Seawater Culture Increases Omega-3 Long-Chain Polyunsaturated Fatty Acids (N-3 LC-PUFA) Levels in Japanese Sea Bass (Lateolabrax japonicus), Probably by Upregulating Elovl5
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethic Statement
2.2. Experimental Fish and Diets
2.3. Sampling
2.4. Proximate Muscle Composition
2.5. Fatty Acid Composition
2.6. Quantitative Real-Time PCR Analysis
2.7. Cloning of the Elovl5 Promoters
2.8. Bisulfate Genomic Sequencing (BSP)
2.9. Statistical Analyses
3. Results
3.1. Proximate Composition of the Muscle
3.2. Fatty Acid Composition of the Muscle and of the Liver
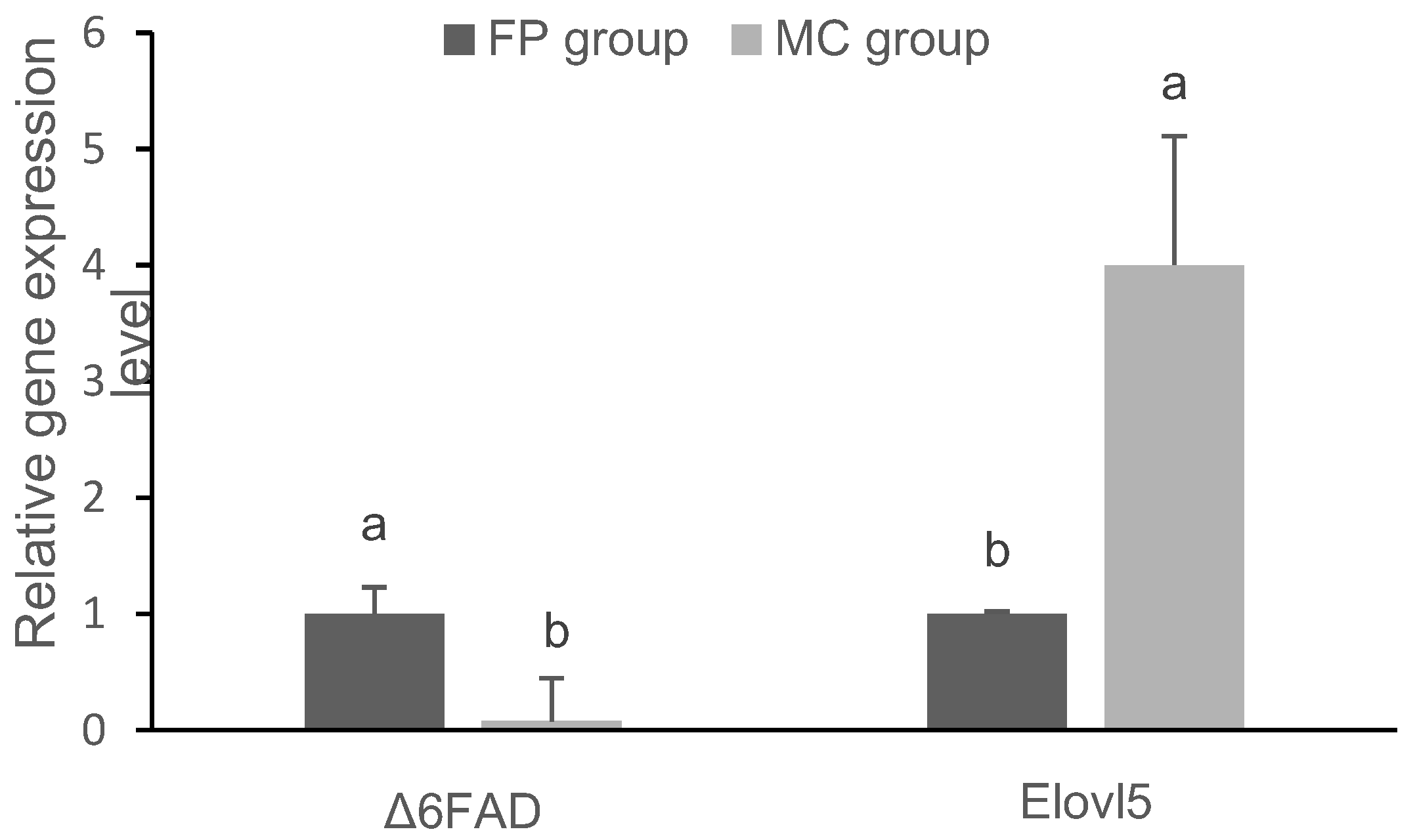
3.3. Gene Expression in the Liver
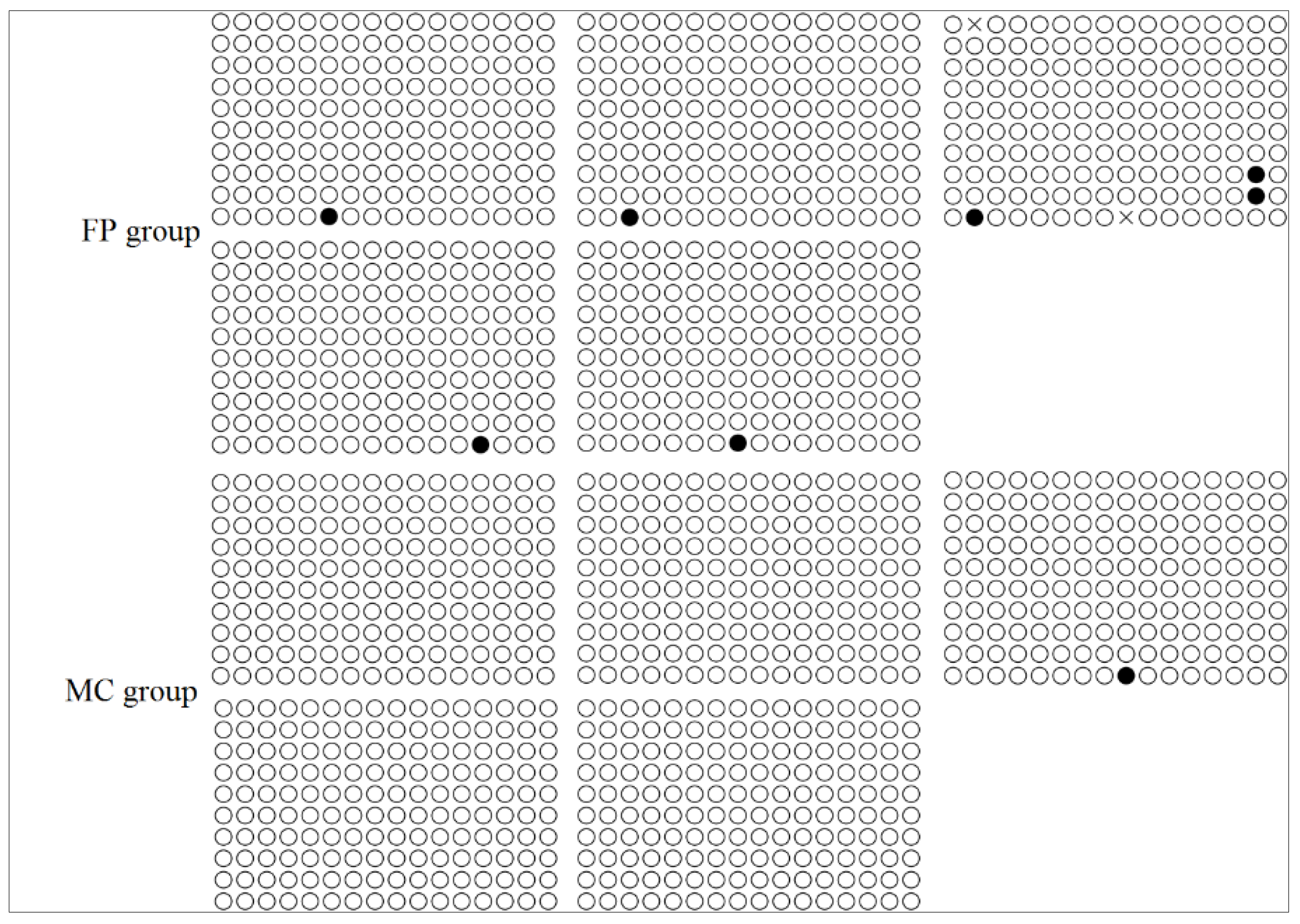
3.4. CpG Methylation of the FADS2 and Elovl5 Promoters
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Cordier, M.; Brichon, G.; Weber, J.-M.; Zwingelstein, G. Changes in the fatty acid composition of phospholipids in tissues of farmed sea bass (Dicentrarchus labrax) during an annual cycle. Roles of environmental temperature and salinity. Comp. Biochem. Physiol. Part B Biochem. Mol. Biol. 2002, 133, 281–288. [Google Scholar] [CrossRef]
- Khériji, S.; El Cafsi, M.; Masmoudi, W.; Castell, J.; Romdhane, M. Salinity and Temperature Effects on the Lipid Composition of Mullet Sea Fry (Mugil cephalus, Linne, 1758). Aquac. Int. 2003, 11, 571–582. [Google Scholar] [CrossRef]
- Haliloǧlu, H.I.; Bayır, A.; Sirkecioǧlu, A.N.; Aras, N.M.; Atamanalp, M.; Haliloğlu, H.I.; Bayir, A.; Sirkecioğlu, A.N. Comparison of fatty acid composition in some tissues of rainbow trout (Oncorhynchus mykiss) living in seawater and freshwater. Food Chem. 2004, 86, 55–59. [Google Scholar] [CrossRef]
