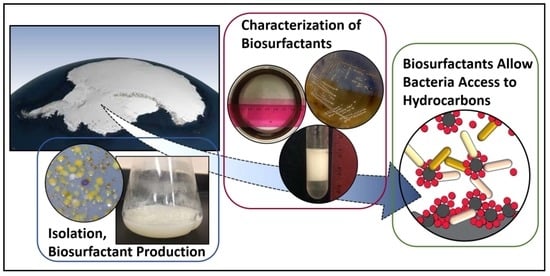
Low-Temperature Biosurfactants from Polar Microbes
Abstract
1. Introduction
2. Materials and Methods
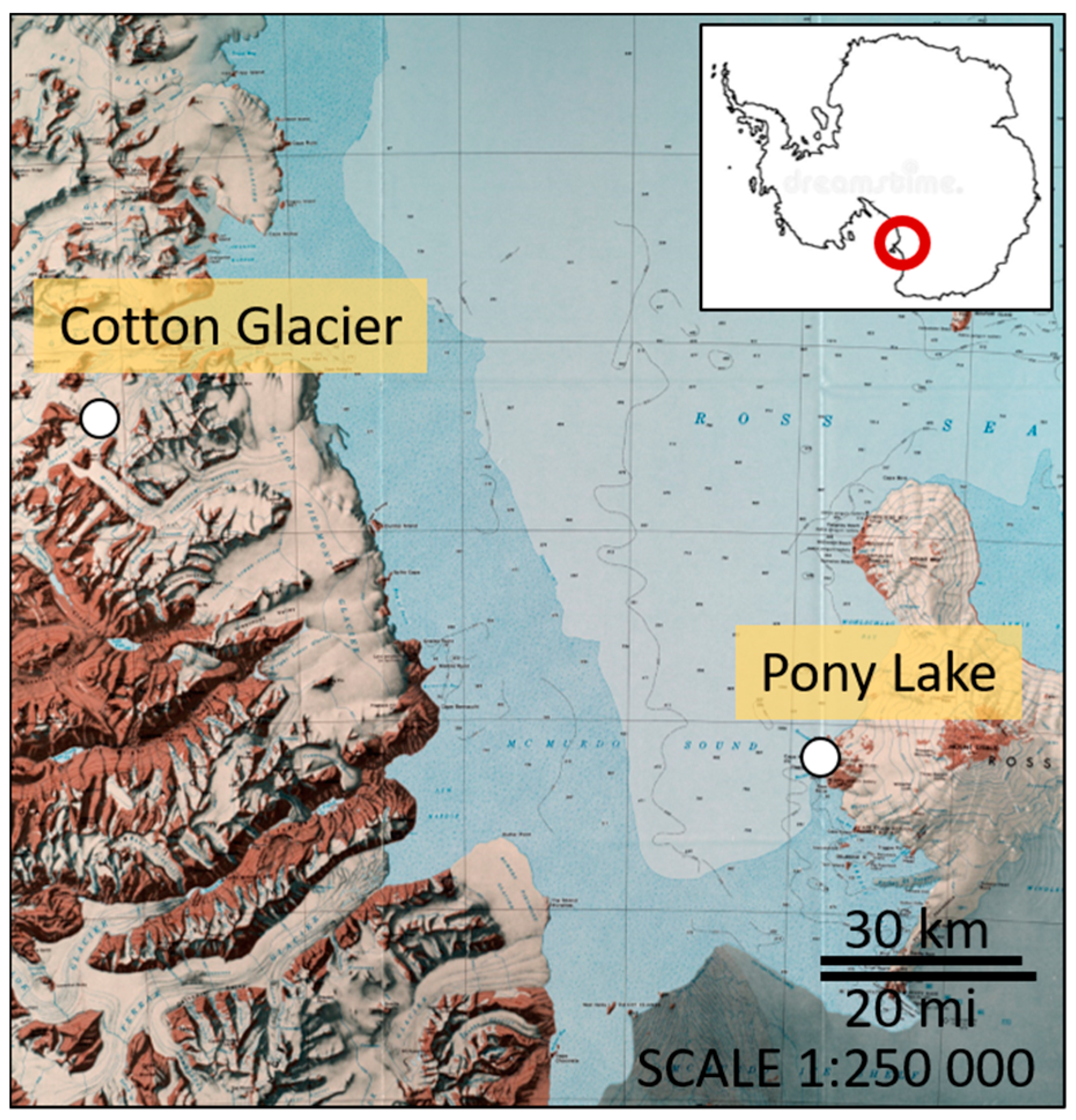
2.1. Sampling Locations of Microbial Isolates
2.2. Bacterial Isolation and Phylogenetic Characterization
2.3. Bacterial Growth
Growth Media for Biosurfactant Production
2.4. Biosurfactant Screening Methods
2.4.1. Oil Displacement Method
2.4.2. Emulsion Index Test (E24)
2.4.3. Growth on Crude Oil
2.5. Biosurfactant Classification
2.5.1. Large Scale Biosurfactant Production
2.5.2. MALDI Mass Spectrometry and Thin-Layer Chromatography
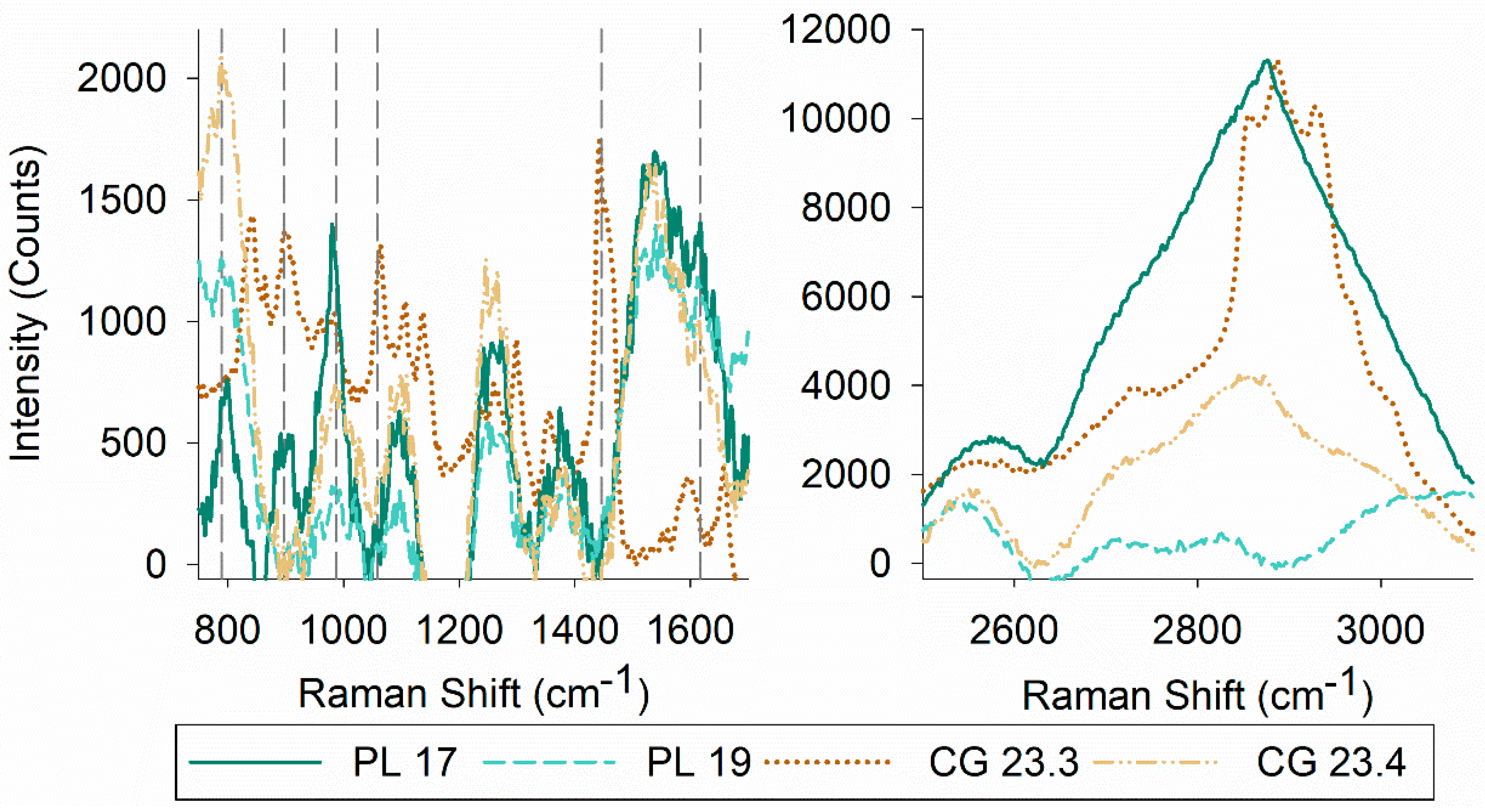
2.5.3. Raman Spectroscopy
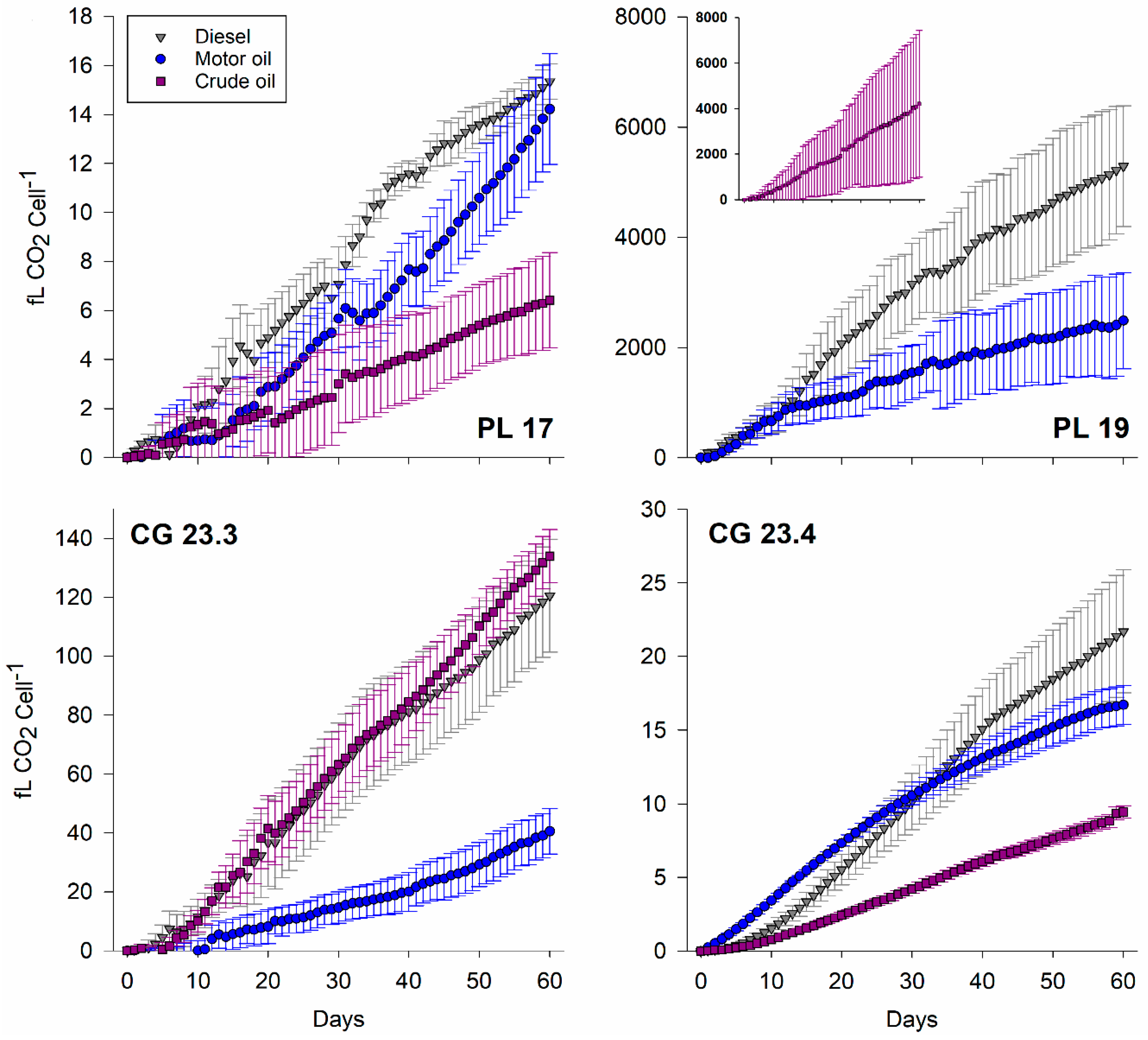
2.6. Respirometry
3. Results
3.1. Phylogeny of Biosurfactants Producing Bacterial Isolates
3.2. Screening for Biosurfactants
3.3. Characteristics of Biosurfactants
3.4. Respiration Activity
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Schramm, L.L.; Stasiuk, E.N.; Marangoni, D.G. 2 Surfactants and their applications. Annu Rep. Prog Chem. Sect. C Phys. Chem 2003, 99, 3–48. [Google Scholar] [CrossRef]
- Negm, N.A.; Aiad, I.A. Synthesis and characterization of multifunctional surfactants in oil-field protection applications. J. Surfactants Detreg 2007, 10, 87–92. [Google Scholar] [CrossRef]
- Barani, H.; Montazer, M. A review on applications of liposomes in textile processing. J. Liposome Res. 2008, 18, 249–262. [Google Scholar] [CrossRef] [PubMed]
- Kumar, N.; Tyagi, R. Industrial applications of dimeric surfactants: A review. J. Disper. Sci. Technol. 2014, 35, 205–214. [Google Scholar] [CrossRef]
- Edwards, K.R.; Lepo, J.E.; Lewis, M.A. Toxicity comparison of biosurfactants and synthetic surfactants used in oil spill remediation to two estuarine species. Mar. Pollut Bull. 2003, 46, 1309–1316. [Google Scholar] [CrossRef]
- Haigh, S.D. A review of the interaction of surfactants with organic contaminants in soil. Sci. Total Environ. 1996, 185, 161–170. [Google Scholar] [CrossRef]
- Palmer, M.; Hatley, H. The role of surfactants in wastewater treatment: Impact, removal and future techniques: A critical review. Water Res. 2018, 147, 60–72. [Google Scholar] [PubMed]
- Rahman, P.K.; Gakpe, E. Production, characterisation and applications of biosurfactants-Review. Biotechnology 2008, 7, 360–370. [Google Scholar] [CrossRef]
