1. Introduction
In the biomedical domain we can find an impressive number of information sources. Traditionally, this information was difficult to access for different reasons, such as that the documents were not publicly available or the content was so complex that only medical specialists could understand the terminology used in the documents. However, with advent of the Internet and open access to all types of data, every day more and more ordinary people like for example students, researchers, patients and relatives try to access this information, as well as medical professionals.
Systems for combining, fusing and retrieving medical information from heterogeneous sources are a great help, not only for professionals who must continuously handle a large number of reports, articles, documents and data but also for ordinary people who want to be empowered with easy and fast access to all kinds of information, including medical information.
In this paper we describe an approach to interactive information retrieval. This approach combines and integrates different semantic resources such as: UMLS (Unified Medical Language System), Google Scholar, SciELO and MedLine. Our main goal is to provide a tool for a better understanding of specialized medical terminology. In addition, our approach implements a semantic search focused on improving the accuracy of the user’s query by providing support to understand the contextual meaning of the terms.
On the one hand, tools for understanding specialized terminology help the normal user to have a better readability of the texts and, on the other hand, help experts to have a more exhaustive analysis using tools that provide additional references on the terms detected. Medical terminology detection systems have always attracted great interest [
1] given the high number of related resources and their good quality, such as the specialized ontologies UMLS [
2] and MeSH [
3]. The identification of terms is not only interesting in specialized texts, it can also be very useful over texts written by patients [
4]. In both cases, systems for identifying terms become the key to accessing bibliographic documentation and related literature, since these terms and their relationships, when identified, allow the connection of knowledge between different sources. The proposed system is flexible enough so that both medical professionals and non-specialized users (for example, patients) can take advantage of it.
Our approach takes a textual document as input and different techniques of Natural Language Processing (NLP) are applied in order to recognize the biomedical entities in the text. It should be noted that our system works on texts written in Spanish, which adds an additional difficulty to the task of term identification. Most of the tools developed to date are oriented toward textual information in English. In fact, there are some good examples of resources that have acceptable accuracy in Named Entity Recognition (NER) in English such as the cTakes tool [
5]. Therefore, our system implements its own NER for recognizing biomedical entities in Spanish including the UMLS codes associated to each recognized term. Then, it connects entities found in the document to a Linked Open Data (LOD) resource, called Linked Life Data (Linked Life Data—A Semantic Data Integration Platform for the Biomedical Domain:
http://linkedlifedata.com/), by using the first UMLS code. Once we have the text marked with all the biomedical entities, the system integrates three different knowledge bases and external sources (MedLine, SciELO and Google Scholar) to perform information retrieval using the entities as query keywords. Finally, the system generates a semantic network relating all the identified entities by taking as central node the most recurrent entity in the original text. The developed system is freely available at
http://sinai.ujaen.es/demo/bsb/. We have called it Buscador Semántico Biomédico (Biomedical Semantic Search Engine).
In order to test the effectiveness of our proposal, we have carried out an evaluation of the system using the MANTRA corpus [
6]. The MANTRA corpus was developed as part of the MANTRA project, aimed at providing multilingual terminologies and semantically annotated multilingual documents in English, French, German, Spanish, and Dutch. We have taken this corpus to test our system comparing the performance with the state-of-the-art biomedical NER tool FreeLing-Med for Spanish. The results obtained are promising and encourage us to continue studying and improving the proposed architecture.
The rest of the paper is organized as follows:
Section 2 describes some related studies.
Section 3 briefly describes the different resources to be integrated into our architecture. The developed system is presented in
Section 4. The experiments and evaluation of our approach using the MANTRA corpus and the results obtained in comparison with FreeLing-Med are described in
Section 5. Finally, conclusions and discussion are presented in
Section 6.
2. Background
The vast volume of biomedical unstructured documents generated on a daily basis is creating interesting challenges for the effective and efficient use of the information and knowledge stored in such texts. Some initiatives have recently emerged and attracted the interest of the research community. For example, the CLEF eHealth Evaluation Lab is focused on combining NLP and information retrieval for clinical care [
7]. This CLEF challenge has been running since 2013 and continues to propose different datasets and tasks every year. Medical information extraction and retrieval have also been relevant topics in the TREC community [
8,
9].
In addition, the automatic or semi-automatic annotation of biomedical texts has gathered considerable attention in the past decade and several tools and resources have been developed mainly for English [
10]. However, for other languages such as Spanish there is a lack of tools with similar levels of maturity imposing an imbalance in the access to health-related information for these languages. Thus, it is even more necessary to study technologies focused on the treatment of documents in Spanish. Actually, some research has begun to be carried out in recent years. For example, [
11] propose a Spanish version of MetaMap combining machine translation and the original MetaMap in order to annotate Spanish texts with UMLS concepts. The Freeling analyzer was extended to develop Freeling-Med integrating ontologies and dictionaries to identify medical entities such as SNOMED-CT [
12]. Finally, [
13] present a prototype for the biomedical term normalization of Spanish electronic health records using UMLS. The approach applies information retrieval technologies to index the Metathesaurus, generating mapping candidates from input text. In addition, some recent medical NLP tasks have been proposed to develop Spanish NER in clinical records (MEDDOCAN [
14] and PharmaCoNER [
15]).
Regarding medical ontologies like MeSH or the wider UMLS, we can consider as profitable and enriched source of semantic content, not only for finding papers in specialized literature [
16], but also for clinical record retrieval [
17,
18]. As can be checked from these references, the most common method for integrating medical ontologies in the retrieval process is by
query expansion [
19]. It involves recognizing entities in the query keywords and adding new concepts according to proximity in meaning or other taxonomic relations like meronymy or holomyny, for example.
Most systems exhibit some sort of “black-box” approach in the retrieval process, i.e., the user enters a query or a sample document and the system retrieves all images, clinical cases, papers or whatever is indexed, trying to maximize the relevance of those retrieved items to the user’s needs. Most evaluation forums currently propose this scenario, whereas user presence is somewhat neglected. In contrast to this understanding of the retrieval process, interactive approaches consider the intervention of the user to refine or fine tune the behavior of the system in order to obtain a better definition of user’s needs, thus, improving the relevance of retrieved items. The system proposed by Mourau and his colleagues [
20] enables the user to add new keywords taken from the MeSH thesaurus before performing query expansion, although in this study keywords are proposed automatically. This idea of allowing the user to refine the query is not new, as
interactive query expansion was proposed more than thirty years ago [
21]. Although the evaluation of interactive systems is a drawback in the organization of evaluation tasks, as interactive systems evaluation is not an easy task some work has been done in this direction [
22]. Nowadays, the interaction of users with medical retrieval systems has been found beneficial, improving user experience and the quality of retrieved items, as is the case for image retrieval [
23] and diagnostic retrieval [
24].
Our proposal goes in the direction of interactive medical retrieval in the belief that enhancing the information search by providing visual tools for understanding specialized medical content helps users to select better query terms. In addition, our system works with Spanish documents although it can easily be extended to other languages, integrating knowledge bases in these different languages. To do this, it will be necessary to have knowledge bases in other languages and to upload the necessary files. Subsequently, pre-processing would be carried out on new knowledge bases and a word matching would be performed to recognize the medical concepts.
