Reducing versus Embracing Variation as Strategies for Reproducibility: The Microbiome of Laboratory Mice
Abstract
Simple Summary
Abstract
1. Introduction
2. Conceptual Framework: Defining and Understanding Reproducibility
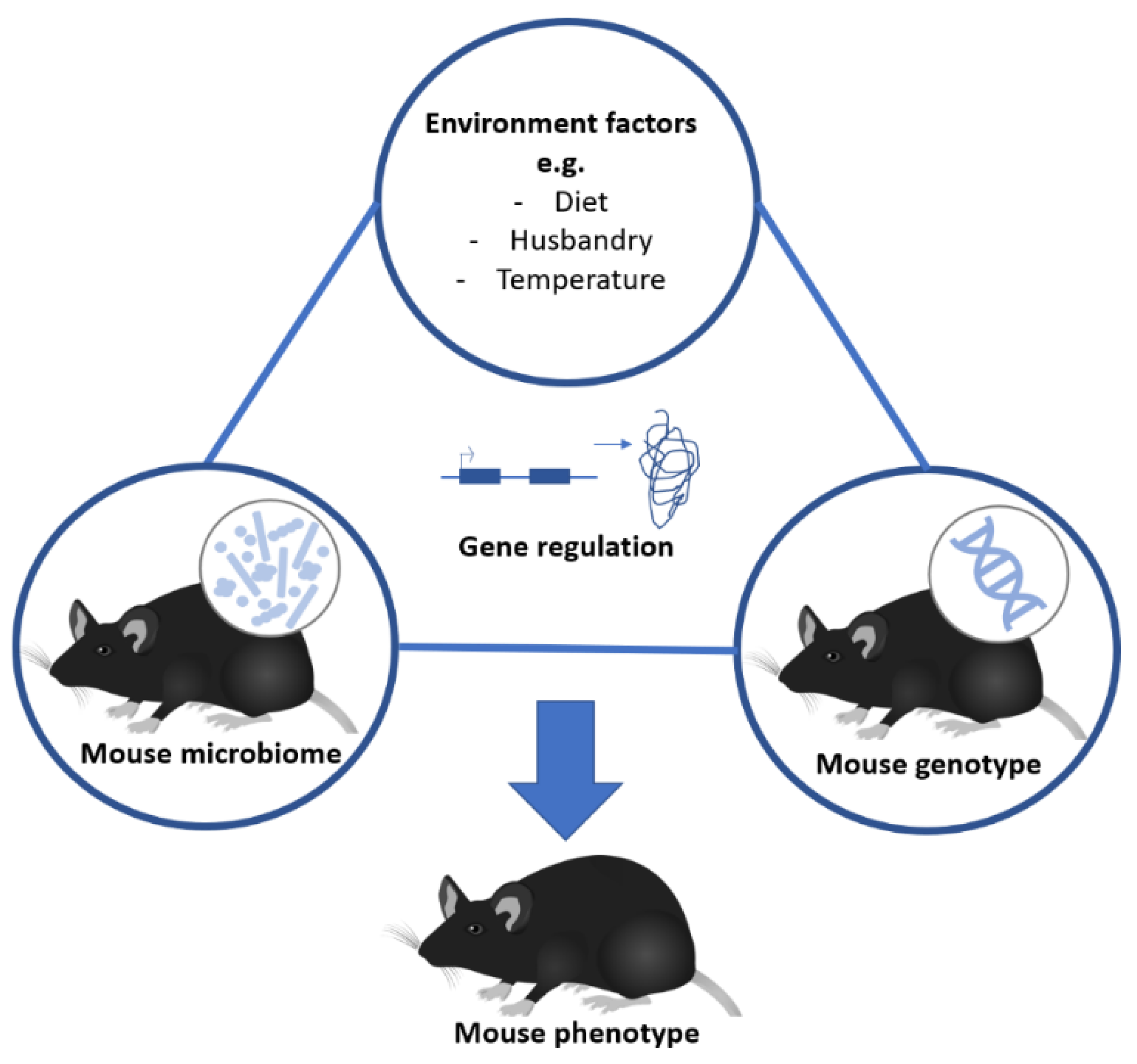
3. Phenotypic Variation: Genotype-Environment/Microbiome
4. The Microbiome Explaining Irreproducibility of Results
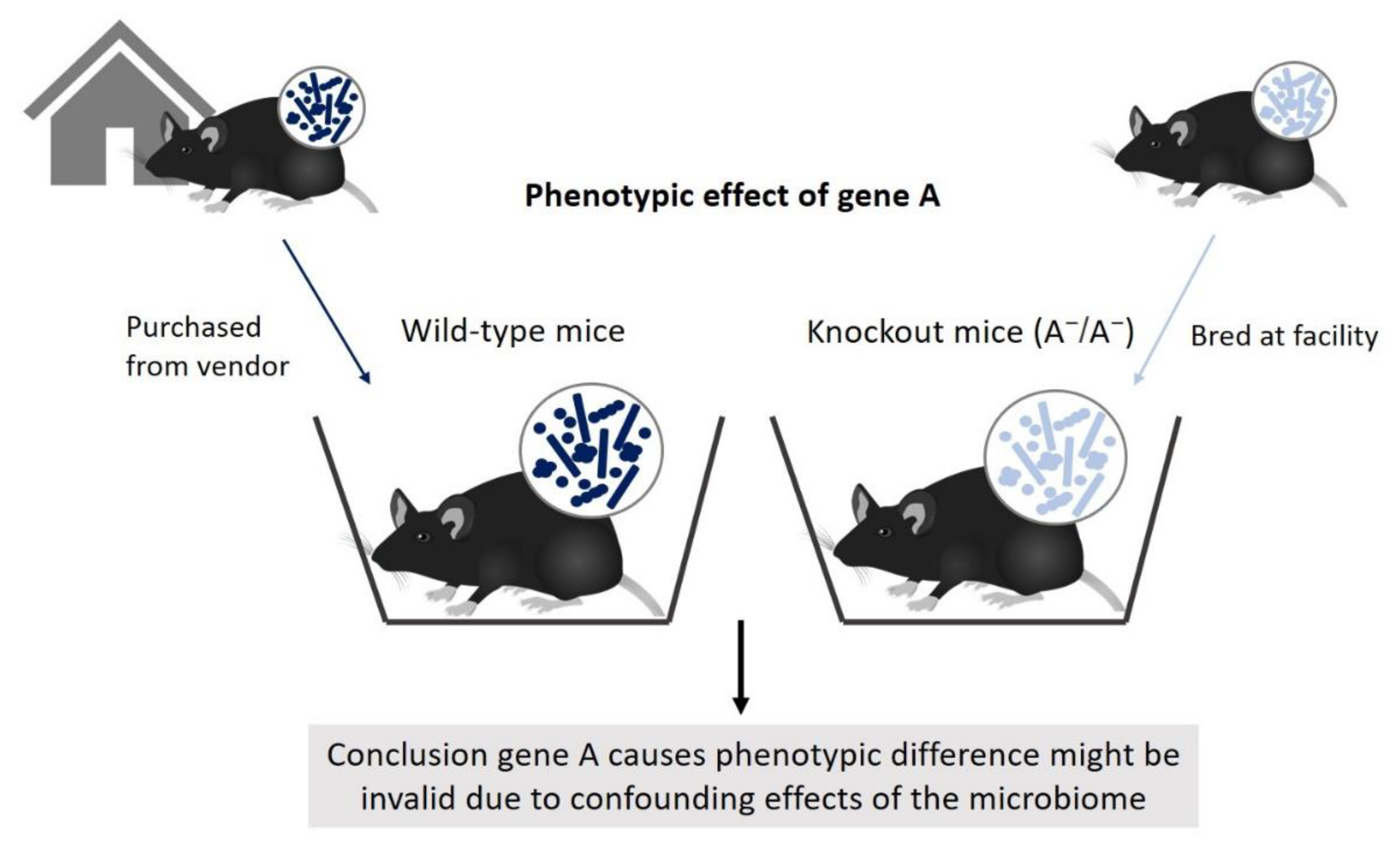
4.1. The Microbiome as a Confounding Factor
4.2. The Result Is not Generalizable to Mice Harboring a Different Microbiome
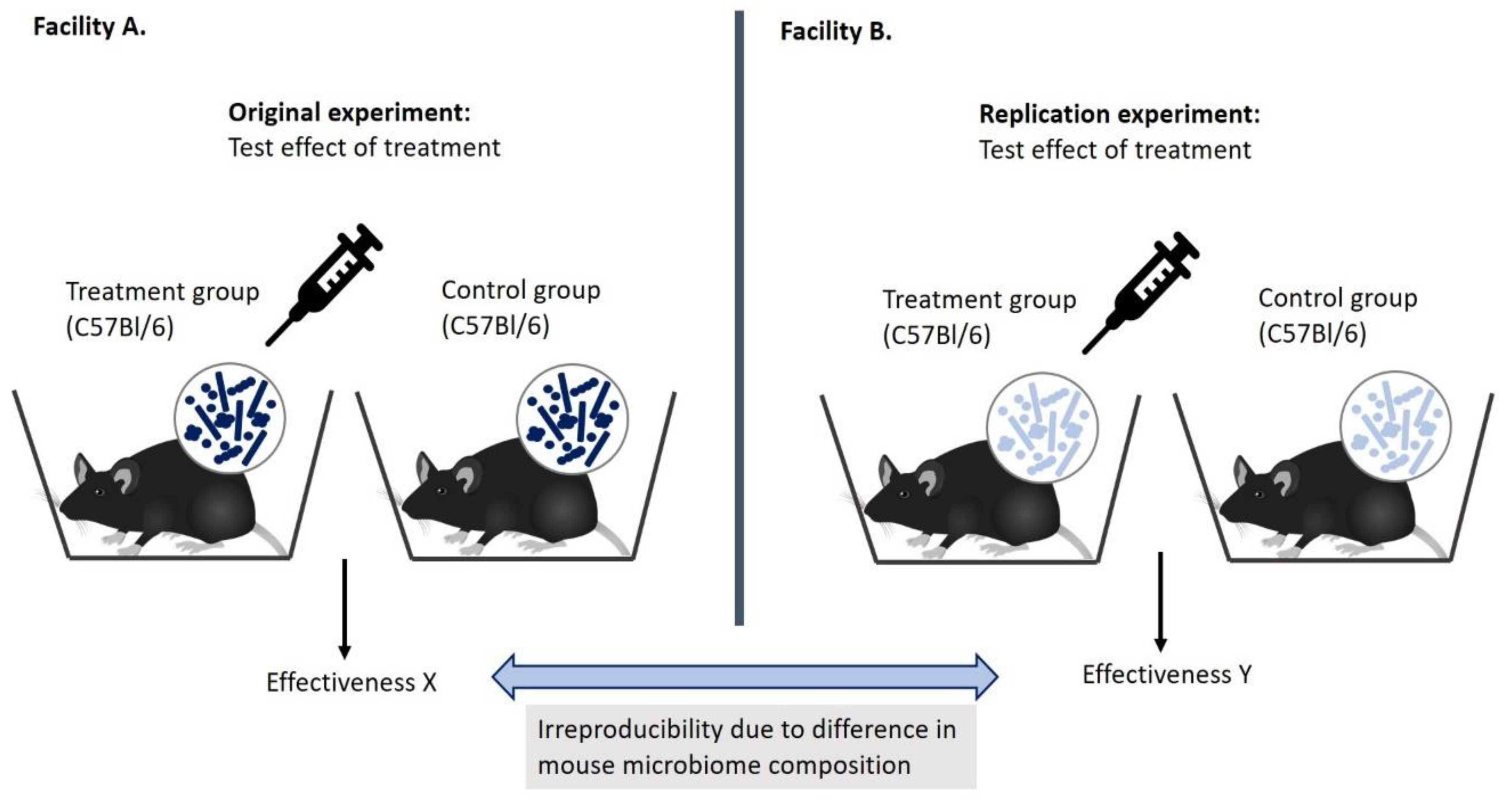
5. The Microbiome and ‘Failed’ Replications: What Does It Mean?
6. Improving Internal Validity
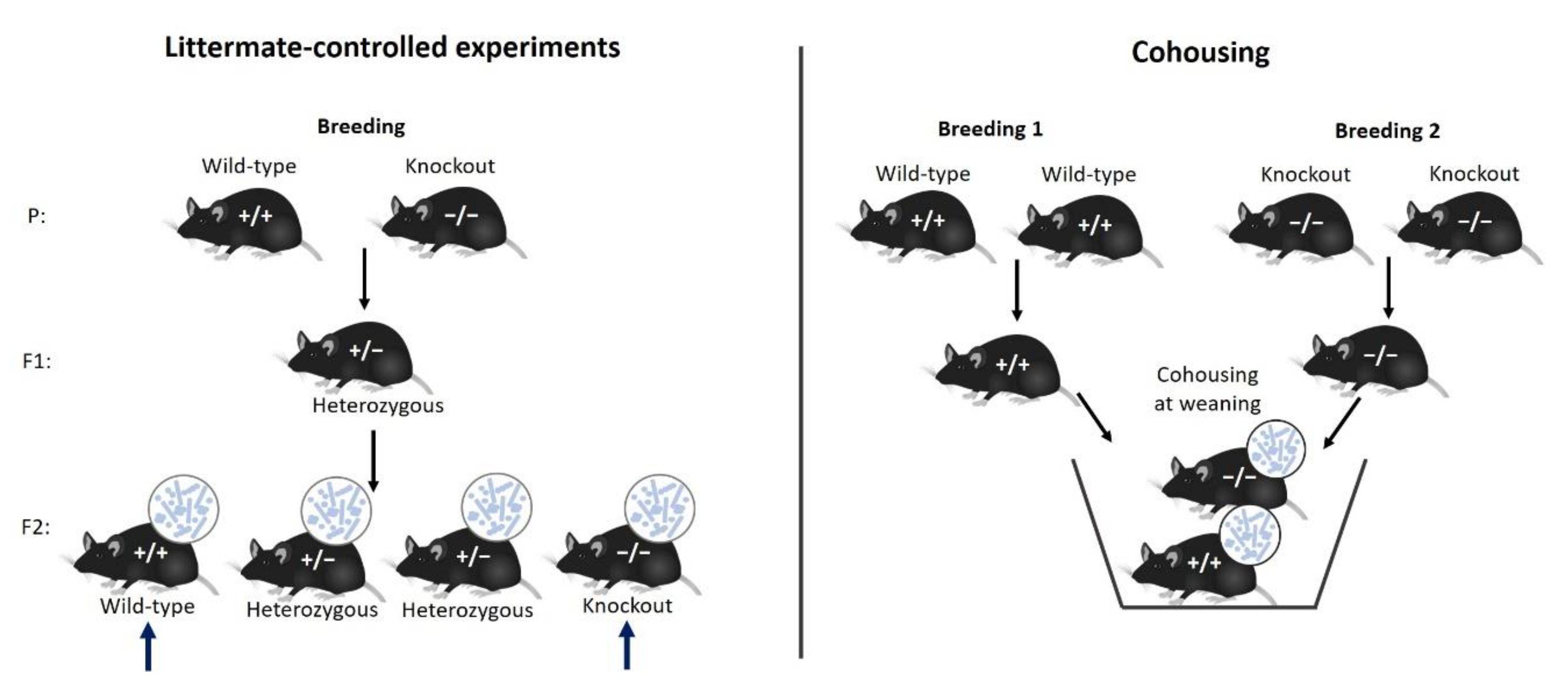
6.1. Cohousing and Littermate Controls: Standardization within Experiments
6.2. Defined Microbial Consortia: Standardization within and across Experiments
7. Are Animal Experiments Over-Standardized?
7.1. The Standardization ‘Fallacy’
7.2. Standardized Clean Laboratories to Model the Dirty World
8. The Future: Reducing Versus Embracing Variation?
9. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- KNAW. Replication Studies—Improving Reproducibility in the Empirical Sciences; KNAW: Amsterdam, The Netherlands, 2018; p. 66. [Google Scholar]
- Baker, M. 1500 scientists lift the lid on reproducibility. Nature 2016, 533, 452–454. [Google Scholar] [CrossRef] [PubMed]
- Romero, F. Philosophy of science and the replicability crisis. Philos. Compass 2019, 14, e12633. [Google Scholar] [CrossRef]
- Kafkafi, N.; Agassi, J.; Chesler, E.J.; Crabbe, J.C.; Crusio, W.E.; Eilam, D.; Gerlai, R.; Golani, I.; Gomez-Marin, A.; Heller, R.; et al. Reproducibility and replicability of rodent phenotyping in preclinical studies. Neurosci. Biobehav. Rev. 2018, 87, 218–232. [Google Scholar] [CrossRef] [PubMed]
- Cheleuitte-Nieves, C.; Lipman, N.S. Improving Replicability, Reproducibility, and Reliability in Preclinical Research: A Shared Responsibility. ILAR J. 2020. [Google Scholar] [CrossRef] [PubMed]
