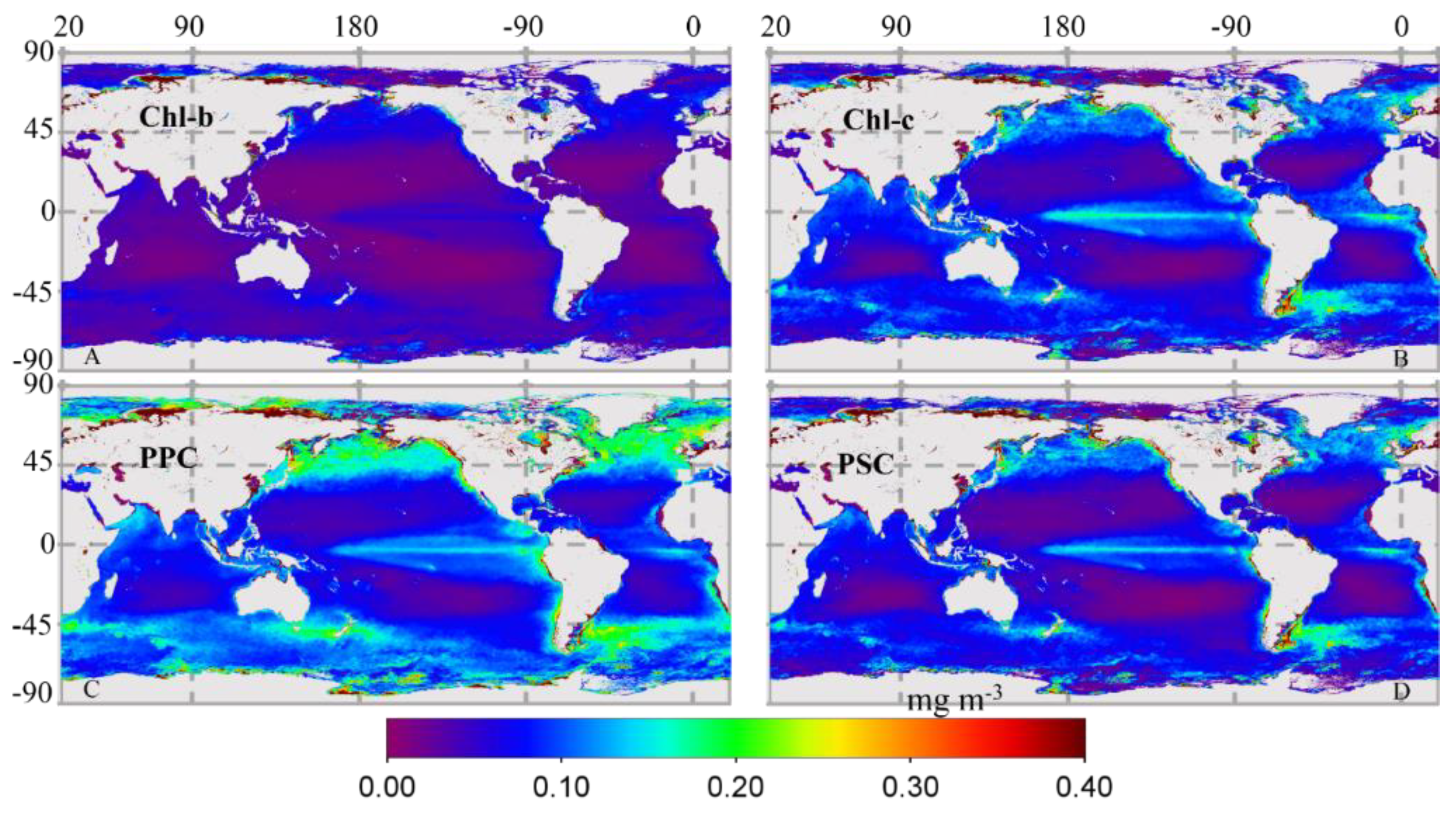
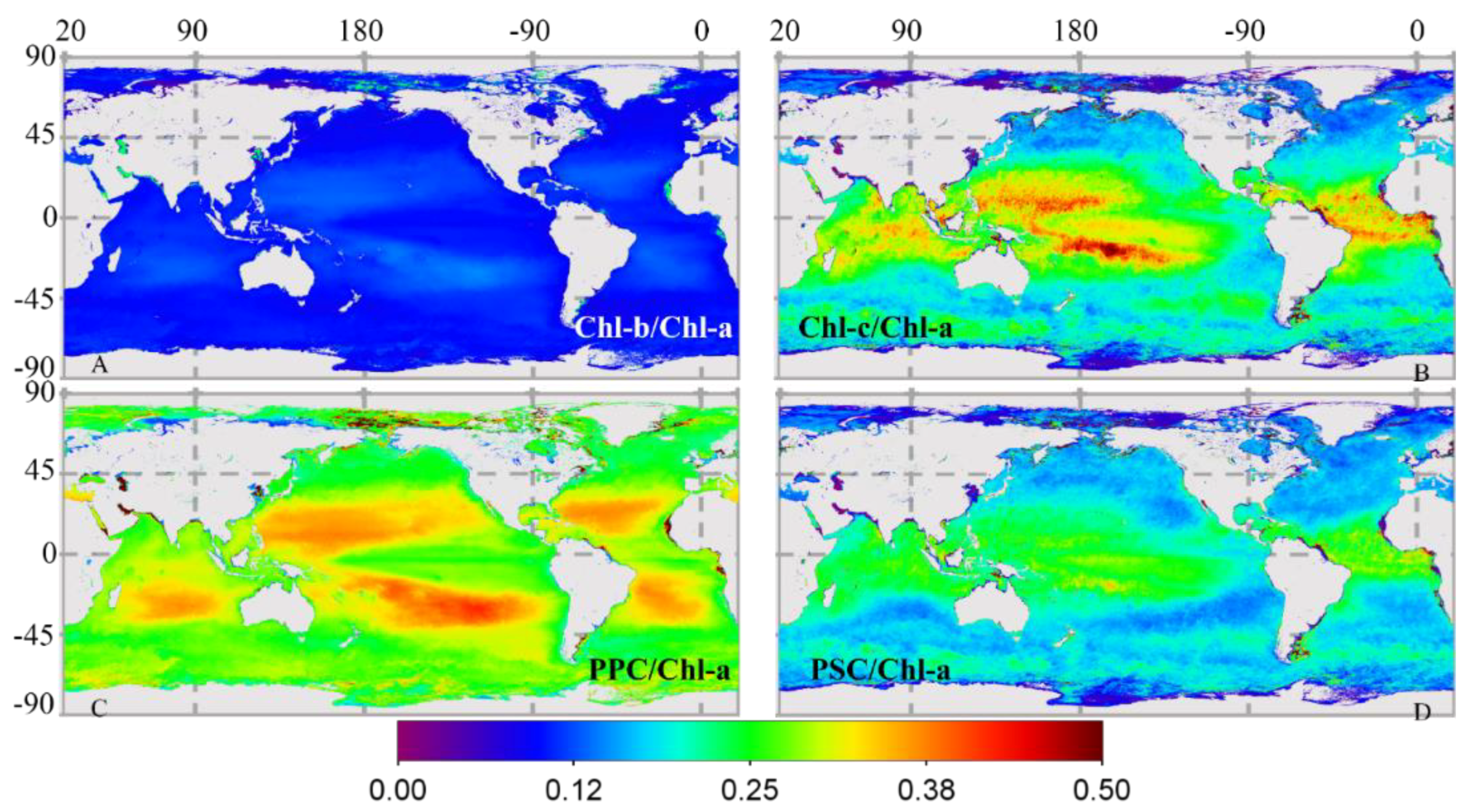
3.2.2. Global Distribution of Cpigs
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Falkowski, P.G. The role of phytoplankton photosynthesis in global biogeochemical cycles. Photosynth. Res. 1994, 39, 235–258. [Google Scholar] [CrossRef] [PubMed]
- Kiørboe, T. Turbulence, phytoplankton cell size, and the structure of pelagic food webs. In Advances in Marine Biology; Academic Press: Cambridge, MA, USA, 1993; Volume 29, pp. 1–72. [Google Scholar]
- Gordon, H.R.; Clark, D.K.; Brown, J.W.; Brown, O.B.; Evans, R.H.; Broenkow, W.W. Phytoplankton pigment concentrations in the Middle Atlantic Bight: Comparison of ship determinations and CZCS estimates. Appl. Opt. 1983, 22, 20–36. [Google Scholar] [CrossRef] [PubMed]
- O’Reilly, J.E.S.; Maritorena, B.G.; Mitchell, D.A.; Siegel, K.L.; Carder, S.A.; Garver, M.; Kahru, C.R. McClain, Ocean color chlorophyll algorithms for SeaWiFS. J. Geophys. Res. 1998, 103, 24937–24953. [Google Scholar] [CrossRef]
- Hu, C.; Lee, Z.; Franz, B. Chlorophyll algorithms for oligotrophic oceans: A novel approach based on three-band reflectance difference. J. Geophys. Res. 2012, 117. [Google Scholar] [CrossRef]
- Alvain, S.; Moulin, C.; Dandonneau, Y.; Bréon, F.M. Remote sensing of phytoplankton groups in case 1 waters from global SeaWiFS imagery. Deep Sea Res. Part I Oceanogr. Res. Pap. 2005, 52, 1989–2004. [Google Scholar] [CrossRef]
- Bracher, A.; Vountas, M.; Dinter, T.; Burrows, J.P.; Röttgers, R.; Peeken, I. Quantitative observation of cyanobacteria and diatoms from space using PhytoDOAS on SCIAMACHY data. Biogeosciences 2009, 6, 751–764. [Google Scholar] [CrossRef]
- Ciotti, A.M.; Bricaud, A. Retrievals of a size parameter for phytoplankton and spectral light absorption by colored detrital matter from water-leaving radiances at SeaWiFS channels in a continental shelf region off Brazil. Limnol. Oceanogr. Methods 2006, 4, 237–253. [Google Scholar] [CrossRef]
- Brewin, R.J.; Sathyendranath, S.; Hirata, T.; Lavender, S.J.; Barciela, R.M.; Hardman-Mountford, N.J. A three-component model of phytoplankton size class for the Atlantic Ocean. Ecol. Model. 2010, 221, 1472–1483. [Google Scholar] [CrossRef]
- Hirata, T.; Hardman-Mountford, N.J.; Brewin, R.J.W.; Aiken, J.; Barlow, R.; Suzuki, K.; Yamanaka, Y. Synoptic relationships between surface Chlorophyll-a and diagnostic pigments specific to phytoplankton functional types. Biogeosciences 2011, 8, 311–327. [Google Scholar] [CrossRef]
- Mouw, C.B.; Yoder, J.A. Optical determination of phytoplankton size composition from global SeaWiFS imagery. J. Geophys. Res. Oceans 2010, 115. [Google Scholar] [CrossRef]
- Sathyendranath, S.; Cota, G.; Stuart, V.; Maass, H.; Platt, T. Remote sensing of phytoplankton pigments: A comparison of empirical and theoretical approaches. Int. J. Remote Sens. 2001, 22, 249–273. [Google Scholar] [CrossRef]
- Gitelson, A.A.; Schalles, J.F.; Hladik, C.M. Remote chlorophyll-a retrieval in turbid, productive estuaries: Chesapeake Bay case study. Remote Sens. Environ. 2007, 109, 464–472. [Google Scholar] [CrossRef]