3. Knowledge Base Resources
Our system combines several knowledge sources and different tools in order to extract relevant medical information. Specifically, we use two types of information: specialized terminology and document collections. We have integrated UMLS, CIE-10 and MedLine Encyclopedia as medical concept ontologies and we have indexed Google Scholar, MedLine Plus and SciELO as textual collections. In addition, we have linked the identified terms with a Linked Open Data base called Linked Life Data. Below, we briefly describe all the resources integrated into our platform. However, these resources are only some examples because one of the advantage of our approach is that it is possible to easily integrate any kind of sources. Thus for example, we could include textual collections with clinical records or introduce new biomedical knowledge bases.
One way to integrate textual collections with clinical records could be through APIs or libraries that provide the website. These resources allow us to quickly access medicine-related websites where the user could consult information. On the other hand, it is also possible to use a collection of documents related to the biomedical domain and create an Information Retrieval System (IRS). This collection would be indexed and searchable.
3.1. UMLS
The Unified Medical Language System (UMLS) is a compendium of various vocabularies and standards, both health and biomedical, which enables interoperability between computer systems and services. Among other uses, the UMLS allows users to link health information, medical terms, drug names and standard codes through different computer systems. The UMLS integrates over 14 million names in different languages to identify over 3 million concepts from more than 150 sources of biomedical vocabularies. The main languages are English (with 70% of names) and Spanish (with 10% of names). In the UMLS, knowledge is organized by concept; synonymous terms are clustered together to form a concept and concepts are linked to other concepts by means of various types of relationships. Furthermore, each concept is categorized into different semantic types.
3.2. Linked Life Data
The use of UMLS is under license, so we use Linked Life Data (LLD) to show additional information about each concept. LLD is a web platform developed by Ontotext AD that provides access to 25 public biomedical databases, including UMLS. The web service allows us to write complex queries using a SPARQL endpoint or browse the information. We use the LLD Public service that provides free anonymous access for developing proof-of-concept applications.
3.3. MedLine
MedLinePlus (MedlinePlus—Health Information from the National Library of Medicine:
https://medlineplus.gov/) is an online information service provided by the U.S. National Library of Medicine [
25]. MedlinePlus contains links to web portals with information on more than 1000 health topics. In addition, these health topics include links to daily news updates. This resource is of interest to less experienced users as it offers a more informal and familiar vocabulary to the reader. Moreover, MedLine offers information in both English and Spanish. It also gives access to the A.D.A.M. Medical Encyclopedia that includes over 4000 articles about diseases, tests, symptoms, injuries, surgeries, and an extensive library of medical photographs and illustrations. We use this knowledge base in two different ways: first, we obtain a list of medical terms from the medical encyclopedia to use in our entity recognition system. Later, in our platform, we give MedLinePlus links to the most relevant articles for the recognized terms.
3.4. CIE-10/ICD-10
CIE-10 is the acronym for the Clasificación Internacional de Enfermedades, 10a versión, and it is the Spanish version of the International Statistical Classification of Diseases and Related Health Problems (ICD). CIE-10 determines the classification and coding of diseases and a wide variety of signs, symptoms, abnormal findings, complaints, social circumstances and external causes of damage and/or illness. Each medical condition is assigned to a category and receives a code up to six characters in length. These categories and their subcategories group together similar diseases.
3.5. Google Scholar
Google Scholar is a Google search engine specialized in academic literature for a variety of disciplines and publication formats. The Google Scholar Index includes most peer-reviewed online academic journals and books, conference papers, theses and dissertations, pre-prints, abstracts, technical reports, and other academic literature, including court opinions and patents.
3.6. SciELO
SciELO (SciELO—Scientific Electronic Library Online:
http://www.scielo.org/) is a virtual library made up of a collection of Spanish health science journals selected according to pre-established quality criteria. The SciELO project in Spain is being developed by the
Biblioteca Nacional de Ciencias de la Salud (National Library of Health Sciences), thanks to the collaboration agreement established between World Health Organization (WHO) and the
Instituto de Salud Carlos III (Carlos III Health Institute).
For our system we have obtained a parallel version of the SciELO corpus [
26], which has been indexed with Elasticsearch (Open Source Search & Analytics · Elasticsearch—Elastic:
https://www.elastic.co/). This retrieval system is a powerful tool that allows us to index a large volume of data and then make queries with advanced functionalities like approximate searches, faceted searches and highlighted results.
4. An Architecture for Information Exploration
In specialized environments like in the biomedical domain, controlled vocabularies are mandatory in order to unify the terminology present in notes, clinical stories, reports and research papers. In Clinical Decision Support (CDS) systems, the precision offered by the a retrieval system to find relevant literature related to a case is paired with a better treatment and clinical intervention [
27]. Traditional query-based retrieval engines forces the expert to carefully define the query. As an alternative to that formulation, we propose to initiate the retrieval process by means of a complete text, from which query terms are extracted thanks to a highly performing process of specialized terminology extraction. Besides CDS, clinical research demands for more accurate and agile ways to crawling a growing volume of scientific literature.
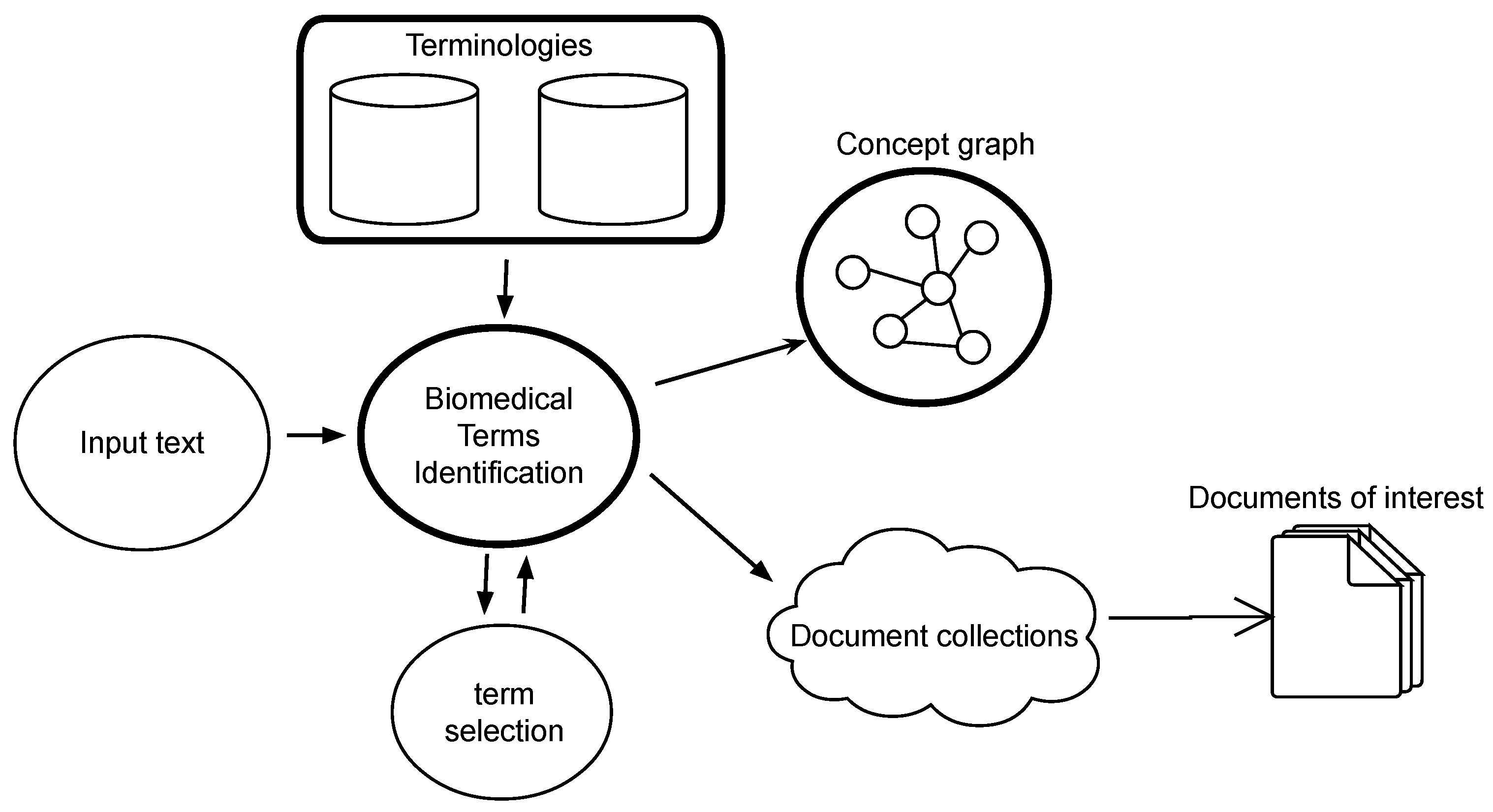
Our approach proposes an integrated architecture with a core focused on terminology, yet provides continuous feedback and allows interactive retrieval. This architecture is composed of four main components and depicted in
Figure 1.