- Begley, C.G.; Ioannidis, J.P. Reproducibility in science: Improving the standard for basic and preclinical research. Circ. Res. 2015, 116, 116–126. [Google Scholar] [CrossRef] [PubMed]
- Freedman, L.P.; Cockburn, I.M.; Simcoe, T.S. The Economics of Reproducibility in Preclinical Research. PLoS Biol. 2015, 13, e1002165. [Google Scholar] [CrossRef] [PubMed]
- Russell, W.M.S.; Burch, R.L. The Principles of Humane Experimental Technique; Methuen: London, UK, 1959. [Google Scholar]
- Osborne, N.; Avey, M.T.; Anestidou, L.; Ritskes-Hoitinga, M.; Griffin, G. Improving animal research reporting standards: HARRP, the first step of a unified approach by ICLAS to improve animal research reporting standards worldwide. EMBO Rep. 2018, 19. [Google Scholar] [CrossRef]
- Ritskes-Hoitinga, M.; Wever, K. Improving the conduct, reporting, and appraisal of animal research. BMJ 2018, 360, j4935. [Google Scholar] [CrossRef]
- Svendsen, O.; Hansen, A.K. Biological variation, reproducibility, and predictability of experimental research in animals. In Trends in Contrast Media; Springer: Berlin/Heidelberg, Germany, 1999; pp. 31–41. [Google Scholar]
- Richter, S.H.; Garner, J.P.; Würbel, H. Environmental standardization: Cure or cause of poor reproducibility in animal experiments? Nat. Methods 2009, 6, 257–261. [Google Scholar] [CrossRef]
- Champy, M.F.; Selloum, M.; Piard, L.; Zeitler, V.; Caradec, C.; Chambon, P.; Auwerx, J. Mouse functional genomics requires standardization of mouse handling and housing conditions. Mamm. Genome 2004, 15, 768–783. [Google Scholar] [CrossRef]
- Voelkl, B.; Altman, N.S.; Forsman, A.; Forstmeier, W.; Gurevitch, J.; Jaric, I.; Karp, N.A.; Kas, M.J.; Schielzeth, H.; Van de Casteele, T.; et al. Reproducibility of animal research in light of biological variation. Nat. Rev. Neurosci. 2020, 21, 384–393. [Google Scholar] [CrossRef] [PubMed]
