Simple Summary
Yaks (Bos grunniens) inhabit the Qinghai-Tibetan Plateau and adjacent highlands at elevations between 2000 and 5000 m, where they are important domestic animals, as they provide meat, milk, fuel, and other necessities for Tibetans and nomads in China. Yak meat is fine in texture and high in protein, yet poor in muscular marbling and tenderness. Diacylglycerol acyltransferase-2 (DGAT2), which regulates fat deposition in animals, is a candidate gene for meat quality and quantity traits. However, there have been few reports on the effects of the DGAT2 gene on the meat quality of yak. Our study elucidated tissue expression of the yak DGAT2 gene and association of variation in the gene with Warner–Bratzler shear force of longissimus muscle. The results provide guidance for the molecular-assisted selection of meat tenderness in yak.
Abstract
Diacylglycerol acyltransferase-2 (DGAT2) plays a key role in the synthesis of animal triglycerides (TGs). This study investigated the relative expression of the DGAT2 gene in tissues, variation in the gene, and its association with carcass and meat quality traits in yaks (Bos grunniens). DGAT2 was found to be expressed in twelve tissues investigated, but the highest expression was detected in subcutaneous fat, and moderate levels were observed in the liver, heart, longissimus dorsi muscle, and abomasum. Three variants (A1 to C1) were found in intron 5 and another three variants (A2 to C2) were found in intron 6, with two single-nucleotide polymorphisms (SNPs) being identified in each region in 694 Gannan yaks. Variants B1 and C2 were associated with a decrease in Warner–Bratzler shear force (WBSF) (p = 0.0020 and p = 0.0441, respectively), and variant C1 was associated with an increase in WBSF (p = 0.0434) and a decrease in drip loss rate (p = 0.0271), whereas variant B2 was associated with a decrease in cooking loss rate (p = 0.0142). Haplotypes A1-A2 and B1-A2 were found to be, respectively, associated with an increase and a decrease in WBSF (p = 0.0191 and p = 0.0010, respectively). These results indicate that DGAT2 could be a useful gene marker for improving meat tenderness in yaks.
1. Introduction
Triglycerides (triacylglycerols, TGs) are the major energy storage molecules in most eukaryotic cells. The enzyme Acyl-CoA: diacylglycerol acyltransferase (DGAT), which was first reported in chicken liver in 1960 [1], plays a predominant role in catalyzing the final, rate-limiting step of triglyceride synthesis in mammals [2]. Two DGAT enzymes, DGAT1 and DGAT2, which are involved in different pathways, have been identified in a wide variety of eukaryotes [3,4,5].
The existence of DGAT2 was predicted from the finding that DGAT1-deficient mice (Dgat1−/−) were viable and still able to synthesize TGs [6]. DGAT2 belongs to an acyltransferase gene family that has no homology with DGAT1 [5] and is highly expressed in tissues that make large amounts of TGs, including the liver, white adipose tissues, and mammary glands [7,8], and appears to be predominantly responsible for TG homeostasis in vivo [9]. Therefore, DGAT2 is a promising candidate gene for traits related to lipid synthesis and storage in farm animals.
The bovine DGAT2 maps to chromosome 15q25–q26 [10] and numerous variations have been detected in cattle [10,11,12]. The expression of DGAT2 is positively correlated to intramuscular fat (IMF) content in longissimus dorsi muscles of Korean steers [13] and pigs [14]. Variation in DGAT2 has been reported to affect carcass muscle and milk quality traits in various animals, such as fat yield and retail cut weight percentage in commercial feedlot steers [11], backfat thickness and lean percentage in pigs [15,16], milk yield and fat percentage in goats [17], and carcass weight, shear force, and IMF in domestic pigeons [18].
Yak (Bos grunniens) is indigenous to the area of Central Asia highlands and has been regarded as one of the world’s most remarkable domestic animal species, because it thrives in conditions of extreme harshness while providing meat, milk, fuel, and other necessities for local people. There are an estimated 15 million yaks in China, accounting for over 90% of the world’s yak population [19]. Yak meat, which is the staple animal protein food and the major source of income for the local herdsmen, is fine textured and high in protein and minerals [20]. Yak meat is regarded as being very palatable, but it is poor in IMF content and intramuscular marbling [21]. To date, little has been done to investigate the effect of the DGAT2 gene on carcass muscle traits in yaks. The objective of this study was to screen genetic variation in yak DGAT2 and to investigate whether the variation is associated with carcass and meat quality traits.