- Marchant, R.; Banat, I.M. Biosurfactants: A sustainable replacement for chemical surfactants? Biotechnol. Lett. 2012, 34, 1597–1605. [Google Scholar] [CrossRef]
- Cheok, C.Y.; Salman, H.A.; Sulaiman, R. Extraction and quantification of saponins: A review. Food Res. Int. 2014, 59, 16–40. [Google Scholar] [CrossRef]
- Santos, D.K.; Rufino, R.D.; Luna, J.M.; Santos, V.A.; Sarubbo, L.A. Biosurfactants: Multifunctional biomolecules of the 21st century. Int. J. Mol. Sci. 2016, 17, 401. [Google Scholar] [CrossRef] [PubMed]
- Pacwa-Płociniczak, M.; Płaza, G.A.; Piotrowska-Seget, Z.; Cameotra, S.S. Environmental applications of biosurfactants: Recent advances. Int. J. Mol. Sci. 2011, 12, 633–654. [Google Scholar] [CrossRef] [PubMed]
- Vijayakumar, S.; Saravanan, V. Biosurfactants-types; sources and applications. Res. J. Microbiol. 2015, 10, 181. [Google Scholar]
- Akbari, S.; Abdurahman, N.H.; Yunus, R.M.; Fayaz, F.; Alara, O.R. Biosurfactants-a new frontier for social and environmental safety: A mini review. Biotechnol. Res. Innov. 2018, 2, 81–90. [Google Scholar] [CrossRef]
- Khademolhosseini, R.; Jafari, A.; Mousavi, S.M.; Hajfarajollah, H.; Noghabi, K.A.; Manteghian, M. Physicochemical characterization and optimization of glycolipid biosurfactant production by a native strain of Pseudomonas aeruginosa HAK01 and its performance evaluation for the MEOR process. RSC Adv. 2019, 9, 7932–7947. [Google Scholar] [CrossRef]
- Płaza, G.; Achal, V. Biosurfactants: Eco-friendly and innovative biocides against biocorrosion. Int. J. Mol. Sci. 2020, 21, 2152. [Google Scholar] [CrossRef]
- Shekhar, S.; Sundaramanickam, A.; Balasubramanian, T. Biosurfactant producing microbes and their potential applications: A review. Crit. Rev. Environ. Sci. Technol. 2015, 45, 1522–1554. [Google Scholar] [CrossRef]
- Singh, P.; Cameotra, S.S. Potential applications of microbial surfactants in biomedical sciences. Trends Biotechnol. 2004, 22, 142–146. [Google Scholar] [CrossRef]
- Nitschke, M.; Costa, S.G. Biosurfactants in food industry. Trends Food Sci. Technol. 2007, 18, 252–259. [Google Scholar] [CrossRef]
- Banat, I.M.; Franzetti, A.; Gandolfi, I.; Bestetti, G.; Martinotti, M.G.; Fracchia, L.; Smyth, T.J.; Marchant, R. Microbial biosurfactants production; applications and future potential. Appl. Microbiol. Biotechnol. 2010, 87, 427–444. [Google Scholar] [CrossRef]
- Bhadoriya, S.S.; Madoriya, N.; Shukla, K.; Parihar, M.S. Biosurfactants: A new pharmaceutical additive for solubility enhancement and pharmaceutical development. Biochem. Pharmacol. 2013, 2, 113. [Google Scholar] [CrossRef]