- Alegre, M.-L. Mouse microbiomes: Overlooked culprits of experimental variability. Genome Biol. 2019, 20, 1–3. [Google Scholar] [CrossRef] [PubMed]
- Stappenbeck, T.S.; Virgin, H.W. Accounting for reciprocal host–microbiome interactions in experimental science. Nature 2016, 534, 191–199. [Google Scholar] [CrossRef] [PubMed]
- Dirnagl, U. Rethinking research reproducibility. EMBO J. 2019, 38, e101117. [Google Scholar] [CrossRef]
- Macleod, M.; Mohan, S. Reproducibility and Rigor in Animal-Based Research. ILAR J. 2019, 60, 17–23. [Google Scholar] [CrossRef]
- Goodman, S.N.; Fanelli, D.; Ioannidis, J.P. What does research reproducibility mean? Sci. Transl. Med. 2016, 8, 341ps12. [Google Scholar] [CrossRef]
- Leonelli, S. Rethinking Reproducibility as a Criterion for Research Quality. In Including a Symposium on Mary Morgan: Curiosity, Imagination, and Surprise; Emerald Publishing Limited: Bingley, UK, 2018; Volume 36B, pp. 129–146. [Google Scholar]
- Voelkl, B.; Würbel, H. Reproducibility Crisis: Are We Ignoring Reaction Norms? Trends Pharmacol. Sci. 2016, 37, 509–510. [Google Scholar] [CrossRef]
- Percie du Sert, N.; Ahluwalia, A.; Alam, S.; Avey, M.T.; Baker, M.; Browne, W.J.; Clark, A.; Cuthill, I.C.; Dirnagl, U.; Emerson, M.; et al. Reporting animal research: Explanation and elaboration for the ARRIVE guidelines 2.0. PLoS Biol. 2020, 18, e3000411. [Google Scholar] [CrossRef]
- Earp, B.D.; Trafimow, D. Replication, falsification, and the crisis of confidence in social psychology. Front. Psychol. 2015, 6. [Google Scholar] [CrossRef]
- Meehl, P.E. Why Summaries of Research on Psychological Theories are Often Uninterpretable. Psychol. Rep. 1990, 66, 195–244. [Google Scholar] [CrossRef]
- Byers, D. Components of phenotypic variance. Nat. Educ. 2008, 1, 161. [Google Scholar]
- Via, S.; Lande, R. Genotype-Environment Interaction and the Evolution of Phenotypic Plasticity. Evolution 1985, 39, 505–522. [Google Scholar] [CrossRef] [PubMed]