- Xu, J.; Yan, B.; Teng, Y.; Lou, G.; Lu, Z. Analysis of nutrient composition and fatty acid profiles of Japanese sea bass Lateolabrax japonicus (Cuvier) reared in seawater and freshwater. J. Food Compos. Anal. 2010, 23, 401–405. [Google Scholar] [CrossRef]
- Hunt, A.Ö.; Özkan, F.; Engin, K.; Tekelioğlu, N. The effects of freshwater rearing on the whole body and muscle tissue fatty acid profile of the European sea bass (Dicentrarchus labrax). Aquac. Int. 2010, 19, 51–61. [Google Scholar] [CrossRef]
- Martínez-Álvarez, R.M.; Sanz, A.; García-Gallego, M.; Domezain, A.; Domezain, J.; Carmona, R.; Ostos-Garrido, M.D.V.; Morales, A.E. Adaptive branchial mechanisms in the sturgeon Acipenser naccarii during acclimation to saltwater. Comp. Biochem. Physiol. Part A Mol. Integr. Physiol. 2005, 141, 183–190. [Google Scholar] [CrossRef]
- Dantagnan, P.; Bórquez, A.S.; Valdebenito, I.N.; Salgado, I.A.; Serrano, E.A.; Izquierdo, M. Lipid and fatty acid composition during embryo and larval development of puye Galaxias maculatus Jenyns, 1842, obtained from estuarine, freshwater and cultured populations. J. Fish Biol. 2007, 70, 770–781. [Google Scholar] [CrossRef]
- Navarro, J.; Perez, M.; Acosta, N.; Viciano, E.; Varó, I.; Urena, R.; Torreblanca, A.; Rodriguez, C. The fatty acid profiles of polar and neutral lipids from enterocytes and hepatocytes of gilthead seabream (Sparus aurata) cultured at two salinities: A chemometrics approach. Comp. Biochem. Physiol. Part A Mol. Integr. Physiol. 2009, 154, S20. [Google Scholar] [CrossRef]
