Abstract
Background: The emergence of multidrug-resistant bacteria remains poorly understood in the wild ecosystem and at the interface of habitats. Here, we explored the spread of Escherichia coli containing IncI1-ST3 plasmid encoding resistance gene cefotaximase-Munich-1 (blaCTX-M-1) in human-influenced habitats and wild fauna using a genomic approach. Methods. Multilocus sequence typing (MLST), single-nucleotide polymorphism comparison, synteny-based analysis and data mining approaches were used to analyse a dataset of genomes and circularised plasmids. Results. CTX-M-1 E. coli sequence types (STs) were preferentially associated with ecosystems. Few STs were shared by distinct habitats. IncI1-ST3-blaCTX-M-1 plasmids are disseminated among all E. coli phylogroups. The main divergences in plasmids were located in a shuffling zone including blaCTX-M-1 inserted in a conserved site. This insertion hot spot exhibited diverse positions and orientations in a zone-modulating conjugation, and the resulting synteny was associated with geographic and biological sources. Conclusions. The ecological success of IncI1-ST3-blaCTX-M-1 appears less linked to the spread of their bacterial recipients than to their ability to transfer in a broad spectrum of bacterial lineages. This feature is associated with the diversity of their shuffling conjugation region that contain blaCTX-M-1. These might be involved in the resistance to antimicrobials, but also in their spread.
1. Introduction
In recent decades, the consumption of antimicrobials has been rising in both humans and animals, and as a result, so has the prevalence of plasmid-mediated extended-spectrum β-lactamases (ESBLs) [1]. However, ESBLs confer resistance to penicillins and cephalosporins, including last-generation cephalosporins, which are key molecules for treating infections caused by Gram-negative bacteria in hospitals [1]. Consequently, the last-generation cephalosporins are classified by the World Health Organization (WHO) as critically important antimicrobial agents in human medicine [2]. The ESBLs are inhibited by clavulanic acid, sulbactam and tazobactam, and they are not efficient against carbapenem antimicrobials. Their main reservoir is Enterobacterales, especially the widespread and versatile species Escherichia coli, which is one of the intestinal microbiota and a major pathogen in humans and animals.
Antimicrobial resistance (AMR) is a complex and multifaceted threat to humans, animals, and the environment. A major cause of the AMR burden is the capability of resistant bacteria such as E. coli and AMR-encoding genes to spread between individuals, including across sectors by horizontal gene transfer. Plasmids are extra-chromosomal mobile genetic elements that play an essential role in bacterial ecology and evolution and they help their hosts adapt to a multitude of environments [3]. Plasmids carry accessory genes, including most clinically relevant resistance genes, such as those encoding carbapenemases [4], cephalosporinases [5] and the widespread ESBLs [6,7,8] that can spread across high-risk bacterial clones [9,10]. The most frequently detected ESBLs are class A β-lactamases. They represented by three major types: cefotaximase-Munich (CTX-M), temoneira (TEM) and sulfhydryl variable (SHV) and they include more than 400 variants reported today. These corresponding genes are often associated with other genes that confer resistance to beta-lactams and other antimicrobial agents such as quinolones, aminoglycosides and sulfonamides [7,8].
Initially, ESBLs were variants of TEM- and SHV-type penicillinases that acquired hydrolytic activity against oxyimino cephalosporins, also called third- and fourth-generation cephalosporins (C3G/C4G) through 1- to 4-point mutations. These enzymes were mainly observed during the 1980s and the 1990s in nosocomial Enterobacterales, such as Klebsiella pneumoniae and Enterobacter cloacae, which are mainly responsible for infections in immunocompromised patients in intensive care units [6,7,8]. Since the early 2000s, CTX-M-type ESBLs have been the dominant ESBLs all over the world, owing to their strong association with the species E. coli. This recipient, which is a major pathobiont of the mammal gut, favours the spread not only in intensive care units, as observed for TEM- and SHV-type ESBLs, but also in all other care units of hospitals and the community [5,6,7]. Consequently, the CTX-M-type ESBLs, especially variants CTX-M-15 and CTX-M-1, are community-acquired ESBLs, which have almost substituted for the TEM- and SHV-type ESBLs, and they are the most common plasmid-mediated ESBL among Enterobacterales isolates of human and veterinary origin worldwide [11,12,13,14,15]. CTX-M-15 is encoded by genes located in IncF plasmids harboured by E. coli ST131 clade C, a clade strongly associated with human hosts. CTX-M-1 is observed in E. coli strains collected from humans and animals, and its gene blaCTX-M-1 has been associated mainly with the broad host range IncN plasmids, and much more frequently in the narrow host range IncI1 [16,17,18,19,20,21,22,23,24,25].
The IncI plasmids belong to the I-complex plasmid family including the incompatibility groups IncI1, IncIγ, IncB, IncZ and IncK [26]. The IncI1 plasmid backbone is organised into four major conserved regions encoding replication, conjugative transfer, stability and leading [27,28], in addition to variable regions encoding accessory functions such as antimicrobial gene resistance.
There is a great concern that contacts with animals may enhance the risk of acquiring ESBL-encoding plasmids by humans [29,30]. IncI1-ST3 plasmids are one of the most prevalent plasmids in ESBL CTX-M-1 in Enterobacterales isolated from humans, animals and environmental sources [18,19,20,21,22,23,24,25]. However, the relationships at the interface of humans and animals remain elusive, especially for wild animals. This study compared CTX-M-1-producing E. coli isolates and IncI1-ST3 plasmids collected from humans, food-producing animals, and wild animals to best understand the CTX-M-1 spread among these ecosystems.
2. Materials and Methods
Genomic dataset. For this study, we collected 122 E. coli whole genome sequences (WGSs) containing IncI1-ST3 plasmids and blaCTX-M-1 (Supplementary Tables S1 and S2). The dataset includes WGSs sequenced during this study (n = 43) and recovered from GenBank (n = 79) after filtering for quality (ATCG assembly size >4.5 Mb, contigs number <200 and N50 > 60,000) and the availability of metadata. The sources of strains were humans (n = 57), domestic animals (n = 11), food or food-producing animals (n = 37), wild animals (n = 13) [20,31,32,33,34,35,36,37] and municipal wastewater (n = 1). Three strains were from unknown origins. Of this collection, 30 human strains and 13 strains isolated from wild animals were sequenced for this study (Supplementary Table S1). The other data were collected from the NCBI Short Read Archive (SRA) or the European Nucleotide Archive (ENA) by screening of the E. coli genomes of the GenBank database (Supplementary Table S2). The screening for encoding CTX-M-1- and IncI1-ST3-specific alleles was performed with DIAMOND and blastn software, respectively, using 100% identity threshold and 100% coverage threshold.
Likewise, 20,668 non-redundant complete plasmids collected from GenBank were screened for the IncI1-ST3 feature and the presence of blaCTX-M-1. It resulted in a collection of 39 IncI1-ST3-blaCTX-M-1 circularised plasmids (Supplementary Table S3).
Whole genome sequencing (WGS). This was performed using the next-generation sequencing platform of the teaching hospital of Clermont-Ferrand, France. DNA was extracted with a DNeasy UltraClean Microbial kit (Qiagen, Hilden, Germany). The libraries were prepared with a Nextera XT Kit (Illumina, San Diego, CA, USA), and they were sequenced by the Illumina MiSeq system, generating 2 × 301-base pair (bp) paired-end reads. Fastp software v0.19.10 [38] was used for quality filtering of Illumina reads, and SPAdes was used for short reads assembly [39]. The mean sequencing depth was ≥163×; the number of assembled contigs ranged between 51 and 175, the mean contig number was 99.77, the N50 ranged between 63,075 and 383,707, and the mean contig number was 186,502. The genome sizes ranged between 4,667,864 and 5,338,201 nucleotides. The raw reads have been deposited in the European Nucleotide Archive (ENA, https://www.ebi.ac.uk/ena) under project accession number PRJEB36175.
Molecular typing. E. coli phylogroups and multilocus sequence typing (MLST) were determined in silico according to the Clermont Typing method [40] and Achtman’s MLST scheme [41]. The molecular typing of isolates was performed by core genome SNP-based typing (cgSNP). BactSNP was used to perform cgSNP using the E. coli core genome downloaded from the Enterobase website (https://enterobase.warwick.ac.uk) as a reference, as previously described [42,43]. After the filtration of recombination zones detected by Gubbins [44], a phylogenetic tree was inferred from the resulting alignment by maximum likelihood using RAxML [45].