- Osbelt, L.; Thiemann, S.; Smit, N.; Lesker, T.R.; Schröter, M.; Gálvez, E.J.C.; Schmidt-Hohagen, K.; Pils, M.C.; Mühlen, S.; Dersch, P.; et al. Variations in microbiota composition of laboratory mice influence Citrobacter rodentium infection via variable short-chain fatty acid production. PLoS Pathog. 2020, 16, e1008448. [Google Scholar] [CrossRef] [PubMed]
- Velazquez, E.M.; Nguyen, H.; Heasley, K.T.; Saechao, C.H.; Gil, L.M.; Rogers, A.W.L.; Miller, B.M.; Rolston, M.R.; Lopez, C.A.; Litvak, Y.; et al. Endogenous Enterobacteriaceae underlie variation in susceptibility to Salmonella infection. Nat. Microbiol. 2019, 4, 1057–1064. [Google Scholar] [CrossRef] [PubMed]
- Marchesi, J.R.; Ravel, J. The vocabulary of microbiome research: A proposal. Microbiome 2015, 3, 31. [Google Scholar] [CrossRef] [PubMed]
- Lloyd-Price, J.; Mahurkar, A.; Rahnavard, G.; Crabtree, J.; Orvis, J.; Hall, A.B.; Brady, A.; Creasy, H.H.; McCracken, C.; Giglio, M.G.; et al. Strains, functions and dynamics in the expanded Human Microbiome Project. Nature 2017, 550, 61–66. [Google Scholar] [CrossRef]
- Snijders, A.M.; Langley, S.A.; Kim, Y.-M.; Brislawn, C.J.; Noecker, C.; Zink, E.M.; Fansler, S.J.; Casey, C.P.; Miller, D.R.; Huang, Y.; et al. Influence of early life exposure, host genetics and diet on the mouse gut microbiome and metabolome. Nat. Microbiol. 2016, 2, 1–8. [Google Scholar] [CrossRef]
- Marchesi, J.R.; Adams, D.H.; Fava, F.; Hermes, G.D.; Hirschfield, G.M.; Hold, G.; Quraishi, M.N.; Kinross, J.; Smidt, H.; Tuohy, K.M.; et al. The gut microbiota and host health: A new clinical frontier. Gut 2016, 65, 330–339. [Google Scholar] [CrossRef]
- Borges-Canha, M.; Portela-Cidade, J.P.; Dinis-Ribeiro, M.; Leite-Moreira, A.F.; Pimentel-Nunes, P. Role of colonic microbiota in colorectal carcinogenesis: A systematic review. Rev. Esp. Enferm. Dig. 2015, 107, 659–671. [Google Scholar] [CrossRef]