- Campos, J.M.; Montenegro Stamford, T.L.; Sarubbo, L.A.; de Luna, J.M.; Rufino, R.D.; Banat, I.M. Microbial biosurfactants as additives for food industries. Biotechnol. Prog. 2013, 29, 1097–1108. [Google Scholar] [CrossRef] [PubMed]
- Sekhon Randhawa, K.K.; Rahman, P.K. Rhamnolipid biosurfactants-past; present; and future scenario of global market. Front. Microbiol. 2014, 5, 454. [Google Scholar] [CrossRef] [PubMed]
- Souza, E.C.; Vessoni-Penna, T.C.; de Souza Oliveira, R.P. Biosurfactant-enhanced hydrocarbon bioremediation: An overview. Int. Biodeter. Biodegr. 2014, 89, 88–94. [Google Scholar] [CrossRef]
- Fracchia, L.; Banat, J.J.; Cavallo, M.; Banat, I.M. Potential therapeutic applications of microbial surface-active compounds. AIMS Bioeng. 2015, 2, 144–162. [Google Scholar] [CrossRef]
- De Almeida, D.G.; Soares Da Silva, R.D.; Luna, J.M.; Rufino, R.D.; Santos, V.A.; Banat, I.M.; Sarubbo, L.A. Biosurfactants: Promising molecules for petroleum biotechnology advances. Front. Microbiol. 2016, 7, 1718. [Google Scholar] [CrossRef]
- Abdel-Mawgoud, A.M.; Lépine, F.; Déziel, E. Rhamnolipids: Diversity of structures, microbial origins and roles. Appl. Microbiol. Biotechnol. 2010, 86, 1323–1336. [Google Scholar] [CrossRef]
- Perfumo, A.; Banat, I.M.; Marchant, R. Going green and cold: Biosurfactants from low-temperature environments to biotechnology applications. Trends Biotechnol. 2018, 36, 277–289. [Google Scholar] [CrossRef]
- Kitamoto, D.; Yanagishita, H.; Endo, A.; Nakaiwa, M.; Nakane, T.; Akiya, T. Remarkable antiagglomeration effect of a yeast biosurfactant, diacylmannosylerythritol, on ice-water slurry for cold thermal storage. Biotechnol. Prog. 2001, 17, 362–365. [Google Scholar] [CrossRef]
- Borah, D. Microbial Bioremediation of Petroleum Hydrocarbon: An Overview. In Microbial Action on Hydrocarbons; Kumar, V., Kumar, M., Prasad, R., Eds.; Springer: Singapore, 2018; pp. 321–341. [Google Scholar]
- Patel, S.; Homaei, A.; Patil, S.; Daverey, A. Microbial biosurfactants for oil spill remediation: Pitfalls and potentials. Appl. Microbiol. Biotechnol. 2019, 103, 27–37. [Google Scholar] [CrossRef]
- Du, M.; Kessler, J.D. Assessment of the spatial and temporal variability of bulk hydrocarbon respiration following the Deepwater Horizon oil spill. Environ. Sci. Technol. 2012, 46, 10499–10507. [Google Scholar] [CrossRef] [PubMed]