- Behrenfeld, M.J.; Falkowski, P.G. Photosynthetic rates derived from satellite-based chlorophyll concentration. Limnol. Oceanogr. 1997, 42, 1–20. [Google Scholar] [CrossRef]
- Stumpf, R.P.; Culver, M.E.; Tester, P.A.; Tomlinson, M.; Kirkpatrick, G.J.; Pederson, B.A.; Soracco, M. Monitoring Karenia brevis blooms in the Gulf of Mexico using satellite ocean color imagery and other data. Harmful Algae 2003, 2, 147–160. [Google Scholar] [CrossRef]
- Lehman, P.W. Comparison of chlorophyll a and carotenoid pigments as predictors of phytoplankton biomass. Mar. Biol. 1981, 65, 237–244. [Google Scholar] [CrossRef]
- Schitüter, L.; Riemann, B.; Søndergaard, M. Nutrient limitation in relation to phytoplankton carotenoid/chlorophyll a ratios in freshwater mesocosms. J. Plankton Res. 1997, 19, 891–906. [Google Scholar] [CrossRef]
- Breton, E.; Brunet, C.; Sautour, B.; Brylinski, J.M. Annual variations of phytoplankton biomass in the Eastern English Channel: Comparison by pigment signatures and microscopic counts. J. Plankton Res. 2000, 22, 1423–1440. [Google Scholar] [CrossRef]
- Kruskopf, M.; Flynn, K.J. Chlorophyll content and fluorescence responses cannot be used to gauge reliably phytoplankton biomass, nutrient status or growth rate. New Phytol. 2006, 169, 525–536. [Google Scholar] [CrossRef] [PubMed]
- Behrenfeld, M.J.; O’Malley, R.T.; Boss, E.S.; Westberry, T.K.; Graff, J.R.; Halsey, K.H.; Brown, M.B. Revaluating ocean warming impacts on global phytoplankton. Nat. Clim. Chang. 2016, 6, 323. [Google Scholar] [CrossRef]
- Bidigare, R.R.; Morrow, J.H.; Kiefer, D.A. Derivative analysis of spectral absorption by photosynthetic pigments in the western Sargasso Sea. J. Mar. Res. 1989, 47, 323–341. [Google Scholar] [CrossRef]
- Jeffrey, S.W.; Vesk, M. Introduction to marine phytoplankton and their pigment signature. In Phytoplankton Pigments in Oceanography; UNESCO Publishing: Paris, France, 1997; p. 3784. [Google Scholar]
- Kirkpatrick, G.J.; Millie, D.F.; Moline, M.A.; Schofield, O. Optical discrimination of a phytoplankton species in natural mixed populations. Limnol. Oceanogr. 2000, 45, 467–471. [Google Scholar] [CrossRef]
- Van Heukelem, L.; Thomas, C.S. Computer-assisted high-performance liquid chromatography method development with applications to the isolation and analysis of phytoplankton pigments. J. Chromatogr. A 2001, 910, 31–49. [Google Scholar] [CrossRef]
- Roy, S.; Llewellyn, C.A.; Egeland, E.S.; Johnsen, G. (Eds.) Phytoplankton Pigments: Characterization, Chemotaxonomy and Applications in Oceanography; Cambridge University Press: Cambridge, UK, 2011. [Google Scholar]
- Simis, S.G.; Peters, S.W.; Gons, H.J. Remote sensing of the cyanobacterial pigment phycocyanin in turbid inland water. Limnol. Oceanogr. 2005, 50, 237–245. [Google Scholar] [CrossRef]
- Wynne, T.; Stumpf, R.; Tomlinson, M.; Warner, R.; Tester, P.; Dyble, J.; Fahnenstiel, G. Relating spectral shape to cyanobacterial blooms in the Laurentian Great Lakes. Int. J. Remote Sens. 2008, 29, 3665–3672. [Google Scholar] [CrossRef]