2. Materials and Methods
2.1. Animals and Sample Collection
All animal experiments were conducted in accordance with the guidelines for the care and use of experimental animals established by the Ministry of Science and Technology of the People’s Republic of China (Approval number 2006-398) and was approved by the Animal Care Committee of Gansu Agricultural University, Lanzhou, China.
A total of 694 Gannan yaks (one of the twelve officially recognized domestic yak breeds) were investigated. These yaks were randomly selected from the Gannan Tibetan Autonomous Prefecture, Gansu Province, China. All of these yaks grazed on the plateau pasture at an altitude of 2500–4000 m all year round. The gender and age of each yak was recorded before slaughter. At slaughter, a blood sample from each yak was collected onto a Flinders Technology Associates (FTA) card (Whatman BioScience, Middlesex, UK) to isolate genomic DNA using a two-step procedure described in Zhou et al. [22].
Twelve tissues, including heart, liver, spleen, lung, kidney, large intestine (colon), small intestine (jejunum), rumen, abomasum, mammary gland, longissimus dorsi muscle, and subcutaneous fat, were carefully collected from three five-year-old female Gannan yaks within 30 min of slaughter in October, which is considered to be the beginning of the cold season of the Qinghai-Tibetan Plateau. All samples were immediately snap-frozen in liquid nitrogen and then stored at −70 °C.
2.2. Carcass and Meat Quality Measurement
Carcass and meat quality traits were measured in the 694 Gannan yaks from which blood samples were collected. Hot carcass weight (HCW; kg) was measured just after slaughter. At 48 h postmortem, the rib eye area (REA; cm2) was traced on the cross-section of the longissimus dorsi muscle between twelfth and thirteenth rib of the right carcass side using sulfate paper, and the tracing area was determined by planimeter. The portion of longissimus muscle, taken from eleventh to thirteenth rib, was used to assess the meat quality. The Warner–Bratzler shear force (WBSF; kg), which represents meat tenderness, was estimated using a Digital Muscle Tenderometer (Model C-LM3, Northeast Agricultural University, Harbin, China), according to the methods of Shackelford et al. [23]. Cooking loss rate (CLR; %) and drip loss rate (DLR; %) were determined using the methods described by Honikel [24] and Liu et al. [25], respectively.
2.3. PCR-SSCP Analysis
Polymerase chain reaction (PCR) primers (see Table 1) were designed based on the bovine DGAT2 sequence (GenBank Accession No. AC_000172.1) to amplify four fragments within intron 3–exon 4, intron 5, intron 6, and exon 8 regions in yak DGAT2. The primers were synthesized by Takara Biotechnology Co., Ltd. (Dalian, China).

Table 1.
PCR primers sequences and SSCP conditions for yak DGAT2.
PCR amplifications were run in a 20-μL reaction mixture, with the genome DNA on one 1.2-mm punch of blood spot on an FTA card as a template, 0.25 μM of each primer, 150 μM dNTP (Takara, Dalian, China), 2.5 mM Mg2+ and 0.5 U of Tαq DNA polymerase (Takara, Dalian, China), and double-distilled water to a final volume of 20 μL. The cycling protocols were 2 min at 94 °C, followed by 35 cycles of 30 s at 94 °C, 30 s at the annealing temperatures (Table 1), and 30 s at 72 °C, and a final extension of 5 min at 72 °C.
A single-stranded conformational polymorphism (SSCP) method was used to screen variation in each of the regions amplified. An aliquot of 2-μL amplicon was mixed with 8 μL of loading dye (98% formamide, 10 mM ethylenediaminetetraacetic acid (EDTA), 0.025% bromophenol blue, 0.025% xylene cyanol). The mixtures were denatured at 98 °C for 5 min, then chilled rapidly on wet ice for 5 min and subjected to 14% polyacrylamide (39:1) gels. Samples were electrophoresed in 0.5 × Tris-boric-acid-EDTA (TBE) buffer for 18 h under conditions described in Table 1. The gels were silver-stained according to Byun et al. [26].
2.4. Variant Sequencing and Analysis
Amplicons that were confirmed as homozygous by SSCP were directly sequenced in both directions at Sangon Biotech (Shanghai, China). Variants that were only found in heterozygous yaks were sequenced using a rapid sequencing method described by Hu et al. [27]. Sequence alignments were performed by DNAMAN (version 5.2.10, Lynnon BioSoft, Vaudreuil, QC, Canada).
2.5. Haplotype Determination
Haplotypes were constructed across introns 5 and 6 of DGAT2, as variations were only found in these two regions. For yaks that were identified as homozygous in either region, it was possible to directly ascertain their haplotypes based on the co-inheritance of sequences. For example, if a yak presented with genotype A1A1 in intron 5 and genotype A2B2 in intron 6, the presence of haplotypes A1–A2 and A1–B2 could be directly inferred. For those yaks that were heterozygous in both regions, the haplotypes could not be inferred because their parents’ genotypes of DGAT2 were not available for analysis in this study.