Terminology knowledge bases. They are the main source of knowledge in our proposal. Controlled vocabularies are used by this architecture to leverage information retrieval and browsing. These are valuable resources developed intensively across decades in the biomedical domain, and the architecture profits from this polished knowledge to put forward a new framework for digital searches.
The term identification engine. This is the central part of our system. Starting from any free text, the system identifies the specialized concepts and uses them to retrieve relevant information and to allow semantic exploration on the other two modules considered.
The information retrieval module. This component retrieves from different sources and collections those documents closer to the concepts identified. These sources can be databases of research papers, articles or even clinical reports. Depending on the orientation of the final system, the architecture would “plug” certain databases that the final user is interested in.
Concept exploration. Semantic links between concepts provided in terminology thesauri and concept graphs, like UMLS, allow for term navigation and gathering. This is an important tool when looking for information, as semantic search is enabled by means of concept exploration wandering the graph of semantic relationships. This tool provides a general overview on the terms of our concern and how they are related. By interacting with the graph of terms, the user would be able to understand how they are related and discover new terms as potential keywords for enhanced query composition.
This work focuses specifically on the search and retrieval of information in order to obtain complete, correct and appropriate answers to the information needs of users. Our approach identifies specialized medical terminology automatically and uses it in a meta-search process. By meta-search we mean that the system uses its own knowledge bases and other search engines showing a combination of the best pages that each one has returned. The knowledge bases semantically enrich the results to obtain a higher precision.
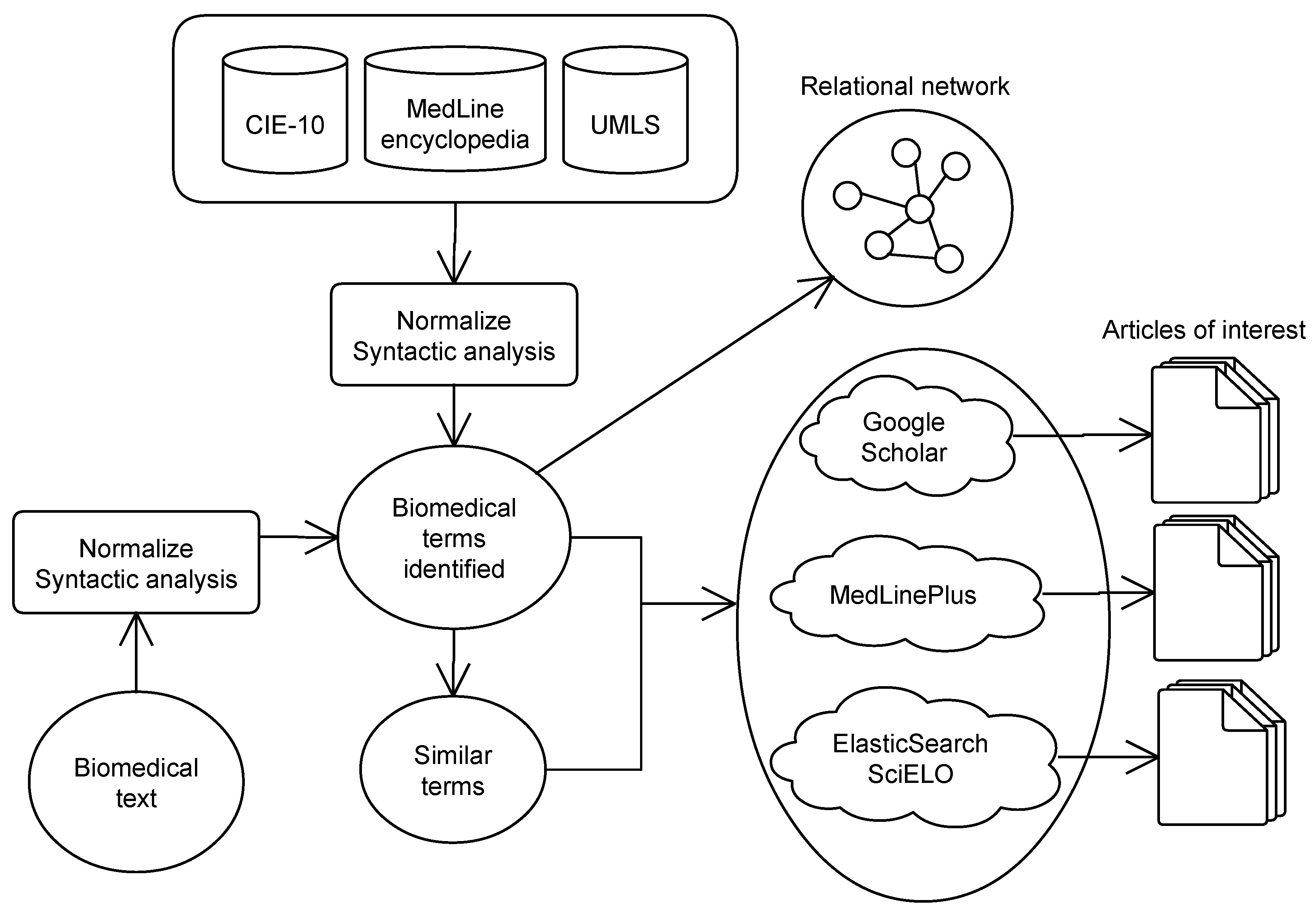
Figure 2 shows the operating structure of the system. The system receives an input text and different normalization processes are applied (see
Section 4.1) in order to recognize biomedical entities such as diseases, symptoms, treatments, parts of the human body, etc.
The system shows the concepts detected along with a list of different ways of writing each concept (synonyms and alternative expressions). Next, using only these concepts, the system searches in several services (see
Section 4.3): academic articles in Google Scholar, health information in MedLine and Spanish scientific articles in SciELO. In the last step, the system shows a semantic graph with the relation between these concepts (see
Section 4.4).
The initial query introduced to search services includes only the words that are part of a concept found in the source text, however, the user can choose alternative terms by selecting different synonyms form a concept list.
The tools described in
Section 3 are integrated in a modular way, so it is possible to add further services and include them in more powerful system to obtain higher performance and wider coverage. It must be noted that, in our case, we have used Spanish knowledge bases but it can be extended to any language, whenever similar resources are available for that language.