- Mackey, M.D.; Mackey, D.J.; Higgins, H.W.; Wright, S.W. CHEMTAX—A program for estimating class abundances from chemical markers: Application to HPLC measurements of phytoplankton. Mar. Ecol. Prog. Ser. 1996, 144, 265–283. [Google Scholar] [CrossRef]
- Vidussi, F.; Claustre, H.; Manca, B.B.; Luchetta, A.; Marty, J.C. Phytoplankton pigment distribution in relation to upper thermocline circulation in the eastern Mediterranean Sea during winter. J. Geophys. Res. Oceans 2001, 106, 19939–19956. [Google Scholar] [CrossRef]
- Uitz, J.; Claustre, H.; Morel, A.; Hooker, S.B. Vertical distribution of phytoplankton communities in open ocean: An assessment based on surface chlorophyll. J. Geophys. Res. Oceans 2006, 111. [Google Scholar] [CrossRef]
- Sathyendranath, S.; Aiken, J.; Alvain, S.; Barlow, R.; Bouman, H.; Bracher, A.; Clementson, L.A. Phytoplankton functional types from Space. In Reports of the International Ocean-Colour Coordinating Group (IOCCG); International Ocean-Colour Coordinating Group: Dartmouth, Canada, 2014; pp. 1–156. [Google Scholar]
- Uitz, J.; Stramski, D.; Reynolds, R.A.; Dubranna, J. Assessing phytoplankton community composition from hyperspectral measurements of phytoplankton absorption coefficient and remote-sensing reflectance in open-ocean environments. Remote Sens. Environ. 2015, 171, 58–74. [Google Scholar] [CrossRef]
- Catlett, D.; Siegel, D.A. Phytoplankton pigment communities can be modeled using unique relationships with spectral absorption signatures in a dynamic coastal environment. J. Geophys. Res. Oceans 2018, 123, 246–264. [Google Scholar] [CrossRef]
- Bracher, A.; Bouman, H.A.; Brewin, R.J.; Bricaud, A.; Brotas, V.; Ciotti, A.M.; Hardman-Mountford, N.J. Obtaining phytoplankton diversity from ocean color: A scientific roadmap for future development. Front. Mar. Sci. 2017, 4, 55. [Google Scholar] [CrossRef]
- Bricaud, A.; Mejia, C.; Blondeau-Patissier, D.; Claustre, H.; Crepon, M.; Thiria, S. Retrieval of pigment concentrations and size structure of algal populations from their absorption spectra using multilayered perceptrons. Appl. Opt. 2007, 46, 1251–1260. [Google Scholar] [CrossRef] [PubMed]
- Ciotti, A.M.; Lewis, M.R.; Cullen, J.J. Assessment of the relationships between dominant cell size in natural phytoplankton communities and the spectral shape of the absorption coefficient. Limnol. Oceanogr. 2002, 47, 404–417. [Google Scholar] [CrossRef]
- Devred, E.; Sathyendranath, S.; Stuart, V.; Platt, T. A three component classification of phytoplankton absorption spectra: Application to ocean-color data. Remote Sens. Environ. 2011, 115, 2255–2266. [Google Scholar] [CrossRef]
- Hoepffner, N.; Sathyendranath, S. Determination of the major groups of phytoplankton pigments from the absorption spectra of total particulate matter. J. Geophys. Res. Oceans 1993, 98, 22789–22803. [Google Scholar] [CrossRef]
- Hirata, T.; Aiken, J.; Hardman-Mountford, N.; Smyth, T.J.; Barlow, R.G. An absorption model to determine phytoplankton size classes from satellite ocean colour. Remote Sens. Environ. 2008, 112, 3153–3159. [Google Scholar] [CrossRef]