Antimicrobial gene detection. The antimicrobial-resistant genes were identified by alignment against a database including the online databases CARD [46], Resfinder [47], and the NCBI National Database of Antibiotic Resistant Organisms (https://www.ncbi.nlm.nih.gov/pathogens/antimicrobial-resistance/ (accessed on 1 April 2021)) using a 95% minimum threshold for the breadth of coverage and identity percentage, as previously described [48].
Synteny analysis. This was performed with the Sibelia package [49]. The presence/absence matrix inferring from the resulting synteny blocks was analysed by multiple correspondence analysis (MCA) and hierarchical clustering (HC) in R with package FactoMiner (https://www.r-project.org).
3. Results
3.1. Whole Genome Typing of E. coli Harbouring Plasmids IncI1-ST3 and blaCTX-M-1
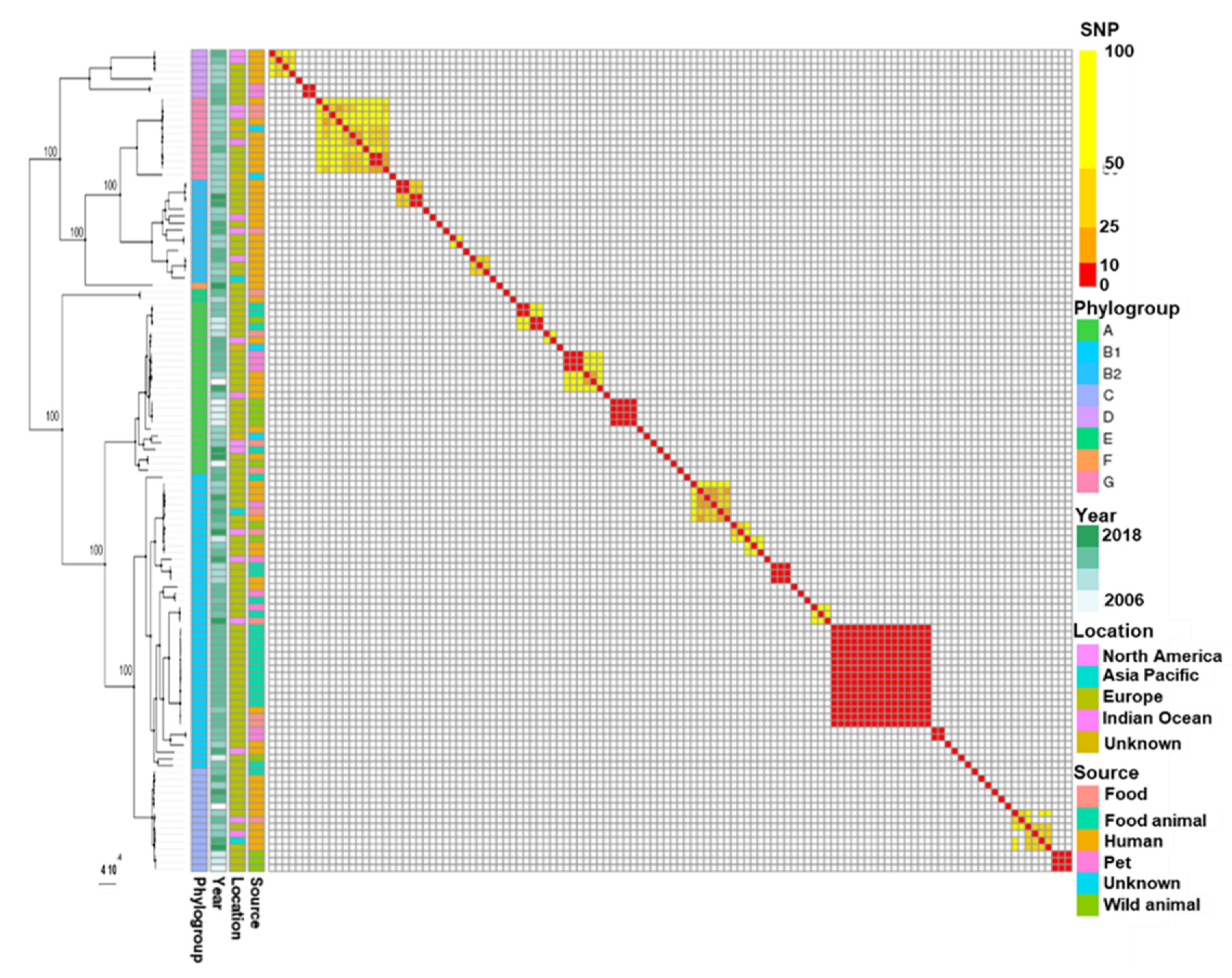
To best understand the large diffusion of the CTX-M-1 ESBL, we collected a dataset comprising 122 E. coli WGSs containing IncI1-ST3 replicon and blaCTX-M-1 isolated from humans, human-influenced habitats and wild fauna. The corresponding genomes were classified into eight major phylogenetic branches by SNP-based core genome typing. These major branches corresponded to the E. coli phylogroups (Figure 1).
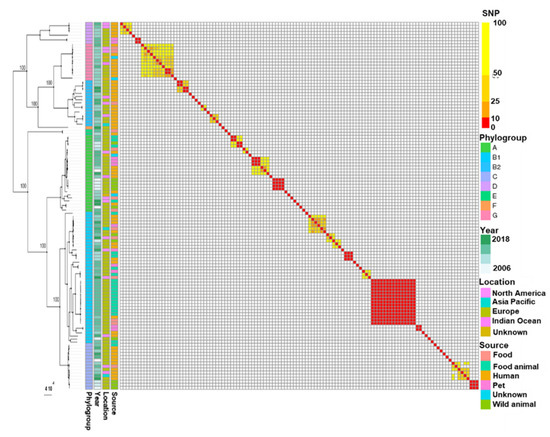
Figure 1.
Phylogenetic tree and corresponding distance matrix based on SNPs of E. coli WGSs containing plasmids IncI1-ST3 encoding blaCTX-M-1. SNP calling, SNP filtering and tree inferring were performed with bactSNP, Gubbins and RAxML, respectively. The bootstrap values are indicated for the major phylogenetic branches as percentages for 500 replications.
Most genomes belonged to phylogroups B1 (35.8%, n = 43), A (20.8%, n = 25), B2 (12.5%, n = 15), C (12.5%, n = 15) and G (10%, n = 12). The distribution of genomes among the phylogroups significantly differed depending on the originating source (Fisher test, p-value < 0.001). Since the genomes of phylogroups A and B1 frequently originated from food or food-producing animals (50% and 67%, respectively), the B2 genomes preferentially originated from humans (93.3% versus 6.7% from food and food-producing animals).