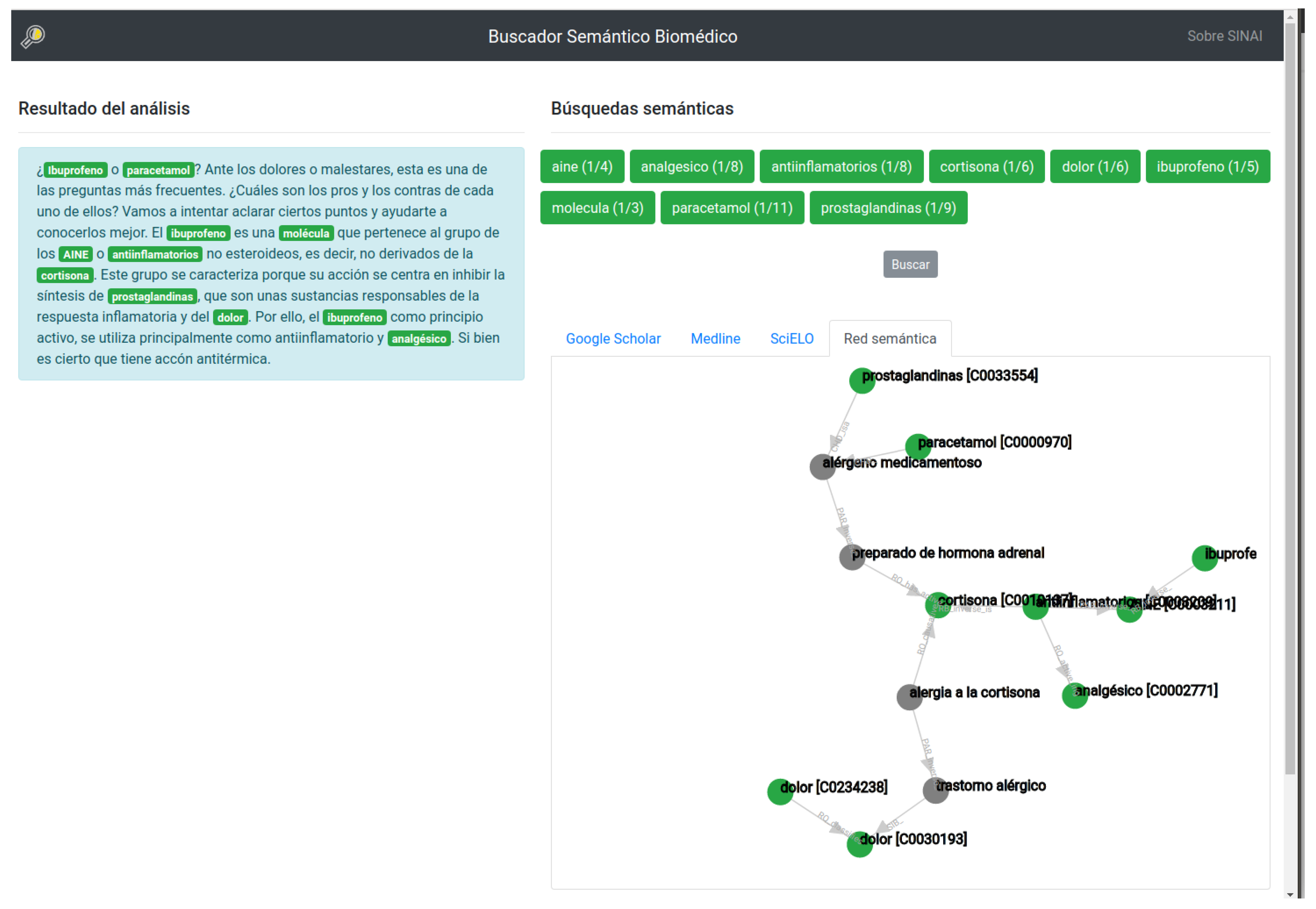
Figure 3 shows the final appearance of the system. The following subsections describe in detail the procedure followed for biomedical named entity recognition, the identification of similar concepts using linked open data, the search engine system and the generation of the semantic relational network.
4.1. Biomedical Named Entity Recognition
Name Entity Recognition (NER) in NLP is an important area of interest initiated many years ago. In the biomedical domain, we can also find some interesting tools for English. For example, MetaMAP is a tool developed by the National Library of Medicine (NLM) focused on discovering UMLS Metathesaurus concepts [
28,
29]; EDGAR is an NLP system for extracting information about drugs and genes relevant to cancer from the biomedical literature [
30]; in the radiology domain, the research conducted by [
31] identifies clinical information in narrative reports and maps that information into a structured representation containing clinical terms. More recently, cTAKES provides concept identification and normalization to UMLS in clinical texts [
5].
On the other hand, tools and resources focused on another language different from English, like for example Spanish, are scarce. For example, there exists a Spanish version of MetaMap where automatic translation techniques were combined with biomedical ontologies and the existing English MetaMap [
11]. Another example is Freeling-Med, which incorporates ontologies and dictionaries to identify medical entities such as SNOMED-CT and UMLS [
12]. However, the recall of these resources is very poor so we decided to develop our own system.
We have applied some NLP technologies in order to develop the Biomedical NER. First, we perform a normalization process that mainly involves:
This process has been carried out for both the input text and the vocabularies and knowledge bases indexed by the system as described in
Section 3. The Spanish dictionaries taken into account for this task are UMLS, MedLine and CIE-10.
The next step in the recognition of entities is the tokenization of sentences. Tokenization is necessary in many natural language processing tasks, such as word counting, parsing, spell checking, corpus generation, and statistical analysis of text. It converts text strings to streams of token objects, where each token object is a separate word, punctuation sign, number/amount, date, e-mail, URL/URI, etc.The system uses the tokenize module included in Python 3. In order to match tokens between the input string and the dictionary used, the tokenization process is applied to both texts (the input and the dictionary).
To increase the accuracy in the identification of specialized terminology we have used Part-Of-Speech Tagger included in CoreNLP tool developed by Stanford University and available for Spanish [
32] and NLTK toolkit [
33].
A Part-Of-Speech Tagger (POS Tagger) is a piece of software that reads text in different languages and tags each word with a part of speech, such as
noun,
verb,
adjective, and so on [
34].
The biomedical dictionaries include entities, procedures and actions. Procedures and actions are mainly made up of verbs, therefore this process is necessary to discard those words that have been identified as a medical concept but are not an adjective, a name or a numeric value.
We subsequently found that the system recognized all possible terms in each sentence, in other words, it found concepts inside other concepts. For example, in the phrase ‘cáncer de mama’, the system identified: ‘cáncer de mama’, ‘cáncer’ and ‘mama’. For a better understanding of the results, we decided to eliminate those identified concepts that were contained within another concept, prioritizing and keeping the longest form.
Figure 4 shows an example of how the system marks the entities detected in a text. In this example we can see how the system prioritizes the detection of entities formed of several words (‘trombosis venosa profunda’) over single term entities (‘trombosis’ or ‘trombosis venosa’).
4.2. Linking Identified Entities with Concepts in LOD
The importance of exchanging information in the clinical domain comes from the new requirements of health care services. These organizations must continue to deliver their services effectively, efficiently and sustainably.
The ability to exchange information and the possibility for it to be automatically interpreted and reused in different applications is known as semantic interoperability. This implies that the systems understand the information they are processing.
The intelligence needed to understand the message is a complex task and it requires establishment of agreements between the different institutions regarding how to represent, describe and contextualize the information to be exchanged. This is made possible by the use of medical terminologies and ontologies that contain internationally standardized catalogs that unify the data used [
35]. We have used and applied Linked Open Data technologies in order to take advantage of the knowledge available in the medical domain.