- Lohrenz, S.E.; Weidemann, A.D.; Tuel, M. Phytoplankton spectral absorption as influenced by community size structure and pigment composition. J. Plankton Res. 2003, 25, 35–61. [Google Scholar] [CrossRef]
- Moisan, J.R.; Moisan, T.A.; Linkswiler, M.A. An inverse modeling approach to estimating phytoplankton pigment concentrations from phytoplankton absorption spectra. J. Geophys. Res. Oceans 2011, 116. [Google Scholar] [CrossRef]
- Organelli, E.; Bricaud, A.; Antoine, D.; Uitz, J. Multivariate approach for the retrieval of phytoplankton size structure from measured light absorption spectra in the Mediterranean Sea (BOUSSOLE site). Appl. Opt. 2013, 52, 2257–2273. [Google Scholar] [CrossRef] [PubMed]
- Mouw, C.B.; Hardman-Mountford, N.J.; Alvain, S.; Bracher, A.; Brewin, R.J.; Bricaud, A.; Hirawake, T. A consumer’s guide to satellite remote sensing of multiple phytoplankton groups in the global ocean. Front. Mar. Sci. 2017, 4, 41. [Google Scholar] [CrossRef]
- Jensen, A.; Sakshaug, E. Studies on the phytoplankton ecology of the trondheemsfjord. II. Chloroplast pigments in relation to abundance and physiological state of the phytoplankton. J. Exp. Mar. Biol. Ecol. 1973, 11, 137–155. [Google Scholar] [CrossRef]
- Suggett, D.J.; Moore, C.M.; Hickman, A.E.; Geider, R.J. Interpretation of fast repetition rate (FRR) fluorescence: Signatures of phytoplankton community structure versus physiological state. Mar. Ecol. Prog. Ser. 2009, 376, 1–19. [Google Scholar] [CrossRef]
- Pan, X.; Mannino, A.; Russ, M.E.; Hooker, S.B.; Harding, L.W., Jr. Remote sensing of phytoplankton pigment distribution in the United States northeast coast. Remote Sens. Environ. 2010, 114, 2403–2416. [Google Scholar] [CrossRef]
- Moisan, T.A.; Rufty, K.M.; Moisan, J.R.; Linkswiler, M.A. Satellite observations of phytoplankton functional type spatial distributions, phenology, diversity, and ecotones. Front. Mar. Sci. 2017, 4, 189. [Google Scholar] [CrossRef]
- Gordon, H.R.; Brown, O.B.; Evans, R.H.; Brown, J.W.; Smith, R.C.; Baker, K.S.; Clark, D.K. A semianalytic radiance model of ocean color. J. Geophys. Res. Atmos. 1988, 93, 10909–10924. [Google Scholar] [CrossRef]
- Lee, Z.; Carder, K.L.; Mobley, C.D.; Steward, R.G.; Patch, J.S. Hyperspectral remote sensing for shallow waters. I. A semianalytical model. Appl. Opt. 1998, 37, 6329–6338. [Google Scholar] [CrossRef] [PubMed]
- Lee, Z.; Carder, K.L.; Arnone, R.A. Deriving inherent optical properties from water color: A multiband quasi-analytical algorithm for optically deep waters. Appl. Opt. 2002, 41, 5755–5772. [Google Scholar] [CrossRef] [PubMed]
- Lee, Z. Remote Sensing of Inherent Optical Properties: Fundamentals, Tests of Algorithms, and Applications; International Ocean-Colour Coordinating Group: Dartmouth, Canada, 2006; Volume 5. [Google Scholar]
- Maritorena, S.; Siegel, D.A.; Peterson, A.R. Optimization of a semianalytical ocean color model for global-scale applications. Appl. Opt. 2002, 41, 2705–2714. [Google Scholar] [CrossRef] [PubMed]