- Washburn, T.W.; Yoskowitz, D.W.; Montagna, P.A. Valuing nature waste removal in the offshore environment following the Deepwater Horizon oil spill. Front. Mar. Sci. 2018, 5, 477. [Google Scholar] [CrossRef]
- Thavasi, R.; Jayalakshmi, S.; Balasubramanian, T.; Banat, I.M. Production and characterization of a glycolipid biosurfactant from Bacillus megaterium using economically cheaper sources. World J. Microb. Biotechnol. 2008, 24, 917–925. [Google Scholar] [CrossRef]
- Dadrasnia, A.; Ismail, S. Biosurfactant production by Bacillus salmalaya for lubricating oil solubilization and biodegradation. Int. J. Environ. Res. Public Health 2015, 12, 9848–9863. [Google Scholar] [CrossRef]
- Kumar, R.; Das, A.J. Rhamnolipid Biosurfactant: Recent Trends in Production and Application; Springer: Singapore, 2018. [Google Scholar]
- Martinez-Toledo, A.; Rodriguez-Vazquez, R. Culture media formulation and growth conditions for biosurfactants production by bacteria. Int. J. Environ. Sci. Nat. Resour. 2018, 10, 117–125. [Google Scholar]
- Tripathi, L.; Irorere, V.U.; Marchant, R.; Banat, I.M. Marine derived biosurfactants: A vast potential future resource. Biotechnol. Lett. 2018, 40, 1441–1457. [Google Scholar] [CrossRef] [PubMed]
- Schultz, J.; Rosado, A.S. Extreme environments: A source of biosurfactants for biotechnological applications. Extremophiles 2020, 24, 189–206. [Google Scholar] [CrossRef]
- Dieser, M.; Foreman, C.M.; Jaros, C.; Lisle, J.T.; Greenwood, M.C.; Laybourn-Parry, J.; Miller, P.L.; Chin, Y.P.; Mcknight, D.M. Physicochemical and biological dynamics in a coastal Antarctic lake as it transitions from frozen to open water. Antarct. Sci. 2013, 25, 663–675. [Google Scholar] [CrossRef][Green Version]
- Foreman, C.M.; Cory, R.M.; Morris, C.E.; SanClements, M.D.; Smith, H.J.; Lisle, J.T.; Miller, P.L.; Chin, Y.P.; McKnight, D.M. Microbial growth under humic-free conditions in a supraglacial stream system on the Cotton Glacier; Antarctica. Environ. Res. Lett. 2013, 8, 035022. [Google Scholar] [CrossRef]
- Stackebrandt, E.; Liesack, W.; Goebel, B.M. Bacterial diversity in a soil sample from a subtropical Australian environment as determined by 16S rDNA analysis. Faseb. J. 1993, 7, 232–236. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glöckner, F.O. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 2012, 41, D590–D596. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef] [PubMed]