2.6. Statistical Analyses
Association analyses of DGAT2 variation in relation to carcass and meat quality traits of yak were performed with the general linear mixed models (GLMMs) in IBM SPSS 24.0 (IBM Corp., Armonk, NY, USA). Initially, single-variant/haplotype models were conducted to confirm which variants/haplotypes should be factored into subsequent multi-variant models that would include any variant/haplotype that had an association with a trait in the single-variant/haplotype models with p < 0.2, and which might therefore potentially impact on the carcass and meat quality traits being tested.
Gender, age, and group were found to affect carcass and meat quality traits, and they were therefore fitted into the models. Unless otherwise indicated, all p-values less than 0.05 were considered to be significantly different.
2.7. RNA Extraction and RT-qPCR Analysis
The reverse transcription-qPCR (RT-qPCR) was performed to ascertain the yak DGAT2 expression in tissues using yak β-actin as an internal control. Two pairs of qPCR primers were designed to amplify a 149-bp fragment of yak DGAT2 and a 133-bp fragment of β-actin based on the mRNA sequences (GenBank Accession Nos. XM_005902498 and DQ838049.1, respectively). The DGAT2 qPCR primers were 5’-GGCCTCTTCTCCTCTGACACC-3’ and 5’-CACCAGGGCTTGCACGTACA-3’, and the β-actin primers were 5’-AGCCTTCCTTCCTGGGCATGGA -3’ and 5’- GGACAGCACCGTGTTGGCGTAGA -3’. Total RNA was extracted from tissue samples of yak using TRIzol reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s instructions. The integrity and concentration of total RNA was assessed by 2% agarose gels in electrophoresis and UV spectrophotometry. The cDNA was synthesized by reverse transcription from total RNA using the PrimeScriptTM RT Reagent Kit with the gDNA Eraser (Takara) following the manufacturer’s instructions. The amplification of the cDNA was conducted in 20-μL reaction mixture consisting of 100 ng cDNA, 0.25 μM of each primer, 10.0 μL AceQ qPCR SYBR® Green Master Mix (Vazyme, Nanjing, China), 0.4 μL ROX Reference Dye 2, and supplemented with ddH2O to a volume of 20 μL. The thermal profiles were one cycle of 5 min at 95 °C, followed by 40 PCR cycles of 10 s at 95 °C, 30 s at 60 °C, and 30 s at 72 °C. The 2−ΔΔCT method was used to analyze the relative expression data [28]. Amplification was performed in Applied Biosystems Quant Studio®6 Flexq (Applied Biosystems, Carlsbad, CA, USA).
3. Results
3.1. Identification of Variation in Yak DGAT2
The four pairs of PCR primers, which amplified four fragments of yak DGAT2, including the intron 3–exon 4, intron 5, intron 6, and exon 8 fragments, worked well on all of the DNA samples under the conditions established. In the 694 yaks investigated, three PCR-SSCP patterns representing three variant sequences were detected for both introns 5 and 6, respectively (Figure 1A). Four single-nucleotide polymorphisms (SNPs) were detected among those sequences (Figure 1B). Of these, SNPs c.612 + 350C/A and c.612 + 228A/G were located in intron 5 and c.787 + 616G/A and c.787 + 691T/G in intron 6. No variations were detected for intron 3–exon 4 and exon 8 fragments (Figure 1A).
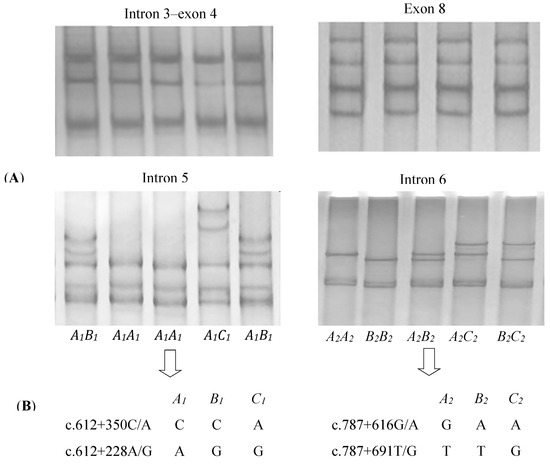
Figure 1.