- Werdell, P.J.; Franz, B.A.; Bailey, S.W.; Feldman, G.C.; Boss, E.; Brando, V.E.; Mangin, A. Generalized ocean color inversion model for retrieving marine inherent optical properties. Appl. Opt. 2013, 52, 2019–2037. [Google Scholar] [CrossRef] [PubMed]
- Werdell, P.J.; McKinna, L.I.; Boss, E.; Ackleson, S.G.; Craig, S.E.; Gregg, W.W.; Stramski, D. An overview of approaches and challenges for retrieving marine inherent optical properties from ocean color remote sensing. Prog. Oceanogr. 2018. [Google Scholar] [CrossRef]
- Brando, V.E.; Dekker, A.G.; Park, Y.J.; Schroeder, T. Adaptive semianalytical inversion of ocean color radiometry in optically complex waters. Appl. Opt. 2012, 51, 2808–2833. [Google Scholar] [CrossRef] [PubMed]
- Brewin, R.J.; Raitsos, D.E.; Dall’Olmo, G.; Zarokanellos, N.; Jackson, T.; Racault, M.F.; Hoteit, I. Regional ocean-colour chlorophyll algorithms for the Red Sea. Remote Sens. Environ. 2015, 165, 64–85. [Google Scholar] [CrossRef]
- Bukata, R.P.; Jerome, J.H.; Kondratyev, A.S.; Pozdnyakov, D.V. Optical Properties and Remote Sensing of Inland and Coastal Waters; CRC Press: Boca Raton, FL, USA, 2018. [Google Scholar]
- Devred, E.; Sathyendranath, S.; Stuart, V.; Maass, H.; Ulloa, O.; Platt, T. A two-component model of phytoplankton absorption in the open ocean: Theory and applications. J. Geophys. Res. Oceans 2006, 111. [Google Scholar] [CrossRef]
- Garver, S.A.; Siegel, D.A. Inherent optical property inversion of ocean color spectra and its biogeochemical interpretation: 1. Time series from the Sargasso Sea. J. Geophys. Res. Oceans 1997, 102, 18607–18625. [Google Scholar] [CrossRef]
- Hoge, F.E.; Lyon, P.E. Satellite retrieval of inherent optical properties by linear matrix inversion of oceanic radiance models: An analysis of model and radiance measurement errors. J. Geophys. Res. Oceans 1996, 101, 16631–16648. [Google Scholar] [CrossRef]
- Roesler, C.S.; Perry, M.J. In situ phytoplankton absorption, fluorescence emission, and particulate backscattering spectra determined from reflectance. J. Geophys. Res. Oceans 1995, 100, 13279–13294. [Google Scholar] [CrossRef]
- Wang, P.; Boss, E.S.; Roesler, C. Uncertainties of inherent optical properties obtained from semianalytical inversions of ocean color. Appl. Opt. 2005, 44, 4074–4085. [Google Scholar] [CrossRef] [PubMed]
- Loisel, H.; Stramski, D.; Dessailly, D.; Jamet, C.; Li, L.; Reynolds, R.A. An Inverse Model for Estimating the Optical Absorption and Backscattering Coefficients of Seawater from Remote-Sensing Reflectance over a Broad Range of Oceanic and Coastal Marine Environments. J. Geophys. Res. Oceans 2018, 123, 2141–2171. [Google Scholar] [CrossRef]
- Wang, G.; Lee, Z.; Mishra, D.R.; Ma, R. Retrieving absorption coefficients of multiple phytoplankton pigments from hyperspectral remote sensing reflectance measured over cyanobacteria bloom waters. Limnol. Oceanogr. Methods 2016, 14, 432–447. [Google Scholar] [CrossRef]