- Parthipan, P.; Preetham, E.; Machuca, L.L.; Rahman, P.K.; Murugan, K.; Rajasekar, A. Biosurfactant and degradative enzymes mediated crude oil degradation by bacterium Bacillus subtilis A1. Front. Microbiol. 2017, 8, 193. [Google Scholar] [CrossRef] [PubMed]
- Cooper, D.G.; Goldenberg, B.G. Surface-active agents from two Bacillus species. Appl. Environ. Microbiol. 1987, 53, 224–229. [Google Scholar] [CrossRef] [PubMed]
- Nozomu, O.; Makoto, I. GlycoPOD 2014. Available online: https://jcggdb.jp/GlycoPOD (accessed on 28 July 2020).
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2018. [Google Scholar]
- Smith, H.; Akiyama, T.; Foreman, C.; Franklin, M.; Woyke, T.; Teshima, H.; Davenport, K.; Daligault, H.; Erkkila, T.; Goodwin, L.; et al. Draft genome sequence and description of Janthinobacterium sp. strain CG3; a psychrotolerant Antarctic supraglacial stream bacterium. Genome Announc. 2013, 1, e00960-13. [Google Scholar] [CrossRef] [PubMed]
- Cho, Y.J.; Jung, Y.J.; Hong, S.G.; Kim, O.S. Complete genome sequence of a psychrotolerant denitrifying bacterium; Janthinobacterium svalbardensis PAMC 27463. Genome Announc. 2017, 5, e01178-17. [Google Scholar] [CrossRef]
- Gong, X.; Skrivergaard, S.; Korsgaard, B.S.; Schreiber, L.; Marshall, I.P.; Finster, K.; Schramm, A. High quality draft genome sequence of Janthinobacterium psychrotolerans sp. nov.; isolated from a frozen freshwater pond. Stand. Genomic Sci. 2017, 12, 8. [Google Scholar] [CrossRef]
- Bakermans, C.; Ayala-del-Río, H.L.; Ponder, M.A.; Vishnivetskaya, T.; Gilichinsky, D.; Thomashow, M.F.; Tiedje, J.M. Psychrobacter cryohalolentis sp. nov. and Psychrobacter arcticus sp. nov.; isolated from Siberian permafrost. Int. J. Syst. Evol. Microbiol. 2006, 56, 1285–1291. [Google Scholar] [CrossRef]
- Spröer, C.; Mendrock, U.; Swiderski, J.; Lang, E.; Stackebrandt, E. The phylogenetic position of Serratia; Buttiauxella and some other genera of the family Enterobacteriaceae. Int. J. Syst. Evol. Microbiol. 1999, 49, 1433–1438. [Google Scholar] [CrossRef]
- Nitschke, M.; Costa, S.G.V.A.O.; Haddad, R.; Gonçalves, L.A.G.; Eberlin, M.N.; Contiero, J. Oil wastes as unconventional substrates for rhamnolipid biosurfactant production by Pseudomonas aeruginosa LBI. Biotechnol. Prog. 2005, 21, 1562–1566. [Google Scholar] [CrossRef]