- Fonseca-Madrigal, J.; Pineda-Delgado, D.; Martínez-Palacios, C.; Rodríguez, C.; Tocher, D.R. Effect of salinity on the biosynthesis of n-3 long-chain polyunsaturated fatty acids in silverside Chirostoma estor. Fish Physiol. Biochem. 2012, 38, 1047–1057. [Google Scholar] [CrossRef]
- Tocher, D.R.; Bell, J.; Dick, J.; Henderson, R.; McGhee, F.; Michell, D.; Morris, P.C. Polyunsaturated fatty acid metabolism in Atlantic salmon (Salmo salar) undergoing parr-smolt transformation and the effects of dietary linseed and rapeseed oils. Fish Physiol. Biochem. 2000, 23, 59–73. [Google Scholar] [CrossRef]
- Özogul, Y.; Ozyurt, G.; Ozogul, F.; Kuley, E.; Polat, A. Freshness assessment of European eel (Anguilla anguilla) by sensory, chemical and microbiological methods. Food Chem. 2005, 92, 745–751. [Google Scholar] [CrossRef]
- Shepherd, C.J.; Monroig, Ó.; Tocher, D.R. Future availability of raw materials for salmon feeds and supply chain implications: The case of Scottish farmed salmon. Aquaculture 2017, 467, 49–62. [Google Scholar] [CrossRef]
- Cardoso, C.; Afonso, C.; Bandarra, N.M. Seafood lipids and cardiovascular health. Nutrire 2016, 41, 5. [Google Scholar] [CrossRef]
- Brenna, J.T. Efficiency of conversion of α-linolenic acid to long chain n-3 fatty acids in man. Curr. Opin. Clin. Nutr. Metab. Care 2002, 5, 127–132. [Google Scholar] [CrossRef]
- Luna, S.M.; Capuli, E.E. FishBase. 2019. Available online: https://www.fishbase.de/summary/Lateolabrax-japonicum.html (accessed on 7 December 2019).