The automatic recognition system developed shows the medical concepts detected together with their identifier code according to a predefined thesaurus. This code also includes a reference to a website where the user can obtain information about a specific concept.
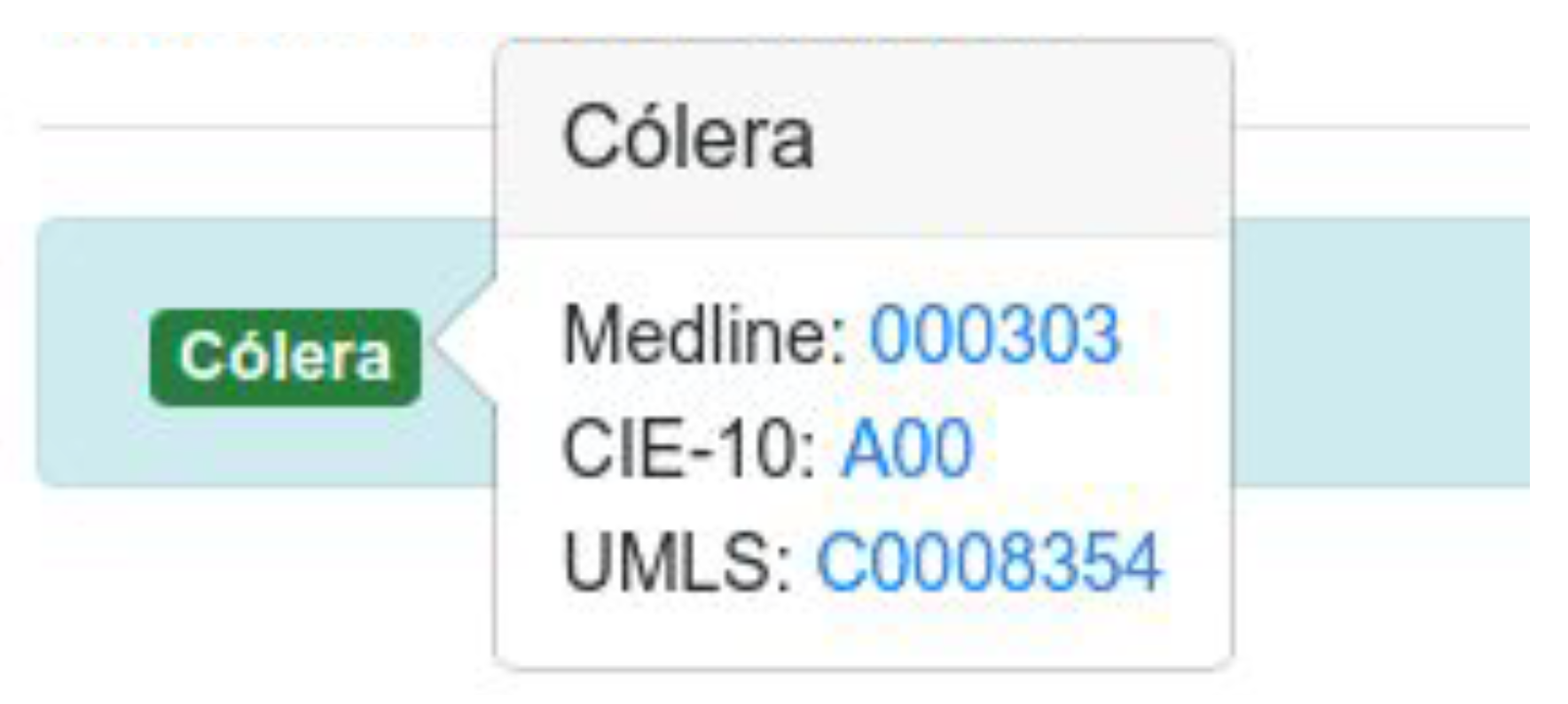
For example, the term
‘cólera’ is associated with the codes
C0008354,
A00 and
000303 in UMLS, CIE-10 and Medline’s encyclopedia respectively (
Figure 5). Through their codes, we can access different websites and obtain relevant information from each one:
UMLS:
http://linkedlifedata.com/resource/umls-concept/C0008354 In this website we can find the description of the concept, the semantic type, the relationships with other concepts, related documents and alternative labels, etc. Another important feature of this website is the possibility of downloading the concept information in different formats (RDF, JSON, N3/Turtle and N-Triples).
4.3. Semantic Search
UMLS is composed of several vocabularies, therefore, a UMLS concept can come from one or more sources. A concept is the fundamental unit in UMLS, and is formed by atoms. All the atoms of a concept are called synonyms or similar terms. Concepts also contain a Unique Concept Identifier (CUI) that uniquely identifies that meaning. Therefore, we conclude that all atoms with the same CUI are synonyms.
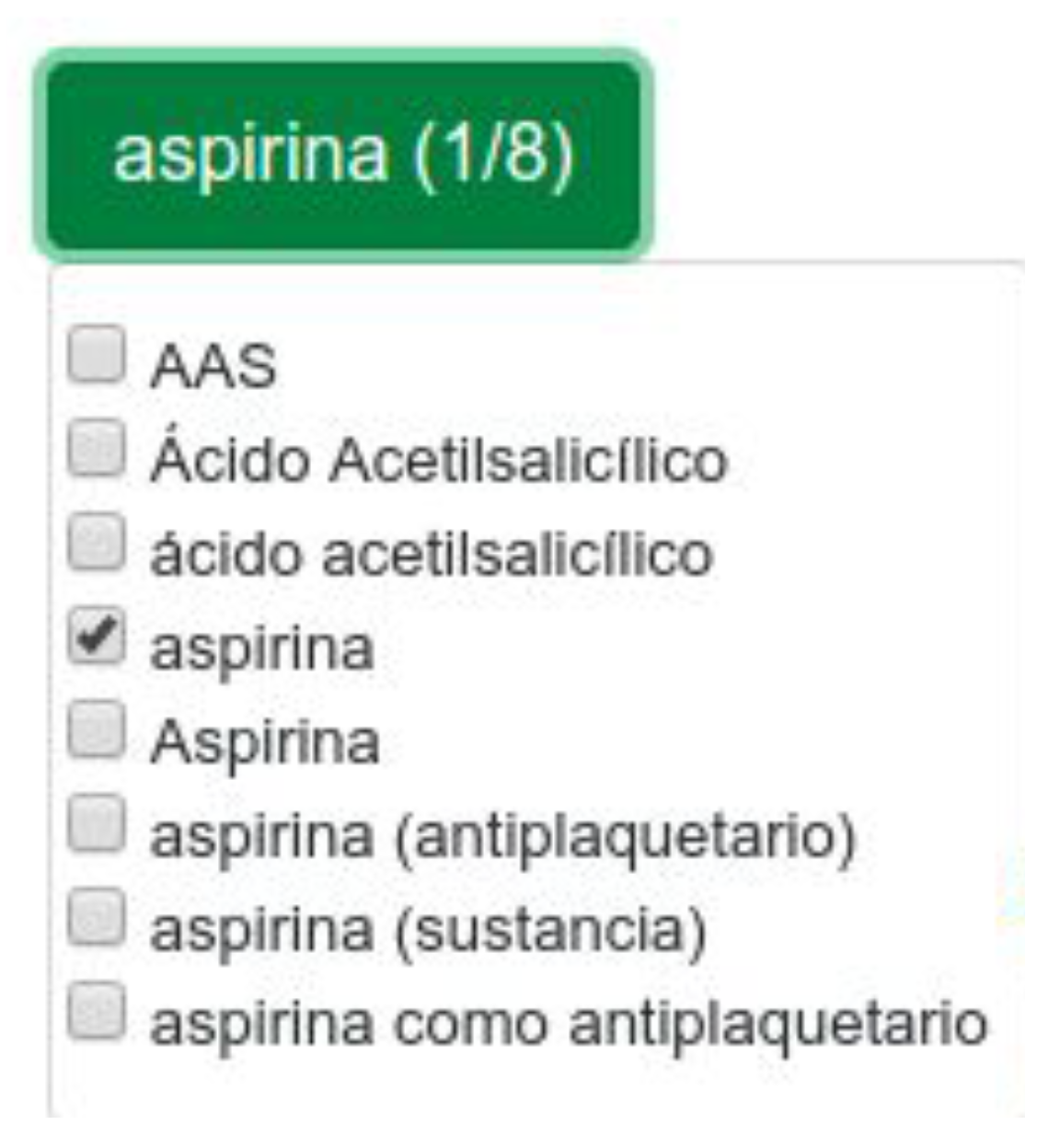
Our system includes a section with all the biomedical terms detected, each one showing its possible synonyms or other ways of naming the concept. In
Figure 6 we can see an example with the concept
‘aspirina’.