- Lindum, P.W.; Anthoni, U.; Christophersen, C.; Eberl, L.; Molin, S.; Givskov, M. N-Acyl-L-homoserine lactone autoinducers control production of an extracellular lipopeptide biosurfactant required for swarming motility of Serratia liquefaciens MG1. J. Bacteriol. 1998, 180, 6384–6388. [Google Scholar] [CrossRef] [PubMed]
- San Keskin, N.O.; Han, D.; Ozkan, A.D.; Angun, P.; Umu, O.C.; Tekinay, T. Production and structural characterization of biosurfactant produced by newly isolated Staphylococcus xylosus STF1 from petroleum contaminated soil. J. Petrol. Sci. Eng. 2015, 133, 689–694. [Google Scholar] [CrossRef]
- Pacheco, G.J.; Reis, R.S.; Fernandes, A.C.; da Rocha, S.L.; Pereira, M.D.; Perales, J.; Freire, D.M. Rhamnolipid production: Effect of oxidative stress on virulence factors and proteome of Pseudomonas aeruginosa PA1. Appl. Microbiol. Biotechnol. 2012, 95, 1519–1529. [Google Scholar] [CrossRef] [PubMed]
- Chong, H.; Li, Q. Microbial production of rhamnolipids: Opportunities; challenges and strategies. Microb. Cell Fact. 2017, 16, 137. [Google Scholar] [CrossRef] [PubMed]
- Gong, Z.; Peng, Y.; Wang, Q. Rhamnolipid production; characterization and fermentation scale-up by Pseudomonas aeruginosa with plant oils. Biotechnol. Lett. 2015, 37, 2033–2038. [Google Scholar] [CrossRef]
- Li, Z.; Zhang, Y.; Lin, J.; Wang, W.; Li, S. High-yield di-rhamnolipid production by Pseudomonas aeruginosa YM4 and its potential application in MEOR. Molecules 2019, 24, 1433. [Google Scholar] [CrossRef]
- Cai, Q.; Zhang, B.; Chen, B.; Zhu, Z.; Lin, W.; Cao, T. Screening of biosurfactant producers from petroleum hydrocarbon contaminated sources in cold marine environments. Mar. Pollut. Bull. 2014, 86, 402–410. [Google Scholar] [CrossRef]
- Malavenda, R.; Rizzo, C.; Michaud, L.; Gerçe, B.; Bruni, V.; Syldatk, C.; Hausmann, R.; Giudice, A.L. Biosurfactant production by Arctic and Antarctic bacteria growing on hydrocarbons. Polar Biol. 2015, 38, 1565–1574. [Google Scholar] [CrossRef]
- Amodu, O.S.; Ntwampe, S.K.; Ojumu, T.V. Emulsification of hydrocarbons by biosurfactant: Exclusive use of agrowaste. BioResources 2014, 9, 3508–3525. [Google Scholar] [CrossRef]
- Sugiyama, J.; Sugiyama, Y.; Iizuka, H.; Torii, T. Report of the Japanese summer parties in dry valleys, Victoria Land, 1963–1965; IV–Mycological studies of the Antarctic fungi; Part 2–Mycoflora of Lake Vanda, an ice-free lake. Antarct. Rec. 1967, 28, 23–32. [Google Scholar]