- Hoepffner, N.; Sathyendranath, S. Effect of pigment composition on absorption properties of phytoplankton. Mar. Ecol. Prog. Ser. 1991, 73, l–23. [Google Scholar] [CrossRef]
- Chase, A.P.; Boss, E.; Cetinić, I.; Slade, W. Estimation of phytoplankton accessory pigments from hyperspectral reflectance spectra: Toward a global algorithm. J. Geophys. Res. Oceans 2017. [Google Scholar] [CrossRef]
- Lutz, V.A.; Sathyendranath, S.; Head, E.J.H. Absorption coefficient of phytoplankton: Regional variations in the North Atlantic. Mar. Ecol. Prog. Ser. 1996, 197–213. [Google Scholar] [CrossRef]
- Mitchell, B.G. Algorithms for determining the absorption coefficient for aquatic particulates using the quantitative filter technique. In Ocean Optics X; International Society for Optics and Photonics: Bellingham, WA, USA, 1990; Volume 1302, pp. 137–149. [Google Scholar]
- Mobley, C.D. Light and Water: Radiative Transfer in Natural Waters; Academic Press: Cambridge, MA, USA, 1994. [Google Scholar]
- Lee, Z.; Pahlevan, N.; Ahn, Y.H.; Greb, S.; O’Donnell, D. Robust approach to directly measuring water-leaving radiance in the field. Appl. Opt. 2013, 52, 1693–1701. [Google Scholar] [CrossRef] [PubMed]
- Mueller, J.L. Ocean Optics Protocols for Satellite Ocean Color Sensor Validation, Revision 4: Radiometric Measurements and Data Analysis Protocols; Goddard Space Flight Center: Greenbelt, MD, USA, 2003; Volume 3. [Google Scholar]
- Bailey, S.W.; Werdell, P.J. A multi-sensor approach for the on-orbit validation of ocean color satellite data products. Remote Sens. Environ. 2006, 102, 12–23. [Google Scholar] [CrossRef]
- Mitchell, B.G.; Kahru, M.; Wieland, J.; Stramska, M. Determination of spectral absorption coefficients of particles, dissolved material and phytoplankton for discrete water samples. In Ocean Optics Protocols for Satellite Ocean Color Sensor Validation, Revision 4, Volume IV: Inherent Optical Properties: Instruments, Characterizations, Field Measurements and Data Analysis Protocols; NASA/TM-2003-211621; Mueller, J.L., Fargion, G.S., McClain, C.R., Eds.; NASA Goddard Space Flight Center: Greenbelt, MD, USA, 2003; pp. 39–64. [Google Scholar]
- Wang, G.; Lee, Z.; Mouw, C. Multi-spectral remote sensing of phytoplankton pigment absorption properties in cyanobacteria bloom waters: A regional example in the western basin of Lake Erie. Remote Sens. 2017, 9, 1309. [Google Scholar] [CrossRef]
- Morel, A. Optical properties of pure water and pure sea water. Opt. Asp. Oceanogr. 1974, 1, 22. [Google Scholar]
- Pope, R.M.; Fry, E.S. Absorption spectrum (380–700 nm) of pure water. II. Integrating cavity measurements. Appl. Opt. 1997, 36, 8710–8723. [Google Scholar] [CrossRef] [PubMed]
- Lee, Z.; Wei, J.; Voss, K.; Lewis, M.; Bricaud, A.; Huot, Y. Hyperspectral absorption coefficient of “pure” seawater in the range of 350–550 nm inverted from remote sensing reflectance. Appl. Opt. 2015, 54, 546–558. [Google Scholar] [CrossRef]
- Nelson, N.B.; Siegel, D.A.; Michaels, A.F. Seasonal dynamics of colored dissolved material in the Sargasso Sea. Deep Sea Res. Part. I Oceanogr. Res. Pap. 1998, 45, 931–957. [Google Scholar] [CrossRef]