SSCP (single-stranded conformational polymorphism) banding patterns for the four regions of yak DGAT2 amplified (A) and single-nucleotide polymorphisms (SNPs) detected in introns 5 and 6 of yak DGAT2 (B).
3.2. Polymorphisms in Yak DGAT2
In the 694 yaks, all genotype and variant frequencies in each region were >5% (Table 2). Variants A1 (in intron 5) and A2 (in intron 6) were the most common, with frequencies of 69.96% and 65.49%, respectively.

Table 2.
Genotype and variant frequency of yak DGAT2.
Seven haplotypes, spanning introns 5 and 6 of DGAT2, could be determined in the 469 individuals that were a subset of the 694 yaks in which the variants were identified. Of these haplotypes, A1-A2, A1-B2, and B1-A2 were the most common, with frequencies of 64.71%, 11.62%, and 14.18%, respectively. The other four haplotypes, A1-C2, B1-B2, C1-A2, and C1-B2 were rare, with frequencies of 3.30%, 0.64%, 4.37%, and 1.17%, respectively.
3.3. Associations of DGAT2 Variation with Carcass and Meat Quality Traits in Yak
In the single-variant (presence/absence) model (Table 3), the presence of variant B1 and the absence of C1 in intron 5 were associated with a decrease in WBSF (p = 0.0020 and p = 0.0434, respectively), and the presence of C1 was associated with decreased DLR (p = 0.0271). The presence of C2 and B2 in intron 6 were found to be associated with a decrease in WBSF (p = 0.0441) and CLR (p = 0.0142), respectively. These associations with WBSF remained significant when the other variants were factored into the models (p < 0.2). No associations with REA and HCW were detected for any variant in both regions of yak DGAT2.

Table 3.
Association of DGAT2 variants with carcass and meat quality traits (mean ± SE) 1 in yaks.
In the single-haplotype (presence/absence) model (Table 4), the absence of haplotype A1-A2 and the presence of haplotype B1-A2 were associated with a decrease in WBSF (p = 0.0191 and p = 0.0010, respectively). These associations persisted in the multi-variant models. No effects of DGAT2 haplotypes were found on HCW, REA, DLR and CLR.

Table 4.
Association of DGAT2 haplotypes with carcass and meat quality traits (mean ± SE) 1 in yaks.
3.4. Tissue Expression of Yak DGAT2
The DGAT2 expression level was significantly different among twelve tissues of yak (p < 0.05; Figure 2). The highest expression was found in subcutaneous fat, whereas moderate expression was detected in the liver, heart, longissimus dorsi muscle, and abomasum, and weak expression was observed in other tissues, including the large intestine, small intestine, mammary gland, lungs, kidney, spleen, and rumen.
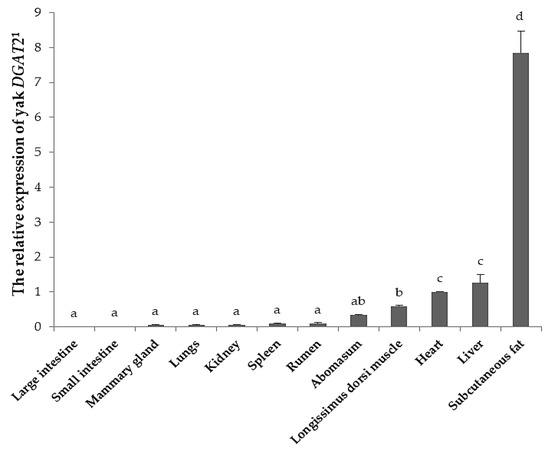
Figure 2.
The relative expression of DGAT2 in twelve tissues of yak. 1 Heart represents control tissue. Error bars indicates standard deviation. The different lower-case letters on columns indicate significant difference (p < 0.05).
4. Discussion
This is the first report to elucidate the associations between DGAT2 variation and meat tenderness in yak. Meat tenderness, which is considered to be the most important component of beef quality identified by consumers, is positively influenced by IMF content [29,30]. IMF could improve meat tenderness by weakening structures of intramuscular connective tissue [31]. Hocquette et al. [29] found that the IMF content had a positive genetic correlation with tenderness (rG = 0.41) and a negative correlation with WBSF (rG = −0.50) in cattle. Given that the deposition of IMF is mainly determined by lipid metabolism, many lipid metabolic genes may be involved in fat deposition in muscle [13]. DGAT2 plays an important role in fat deposition, as it is a key enzyme that catalyzes the final and rate-limited step of TG synthesis. Wakimoto et al. [32] found that DGAT2 is associated with adipose assembly in adipose tissue and skeletal muscle, and Winter et al. [10] reported that DGAT2 could be a priority candidate gene for quantitative traits related to TG synthesis and storage in farm animals.