- Morita, T.; Konishi, M.; Fukuoka, T.; Imura, T.; Kitamoto, H.K.; Kitamoto, D. Characterization of the genus Pseudozyma by the formation of glycolipid biosurfactants; mannosylerythritol lipids. FEMS Yeast Res. 2007, 7, 286–292. [Google Scholar] [CrossRef] [PubMed]
- Irorere, V.U.; Tripathi, L.; Marchant, R.; McClean, S.; Banat, I.M. Microbial rhamnolipid production: A critical re-evaluation of published data and suggested future publication criteria. Appl. Microbiol. Biotechnol. 2017, 101, 3941–3951. [Google Scholar] [CrossRef] [PubMed]
- Prabagaran, S.R.; Manorama, R.; Delille, D.; Shivaji, S. Predominance of Roseobacter; Sulfitobacter; Glaciecola and psychrobacter in seawater collected off Ushuaia; Argentina; Sub-Antarctica. FEMS Microbiol. Ecol. 2007, 59, 342–355. [Google Scholar] [CrossRef]
- Lo Giudice, A.; Casella, P.; Caruso, C.; Mangano, S.; Bruni, V.; De Domenico, M.; Michaud, L. Occurrence and characterization of psychrotolerant hydrocarbon-oxidizing bacteria from surface seawater along the Victoria Land coast (Antarctica). Polar Biol. 2010, 33, 929–943. [Google Scholar] [CrossRef]
- Azevedo, J.S.; Correia, A.; Henriques, I. Molecular analysis of the diversity of genus Psychrobacter present within a temperate estuary. FEMS Microbiol. Ecol. 2013, 84, 451–460. [Google Scholar] [CrossRef][Green Version]
- Révész, F.; Figueroa-Gonzalez, P.A.; Probst, A.J.; Kriszt, B.; Banerjee, S.; Szoboszlay, S.; Maróti, G.; Táncsics, A. Microaerobic conditions caused the overwhelming dominance of Acinetobacter spp. and the marginalization of Rhodococcus spp. in diesel fuel/crude oil mixture-amended enrichment cultures. Arch. Microbiol. 2020, 202, 329–342. [Google Scholar] [CrossRef]
- Bodour, A.A.; Wang, J.M.; Brusseau, M.L.; Maier, R.M. Temporal change in culturable phenanthrene degraders in response to long-term exposure to phenanthrene in a soil column system. Environ. Microbiol 2003, 5, 888–895. [Google Scholar] [CrossRef]
- Guo, G.; Tian, F.; Ding, K.; Wang, L.; Liu, T.; Yang, F. Effect of a bacterial consortium on the degradation of polycyclic aromatic hydrocarbons and bacterial community composition in Chinese soils. Int. Biodeter. Biodegr. 2017, 123, 56–62. [Google Scholar] [CrossRef]
- Wongsa, P.; Tanaka, M.; Ueno, A.; Hasanuzzaman, M.; Yumoto, I.; Okuyama, H. Isolation and characterization of novel strains of Pseudomonas aeruginosa and Serratia marcescens possessing high efficiency to degrade gasoline; kerosene; diesel oil; and lubricating oil. Curr. Microbiol. 2004, 49, 415–422. [Google Scholar] [CrossRef]