- Fisheries and Fisheries Administration Bureau of the Ministry of Agriculture and Rural Affairs. China Fishery Statistical Yearbook; China Agriculture Press: Beijing, China, 2018.
- Geay, F.; Culi, E.S.I.; Corporeau, C.; Boudry, P.; Dreano, Y.; Corcos, L.; Bodin, N.; Vandeputte, M.; Zambonino-Infante, J.-L.; Mazurais, D.; et al. Regulation of FADS2 expression and activity in European sea bass (Dicentrarchus labrax L.) fed a vegetable diet. Comp. Biochem. Physiol. Part B Biochem. Mol. Biol. 2010, 156, 237–243. [Google Scholar] [CrossRef]
- Xu, H.; Dong, X.; Ai, Q.; Mai, K.; Xu, W.; Zhang, Y.; Zuo, R. Regulation of Tissue LC-PUFA Contents, Δ6 Fatty Acyl Desaturase (FADS2) Gene Expression and the Methylation of the Putative FADS2 Gene Promoter by Different Dietary Fatty Acid Profiles in Japanese Seabass (Lateolabrax japonicus). PLoS ONE 2014, 9, e87726. [Google Scholar] [CrossRef]
- Tocher, D.R. Omega-3 long-chain polyunsaturated fatty acids and aquaculture in perspective. Aquaculture 2015, 449, 94–107. [Google Scholar] [CrossRef]
- Morais, S.; Monroig, Ó.; Zheng, X.; Leaver, M.J.; Tocher, D.R. Highly Unsaturated Fatty Acid Synthesis in Atlantic Salmon: Characterization of ELOVL5- and ELOVL2-like Elongases. Mar. Biotechnol. 2009, 11, 627–639. [Google Scholar] [CrossRef]
- Fonseca-Madrigal, J.; Bell, J.G.; Tocher, D.R. Nutritional and environmental regulation of the synthesis of highly unsaturated fatty acids and of fatty-acid oxidation in Atlantic salmon (Salmo salar L.) enterocytes and hepatocytes. Fish Physiol. Biochem. 2006, 32, 317–328. [Google Scholar] [CrossRef]
- Vagner, M.; Santigosa, E. Characterization and modulation of gene expression and enzymatic activity of delta-6 desaturase in teleosts: A review. Aquaculture 2011, 315, 131–143. [Google Scholar] [CrossRef]
- Teoh, C.-Y.; Ng, W.-K. The implications of substituting dietary fish oil with vegetable oils on the growth performance, fillet fatty acid profile and modulation of the fatty acid elongase, desaturase and oxidation activities of red hybrid tilapia, Oreochromis sp. Aquaculture 2016, 465, 311–322. [Google Scholar] [CrossRef]
- Dong, X.; Tan, P.; Cai, Z.; Xu, H.; Li, J.; Ren, W.; Xu, H.; Zuo, R.; Zhou, J.; Mai, K.; et al. Regulation of FADS2 transcription by SREBP-1 and PPAR-α influences LC-PUFA biosynthesis in fish. Sci. Rep. 2017, 7, 40024. [Google Scholar] [CrossRef] [PubMed]
- Zhu, K.-C.; Song, L.; Zhao, C.-P.; Guo, H.-Y.; Zhang, N.; Guo, L.; Liu, B.-S.; Jiang, S.-G.; Zhang, D.-C. The Transcriptional Factor PPARαb Positively Regulates Elovl5 Elongase in Golden Pompano Trachinotus ovatus (Linnaeus 1758). Front. Physiol. 2018, 9, 1340. [Google Scholar] [CrossRef] [PubMed]