This example shows how the concept with CUI ‘C0004057’ contains atoms such as ‘Aspirina’, ‘Ácido acetilsalicílico’, ‘AAS’ or ‘aspirina como antiplaquetario’, among others. As can be seen, UMLS also includes abbreviations, so it offers extra help to the user inexperienced in the area of medicine. From the eight similar terms (or atoms) in the example, only one is marked, which means that only this word is used in the web search.
Through all these concepts, the user can mark and unmark atoms to in order to refine the search and re-launch the new query on Google Scholar, MedLine and SciELO, obtaining different results.
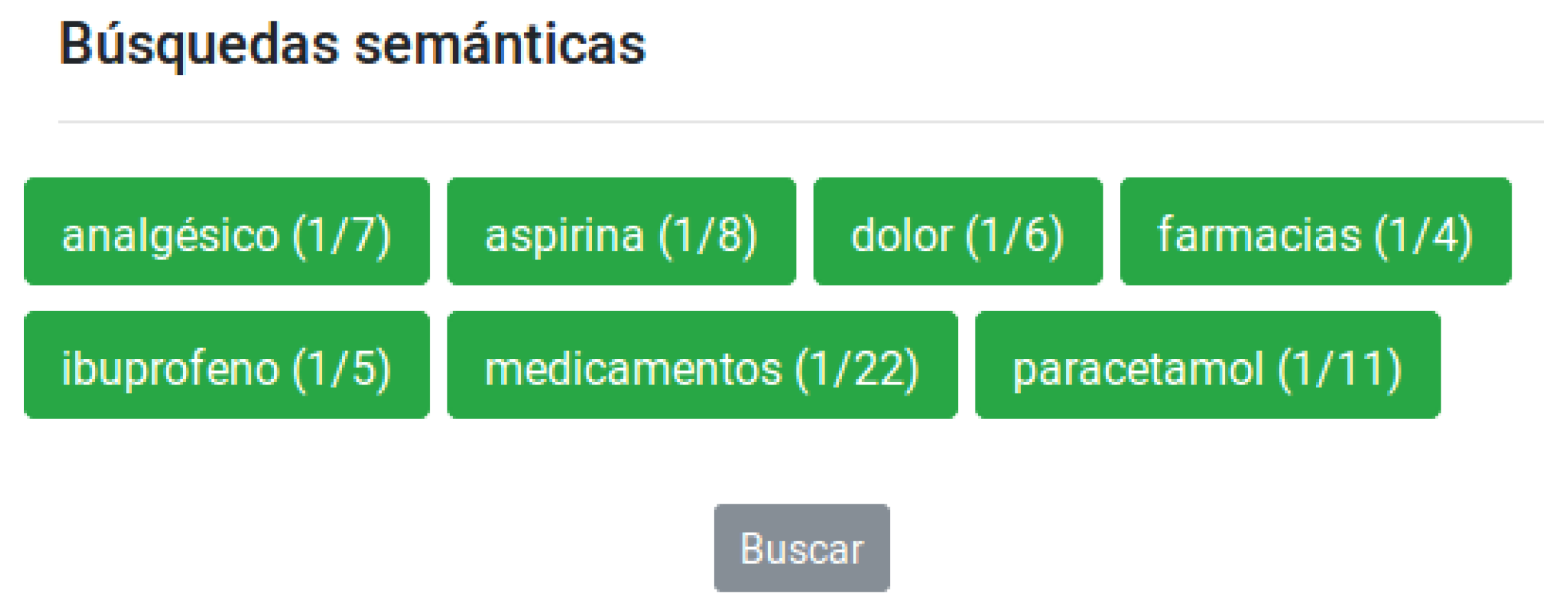
In
Figure 7 we can see seven concepts and only one atom is marked per concept. If we click on the search bottom
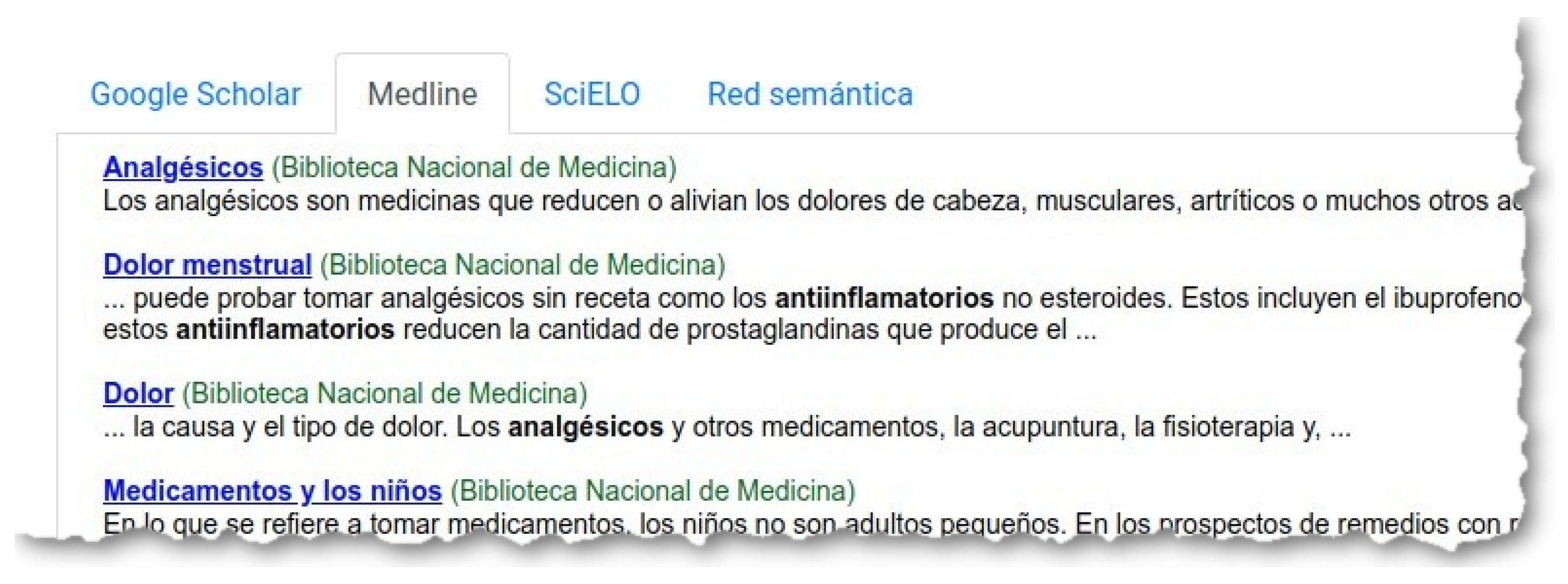
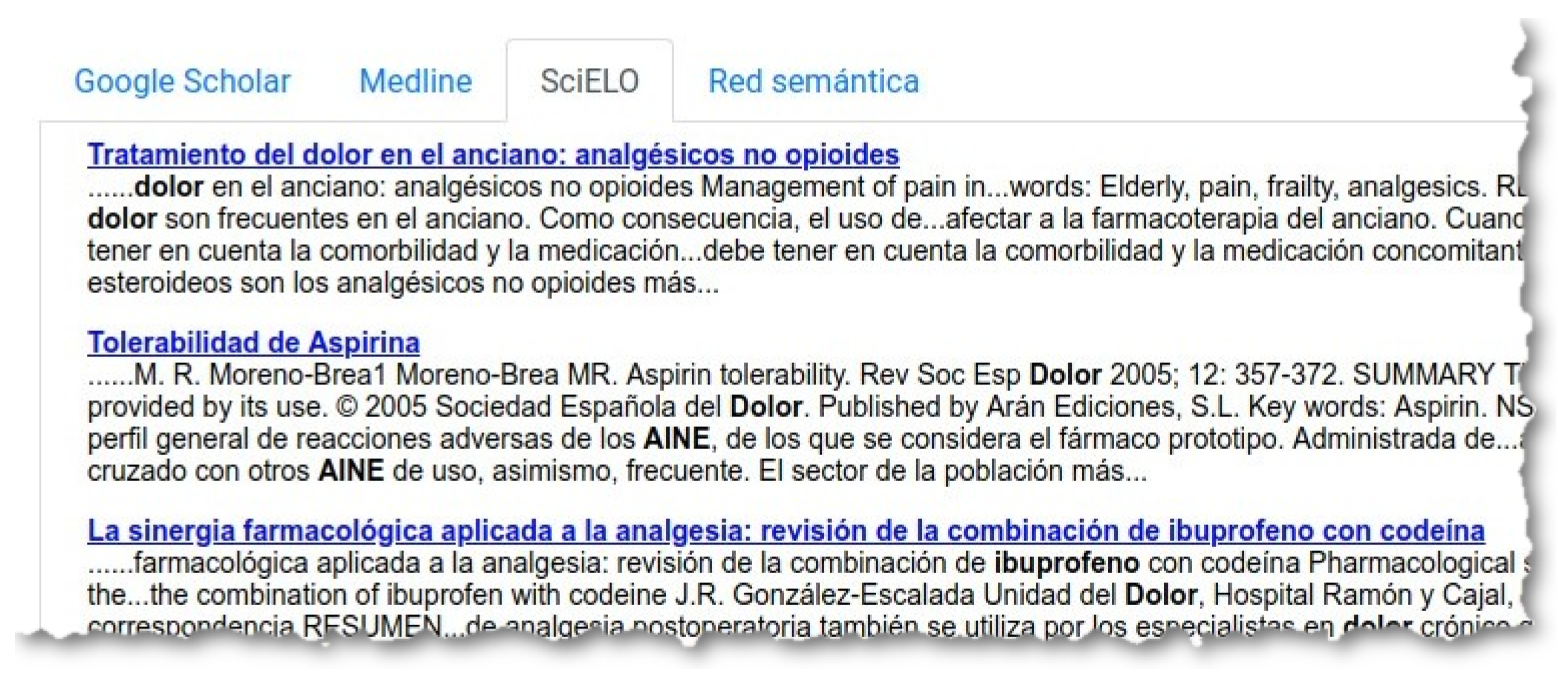
‘Buscar’ (Search), the system creates a query with the words in all the selected atoms, and launches this query through three search engines: Google Scholar (
Figure 8), Medline (
Figure 9) and SciELO (
Figure 10).
4.4. Semantic Relational Network/conceptual Graph
Due to the rich semantic content available after the recognition process, another functionality supported by the platform is the generation of a semantic graph over medical terms. In UMLS, the synonyms are grouped under a unique concept, and these concepts are linked with different types of relationships. Thus, UMLS is also a directed and labeled graph.
For every medical term extracted from the graph it is possible to find the related UMLS concept and find the relations with the rest of concepts in the text by finding paths through the UMLS network. In this way, a subgraph of UMLS is generated, applying the Dijsktra algorithm to find the minimal path between a central concept and the rest. The central node is, by default, selected according to its frequency of occurrence in the source text.
This graph is interactive, so the user can select any other node as central. This selection will trigger the regeneration of the graph, so minimal paths between this new central node and the rest are computed and rendered in a graphical representation on-the-fly. Due to the high number of concepts and relationships in UMLS, this is a very convenient approach for understanding how different concepts are interrelated and exploring UMLS relationships visually in an interactive manner.
Furthermore, in order to increase the readiness of the visual graph we have relabeled these relationships. There are two main types of semantic relations in UMLS:
hierarchical (e.g., ‘
is a kind of’, ‘
is a’, ‘
part of’) or
associative (e.g., ‘
location of’, ‘
caused by’). UMLS also assigns relationship types (UMLS abbreviations:
https://www.nlm.nih.gov/research/umls/knowledge_sources/metathesaurus/release/abbreviations.html) between concepts, for instance: narrower relationship (RN), synonymy (SY), parent relationship (PAR), among others.