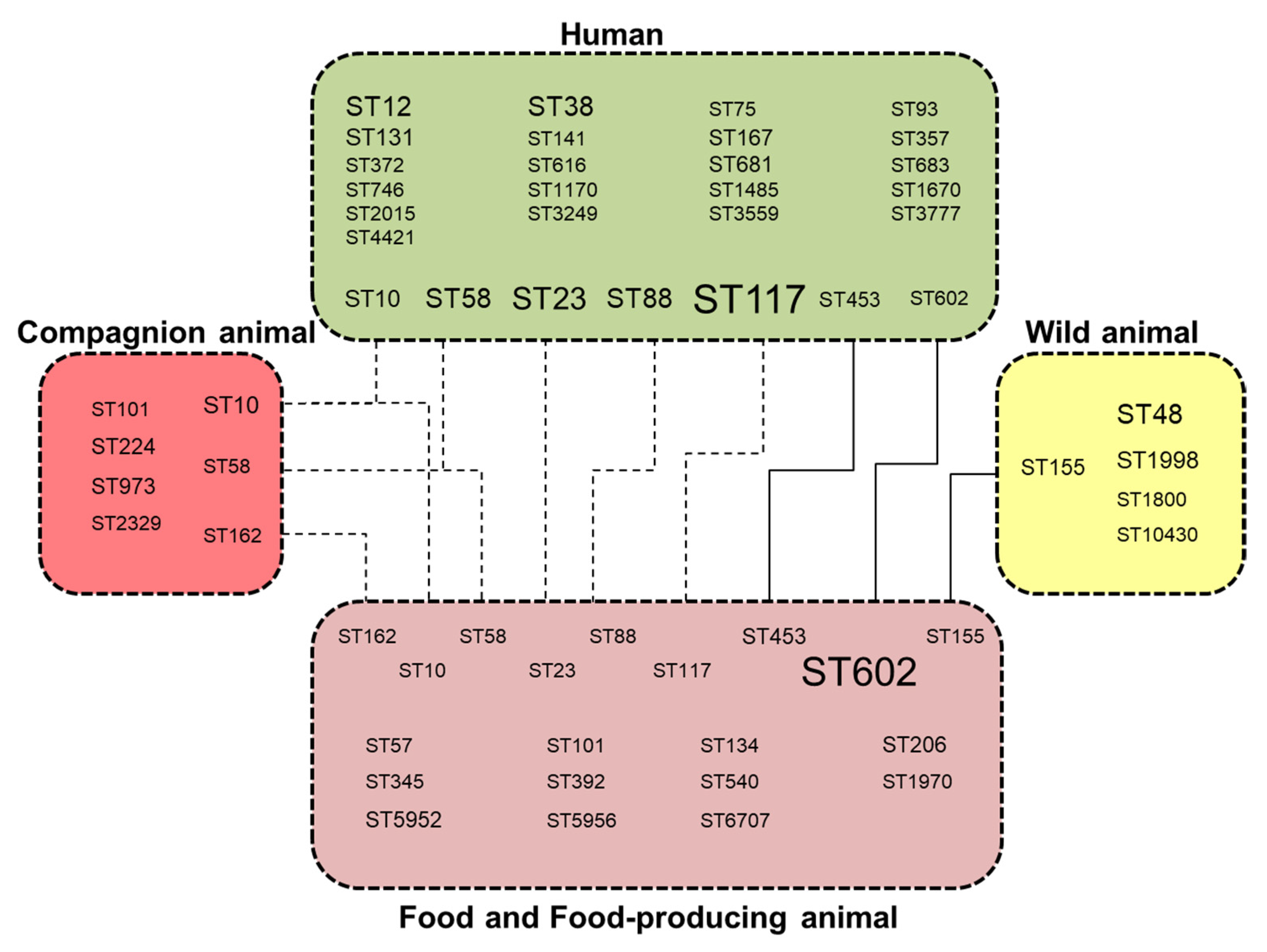
Human E. coli strains (n = 51/57) were distantly related, except for three clusters of two strains diverging by ≤10 SNPs and belonging to ST117 (n = 2) and ST12 (n = 2 × 2). The clonal isolates (divergence ≤ 10 SNPs) mainly clustered isolates from food and animals including wild animals. Few clonal clusters and STs contained isolates from different habitats (Figure 2), with possible cross-transmissions between humans and human-influenced habitats, and between wild and food-producing animals.
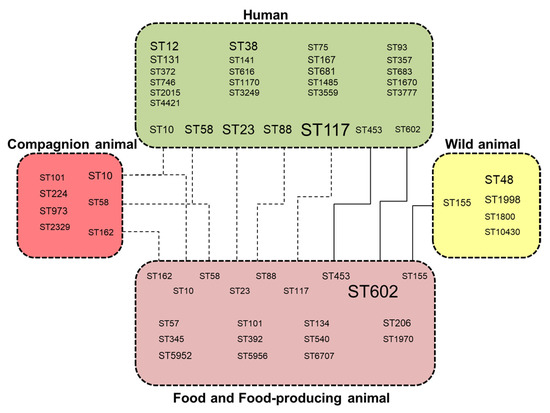
Figure 2.
Distribution of E. coli ST lineages in human, human-influenced habitats and wild fauna showing deduced transmission pathways between these ecosystems. Dashed lines indicate STs shared by different habitats, and solid lines indicate closely related isolates diverging by ≤10 SNPs.
3.2. Antimicrobial Resistance Genes
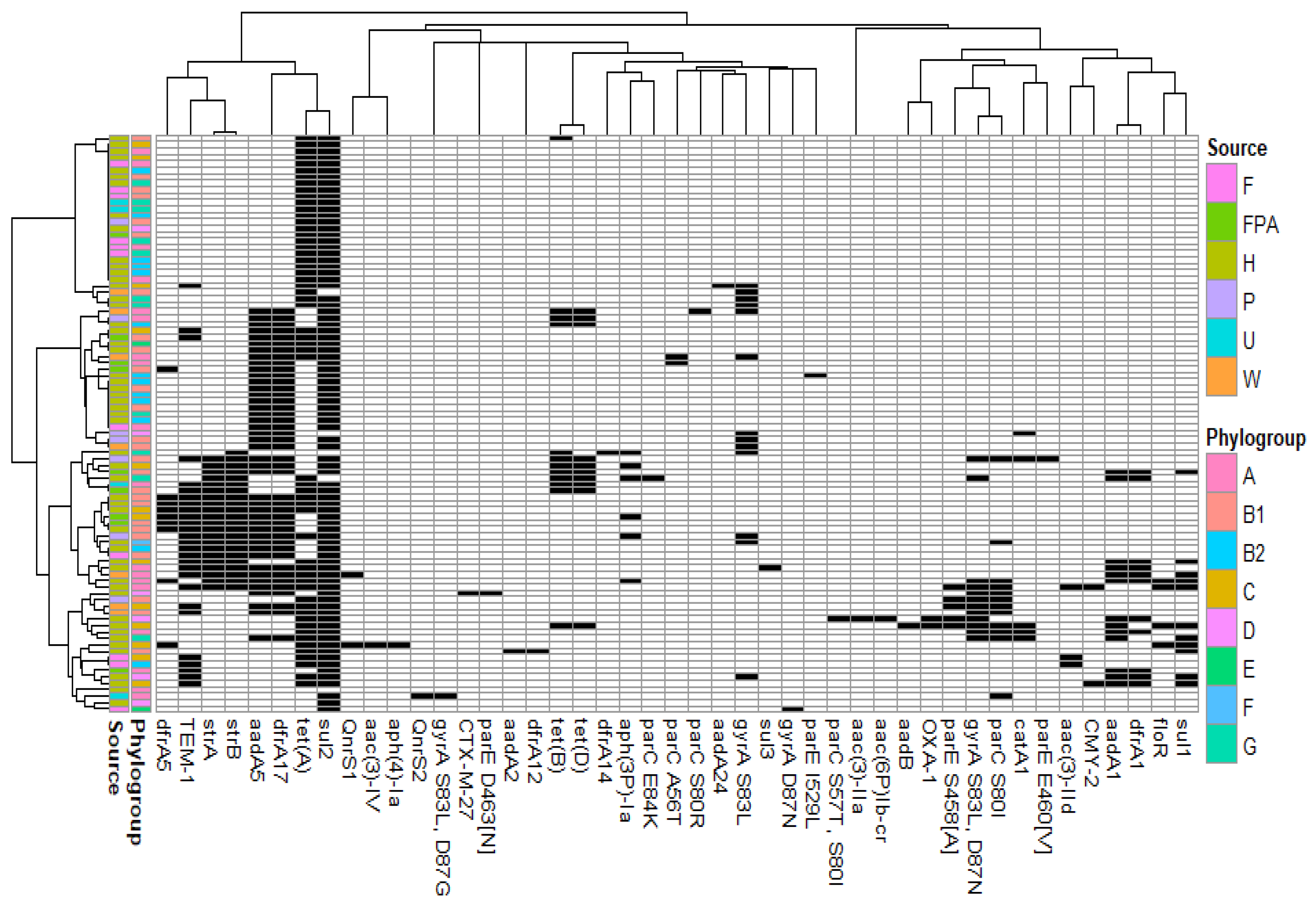
In addition to the chromosome-mediated ampC gene encoding cephalosporinase, 45 acquired antimicrobial resistance mechanisms were associated with blaCTX-M-1 (Figure 3). None of the genes were strictly conserved.
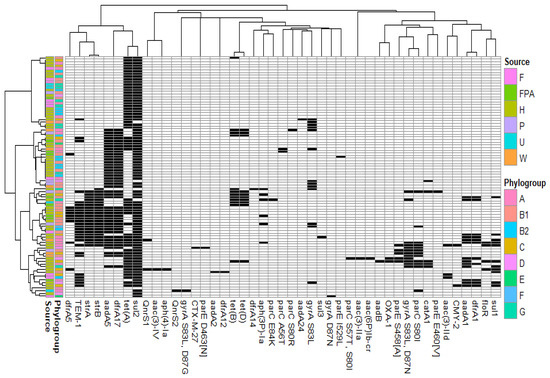
Figure 3.
Antimicrobial resistance mechanisms associated with non-redundant E. coli harbouring blaCTX-M-1 and IncI1-ST3 plasmids (black: presence; white: absence). Source: F, food; FPA, food-producing animals; H, human; P, pet; U: unknown and W, wild animal.