- Babin, M.; Stramski, D.; Ferrari, G.M.; Claustre, H.; Bricaud, A.; Obolensky, G.; Hoepffner, N. Variations in the light absorption coefficients of phytoplankton, nonalgal particles, and dissolved organic matter in coastal waters around Europe. J. Geophys. Res. Oceans 2003, 108. [Google Scholar] [CrossRef]
- Lasdon, L.S.; Waren, A.D.; Jain, A.; Ratner, M. Design and testing of a generalized reduced gradient code for nonlinear programming. ACM Trans. Math. Softw. (TOMS) 1978, 4, 34–50. [Google Scholar] [CrossRef]
- Gilerson, A.A.; Gitelson, A.A.; Zhou, J.; Gurlin, D.; Moses, W.; Ioannou, I.; Ahmed, S.A. Algorithms for remote estimation of chlorophyll-a in coastal and inland waters using red and near infrared bands. Opt. Express 2010, 18, 24109–24125. [Google Scholar] [CrossRef] [PubMed]
- Ruddick, K.; Park, Y.; Astoreca, R.; Neukermans, G.; Van Mol, B. Validation of MERIS water products in the Southern North Sea. In Proceedings of the 2nd MERIS—(A) ATSR Workshop; ESA Publications Office Frascati: Frascati, Spain, 2008. [Google Scholar]
- Claustre, H. The trophic status of various oceanic provinces as revealed by phytoplankton pigment signatures. Limnol. Oceanogr. 1994, 39, 1206–1210. [Google Scholar] [CrossRef]
- Aiken, J.; Pradhan, Y.; Barlow, R.; Lavender, S.; Poulton, A.; Holligan, P.; Hardman-Mountford, N. Phytoplankton pigments and functional types in the Atlantic Ocean: A decadal assessment, 1995–2005. Deep Sea Res. Part II Top. Stud. Oceanogr. 2009, 56, 899–917. [Google Scholar] [CrossRef]
- Descy, J.P.; Sarmento, H.; Higgins, H.W. Variability of phytoplankton pigment ratios across aquatic environments. Eur. J. Phycol. 2009, 44, 319–330. [Google Scholar] [CrossRef]
- Behrenfeld, M.J.; O’Malley, R.T.; Siegel, D.A.; McClain, C.R.; Sarmiento, J.L.; Feldman, G.C.; Boss, E.S. Climate-driven trends in contemporary ocean productivity. Nature 2006, 444, 752. [Google Scholar] [CrossRef] [PubMed]
- Bricaud, A.; Claustre, H.; Ras, J.; Oubelkheir, K. Natural variability of phytoplanktonic absorption in oceanic waters: Influence of the size structure of algal populations. J. Geophys. Res. Oceans 2004, 109. [Google Scholar] [CrossRef]
| Datasets/Cruises | Time | Size (N) | Measurements | Chl-a (mg∙m−3) | Usage |
|---|---|---|---|---|---|
| SeaBASS | 2001–2012 | 1619 | aph(λ) | NA | Gaussian curves |
| 1991–2007 | 430 | aph(λ), HPLC | 0.02–13.2 | aGau(λ) vs. Cpigs relationships | |
| IOCCG | NA | 500 | Rrs(λ), aph(λ), adg(λ), bbp(λ) | 0.03–30 | aGau(λ) and Cpigs validation |
| Tara Oceans expedition | 2010–2012 | 23 | Rrs(λ), aph(λ), HPLC | 0.02–0.95 | |
| VIIRS cal/val | 2014–2015 | 21 | Rrs(λ), aph(λ), HPLC | 0.15–1.5 | |
| BIOSOPE | 2004 | 31 | Rrs(λ), aph(λ), HPLC | 0.00036–3.06 | |
| BATS | 2002–2012 | 148 | HPLC | 0.002–0.486 | Cpigs variation |
| MERIS | 2002–2012 | 148 | Rrs(λ) | 0.037–0.325 |
| Peak | Pigments | Peak_loc (nm) | Width(FWHM) (nm) | Relationships | R2 |
|---|---|---|---|---|---|
| 1 | Chl-a | 406 | 16 | 1.13x11.01 | 0.98 |
| 2 | Chl-a | 434 | 12 | x1 | -- |
| 3 | Chl-c | 453 | 12 | 0.60x10.95 | 0.99 |