- Rajasekar, A.; Balasubramanian, R.; VM Kuma, J. Role of hydrocarbon degrading bacteria Serratia marcescens ACE2 and Bacillus cereus ACE4 on corrosion of carbon steel API 5LX. Ind. Eng. Chem. Res. 2011, 50, 10041–10046. [Google Scholar] [CrossRef]
| Isolate | Location | Closest Relative | % ID | Accession # |
|---|---|---|---|---|
| PL 17 | Pony Lake | Serratia quinivorans 4364 (NR_037112.1) | 99 | MT594460 |
| PL 19 | Pony Lake | Psychrobacter arcticus strain 273-4 (NR_075054.1) | 98 | MT594461 |
| CG 23.3 | Cotton Glacier | Janthinobacterium svalbardensis JA-1 (NR_132608.1) | 98 | MT594462 |
| CG 23.4 | Cotton Glacier | Janthinobacterium svalbardensis JA-1 (NR_132608.1) | 99 | MT594463 |
| Isolate | E24 (%) | Oil Displacement (cm) | Dried Supernatant (g/L) | Growth on Crude Oil | MALDI Peak Matches (m/z Ratio) |
|---|---|---|---|---|---|
| PL 17 | 36.4 | 6.4 ± 0.4 | 8.5 |  | Sophorolipid (621, 663) i Di-rhamnolipid (649, 689) i,ii |
| PL 19 | 58.3 | 3.5 ± 0.1 | 9.9 |  | Sophorolipid (663) Di-rhamnolipid (689) |
| CG 23.3 | 66.7 | 4.6 ± 1.1 | 17.3 |  | Sophorolipid (647) iii Di-rhamnolipid (689) |
| CG 23.4 | 46.2 | 3.5 ± 2.2 | 13.3 |  | Sophorolipid (663) Di-rhamnolipid (689) |
| Isolate | Diesel (fg CO2 d−1 cell−1) | Motor Oil (fg CO2 d−1 cell−1) | Crude Oil (fg CO2 d−1 cell−1) |
|---|---|---|---|
| PL 17 | 1.0 ± 0.1 | 0.9 ± 0.2 | 0.0 ± 0.1 |
| PL 19 | 344.0 ± 81.5 | 144.8 ± 59.7 | 271.6 ± 199.1 |
| CG 23.3 | 7.8 ± 1.2 | 2.5 ± 0.5 | 8.7 ± 0.5 |
| CG 23.4 | 1.5 ± 0.3 | 1.1 ± 0.1 | 0.6 ± 0.04 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Trudgeon, B.; Dieser, M.; Balasubramanian, N.; Messmer, M.; Foreman, C.M. Low-Temperature Biosurfactants from Polar Microbes. Microorganisms 2020, 8, 1183. https://doi.org/10.3390/microorganisms8081183
Trudgeon B, Dieser M, Balasubramanian N, Messmer M, Foreman CM. Low-Temperature Biosurfactants from Polar Microbes. Microorganisms. 2020; 8(8):1183. https://doi.org/10.3390/microorganisms8081183
Chicago/Turabian StyleTrudgeon, Benjamin, Markus Dieser, Narayanaganesh Balasubramanian, Mitch Messmer, and Christine M. Foreman. 2020. "Low-Temperature Biosurfactants from Polar Microbes" Microorganisms 8, no. 8: 1183. https://doi.org/10.3390/microorganisms8081183
APA StyleTrudgeon, B., Dieser, M., Balasubramanian, N., Messmer, M., & Foreman, C. M. (2020). Low-Temperature Biosurfactants from Polar Microbes. Microorganisms, 8(8), 1183. https://doi.org/10.3390/microorganisms8081183