Excluding the redundant isolates corresponding to clonal isolates (n = 30), the most frequent genes were sulphonamide resistance gene sul2 (94.4%), tetracycline resistance gene tet(A) (63.3%), streptomycin/spectinomycin resistance genes aadA5 (47.8%), strB (24.4%) and strA (23.3%), trimethoprim resistance gene dfrA17 (47.8%), and penicillinase-encoding gene blaTEM-1 (30.0%) (Supplementary Figure S1a). The investigation of resistance gene co-occurrence revealed preferential associations. Among the most frequent genes, aadA5, dfrA17, strA, strB and blaTEM-1 exhibited a strong association index. This suggests their frequent coexistence with blaCTX-M-1 probably in the same plasmid IncI1-ST3 (Supplementary Figure S1b).
3.3. SNP Analysis of blaCTX-M-1-Encoding Plasmids IncI1-ST3
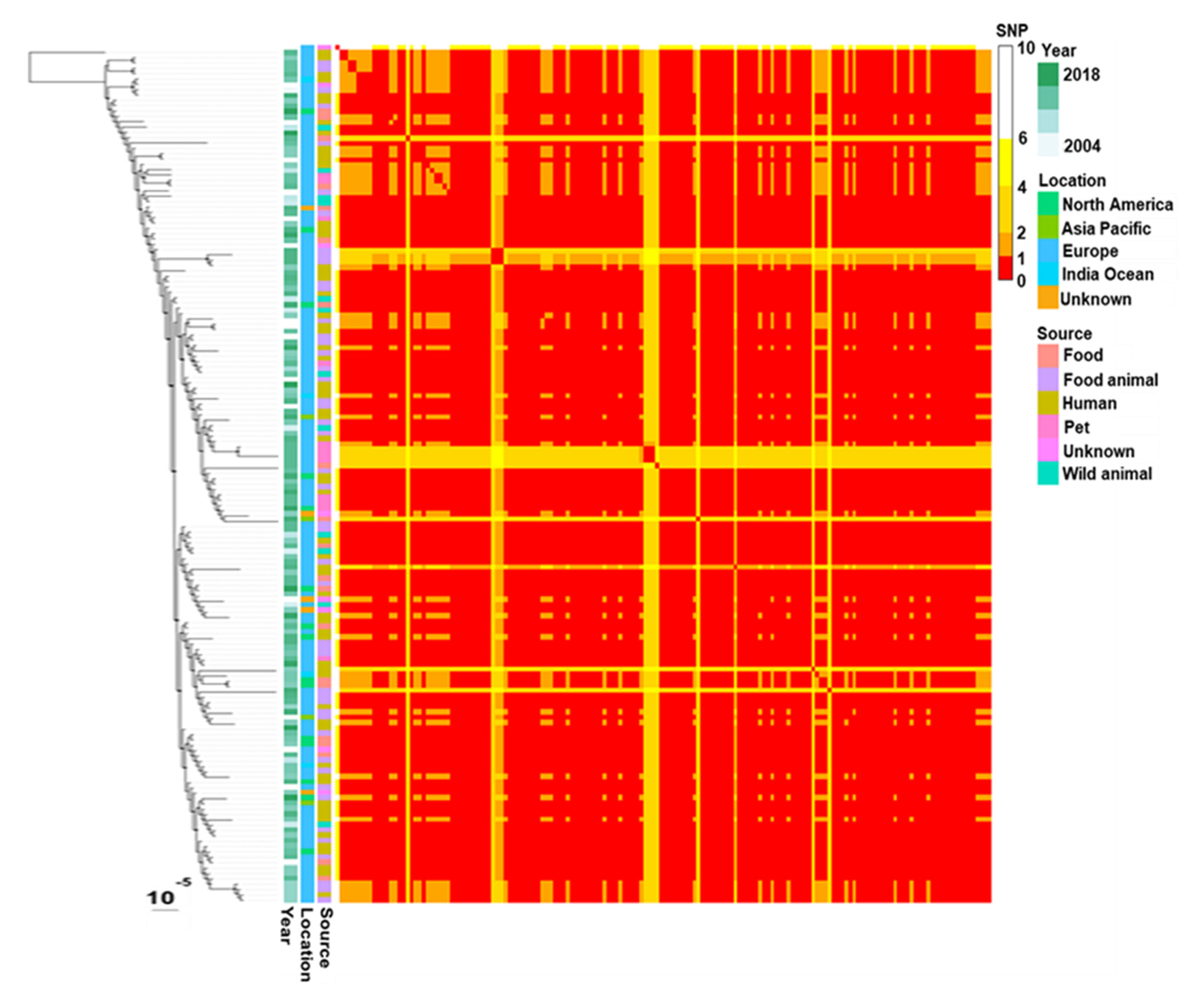
A total of 117 assemblies (96%) contained a contig harbouring blaCTX-M-1, ISEcp1 and at least the B segment of a region previously called shufflon that is specific to IncI1 plasmids [50]. The well-known mobile block ISEcp1-blaCTX-M-1 [51,52] was located 333 pb upstream of the B segment of the shufflon. In four cases, mobile element ISKpn26 was inserted between blaCTX-M-1 and ISEcp1. The blaCTX-M-1 gene was encoded by plasmids IncI1 in most E. coli harbouring this family of plasmids. SNP analysis of IncI1-ST3 plasmids encoding blaCTX-M-1 showed that they differ by <10 SNPs and most often by 1–2 SNPs. The resulting tree had a comb-like shape constituting a unique major clade (Figure 4).
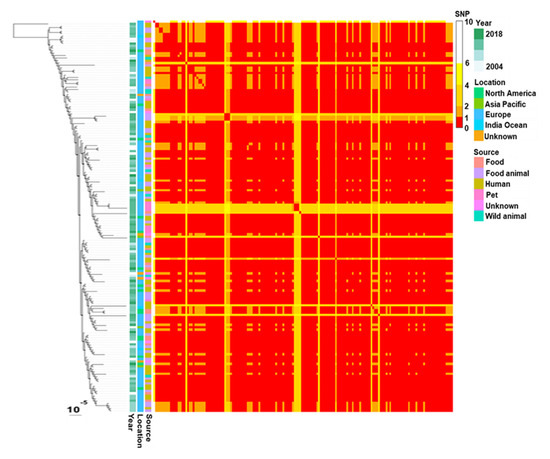
Figure 4.
Phylogenetic tree and corresponding distance matrix based on SNPs of plasmids IncI1-ST3 encoding blaCTX-M-1. SNP calling, SNP filtering and tree inferring were performed with bactSNP, Gubbins and RAxML, respectively.