- Iglesias-Vázquez, L.; Van Ginkel Riba, G.; Arija, V.; Canals, J. Composition of Gut Microbiota in Children with Autism Spectrum Disorder: A Systematic Review and Meta-Analysis. Nutrients 2020, 12, 792. [Google Scholar] [CrossRef]
- Jamshidi, P.; Hasanzadeh, S.; Tahvildari, A.; Farsi, Y.; Arbabi, M.; Mota, J.F.; Sechi, L.A.; Nasiri, M.J. Is there any association between gut microbiota and type 1 diabetes? A systematic review. Gut Pathog. 2019, 11, 49. [Google Scholar] [CrossRef] [PubMed]
- Hooper, L.V.; Gordon, J.I. Commensal host-bacterial relationships in the gut. Science 2001, 292, 1115–1118. [Google Scholar] [CrossRef] [PubMed]
- Glowacki, R.W.P.; Martens, E.C. In sickness and health: Effects of gut microbial metabolites on human physiology. PLoS Pathog. 2020, 16, e1008370. [Google Scholar] [CrossRef]
- Franklin, C.L.; Ericsson, A.C. Microbiota and reproducibility of rodent models. Lab Anim. 2017, 46, 114–122. [Google Scholar] [CrossRef]
- Rausch, P.; Basic, M.; Batra, A.; Bischoff, S.C.; Blaut, M.; Clavel, T.; Gläsner, J.; Gopalakrishnan, S.; Grassl, G.A.; Günther, C.; et al. Analysis of factors contributing to variation in the C57BL/6J fecal microbiota across German animal facilities. Int. J. Med. Microbiol. 2016, 306, 343–355. [Google Scholar] [CrossRef] [PubMed]
- Hufeldt, M.R.; Nielsen, D.S.; Vogensen, F.K.; Midtvedt, T.; Hansen, A.K. Variation in the gut microbiota of laboratory mice is related to both genetic and environmental factors. Comp. Med. 2010, 60, 336–347. [Google Scholar]
- Adair, K.L.; Douglas, A.E. Making a microbiome: The many determinants of host-associated microbial community composition. Curr. Opin. Microbiol. 2017, 35, 23–29. [Google Scholar] [CrossRef]
- McCafferty, J.; Mühlbauer, M.; Gharaibeh, R.Z.; Arthur, J.C.; Perez-Chanona, E.; Sha, W.; Jobin, C.; Fodor, A.A. Stochastic changes over time and not founder effects drive cage effects in microbial community assembly in a mouse model. ISME J. 2013, 7, 2116–2125. [Google Scholar] [CrossRef]