- Zheng, X.; Torstensen, B.E.; Tocher, D.R.; Dick, J.R.; Henderson, R.J.; Bell, J.G. Environmental and dietary influences on highly unsaturated fatty acid biosynthesis and expression of fatty acyl desaturase and elongase genes in liver of Atlantic salmon (Salmo salar). Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2005, 1734, 13–24. [Google Scholar] [CrossRef]
- Morán, P.; Marco-Rius, F.; Megías, M.; Covelo-Soto, L.; Pérez-Figueroa, A. Environmental induced methylation changes associated with seawater adaptation in brown trout. Aquaculture 2013, 392, 77–83. [Google Scholar] [CrossRef]
- Kong, N.; Liu, X.; Li, J.; Mu, W.; Lian, J.; Xue, Y. Effects of temperature and salinity on survival, growth and DNA methylation of juvenile Pacific abalone, Haliotis discus hannai Ino. Chin. J. Oceanol. Limnol. 2017, 337, 1248–1258. [Google Scholar] [CrossRef]
- Strömqvist, M.; Tooke, N.; Brunström, B. DNA methylation levels in the 5′ flanking region of the vitellogenin I gene in liver and brain of adult zebrafish (Danio rerio)—Sex and tissue differences and effects of 17α-ethinylestradiol exposure. Aquat. Toxicol. 2010, 98, 275–281. [Google Scholar] [CrossRef]
- Li, S.; He, F.; Wen, H.-S.; Li, J.; Si, Y.; Liu, M.; He, H.; Huang, Z. Analysis of DNA methylation level by methylation-sensitive amplification polymorphism in half smooth tongue sole (Cynoglossus semilaevis) subjected to salinity stress. J. Ocean Univ. China 2016, 16, 269–278. [Google Scholar] [CrossRef]
- Dong, X.; Wu, J.; Shen, Y.; Chen, J.Y.; Miao, S.Y.; Zhang, X.J.; Sun, L. Effects of different arginine/lysine level on growth performance, body composition and digestive enzyme activity of Macrobrachium rosenbergii. Aquac. Nutr. 2018, 24, 1101–1111. [Google Scholar] [CrossRef]
- Association of Official Agricultural Chemists. Official Methods of Analysis, 16th ed.; Association of Official Analytical Chemists: Arlington, VA, USA, 1995. [Google Scholar]
- Dong, X.; Xue, W.; Hua, J.; Hang, Y.; Sun, L.; Miao, S.; Wei, W.; Wu, X.; Du, X. Effects of dietary betaine in allogynogenetic gibel carp (Carassius auratus gibelio): Enhanced growth, reduced lipid deposition and depressed lipogenic gene expression. Aquac. Res. 2018, 49, 1967–1972. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Li, L.-C.; Dahiya, R. MethPrimer: Designing primers for methylation PCRs. Bioinformatics 2002, 18, 1427–1431. [Google Scholar] [CrossRef] [PubMed]
- Jana, S.N.; Garg, S.K.; Patra, B.C. Effect of inland water salinity on growth performance and nutritional physiology in growing milkfish, Chanos chanos (Forsskal): Field and laboratory studies. J. Appl. Ichthyol. 2006, 22, 25–34. [Google Scholar] [CrossRef]
- Jiang, Z.; Liu, G.; Jin, B. Effects of salinity on feeding and growth of juvenile red drum Sciaenops ocellatus. J. Dalian Fish. Univ. 2005, 20, 91–94. [Google Scholar]