3.4. Synteny Variation in blaCTX-M-1-Encoding Plasmids IncI1-ST3
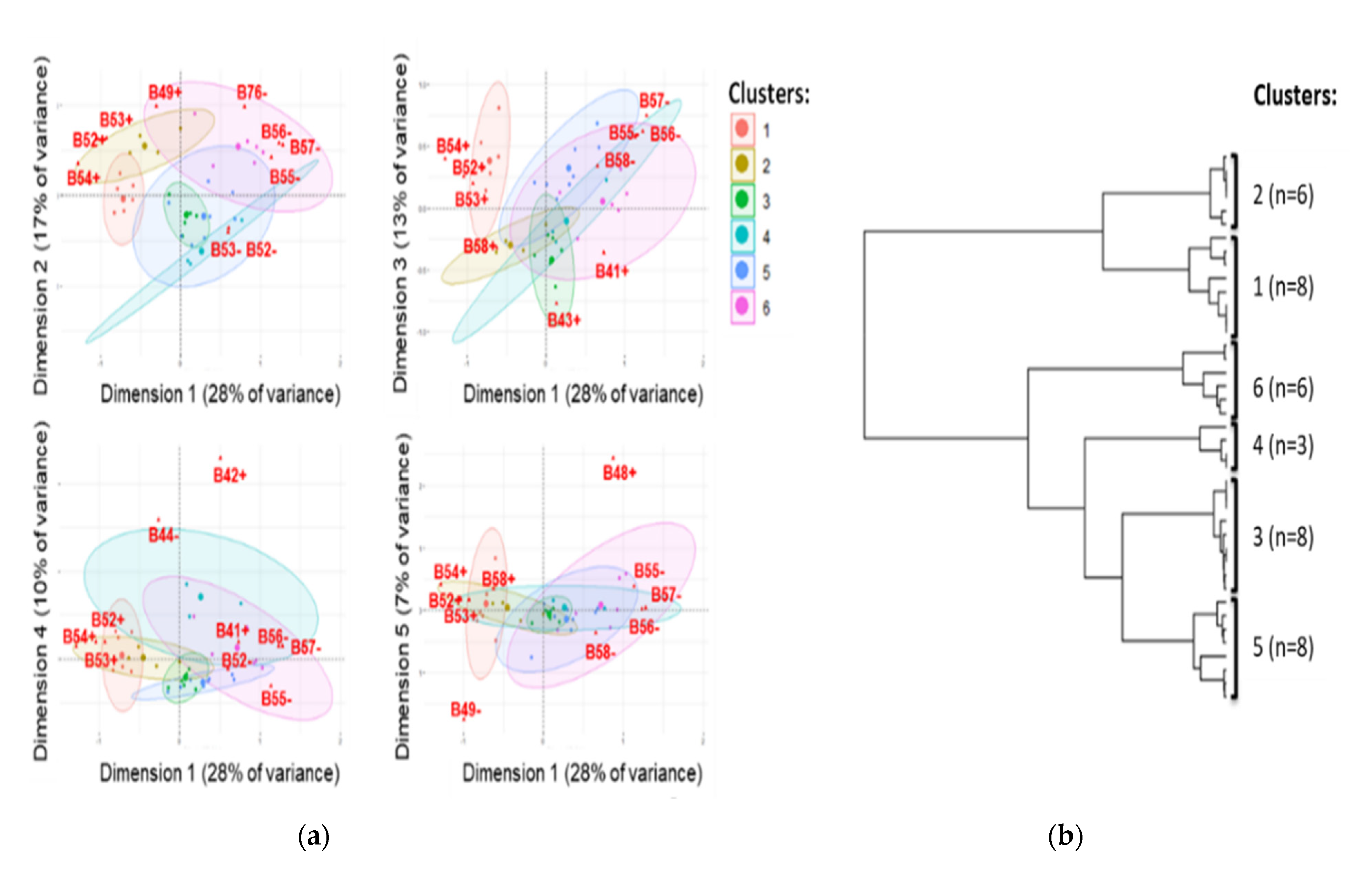
Although not explored for epidemiologic investigations, genetic rearrangements are a major driving force of plasmid evolution. Therefore, we investigated synteny variations in 39 circularised IncI1-ST3-blaCTX-M-1 plasmids. The synteny analysis by multiple correspondence analysis (MCA) and hierarchical clustering (HC) classified the plasmids into six major clusters (Figure 5). The clusters are supported by statistical tests (Adonis test’s p-value: 0.001 and R2: 0.75; Dispersion permutation test’s p-value: 0.1).
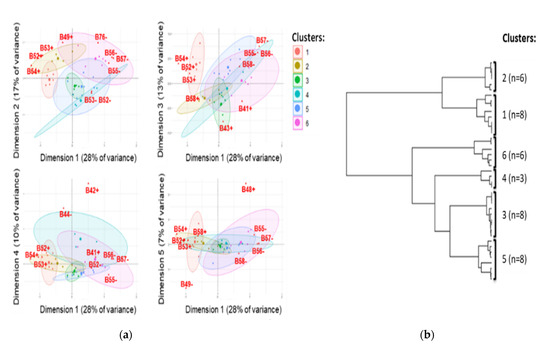
Figure 5.
Multiple corresponding analysis (a) and hierarchical clustering (b) inferring synteny analysis of 39 circularised plasmids IncI1-ST3 encoding blaCTX-M-1. The clusters are surrounded by 95% confidence ellipses, and the 10 most contributory synteny blocks of sequences are indicated in red (B#: shuffling segment B associated with ISEcp1 and blaCTX-M-1; #: position of the genetic feature, rv: reverse and fd: forward).
Among the 14 synteny blocks that were significantly associated with the clusters (FDR-adjusted X2-test’s p-values, 1.4 × 10−6 to 5.5 × 10−3), 13 were in the unique shufflon region between the conserved genes rci and pilV. The genetic features in region rci-pilV are specific to IncI1 plasmids, and they comprise up to four DNA segments A to D, previously identified as randomly rearranged by recombinase Rci [50]. This region harboured the most synteny variations.
The synteny of rci-pilV was also investigated from E. coli WGSs. A total of 77 WGS-encoding plasmids IncI1-ST3 exhibited a complete shufflon assembly, including blaCTX-M-1. MCA and HC analyses of synteny variants from WGSs confirmed the classification of plasmids in six major clusters (Supplementary Figures S2–S4). Segment D was absent in all plasmids, and mobile element ISEcp1-blaCTX-M-1 was always located downstream from segment B to form a conserved block exhibiting different positions in the shufflon. This block can affect the shuffling process, and it was paradoxically the feature that contributed most to diversity (Supplementary Figure S2). The shufflon segments are involved in the synthesis of PilV adhesins of the conjugative pilus [53]. Therefore, the shuffling of segments associated with ISEcp1-blaCTX-M-1 insertion generates diversity in the PilV-encoding region. This can modulate the recognition of recipient cells during IncI1-ST3-blaCTX-M-1 conjugation and therefore probably their dissemination.
At the highest level of classification resolution, synteny analysis revealed 16 clusters of two to seven plasmids sharing identical synteny (Supplementary Figures S3 and S4). Ten of these clusters included plasmids isolated from the same country and the same source. Seven clusters were specific to the source. Nine clusters were only observed in human-influenced habitats. The plasmids isolated from wild animals were included in four clusters; two were specific to wild fauna, and the two others supported a possible spread between human-influenced animals and wild fauna.
4. Discussion
The increase in antimicrobial resistance worldwide is a result of inappropriate use of antimicrobials during the last decades, including those used for human medication and animal husbandry. This broad use increases the selective pressure on both commensal and pathogenic bacteria, which can spread between different ecosystems [1,54]. Livestock animals may act as reservoirs of AMR and multidrug resistant bacteria. This can lead to dissemination of AMR into humans directly by contact and the food chain or indirectly from the environment [1,54].
In this study, we analysed genomic data belonging to E. coli isolates collected from humans, animals (food-producing animals, companion animals, wildlife), and food samples to understand the interaction between these ecosystems in the diffusion of IncI1-ST3 plasmids encoding blaCTX-M-1. The genomic data analysis showed that the E. coli phylogroups harbouring IncI1-ST3 plasmids encoding blaCTX-M-1 significantly differed depending on the originating source. Of the isolates, 68% belonged to the phylogenetic A, B1 and C, which are associated with multiple antimicrobial resistance genes especially those encoding sulphonamide and tetracycline resistance. MLST revealed 50 sequence types. The most abundant sequence type was ST602, followed by ST117 and ST10. The correlation between ST602, which was the most abundant sequence type detected in E. coli phylogenetic group B1 isolates in this study, and food-producing animals was pointed out in recent reports [55,56,57].
Human contamination by ESBL-producing Enterobacterales from animals is often supposed, and food is considered a direct transmission vehicle. ESBL gene blaCTX-M-1 and IncI1-ST3 plasmids were widespread in humans, human-influenced habitats and wild fauna, as previously observed [21,22,23,24,25]. Here, we observed that IncI1-ST3-blaCTX-M-1 plasmids also have disseminated into all E. coli lineages, which cover a broad diversity of bacteria and different lifestyles, including commensal and pathogenic strains. Few clonal clusters and STs were shared by different habitats, suggesting E. coli lineages have a preferential habitat and few of them are involved in cross-sector spread. As shuttles, these subgroups may be risk factors for spreading antimicrobial resistance and they might be preferential targets for strategies to prevent the spread of antimicrobial resistance.
The SNP-based comparison of blaCTX-M-1 IncI1-ST3 plasmids originating from several continents revealed a core genome highly conserved. This suggests that the dissemination of these plasmids across all sources over distant areas took decades. However, the evolutionary rate of bacterial genomes may not generate enough variations to resolve recent epidemiological processes involving small genetic elements such as IncI1-ST3-blaCTX-M-1 plasmids. Recombination, gain and loss of DNA fragments are key processes of evolution [58]. They affect synteny and are not investigated by comparisons based on core genome SNPs. The analysis of synteny in complete blaCTX-M-1 IncI1-ST3 plasmids revealed more diversity than SNP analysis. Synteny-based clusters were associated with sampling sources and geographic origins. This suggests that synteny analysis can be a useful approach for monitoring IncI1-ST3 plasmid spread over short periods, and it might help to analyse transmission chains.