- Ericsson, A.C.; Davis, J.W.; Spollen, W.; Bivens, N.; Givan, S.; Hagan, C.E.; McIntosh, M.; Franklin, C.L. Effects of Vendor and Genetic Background on the Composition of the Fecal Microbiota of Inbred Mice. PLoS ONE 2015, 10, e0116704. [Google Scholar] [CrossRef]
- Laukens, D.; Brinkman, B.M.; Raes, J.; De Vos, M.; Vandenabeele, P. Heterogeneity of the gut microbiome in mice: Guidelines for optimizing experimental design. FEMS Microbiol. Rev. 2015, 40, 117–132. [Google Scholar] [CrossRef]
- Oyserman, B.O.; Cordovez, V.; Sarango Flores, S.W.; Nijveen, H.; Medema, M.H.; Raaijmakers, J.M. Extracting the GEMs: Genotype, Environment and Microbiome interactions shaping host phenotypes. bioRxiv 2019, 863399. [Google Scholar] [CrossRef]
- Henry, L.P.; Bruijning, M.; Forsberg, S.K.G.; Ayroles, J.F. Can the microbiome influence host evolutionary trajectories? bioRxiv 2019, 700237. [Google Scholar] [CrossRef]
- Awany, D.; Allali, I.; Dalvie, S.; Hemmings, S.; Mwaikono, K.S.; Thomford, N.E.; Gomez, A.; Mulder, N.; Chimusa, E.R. Host and Microbiome Genome-Wide Association Studies: Current State and Challenges. Front. Genet. 2019, 9. [Google Scholar] [CrossRef] [PubMed]
- Davisson, M.T. Genetic and phenotypic definition of laboratory mice and rats/what constitutes an acceptable genetic-phenotypic definition. In Proceedings of the Microbial and Phenotypic Definition of Rats and Mice: Proceedings of the 1998 US/Japan Conference, Kawasaki, Japan, 23 October 1998; p. 63. [Google Scholar]
- Zutphen, L.F.M.; Baumans, V.; Ohl, F. Standaardisatie van dierproeven. In Handboek Proefdierkunde; Elsevier Gezondheidszorg: Maarssen, The Netherlands, 2009; p. 404. [Google Scholar]
- Yasinki, E. Microbes May Take Some of the Blame for the Reproducibility Crisis. Scientist 2019. Available online: https://www.the-scientist.com/news-opinion/microbes-may-take-some-of-the-blame-for-the-reproducibility-crisis-65707 (accessed on 19 February 2020).