In this study, a total of four SNPs were detected in introns 5 and 6 of the yak DGAT2 gene. The level of variation detected in yaks appears to be lower than that reported in cattle in which 13 SNPs and 2 ins/del variations have been found in introns 5 and 6 [10]. SNPs in introns have been shown to affect gene expression by regulating the rate of transcription, nuclear export, transcript stability, and the efficiency of mRNA translation in many eukaryotes [33]. Moreover, the optimal expression of many genes also requires the presence of one or more introns [34]. Although these SNPs in yak DGAT2 were not located in the coding regions, they may influence the gene expression, and thus affect fat-related traits.
The association between variation in DGAT2 and the various quantitative traits related to TG synthesis and storage has been recently reported in a number of other animal species. Li et al. [11] found that SNPs in intron 3 of DGAT2 were associated with the body weight and fat yield in commercial feedlot steers. Yin et al. [15] and Li et al. [16] reported that a SNP and 13 bp indel in the 3′-UTR of DGAT2 were associated with backfat thickness and lean percentage in pigs. Mao et al. [18] found that SNPs in exons 5 and 6 of DGAT2 had an effect on carcass and meat quality traits in domestic pigeons. In goats, variations in exon 3 of DGAT2 were found to be associated with growth traits [35], while variations in exon 4 and intron 5 had an influence on the milk yield and fat percentage [17]. In current study, variations in introns 5 and 6 of yak DGAT2 were identified and showed that the presence of variant B1 and absence of C1 in intron 5, as well as the presence of C2 in intron 6, was associated with a decrease in WBSF. Despite there being few reports on the associations between DGAT2 variation and WBSF in other species, the level of DGAT2 expression was found to be positively correlated (p < 0.05) with IMF content in cattle and pigs [13,14]. DGAT2 was found to be moderately expressed in longissimus dorsi muscle of yak in this study. Given the positive influence of the IMF content on meat tenderness, variation in yak DGAT2 could affect meat tenderness by regulating IMF deposition in muscle.
In this study, the absence of haplotype A1-A2 and the presence of B1-A2 were associated with a decrease in WBSF of yak meat. Moreover, no haplotype was found to be associated with DLR and CLR, even though the presence of variant C1 in intron 5 and B2 in intron 6 was associated with decreased DLR and increased CLR, respectively. Haplotypes are particular combinations of alleles or variant sequences on a single chromosome. Their information, which is composed of multiple markers, would be more powerful than individual and unorganized markers, as they may improve the power of genome-wide association studies [36,37]. In general, molecular markers could be accurately identified when integrating the haplotypes and SNPs into association analysis [38]. For instance, Zollner et al. [39] reported that markers with more variants were much more efficient in detecting any significant associations. The associations identified here suggest that variations in yak DGAT2 affect meat tenderness. Further investigations on more yaks from different farms and breeds are needed to confirm this.
DGAT2 is expressed in a variety of tissues in mammals. In this study, we found that the highest expression level of yak DGAT2 was in subcutaneous fat, followed by the liver, heart, longissimus dorsi muscle, and other tissues. High expression of DGAT2 in adipose tissue indicates an important function in triglyceride storage [4]. Yaks are indigenous to the Qinghai-Tibet Plateau and adjacent highlands at an altitude from 2000 m to 5000 m with cold, semi-humid climates, and they deposit more subcutaneous fat prior to winter in order to cope with the severe cold [21]. In addition, previous studies have showed that the TG synthesis pathway is not an important step for hepatic fat accumulation in cattle [40,41]. Therefore, we speculate that the higher DGAT2 expression in subcutaneous fat than in liver is propitious to more accumulation of subcutaneous fat, which would provide more energy reserve that can used over the seven-month long cold season in the Qinghai-Tibetan Plateau. However, some reports have showed that DGAT2 expression abundance might relate to the species and developmental stage of tissues. The highest expression of DGAT2 is found in adipose tissues in mice, but in pigs and humans the highest level of expression is in the liver [4,14]. In sheep, different expression levels are observed in various tissues at different developmental stages, such as the liver, rumen, and jejunum [42].This discrepancy may be due to the biological function of DGAT2, and it is speculated that the post-transcriptional regulatory mechanism plays a role in DGAT2 expression in different species and developmental stages. Further studies are required to detect DGAT2 expression in tissues from yaks at different development stages and with different DGAT2 variants in order to understand whether genetic variations and developmental stages affect gene expression.