Most variations in synteny were observed in a single region, which was previously designated shufflon and encoding a recombinase and targeted DNA segments [50]. Most diversity resulted from the positioning and orientation of a conserved block including the shufflon segment B, ISEcp1 and blaCTX-M-1. Shufflon B appears, therefore, as a hot spot for the insertion of mobile element ISEcp1 and associated gene blaCTX-M-1. Since ISEcp1 is involved in the mobilisation of ESBL- and cephalosporinase-encoding genes [51,59], shufflon B of IncI1 explains the key role of these plasmids in the spread of resistance to last-generation cephalosporins.
The assembly of blaCTX-M-1 from short reads suggest a certain stability and/or the preponderance of a shufflon synteny within a bacterial clone. This contrasts with plasmids IncI2 harbouring active shufflons [60]. This stability was confirmed by assembly from long-read sequencing, which did not reveal alternative conformation of shufflons [61] and may be explained by the insertion of ISEcp1-blaCTX-M-1 in the shuffling zone. The shufflon is involved in the synthesis of PilV adhesins, which are responsible for recipient recognition in the conjugation process [53]. The insertion of ISEcp1-blaCTX-M-1 associated with the shuffling of segments can affect PilV and consequently modulate the recognition of recipient cells during IncI1-ST3-blaCTX-M-1 conjugation. Resistance gene blaCTX-M-1 can, therefore, be involved in both antimicrobial resistance and plasmid spread, two synergic functions that may explain the ecological success of blaCTX-M-1 IncI1-ST3 plasmids.
5. Conclusions
Although additional animal and environmental sources of CTX-M-1-producing E. coli should be investigated, the results showed there was broad dissemination of IncI1-ST3-blaCTX-M-1 plasmids. Their bacterial recipients differ by habitats, with a few of them playing the role of disseminating shuttles. The sequence of IncI1-ST3-blaCTX-M-1 plasmids is highly conserved except in the shufflon zone. Their broad ecological success does not seem to be linked to their ability to transfer a broad spectrum of bacterial lineages, a feature associated with the diversity of their shuffling conjugation region.
Supplementary Materials
The following are available online at https://www.mdpi.com/article/10.3390/microorganisms9071471/s1, Figure S1: Frequency (a) and correlation index (b) of antimicrobial resistance genes in non-redundant E. coli harbouring blaCTX-M-1 and IncI1-ST3 plasmids, Figure S2: Multiple corresponding analysis performed from the synteny of shuffling region of blaCTX-M-1 IncI1-ST3 plasmids, Figure S3: Heatmap showing the presence (red) and the absence (blue) of synteny blocks in the shuffling region of plasmids IncI1-ST3-blaCTX-M-1, Figure S4: Hierarchical classification of plasmids IncI1-ST3-blaCTX-M-1 based on the synteny of the shufflon region, Table S1: Metadata associated with the Escherichia coli isolates sequenced during this study, Table S2: Dataset of Escherichia coli whole genome sequences (WGSs) containing replicon IncI1-ST3 encoding blaCTX-M-1, Table S3: Circularised blaCTX-M-1-encoding plasmids IncI1-ST3 included in this study.
Author Contributions
Conceptualisation, R.B. (Racha Beyrouthy) and R.B. (Richard Bonnet); methodology, R.B. (Racha Beyrouthy), and R.B. (Richard Bonnet); formal analysis, R.B., C.S., F.R., G.I. and P.P.; investigation, R.B. (Racha Beyrouthy), and R.B. (Richard Bonnet); resources, F.R., C.S., F.R., G.I. and P.P.; data curation, R.B. (Racha Beyrouthy), F.R. and R.B. (Richard Bonnet); writing—original draft preparation, R.B. (Racha Beyrouthy) and R.B. (Richard Bonnet); writing—review and editing, R.B. (Racha Beyrouthy) F.R., G.I., and R.B. (Richard Bonnet); supervision, R.B. (Racha Beyrouthy) and R.B. (Richard Bonnet); project administration, R.B. (Richard Bonnet); funding acquisition, R.B. (Richard Bonnet). All authors have read and agreed to the published version of the manuscript.
Funding
This study was funded by the Ministère de la Recherche et de la Technologie, Inserm (UMR 1071), INRAe (USC-2018), the French government’s IDEX-ISITE initiative 16-IDEX-0001 (CAP 20-25) and by the JPIAMR program TransComp-ESC-R of the European Commission. We also thanks the “Espectrometria de massa e sequenciação aplicadas à microbiologia clínica moderna: pesquisa e identificação de biomarcadores em espécies bacterianas multiresistentes” (Programa de Acções Universitárias Integradas Luso-Francesas, Ação n.º: TC-14/2017) and by the Associate Laboratory for Green Chemistry-LAQV which is financed by national funds from FCT/MCTES (UID/QUI/50006/2019).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The accession numbers of genomic data are reported in Supplementary Tables S2 and S3.
Acknowledgments
We thank Alexis Pontvianne and Lucie Pourpuech for their technical assistance.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Roca, I.; Akova, M.; Baquero, F.; Carlet, J.; Cavaleri, M.; Coenen, S.; Cohen, J.; Findlay, D.; Gyssens, I.; Heure, O.E.; et al. The global threat of antimicrobial resistance: Science for intervention. New Microbes New Infect. 2015, 6, 22–29. [Google Scholar] [CrossRef]
- WHO. WHO List of Critically Important Antimicrobials for Human Medicine (WHO CIA List); 5th Revision; World Health Organization: Geneva, Switzerland, 2017. [Google Scholar]
- Frost, L.; Leplae, R.; Summers, A.; Toussaint, A. Mobile genetic elements: The agents of open source evolution. Nat. Rev. Genet. 2005, 3, 722–732. [Google Scholar] [CrossRef]
- Queenan, A.M.; Bush, K. Carbapenemases: The versatile β-lactamases. Clin. Microbiol. Rev. 2007, 20, 440–458. [Google Scholar] [CrossRef] [PubMed]
- Philippon, A.; Arlet, G.; Jacoby, G.A. Plasmid-determined AmpC-type β-lactamases. Antimicrob. Agents Chemother. 2002, 46, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Bonnet, R. Growing group of extended-spectrum β-lactamases: The CTX-M enzymes. Antimicrob. Agents Chemother. 2004, 48, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Paterson, D.L.; Bonomo, R.A. Extended spectrum beta lactamases: A critical update. Clin. Microbiol. Rev. 2005, 18, 657–686. [Google Scholar] [CrossRef]
- Pitout, J.D.; Laupland, K.B. Extended-spectrum β-lactamase-producing Enterobacteriaceae: An emerging public-health concern. Lancet Infect. Dis. 2008, 8, 159–166. [Google Scholar] [CrossRef]
- Bonomo, R.A.; Burd, E.M.; Conly, J.; Limbago, B.M.; Poirel, L.; Segre, J.A.; Westblade, L.F. Carbapenemase-producing organisms: A global scourge. Clin. Infect. Dis. 2018, 66, 1290–1297. [Google Scholar] [CrossRef]
- David, S.; Reuter, S.; Harris, S.R.; Glasner, C.; Feltwell, T.; Argimon, S.; Abudahab, K.; Goater, R.; Giani, T.; Errico, G.; et al. Europe PMC funders group europe PMC funders author manuscripts Europe PMC funders author manuscripts epidemic of carbapenem-resistant Klebsiella pneumoniae in Europe is driven by nosocomial spread. Nature 2020, 4, 1919–1929. [Google Scholar]
- Haenni, M.; Beyrouthy, R.; Lupo, A.; Châtre, P.; Madec, J.Y.; Bonnet, R. Epidemic spread of Escherichia coli ST744 isolates carrying mcr-3 and blaCTX-M-55 in cattle in France. J. Antimicrob. Chemother. 2018, 73, 533–536. [Google Scholar] [CrossRef]