- Hildebrand, F.; Nguyen, T.L.; Brinkman, B.; Yunta, R.G.; Cauwe, B.; Vandenabeele, P.; Liston, A.; Raes, J. Inflammation-associated enterotypes, host genotype, cage and inter-individual effects drive gut microbiota variation in common laboratory mice. Genome Biol. 2013, 14, R4. [Google Scholar] [CrossRef]
- Ivanov, I.I.; Frutos Rde, L.; Manel, N.; Yoshinaga, K.; Rifkin, D.B.; Sartor, R.B.; Finlay, B.B.; Littman, D.R. Specific microbiota direct the differentiation of IL-17-producing T-helper cells in the mucosa of the small intestine. Cell Host Microbe 2008, 4, 337–349. [Google Scholar] [CrossRef] [PubMed]
- Sivan, A.; Corrales, L.; Hubert, N.; Williams, J.B.; Aquino-Michaels, K.; Earley, Z.M.; Benyamin, F.W.; Lei, Y.M.; Jabri, B.; Alegre, M.L.; et al. Commensal Bifidobacterium promotes antitumor immunity and facilitates anti-PD-L1 efficacy. Science 2015, 350, 1084–1089. [Google Scholar] [CrossRef]
- Guthrie, L.; Kelly, L. Bringing microbiome-drug interaction research into the clinic. EBioMedicine 2019, 44, 708–715. [Google Scholar] [CrossRef]
- Servick, K. Of mice and microbes. Science 2016, 353, 741–743. [Google Scholar] [CrossRef]
- O’Connor, A.M.; Sargeant, J.M. Critical Appraisal of Studies Using Laboratory Animal Models. ILAR J. 2014, 55, 405–417. [Google Scholar] [CrossRef]
- Caruso, R.; Ono, M.; Bunker, M.E.; Núñez, G.; Inohara, N. Dynamic and Asymmetric Changes of the Microbial Communities after Cohousing in Laboratory Mice. Cell Rep. 2019, 27, 3401–3412.e3. [Google Scholar] [CrossRef]
- Robertson, S.J.; Lemire, P.; Maughan, H.; Goethel, A.; Turpin, W.; Bedrani, L.; Guttman, D.S.; Croitoru, K.; Girardin, S.E.; Philpott, D.J. Comparison of Co-housing and Littermate Methods for Microbiota Standardization in Mouse Models. Cell Rep. 2019, 27, 1910–1919.e2. [Google Scholar] [CrossRef] [PubMed]
- Hufgard, J.R.; Williams, M.T.; Skelton, M.R.; Grubisha, O.; Ferreira, F.M.; Sanger, H.; Wright, M.E.; Reed-Kessler, T.M.; Rasmussen, K.; Duman, R.S.; et al. Phosphodiesterase-1b (Pde1b) knockout mice are resistant to forced swim and tail suspension induced immobility and show upregulation of Pde10a. Psychopharmacology 2017, 234, 1803–1813. [Google Scholar] [CrossRef] [PubMed]
- Lemire, P.; Robertson, S.J.; Maughan, H.; Tattoli, I.; Streutker, C.J.; Platnich, J.M.; Muruve, D.A.; Philpott, D.J.; Girardin, S.E. The NLR Protein NLRP6 Does Not Impact Gut Microbiota Composition. Cell Rep. 2017, 21, 3653–3661. [Google Scholar] [CrossRef] [PubMed]
- Macpherson, A.J.; McCoy, K.D. Standardised animal models of host microbial mutualism. Mucosal Immunol. 2015, 8, 476–486. [Google Scholar] [CrossRef] [PubMed]
- Eberl, G. Addressing the experimental variability associated with the microbiota. Mucosal Immunol. 2015, 8, 487–490. [Google Scholar] [CrossRef]
- Pang, W.; Stradiotto, D.; Krych, L.; Karlskov-Mortensen, P.; Vogensen, F.K.; Nielsen, D.S.; Fredholm, M.; Hansen, A.K. Selective inbreeding does not increase gut microbiota similarity in BALB/c mice. Lab. Anim. 2012, 46, 335–337. [Google Scholar] [CrossRef]
- National Academies of Sciences, Engineering, and Medicine. Animal Models for Microbiome Research: Advancing Basic and Translational Science: Proceedings of a Workshop; The National Academies Press: Washington, DC, USA, 2018; p. 114. [Google Scholar] [CrossRef]
- Wymore Brand, M.; Wannemuehler, M.J.; Phillips, G.J.; Proctor, A.; Overstreet, A.-M.; Jergens, A.E.; Orcutt, R.P.; Fox, J.G. The Altered Schaedler Flora: Continued Applications of a Defined Murine Microbial Community. ILAR J. 2015, 56, 169–178. [Google Scholar] [CrossRef]
- Uchimura, Y.; Wyss, M.; Brugiroux, S.; Limenitakis, J.P.; Stecher, B.; McCoy, K.D.; Macpherson, A.J. Complete Genome Sequences of 12 Species of Stable Defined Moderately Diverse Mouse Microbiota 2. Genome Announc. 2016, 4. [Google Scholar] [CrossRef]