| 4 | Chl-b | 470 | 13 | 0.51x10.97 | 0.98 |
| 5 | PPC | 492 | 16 | x2 | -- |
| 6 | PSC | 523 | 14 | 0.87x21.17 | 0.99 |
| 7 | PE | 550 | 14 | 0.79x21.27 | 0.96 |
| 8 | Chl-c | 584 | 16 | 0.40x21.17 | 0.96 |
| 9 | PC | 617 | 13 | 0.34x11.14 | 0.93 |
| 10 | Chl-c | 638 | 11 | 0.47x21.19 | 0.96 |
| 11 | Chl-b | 660 | 11 | 0.30x21.11 | 0.94 |
| 12 | Chl-a | 675 | 10 | 0.86x11.11 | 0.98 |
| p-Value | Peak 406 | Peak 434 | Peak 453 | Peak 470 | Peak 492 | Peak 523 | Peak 550 | Peak 584 | Peak 617 | Peak 638 | Peak 660 | Peak 675 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Chl-a | 0.01 | 0.00 | 0.00 | 0.01 | 0.03 | 0.00 | 0.00 | 0.00 | 0.00 | 0.00 | ||
| Chl-b | 0.03 | 0.00 | ||||||||||
| Chl-c | 0.04 | 0.03 | 0.00 | 0.04 | ||||||||
| PPC | 0.00 | 0.00 | 0.00 | 0.00 | ||||||||
| PSC | 0.00 | 0.02 | 0.01 | |||||||||
| R2 | 0.80 | 0.87 | 0.83 | 0.87 | 0.83 | 0.78 | 0.64 | 0.68 | 0.76 | 0.73 | 0.81 | 0.91 |
| Pigments | aGau(λ) | Parameters (a0, a1, …, ai) | R2 |
|---|---|---|---|
| Chl-a | 675 | 1.804, 0.975 | 0.89 |
| Chl-b | 434, 453, 470 | −0.066, 2.470, −3.073, 1.379 | 0.72 |
| Chl-c | 470, 492, 523, 675 | 1.334, 2.022, −3.125, 0.745, 1.119 | 0.83 |
| PPC | 453, 470 | 0.734, 1.311, −0.416 | 0.76 |
| PSC | 470, 492, 523 | 1.67, 3.034, −2.670, 0.725 | 0.84 |
| Peak Center | IOCCG | Tara Oceans | BIOSOPE | VIIRS Cruises | ||||
|---|---|---|---|---|---|---|---|---|
| Mea. | Med. | Mea. | Med. | Mea. | Med. | Mea. | Med. | |
| 406 | 45 | 45 | 34 | 27 | 34 | 28 | 28 | 20 |
| 434 | 37 | 36 | 34 | 28 | 26 | 25 | 28 | 13 |
| 453 | 47 | 49 | 28 | 24 | 23 | 18 | 27 | 15 |
| 470 | 35 | 34 | 29 | 25 | 30 | 27 | 31 | 18 |
| 492 | 34 | 31 | 26 | 21 | 22 | 18 | 29 | 18 |
| 523 | 44 | 34 | 38 | 28 | 34 | 29 | 48 | 44 |
| 550 | 45 | 35 | 53 | 41 | 37 | 34 | 41 | 35 |
| 584 | 55 | 48 | 48 | 37 | 38 | 36 | 53 | 57 |
| 617 | 51 | 45 | 47 | 38 | 36 | 29 | 37 | 40 |
| 638 | 54 | 42 | 66 | 68 | 41 | 35 | 41 | 35 |
| 660 | 52 | 48 | 35 | 23 | 32 | 29 | 43 | 34 |
| 675 | 46 | 40 | 30 | 26 | 60 | 56 | 32 | 21 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, G.; Lee, Z.; Mouw, C.B. Concentrations of Multiple Phytoplankton Pigments in the Global Oceans Obtained from Satellite Ocean Color Measurements with MERIS. Appl. Sci. 2018, 8, 2678. https://doi.org/10.3390/app8122678
Wang G, Lee Z, Mouw CB. Concentrations of Multiple Phytoplankton Pigments in the Global Oceans Obtained from Satellite Ocean Color Measurements with MERIS. Applied Sciences. 2018; 8(12):2678. https://doi.org/10.3390/app8122678
Chicago/Turabian StyleWang, Guoqing, Zhongping Lee, and Colleen B. Mouw. 2018. "Concentrations of Multiple Phytoplankton Pigments in the Global Oceans Obtained from Satellite Ocean Color Measurements with MERIS" Applied Sciences 8, no. 12: 2678. https://doi.org/10.3390/app8122678
APA StyleWang, G., Lee, Z., & Mouw, C. B. (2018). Concentrations of Multiple Phytoplankton Pigments in the Global Oceans Obtained from Satellite Ocean Color Measurements with MERIS. Applied Sciences, 8(12), 2678. https://doi.org/10.3390/app8122678