- Robin, F.; Beyrouthy, R.; Bonacorsi, S.; Aissa, N.; Bret, L.; Brieu, N.; Cattoir, V.; Chapuis, A.; Chardon, H.; Degand, N.; et al. Inventory of extended-spectrum-β-lactamase-producing enterobacteriaceae in France as assessed by a multicenter study. Antimicrob. Agents Chemother. 2017, 61, e01911-16. [Google Scholar] [CrossRef] [PubMed]
- Wellington, E.M.; Boxall, A.B.; Cross, P.; Feil, E.J.; Gaze, W.H.; Hawkey, P.M.; Johnson-Rollings, A.S.; Jones, D.L.; Lee, N.M.; Otten, W.; et al. The role of the natural environment in the emergence of antibiotic resistance in Gram-negative bacteria. Lancet Infect. Dis. 2013, 13, 155–165. [Google Scholar] [CrossRef]
- Pietsch, M.; Eller, C.; Wendt, C.; Holfelder, M.; Falgenhauer, L.; Fruth, A.; Grössl, T.; Leistner, R.; Valenza, G.; Werner, G.; et al. Molecular characterisation of extended-spectrum β-lactamase (ESBL)-producing Escherichia coli isolates from hospital and ambulatory patients in Germany. Vet. Microbiol. 2017, 200, 130–137. [Google Scholar] [CrossRef]
- Liu, X.; Thungrat, K.; Boothe, D.M. Occurrence of oxa-48 carbapenemase and other β-lactamase genes in esbl-producing multidrug resistant: Escherichia coli from dogs and cats in the united states, 2009–2013. Front. Microbiol. 2016, 7, 1057. [Google Scholar] [CrossRef]
- Götz, A.; Pukall, R.; Smit, E.; Tietze, E.; Prager, R.; Tschäpe, H.; van Elsas, J.D.; Smalla, K. Detection and characterization of broad-host-range plasmids in environmental bacteria by PCR. Appl. Environ. Microbiol. 1996, 62, 2621–2628. [Google Scholar] [CrossRef] [PubMed]
- Pukall, R.; Tschäpe, H.; Smalla, K. Monitoring the spread of broad host and narrow host range plasmids in soil microcosms. FEMS Microbiol. Ecol. 1996, 20, 53–66. [Google Scholar] [CrossRef]
- Dolejska, M.; Papagiannitsis, C.C. Plasmid-mediated resistance is going wild. Plasmid 2018, 99, 99–111. [Google Scholar] [CrossRef]
- Carattoli, A.; Villa, L.; Fortini, D.; García-Fernández, A. Contemporary IncI1 plasmids involved in the transmission and spread of antimicrobial resistance in Enterobacteriaceae. Plasmid 2018, 102392. [Google Scholar] [CrossRef]
- Radhouani, H.; Pinto, L.; Coelho, C.; Gonçalves, A.; Sargo, R.; Torres, C.; Igrejas, G.; Poeta, P. Detection of Escherichia coli harbouring extended-spectrum β-lactamases of the CTX-M classes in faecal samples of common buzzards (Buteo buteo). J. Antimicrob. Chemother. 2010, 65, 171–173. [Google Scholar] [CrossRef]
- Madec, J.Y.; Haenni, M.; Métayer, V.; Saras, E.; Nicolas-Chanoine, M.H. High prevalence of the animal-associated blaCTX-M-1 IncI1/ST3 plasmid in human Escherichia coli isolates. Antimicrob. Agents Chemother. 2015, 59, 5860–5861. [Google Scholar] [CrossRef]
- Jakobsen, L.; Bortolaia, V.; Bielak, E.; Moodley, A.; Olsen, S.S.; Hansen, D.S.; Frimodt-Møller, N.; Guardabassi, L.; Hasman, H. Limited similarity between plasmids encoding CTX-M-1 β-lactamase in Escherichia coli from humans, pigs, cattle, organic poultry layers and horses in Denmark. J. Glob. Antimicrob. Resist. 2015, 3, 132–136. [Google Scholar] [CrossRef]
- Carattoli, A. Resistance plasmid families in enterobacteriaceae. Antimicrob. Agents Chemother. 2009, 53, 2227–2238. [Google Scholar] [CrossRef]
- Literak, I.; Dolejska, M.; Janoszowska, D.; Hrusakova, J.; Meissner, W.; Rzyska, H.; Bzoma, S.; Cizek, A. Antibiotic-resistant Escherichia coli bacteria, including strains with genes encoding the extended-spectrum beta-lactamase and QnrS, in waterbirds on the baltic sea coast of Poland. Appl. Environ. Microbiol. 2010, 76, 8126–8134. [Google Scholar] [CrossRef]
- Veldman, K.; Van Tulden, P.; Kant, A.; Testerink, J.; Mevius, D. Characteristics of cefotaxime-resistant escherichia coli from wild birds in The Netherlands. Appl. Environ. Microbiol. 2013, 79, 7556–7561. [Google Scholar] [CrossRef]
- Praszkier, J.; Pittard, A.J. Control of replication in I-complex plasmids. Plasmid 2005, 53, 97–112. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, H.; Shao, M.; Furuya, N.; Komano, T. The genome sequence of the incompatibility group Iγ plasmid R621a: Evolution of IncI plasmids. Plasmid 2011, 66, 112–121. [Google Scholar] [CrossRef] [PubMed]
- Johnson, T.J.; Shepard, S.M.; Rivet, B.; Danzeisen, J.L.; Carattoli, A. Comparative genomics and phylogeny of the IncI1 plasmids: A common plasmid type among porcine enterotoxigenic Escherichia coli. Plasmid 2011, 66, 144–151. [Google Scholar] [CrossRef] [PubMed]
- Hernando-Amado, S.; Coque, T.M.; Baquero, F.; Martínez, J.L. Defining and combating antibiotic resistance from One Health and Global Health perspectives. Nat. Microbiol. 2019, 4, 1432–1442. [Google Scholar] [CrossRef]
- Huijbers, P.M.C.; Graat, E.A.M.; Haenen, A.P.J.; van Santen, M.G.; van Essen-Zandbergen, A.; Mevius, D.J.; Van Duijkeren, E.; Van Hoek, A.H.A.M. Extended-spectrum and AmpC β-lactamase-producing Escherichia coli in broilers and people living and/or working on broiler farms: Prevalence, risk factors and molecular characteristics. J. Antimicrob. Chemother. 2014, 69, 2669–2675. [Google Scholar] [CrossRef]
- Ramos, S.; Silva, N.; Dias, D.; Sousa, M.; Capelo-Martínez, J.L.; Brito, F.; Caniça, M.; Igrejas, G.; Poeta, P. Clonal Diversity of ESBL-Producing Escherichia coli in Pigs at Slaughter Level in Portugal. Foodborne Pathog. Dis. 2013, 10, 74–79. [Google Scholar] [CrossRef]
- Poeta, P.; Radhouani, H.; Pinto, L.; Martinho, A.; Rego, V.; Rodrigues, R.; Gonçalves, A.; Rodrigues, J.; Estepa, V.; Torres, C.; et al. Wild boars as reservoirs of extended-spectrum beta-lactamase (ESBL) producing Escherichia coli of different phylogenetic groups. J. Basic Microbiol. 2009, 49, 584–588. [Google Scholar] [CrossRef] [PubMed]
- Poeta, P.; Radhouani, H.; Igrejas, G.; Gonçalves, A.; Carvalho, C.; Rodrigues, J.; Vinue, L.; Somalo, S.; Torres, C. Seagulls of the berlengas natural reserve of portugal as carriers of fecal escherichia coli harboring CTX-M and TEM extended-spectrum beta-lactamases. Appl. Environ. Microbiol. 2008, 74, 7439–7441. [Google Scholar] [CrossRef] [PubMed]
- Pinto, L.; Radhouani, H.; Coelho, C.; Da Costa, P.M.; Simões, R.; Brandão, R.M.L.; Torres, C.; Igrejas, G.; Poeta, P. Genetic detection of extended-spectrum β-lactamase-containing Escherichia coli isolates from birds of prey from Serra da Estrela natural reserve in Portugal. Appl. Environ. Microbiol. 2010, 76, 4118–4120. [Google Scholar] [CrossRef] [PubMed]
- Gonçalves, A.; Igrejas, G.; Radhouani, H.; Estepa, V.; Pacheco, R.; Monteiro, R.; Brito, F.; Guerra, A.; Petrucci-Fonseca, F.; Torres, C.; et al. Iberian wolf as a reservoir of extended-spectrum β-lactamase-producing escherichia coli of the TEM, SHV, and CTX-M groups. Microb. Drug Resist. 2012, 18, 215–219. [Google Scholar] [CrossRef] [PubMed]
- Gonçalves, A.; Igrejas, G.; Radhouani, H.; Estepa, V.; Alcaide, E.; Zorrilla, I.; Serra, R.; Torres, C.; Poeta, P. Detection of extended-spectrum beta-lactamase-producing Escherichia coli isolates in faecal samples of Iberian lynx. Lett. Appl. Microbiol. 2011, 54, 73–77. [Google Scholar] [CrossRef]