5. Conclusions
This study elucidated tissue expression of the yak DGAT2 gene and revealed variation in the gene. The highest expression was found in the subcutaneous fat of yaks. Variants and haplotypes in introns 5 and 6 of yak DGAT2 were associated with the WBSF of longissimus muscle. These results suggest that DGAT2 variations could be used as genetic markers in breeding programs to improve meat tenderness in yaks.
Author Contributions
Conceptualization: J.H. and Y.L.; Methodology: Z.Z.; Validation: J.H. and Y.L.; Formal Analysis: J.W. and X.L.; Investigation: B.S., J.X., S.L., and Z.Z.; Resources: X.L., S.L., and Z.Z.; Data Curation: J.H. and J.X.; Writing—Original Draft Preparation: J.H., B.S., and Z.Z.; Writing—Review and Editing: H.Z. and Y.L.; Visualization: J.H.; Supervision: Y.L.; Project Administration: J.H. and Y.L.; Funding acquisition: J.H. and Y.L.
Funding
This research was funded by the Discipline Construction Fund Project of Gansu Agricultural University (GAU-XKJS-2018-050).
Acknowledgments
We thank Gannan Institute of Animal Science for its part in collecting the samples and LetPub (www.letpub.com) for its linguistic assistance during the preparation of this manuscript.
Conflicts of Interest
The authors have no conflict of interest to declare.
References
- Weiss, S.B.; Kennedy, E.P.; Kiyasu, J.Y. The enzymatic synthesis of triglycerides. J. Biol. Chem. 1960, 235, 40–44. [Google Scholar] [CrossRef] [PubMed]
- Mayorek, N.; Grinstein, I.; Bar-Tana, J. Triacylglycerol synthesis in cultured rat hepatocytes. The rate-limiting role of diacylglycerol acyltransferase. Eur. J. Biochem. 1989, 182, 395–400. [Google Scholar] [CrossRef] [PubMed]
- Cases, S.; Smit, S.J.; Zheng, Y.W.; Myers, H.M.; Lear, S.R.; Sande, E.; Novak, S.; Collins, C.; Welchi, B.C.; Lusisi, A.J.; et al. Identification of a gene encoding an acyl CoA: Diacylglycerol cyltransferase, a key enzyme in triacylglycerol synthesis. Proc. Natl. Acad. Sci. 1998, 95, 13018–13023. [Google Scholar] [CrossRef] [PubMed]
- Cases, S.; Stone, S.J.; Zhou, P.; Yen, E.; Tow, B.; Lardizabal, K.D.; Voelker, T.; Farese, R.V. Cloning of DGAT2, a second mammalian diacylglycerol acyltransferase, and related family members. J. Biol. Chem. 2001, 276, 38870–38876. [Google Scholar] [CrossRef] [PubMed]
- Lardizabal, K.D.; Mai, J.T.; Wagner, N.W.; Wyrick, A.; Voelker, T.; Hawkins, D.J. DGAT2 is a new diacylglycerol acyltransferase gene family: Purification, cloning, and expression in insect cells of two polypeptides from Mortierella ramanniana with diacylglycerol acyltransferase activity. J. Biol. Chem. 2001, 276, 38862–38869. [Google Scholar] [CrossRef] [PubMed]
- Smith, S.J.; Cases, S.; Jensen, D.R.; Chen, H.C.; Sande, E.; Tow, B.; Sanan, D.A.; Raber, J.; Eckel, R.H.; Farese, R.V., Jr. Obesity resistance and multiple mechanisms of triglyceride synthesis in mice lacking DGAT. Nat. Genet. 2000, 25, 87–90. [Google Scholar] [CrossRef] [PubMed]
- Jin, Y.; Mcfie, P.J.; Banman, S.L.; Curtis Brandtm, C.; Stone, S.J. Diacylglycerol acyltransferase-2 (DGAT2) and monoacylglycerol acyltransferase-2 (MGAT2) interact to promote triacylglycerol synthesis. J. Biol. Chem. 2014, 289, 28237–28248. [Google Scholar] [CrossRef] [PubMed]
- Yen, C.L.; Stone, S.J.; Koliwad, S.; Harris, C.; Farese, R.V., Jr. Thematic review series: Glycerolipids. DGAT enzymes and triacylglycerol biosynthesis. J. Lipid Res. 2008, 49, 2283–2301. [Google Scholar] [CrossRef] [PubMed]
- Stone, S.J.; Myers, H.; Brown, B.E.; Watkins, S.M.; Feingold, K.R.; Elias, P.M.; Farese, R.V., Jr. Lipopenia and skin barrier abnormalities in DGAT2-deficient mice. J. Biol. Chem. 2004, 279, 11767–11776. [Google Scholar] [CrossRef] [PubMed]