- Krogdahl, Å.; Sundby, A.; Olli, J.J. Atlantic salmon (Salmo salar) and rainbow trout (Oncorhynchus mykiss) digest and metabolize nutrients differently. Effects of water salinity and dietary starch level. Aquaculture 2004, 229, 335–360. [Google Scholar] [CrossRef]
- Li, Y.; Hu, C.-B.; Zheng, Y.-J.; Xia, X.-A.; Xu, W.-J.; Wang, S.-Q.; Chen, W.-Z.; Sun, Z.-W.; Huang, J.-H. The effects of dietary fatty acids on liver fatty acid composition and Δ6-desaturase expression differ with ambient salinities in Siganus canaliculatus. Comp. Biochem. Physiol. Part B Biochem. Mol. Biol. 2008, 151, 183–190. [Google Scholar] [CrossRef]
- Al-Lawati, A.; Al-Bahry, S.; Victor, R.; Yaish, M.; Yaish, M.W. Salt stress alters DNA methylation levels in alfalfa (Medicago spp). Genet. Mol. Res. 2016, 15. [Google Scholar] [CrossRef]
- Wang, B.; Fu, R.; Zhang, M.; Ding, Z.; Chang, L.; Zhu, X.; Wang, Y.; Fan, B.; Ye, W.-W.; Yuan, Y. Analysis of methylation-sensitive amplified polymorphism in different cotton accessions under salt stress based on capillary electrophoresis. Genes Genom. 2015, 37, 713–724. [Google Scholar] [CrossRef]
- Reik, W.; Dean, W.; Walter, J. Epigenetic Reprogramming in Mammalian Development. Science 2001, 293, 1089–1093. [Google Scholar] [CrossRef]



| Biochemical Composition | Content (%) |
|---|---|
| Moisture | 8.8 |
| Crude protein | 46.2 |
| Crude lipid | 10.5 |
| Ash | 9.5 |
| Primers * | Sequences (5′–3′) | Purpose |
|---|---|---|
| FADS2-F | CATCACGCCAAACCCAACATC | qRT-PCR |
| FADS2-R | AGACATAGACCAAGCCATATCCAC | qRT-PCR |
| Elovl5-F | GCTGAATATCTGGTGGTTCGTTATG | qRT-PCR |
| Elovl5-R | GGCTGGGATGGCTGAGAGG | qRT-PCR |
| β-actin-F | CAACTGGGATGACATGGAGAAG | qRT-PCR |
| β-actin-R | TTGGCTTTGGGGTTCAGG | qRT-PCR |
| SP1 | AGCCGGATGCGGATGGAGAGAT | Promoter cloning |
| SP2 | CCGTGGGCTGCCTACCTTGAGA | Promoter cloning |
| AP1 | GTAATACGACTCACTATAGGGC | Promoter cloning |
| AP2 | TCGACGGCCCGGGCTGGTAGCT | Promoter cloning |
| F-BF | GATTAGTATTTGGATATTGTAAGTTTTT | FADS2 BSP |
| F-BR | ACTTAAAAAAATTACACCATCTTACC | FADS2 BSP |
| E-BF | GGGAAAGTAGAGATAGTTTTGATGT | Elovl5 BSP |
| E-BR | ACTACCTACCTTAAAAAACRAAAAAAC | Elovl5 BSP |
| Composition | Freshwater | Seawater |
|---|---|---|
| Muscle lipid | 2.0 ± 0.1 b | 2.8 ± 0.1 a |
| Muscle protein | 21.0 ± 0.1 b | 21.6 ± 0.2 a |
| Muscle moisture | 77.8 ± 0.2 a | 76.4 ± 0.3 b |
| Fatty Acid | FP Group | MC Group |
|---|---|---|
| C14:0 | 16.2 ± 7.7 b | 112.9 ± 20.6 a |
| C16:0 | 714.1 ± 152.1 b | 1269.9 ± 322.9 a |
| C18:0 | 247.1 ± 37.5 b | 341.4 ± 33.3 a |
| ∑SFA 2 | 977.4 ± 188.0 b | 1724.2 ± 149.4 a |
| C16:1 | 102.4 ± 34.4 b | 296.3 ± 24.5 a |
| C18:1n-9 | 98.7 ± 11.3 b | 187.9 ± 25.8 a |