- Studer, N.; Desharnais, L.; Beutler, M.; Brugiroux, S.; Terrazos, M.A.; Menin, L.; Schürch, C.M.; McCoy, K.D.; Kuehne, S.A.; Minton, N.P.; et al. Functional Intestinal Bile Acid 7α-Dehydroxylation by Clostridium scindens Associated with Protection from Clostridium difficile Infection in a Gnotobiotic Mouse Model. Front. Cell. Infect. Microbiol. 2016, 6. [Google Scholar] [CrossRef]
- Lehmann, F.M.; von Burg, N.; Ivanek, R.; Teufel, C.; Horvath, E.; Peter, A.; Turchinovich, G.; Staehli, D.; Eichlisberger, T.; Gomez de Agüero, M.; et al. Microbiota-induced tissue signals regulate ILC3-mediated antigen presentation. Nat. Commun. 2020, 11, 1–15. [Google Scholar] [CrossRef]
- Van der Staay, F.J.; Arndt, S.S.; Nordquist, R.E. The standardization-generalization dilemma: A way out. Genes Brain Behav. 2010, 9, 849–855. [Google Scholar] [CrossRef] [PubMed]
- Sorge, R.E.; Martin, L.J.; Isbester, K.A.; Sotocinal, S.G.; Rosen, S.; Tuttle, A.H.; Wieskopf, J.S.; Acland, E.L.; Dokova, A.; Kadoura, B.; et al. Olfactory exposure to males, including men, causes stress and related analgesia in rodents. Nat. Methods 2014, 11, 629–632. [Google Scholar] [CrossRef] [PubMed]
- Gouveia, K.; Hurst, J.L. Optimising reliability of mouse performance in behavioural testing: The major role of non-aversive handling. Sci. Rep. 2017, 7, 44999. [Google Scholar] [CrossRef] [PubMed]
- Friese, C. Realizing Potential in Translational Medicine: The Uncanny Emergence of Care as Science. Curr. Anthropol. 2013, 54, S129–S138. [Google Scholar] [CrossRef]
- Würbel, H. Behaviour and the standardization fallacy. Nat. Genet. 2000, 26, 263. [Google Scholar] [CrossRef]
- Voelkl, B.; Vogt, L.; Sena, E.S.; Würbel, H. Reproducibility of preclinical animal research improves with heterogeneity of study samples. PLoS Biol. 2018, 16, e2003693. [Google Scholar] [CrossRef]
- Usui, T.; Macleod, M.R.; McCann, S.K.; Senior, A.M.; Nakagawa, S. Embrace heterogeneity to improve reproducibility: A perspective from meta-analysis of variation in preclinical research. bioRxiv 2020. [Google Scholar] [CrossRef]
- Rosshart, S.P.; Vassallo, B.G.; Angeletti, D.; Hutchinson, D.S.; Morgan, A.P.; Takeda, K.; Hickman, H.D.; McCulloch, J.A.; Badger, J.H.; Ajami, N.J.; et al. Wild Mouse Gut Microbiota Promotes Host Fitness and Improves Disease Resistance. Cell 2017, 171, 1015–1028.e13. [Google Scholar] [CrossRef]
- Abolins, S.; King, E.C.; Lazarou, L.; Weldon, L.; Hughes, L.; Drescher, P.; Raynes, J.G.; Hafalla, J.C.R.; Viney, M.E.; Riley, E.M. The comparative immunology of wild and laboratory mice, Mus musculus domesticus. Nat. Commun. 2017, 8, 1–13. [Google Scholar] [CrossRef]
- Rosshart, S.P.; Herz, J.; Vassallo, B.G.; Hunter, A.; Wall, M.K.; Badger, J.H.; McCulloch, J.A.; Anastasakis, D.G.; Sarshad, A.A.; Leonardi, I.; et al. Laboratory mice born to wild mice have natural microbiota and model human immune responses. Science 2019, 365. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Witjes, V.M.; Boleij, A.; Halffman, W. Reducing versus Embracing Variation as Strategies for Reproducibility: The Microbiome of Laboratory Mice. Animals 2020, 10, 2415. https://doi.org/10.3390/ani10122415
Witjes VM, Boleij A, Halffman W. Reducing versus Embracing Variation as Strategies for Reproducibility: The Microbiome of Laboratory Mice. Animals. 2020; 10(12):2415. https://doi.org/10.3390/ani10122415
Chicago/Turabian StyleWitjes, Vera M., Annemarie Boleij, and Willem Halffman. 2020. "Reducing versus Embracing Variation as Strategies for Reproducibility: The Microbiome of Laboratory Mice" Animals 10, no. 12: 2415. https://doi.org/10.3390/ani10122415
APA StyleWitjes, V. M., Boleij, A., & Halffman, W. (2020). Reducing versus Embracing Variation as Strategies for Reproducibility: The Microbiome of Laboratory Mice. Animals, 10(12), 2415. https://doi.org/10.3390/ani10122415