- Winter, A.; Van, E.M.; Bininda-Emonds, O.R.; Habermann, F.A.; Fries, R. Genomic organization of the DGAT2/MOGAT gene family in cattle (Bos taurus) and other mammals. Cytogenet. Genome Res. 2003, 102, 42–47. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Xu, X.; Zhang, Q.; Wang, X.; Deng, G.; Fang, X.; Gao, X.; Ren, H.; Xu, S. Association between single nucleotide polymorphisms in the DGAT2 gene and beef carcass and quality traits in commercial feedlot steers. Asian Austral. J. Anim. 2009, 22, 943–954. [Google Scholar] [CrossRef]
- Alshuhaib, M.B.S.; Alfihan, R.A.; Alqutbi, A.A.; Althuwaini, T.M. Potential consequences of DGAT2 and BTN genes polymorphism in Iraqi Holstein cattle. Sci. Agric. Bohemica 2017, 48, 127–141. [Google Scholar]
- Jeong, J.; Kwon, E.G.; Im, S.K.; Seo, K.S.; Baik, M. Expression of fat deposition and fat removal genes is associated with intramuscular fat content in longissimus dorsi muscle of Korean cattle steers. J. Anim. Sci. 2012, 90, 2044–2053. [Google Scholar] [CrossRef] [PubMed]
- Cui, J.; Zeng, Y.; Wang, H.; Chen, W.; Du, J.; Chen, Q.; Hu, Y.; Yang, L. The effects of DGAT1 and DGAT2 mRNA expression on fat deposition in fatty and lean breeds of pig. Livest. Sci. 2011, 140, 292–296. [Google Scholar] [CrossRef]
- Yin, Q.; Yang, H.; Han, X.; Fan, B.; Liu, B. Isolation, mapping, SNP detection and association with backfat traits of the porcine CTNNBL1 and DGAT2 genes. Mol. Biol. Rep. 2012, 39, 4485–4490. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Wang, Y.; Sun, B.; Zhang, X.; Yang, C.; Kang, L.; Zhao, Z.; Jiang, Y. Identification of a 13 bp indel polymorphism in the 3′-UTR of DGAT2 gene associated with backfat thickness and lean percentage in pigs. Gene 2016, 576, 729–733. [Google Scholar]
- An, X.; Song, S.; Hou, J.; Zhua, C.; Penga, J.; Liua, X.; Liua, H.; Xiao, W.; Zhao, H.; Baia, L.; et al. Polymorphism identification in goat DGAT2, gene and association analysis with milk yield and fat percentage. Small Rumin. Res. 2011, 100, 107–112. [Google Scholar] [CrossRef]
- Mao, H.; Dong, X.; Cao, H.; Xu, N.; Yin, Z. Association of DGAT2 gene polymorphisms with carcass and meat quality traits in domestic pigeons (Columba livia). Brit. Poultry Sci. 2018, 59, 149–153. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Wang, Z.; Peng, Q.; Tan, C.; Zou, H. Effects of different levels of protein supplementary diet on gene expressions related to intramuscular deposition in early-weaned yaks. Anim. Sci. J. 2014, 85, 411–419. [Google Scholar] [CrossRef]
- Tian, J.; Han, L.; Yu, Q.; Shi, X.; Wang, W. Changes in tenderness and cathepsins activity during post mortem ageing of yak meat. Can. J. Anim. Sci. 2013, 93, 321–328. [Google Scholar] [CrossRef]
- Wiener, G.; Han, J.; Long, R. The Yak, 2nd ed.; FAO Regional Office for Asia and the Pacific Press: Bangkok, Thailand, 2003; p. 5. [Google Scholar]
- Zhou, H.; Hickford, J.G.H.; Fang, Q. A two-step procedure for extracting genomic DNA from dried blood spots on filter paper for polymerase chain reaction amplification. Anal. Biochem. 2006, 354, 159–161. [Google Scholar] [CrossRef]
- Shackelford, S.D.; Wheeler, T.L.; Koohmaraie, M. Evaluation of slice shear force as an objective method of assessing beef longissimus tenderness. J. Anim. Sci. 1999, 77, 2693–2699. [Google Scholar] [CrossRef] [PubMed]
- Honikel, K.O. Reference methods for the assessment of physical characteristics of meat. Meat Sci. 1998, 49, 457–477. [Google Scholar] [CrossRef]