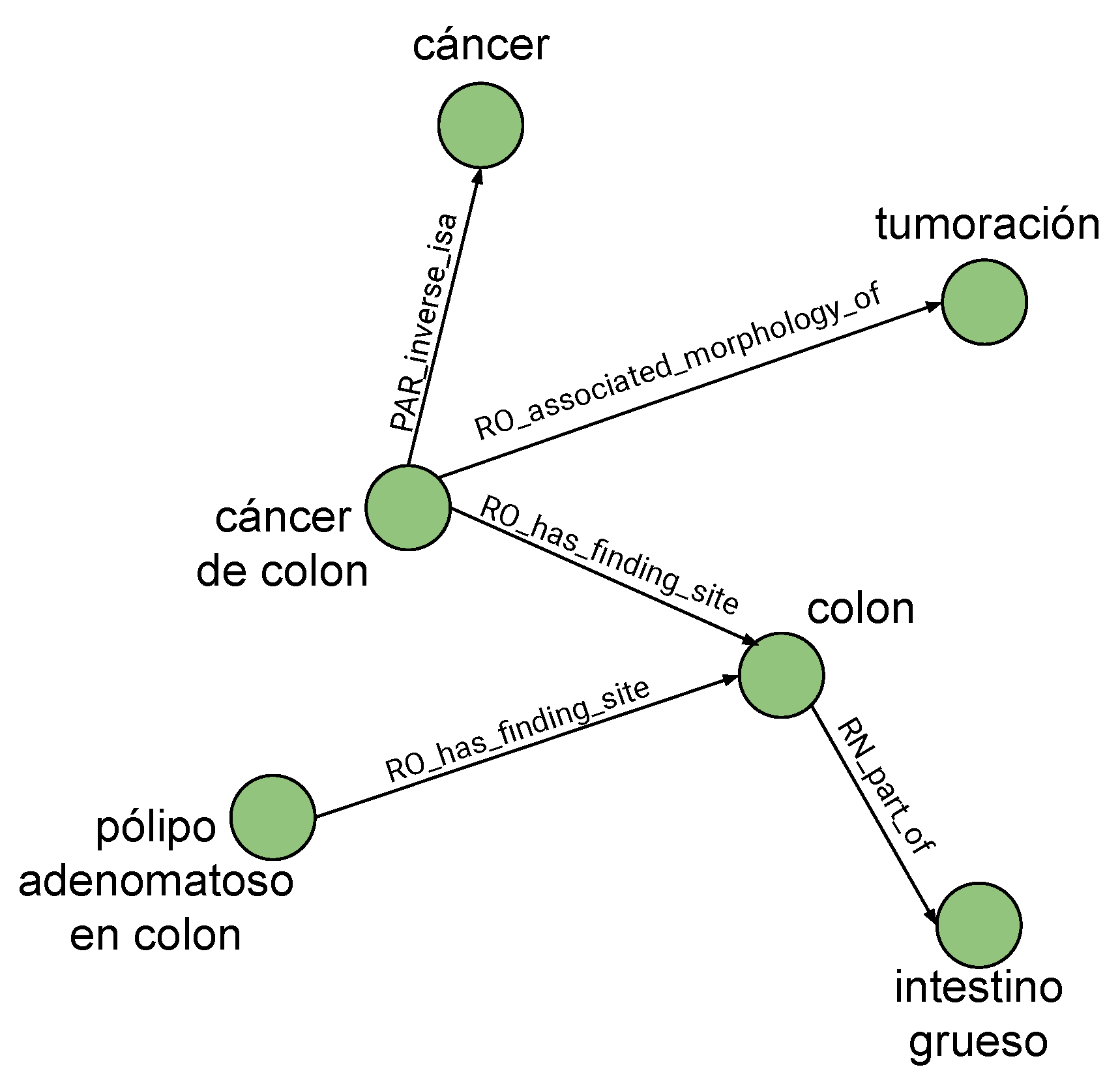
Figure 11 illustrates how, from the term identified as central ‘
cáncer of colon’, the other concepts are related to it. Intermediate nodes like ‘
colon’ are included as they are needed to reach other present and recognized terms like ‘
intestino grueso’ (‘
large intestine’) and ‘
pólipo adenomatoso en colon’ (‘
colon adenomatous polyp’).
As UMLS is a multilingual resource, the graph could be shown in several languages (even different to that from the analyzed text).
5. Evaluation
In order to evaluate our proposal, we have carried out experiments using the MANTRA corpus [
6] that includes biomedical Spanish documents annotated semantically. Later, in
Section 5.3 we perform an error analysis in order to detect and understand our failures.
5.1. The MANTRA Corpus
The MANTRA corpus was developed as part of the MANTRA project, aimed at providing multilingual terminologies and semantically annotated multilingual documents in English, French, German, Spanish, and Dutch. The MANTRA corpus consists of three parallel corpora: Medline titles, sentences from drug labels provided by the European Medicines Agency (EMEA), and sentences from patents made available by the European Patent Office. The Medline and EMEA corpora include parallel texts in English, French, Dutch, German and Spanish, while the patents corpus is available for English, French and German. In the case of the English-Spanish pairs, both Medline and EMEA corpora include 100 textual parallel units (titles or sentences) annotated with a subset of UMLS concepts from MeSH, SNOMED CT and MedDRA.
To test our system, we have used the Spanish subset in MANTRA Medline. We have taken into account the first 2000 documents. Analyzing these 2000 documents of Medline, we have been able to verify that the average number of tokens is of 12 words per document.
Evaluation Metrics
The primary evaluation metrics consisted of standard measures from the NLP community, namely micro-averaged precision, recall, and balanced F-score, the last one being the official evaluation measure:
where
= true positives,
= false positive and
= false negative.
5.2. Experiments and Results
Table 1 shows the results obtained in terminology detection by the two systems tested on the MANTRA Medline corpus (2000 documents): BSB (Buscador Semántico Biomédico—Biomedical Semantic Search Engine) and FreelingMed. We have chosen FreelingMed because it is the most popular system for Spanish biomedical terminology detection.
The recall obtained with BSB is higher because the system recognizes more terms than FreelingMed. Specifically we have checked that only in the the first 200 documents, BSB annotate 346 concepts that are not taken into account in MANTRA Medline. Regarding precision, both systems behave in a similar manner, but thanks to the good recall reported by BSB, the F1 score increases considerably for the BSB system. Finally, on accuracy BSB outperforms FreelingMed by 22%.
5.3. Error Analysis
The main purpose of this section is to carry out an error analysis to identify the weaknesses of our system. To this end, we have obtained some basic statistics for the first 2000 documents of the MANTRA subset used:
The average number of entities annotated in the MANTRA Medline corpus is 4.2 per document.
The average number of entities recognized by the BSB system is 5.8 concepts per document.
The average number of entities recognized by FreelingMed is 4.6 per document.
We have tried to identify which entities are not recognized by the BSB system. For this, we have taken 200 random documents with 483 medical entities and characterized the concepts annotated in MANTRA Medline that BSB was unable to detect.
Table 2 summarizes the situations where concepts were not detected by the BSB system and the main cause.
Below are several examples of missing terms in BSB. In each example, the text of the document is shown with the unidentified entity marked in bold, the CUI of UMLS as annotated in MANTRA and, finally, the atoms that we have found for that CUI in our UMLS dictionary in Spanish.
To finish this study we wanted to perform an analysis of the groups of entities that are classified in MANTRA Medline and how our system fails to address them. All terms are categorized with semantic groups: disorders (DISO), living (LIVB), anatomy (ANAT), phenomena (PHEN), procedures (PROC), devices (DEVI), physiological disorders (DISO-PHYS), geographic areas (GEOG), objects (OBJC), chemical phenomena (CHEM-PHEN), physiology (PHYS) and chemical and drugs (CHEM).
As can be seen in
Table 9, our system is usually 20 percentage point higher in every semantic category compared to FreelingMed, except in the OBJ group where FreelingMed obtains 60.68% and BSB 39.32%.
6. Conclusions
The main goal of this work is to serve as a proof of concept in the development of support systems in medical text analysis and search, for both experts and non-expert user communities. This is achieved by a core automatic detection system on biomedical terminology, which helps in query refinement and text comprehension by concept linking. To evaluate this core component, we have tested the performance of our system on the MANTRA Spanish subset compared to the state-of-the-art FreelingMed tool on this term identification task. The results show that BSB is an accurate system, so we expect that the interactive functionalities provided by this experimental software could be useful in document retrieval and semantic analysis. Query construction by concept selection approach and the semantic graph of specialized concepts are promising features to be integrated into medical information retrieval systems.
As future work we plan to incorporate multilingual support into the tool in order to extrapolate the system to other languages. In addition, and after the issues identified in error analysis, we consider that we need to continue working to develop a more accurate concept detector, for example by extending the vocabulary covered by BSB in order to improve recall. As a last expectation, we plan to test the usability and validity of the system with professional users from the medical domain.
In the era of Artificial Intelligence, Clinical Decision Support systems are arising as autonomous and fully end-to-end automatic systems able to propose a limited number of most confident choices [
36]. We believe that doctors and scientists still can profit from an improved access to document databases, as it augment their knowledge and empower them as experts.