| ∑MUFA 3 | 201.1 ± 93.6 b | 484.2 ± 97.1 a |
| C18:2n-6 | 981.6 ± 179.2 b | 1312.5 ± 169.2 a |
| C18:3n-6 | 30.1 ± 1.4 a | 25.8 ± 0.2 b |
| C20:4n-6 | 92.4 ± 9.9 b | 167.1 ± 50.3 a |
| ∑n-6 PUFA 4 | 1104.2 ± 182.2 b | 1505.4 ± 151.7 a |
| C18:3n-3 | 102.8 ± 12.7 b | 181.2 ± 27.4 a |
| C20:5n-3 | 113.3 ± 14.7 b | 408.4 ± 70.0 a |
| C22:6n-3 | 390.8 ± 54.4 b | 1294.9 ± 292.1 a |
| ∑n-3 PUFA 5 | 607.0 ± 67.5 b | 1884.5 ± 332.3 a |
| ∑n-3/∑n-6 PUFA | 0.56 ± 0.08 b | 1.26 ± 0.25 a |
| ∑n-3 LC-PUFA 6 | 504.1 ± 68.3 b | 1703.3 ± 320.7 a |
| Fatty Acid | FP Group | MC Group |
|---|---|---|
| C14:0 | 1205.9 ± 216.1 a | 902.7 ± 127.5 b |
| C16:0 | 11,957.4 ± 1310.3 a | 9799.0 ± 1437.6 b |
| C18:0 | 2987.9 ± 683.0 a | 1663.1 ± 104.5 b |
| ∑SFA 2 | 16,151.2 ± 992.2 a | 12,364.7 ± 340.9 b |
| C16:1 | 4296.3 ± 586.1 | 3974.3 ± 540.4 |
| C18:1n-9 | 1490.7 ± 111.3 | 1570.8 ± 272.6 |
| ∑MUFA 3 | 5787.0 ± 664.0 | 5555.0 ± 843.6 |
| C18:2n-6 | 7066.9 ± 949.5 a | 2271.2 ± 254.3 b |
| C18:3n-6 | 1211.9 ± 191.6 a | 896.7 ± 184.1 b |
| C20:4n-6 | 605.9 ± 107.4 | 666.5 ± 110.5 |
| ∑n-6 PUFA 4 | 8884.7 ± 994.2 a | 3834.4 ± 263.0 b |
| C18:3n-3 | 1217.2 ± 178.0 a | 741.0 ± 120.9 b |
| C20:5n-3 | 410.5 ± 66.6 b | 986.3 ± 173.5 a |
| C22:6n-3 | 1629.9 ± 226.1 b | 6386.0 ± 705.2 a |
| ∑n-3 PUFA 5 | 3257.5 ± 310.0 b | 8113.4 ± 740.8 a |
| ∑n-3/∑n-6 PUFA | 0.4 ± 0.04 b | 2.1 ± 0.1 a |
| ∑n-3 LC-PUFA 6 | 2040.4 ± 253.1 b | 7372.3 ± 738.7 a |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dong, X.; Wang, J.; Ji, P.; Sun, L.; Miao, S.; Lei, Y.; Du, X. Seawater Culture Increases Omega-3 Long-Chain Polyunsaturated Fatty Acids (N-3 LC-PUFA) Levels in Japanese Sea Bass (Lateolabrax japonicus), Probably by Upregulating Elovl5. Animals 2020, 10, 1681. https://doi.org/10.3390/ani10091681
Dong X, Wang J, Ji P, Sun L, Miao S, Lei Y, Du X. Seawater Culture Increases Omega-3 Long-Chain Polyunsaturated Fatty Acids (N-3 LC-PUFA) Levels in Japanese Sea Bass (Lateolabrax japonicus), Probably by Upregulating Elovl5. Animals. 2020; 10(9):1681. https://doi.org/10.3390/ani10091681
Chicago/Turabian StyleDong, Xiaojing, Jianqiao Wang, Peng Ji, Longsheng Sun, Shuyan Miao, Yanju Lei, and Xuedi Du. 2020. "Seawater Culture Increases Omega-3 Long-Chain Polyunsaturated Fatty Acids (N-3 LC-PUFA) Levels in Japanese Sea Bass (Lateolabrax japonicus), Probably by Upregulating Elovl5" Animals 10, no. 9: 1681. https://doi.org/10.3390/ani10091681
APA StyleDong, X., Wang, J., Ji, P., Sun, L., Miao, S., Lei, Y., & Du, X. (2020). Seawater Culture Increases Omega-3 Long-Chain Polyunsaturated Fatty Acids (N-3 LC-PUFA) Levels in Japanese Sea Bass (Lateolabrax japonicus), Probably by Upregulating Elovl5. Animals, 10(9), 1681. https://doi.org/10.3390/ani10091681