- Liu, M.; Peng, J.; Xu, D.; Zheng, R.; Li, F.; Li, J.; Zuo, B.; Lei, M.; Xiong, Y.; Deng, C.; et al. Association of MYF5 and MYOD1 gene polymorphisms and meat quality traits in large whitex meishan F2 pig populations. Biochem. Genet. 2008, 46, 720–732. [Google Scholar] [CrossRef]
- Byun, S.O.; Fang, Q.; Zhou, H.; Hickford, J.G.H. An effective method for silver-staining DNA in large numbers of polyacrylamide gels. Anal. Biochem. 2009, 385, 174–175. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.; Zhou, H.; Smyth, A.; Luo, Y.; Hickford, J.G.H. Polymorphism of the bovine ADRB3 gene. Mol. Biol. Rep. 2010, 37, 3389–3392. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Hocquette, J.F.; Gondret, F.; Baéza, E.; Médale, F.; Jurie, C.; Pethicket, D.W. Intramuscular fat content in meat-producing animals: Development, genetic and nutritional control, and identification of putative markers. Animal 2010, 4, 303–319. [Google Scholar] [CrossRef] [PubMed]
- Devlin, D.J.; Gault, N.F.S.; Moss, B.W.; Tolland, E.; Tollerton, J.; Farmer, L.J.; Gordon, A.W. Factors affecting eating quality of beef. Adv. Animal Biosci. 2017, 8, s2–s5. [Google Scholar] [CrossRef]
- Nishimura, T.; Hattori, A.; Takahashi, K. Structural changes in intramuscular connective tissue during the fatting of Japanese black cattle: Effect of marbling on beef tenderization. J. Anim. Sci. 1999, 77, 93–104. [Google Scholar] [CrossRef] [PubMed]
- Wakimoto, K.; Chiba, H.; Michibata, H.; Seishima, M.; Kawasaki, S.; Okubo, K.; Mitsui, H.; Torii, H.; Mai, Y.I. A novel diacylglycerol acyltransferase (dgat2) is decreased in human psoriatic skin and increased in diabetic mice. Biochem. Bioph. Res. Commun. 2003, 310, 296–302. [Google Scholar] [CrossRef]
- Shaul, O. How introns enhance gene expression. Int. J. Biochem. Cell Biol. 2017, 91, 145–155. [Google Scholar] [CrossRef] [PubMed]
- Nott, A.; Meislin, S.H.; Moore, M.J. A quantitative analysis of intron effects on mammalian gene expression. RNA 2003, 9, 607–617. [Google Scholar] [CrossRef]
- Fang, X.; Zhang, J.; Xu, H.; Zhang, C.; Du, Y.; Shi, X.; Chen, D.; Sun, J.; Jin, Q.; Lan, X.; et al. Polymorphisms of diacylglycerol acyltransferase 2 gene and their relationship with growth traits in goats. Mol. Bio. Rep. 2012, 39, 1801–1807. [Google Scholar] [CrossRef]
- Snyder, M.W.; Adey, A.; Kitzman, J.O.; Shendure, J. Haplotype-resolved genome sequencing: Experimental methods and applications. Nat. Rev. Genet. 2015, 16, 344–358. [Google Scholar] [CrossRef] [PubMed]
- Akey, J.; Jin, L.; Xiong, M. Haplotypes vs. single marker linkage disequilibrium tests: What do we gain? Eur. J. Hum. Genet. 2001, 9, 291–300. [Google Scholar] [CrossRef]
- Scheike, T.H.; Martinussen, T.; Silver, J.D. Estimating haplotype effects for survival data. Biometrics 2010, 66, 705–715. [Google Scholar] [CrossRef] [PubMed]
- Zollner, S.; von Haeseler, A. Coalescent approach to study linkage disequilibrium between single-nucleotide polymorphisms. Am. J. Hum. Genet. 2000, 66, 615–628. [Google Scholar] [CrossRef]
- Baik, M.; Nguyen, T.H.; Jin, Y.J.; Min, Y.P.; Kang, H.J. Effects of castration on expression of lipid metabolism genes in the liver of Korean cattle. Asian Australas. J. Anim. Sci. 2015, 28, 127–134. [Google Scholar] [CrossRef]
- Van Den Top, A.M.; Wensing, T.; Geelen, M.J.H.; Wentink, G.H.; Van’t Klooster, A.T.; Beynen, A.C. Time trends of plasma lipids and enzymes synthesizing hepatic triacylglycerol during postpartum development of fatty liver in dairy cows. J. Dairy Sci. 1995, 78, 2208–2220. [Google Scholar] [CrossRef]
- BioGPS. Available online: http://biogps.org/sheepatlas/#goto=genereport&id=101110535 (accessed on 28 January 2019).
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).