- Garcês, A.; Correia, S.; Amorim, F.; Pereira, J.; Igrejas, G.; Poeta, P. First report on extended-spectrum beta-lactamase (ESBL) producing Escherichia coli from European free-tailed bats (Tadarida teniotis) in Portugal: A one-health approach of a hidden contamination problem. J. Hazard. Mater. 2019, 370, 219–224. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Zhou, Y.; Chen, Y.; Gu, J. fastp: An ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 2018, 34, i884–i890. [Google Scholar] [CrossRef]
- Nurk, S.; Bankevich, A.; Antipov, D.; Gurevich, A.; Korobeynikov, A.; Lapidus, A.; Prjibelski, A.D.; Pyshkin, A.; Sirotkin, A.; Sirotkin, Y.; et al. Assembling single-cell genomes and mini-metagenomes from chimeric MDA products. J. Comput. Biol. 2013, 20, 714–737. [Google Scholar] [CrossRef] [PubMed]
- Beghain, J.; Bridier-Nahmias, A.; Le Nagard, H.; Denamur, E.; Clermont, O. ClermonTyping: An easy-to-use and accurate in silico method for Escherichia genus strain phylotyping. Microb. Genom. 2018, 4, e000192. [Google Scholar] [CrossRef]
- Wirth, T.; Falush, D.; Lan, R.; Colles, F.; Mensa, P.; Wieler, L.H.; Karch, H.; Reeves, P.R.; Maiden, M.C.J.; Ochman, H.; et al. Sex and virulence in Escherichia coli: An evolutionary perspective. Mol. Microbiol. 2006, 60, 1136–1151. [Google Scholar] [CrossRef]
- Phan, M.D.; Peters, K.M.; Sarkar, S.; Lukowski, S.W.; Allsopp, L.P.; Moriel, D.G.; Achard, M.E.S.; Totsika, M.; Marshall, V.M.; Upton, M.; et al. The serum resistome of a globally disseminated multidrug resistant uropathogenic escherichia coli clone. PLoS Genet. 2013, 9, e1003834. [Google Scholar] [CrossRef] [PubMed]
- Ben Zakour, N.L.; Alsheikh-Hussain, A.S.; Ashcroft, M.M.; Khanh Nhu, N.T.; Roberts, L.W.; Stanton-Cook, M.; Schembri, M.A.; Beatsona, S.A. Sequential acquisition of virulence and fluoroquinolone resistance has shaped the evolution of Escherichia coli ST131. mBio 2016, 7, e00347-16. [Google Scholar] [CrossRef] [PubMed]
- Croucher, N.; Page, A.; Connor, T.; Delaney, A.J.; Keane, J.A.; Bentley, S.D.; Parkhill, J.; Harris, S.R. Rapid phylogenetic analysis of large samples of recombinant bacterial whole genome sequences using Gubbins. Nucleic Acids Res. 2015, 43, e15. [Google Scholar] [CrossRef] [PubMed]
- Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bionformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef]
- Jia, B.; Raphenya, A.R.; Alcock, B.; Waglechner, N.; Guo, P.; Tsang, K.K.; Lago, B.A.; Dave, B.M.; Pereira, S.; Sharma, A.N.; et al. CARD 2017: Expansion and model-centric curation of the comprehensive antibiotic resistance database. Nucleic Acids Res. 2017, 45, D566–D573. [Google Scholar] [CrossRef]
- Zankari, E.; Hasman, H.; Cosentino, S.; Vestergaard, M.; Rasmussen, S.; Lund, O.; Aarestrup, F.; Larsen, M.V. Identification of acquired antimicrobial resistance genes. J. Antimicrob. Chemother. 2012, 67, 2640–2644. [Google Scholar] [CrossRef]
- Beyrouthy, R.; Barets, M.; Marion, E.; Dananché, C.; Dauwalder, O.; Robin, F.; Gauthier, L.; Jousset, A.; Dortet, L.; Guérin, F.; et al. Novel enterobacter lineage as leading cause of nosocomial outbreak involving carbapenemase-producing strains. Emerg. Infect. Dis. 2018, 24, 1505–1515. [Google Scholar] [CrossRef]
- Minkin, I.; Patel, A.; Kolmogorov, M.; Vyahhi, N.; Pham, S. Sibelia: A Scalable and Comprehensive Synteny Block Generation Tool for Closely Related Microbial Genomes. In International Workshop on Algorithms in Bioinformatics; Springer: Berlin/Heidelberg, Germany, 2013; Volume 8126, pp. 215–229. [Google Scholar]
- Gyohda, A.; Furuya, N.; Kogure, N.; Komano, T. Sequence-specific and Non-specific binding of the rci protein to the asymmetric recombination sites of the R64 shufflon. J. Mol. Biol. 2002, 318, 975–983. [Google Scholar] [CrossRef]
- Poirel, L.; Decousser, J.W.; Nordmann, P. Insertion sequence ISEcp1B is involved in expression and mobilization of a blaCTX-M β-lactamase gene. Antimicrob. Agents Chemother. 2003, 47, 2938–2945. [Google Scholar] [CrossRef]
- Poirel, L.; Lartigue, M.-F.; Decousser, J.-W.; Nordmann, P. IS Ecp1B -Mediated Transposition of bla CTX-M in Escherichia coli. Antimicrob. Agents Chemother. 2005, 49, 447–450. [Google Scholar] [CrossRef] [PubMed]
- Ishiwa, A.; Komano, T. Thin pilus Pilv adhesins of plasmid R64 recognize specific structures of the lipopolysaccharide molecules of recipient cells. J. Bacteriol. 2003, 185, 5192–5199. [Google Scholar] [CrossRef]
- Liebana, E.; Carattoli, A.; Coque, T.M.; Hasman, H.; Magiorakos, A.P.; Mevius, D.; Peixe, L.; Poirel, L.; Schuepbach-Regula, G.; Torneke, K.; et al. Public health risks of enterobacterial isolates producing extended-spectrum β-lactamases or AmpC β-lactamases in food and food-producing animals: An EU perspective of epidemiology, analytical methods, risk factors, and control options. Clin. Infect. Dis. 2013, 56, 1030–1037. [Google Scholar] [CrossRef]
- Jouini, A.; Ben Slama, K.; Klibi, N.; Ben Sallem, R.; Estepa, V.; Vinue, L.; Sáenz, Y.; Ruiz-Larrea, F.; Boudabous, A.; Torres, C. Lineages and Virulence Gene Content among Extended-Spectrum β-Lactamase–Producing Escherichia coli Strains of Food Origin in Tunisia. J. Food Prot. 2013, 76, 323–327. [Google Scholar] [CrossRef] [PubMed]
- Day, M.J.; Hopkins, K.L.; Wareham, D.W.; Toleman, M.; Elviss, N.; Randall, L.; Teale, C.; Cleary, P.; Wiuff, C.; Doumith, M.; et al. Extended-spectrum β-lactamase-producing Escherichia coli in human-derived and foodchain-derived samples from England, Wales, and Scotland: An epidemiological surveillance and typing study. Lancet Infect. Dis. 2019, 19, 1325–1335. [Google Scholar] [CrossRef]
- Irrgang, A.; Hammerl, J.A.; Falgenhauer, L.; Guiral, E.; Schmoger, S.; Imirzalioglu, C.; Fischer, J.; Guerra, B.; Chakraborty, T.; Käsbohrer, A. Diversity of CTX-M-1-producing E. coli from German food samples and genetic diversity of the blaCTX-M-1 region on IncI1 ST3 plasmids. Vet. Microbiol. 2018, 221, 98–104. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Beltrán, J.; DelaFuente, J.; León-Sampedro, R.; MacLean, R.C.; Millán, Á.S. Beyond horizontal gene transfer: The role of plasmids in bacterial evolution. Nat. Rev. Genet. 2021, 19, 347–359. [Google Scholar] [CrossRef] [PubMed]
- Giles, W.P.; Benson, A.K.; Olson, M.E.; Hutkins, R.W.; Whichard, J.M.; Winokur, P.; Fey, P.D. DNA sequence analysis of regions surrounding blaCMY-2 from multiple salmonella plasmid backbones. Antimicrob. Agents Chemother. 2004, 48, 2845–2852. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Sekizuka, T.; Kawanishi, M.; Ohnishi, M.; Shima, A.; Kato, K.; Yamashita, A.; Matsui, M.; Suzuki, S.; Kuroda, M. Elucidation of quantitative structural diversity of remarkable rearrangement regions, shufflons, in IncI2 plasmids. Sci. Rep. 2017, 7, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Sabença, C.; Igrejas, G.; Poeta, P.; Robin, F.; Bonnet, R.; Beyrouthy, R. Multidrug Resistance Dissemination in Escherichia coli isolated from wild animals: Bacterial clones and plasmid complicity. Microbiol. Res. 2021, 12, 9. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).