Temporal Dynamics of Bacterial Communities in a Pilot-Scale Vermireactor Fed with Distilled Grape Marc
Abstract
1. Introduction
2. Materials and Methods
2.1. Distilled Grape Marc
2.2. Vermicomposting Set-Up and Sampling Design
2.3. Microbial Activity
2.4. DNA Sequencing and Bioinformatic Analyses
3. Results
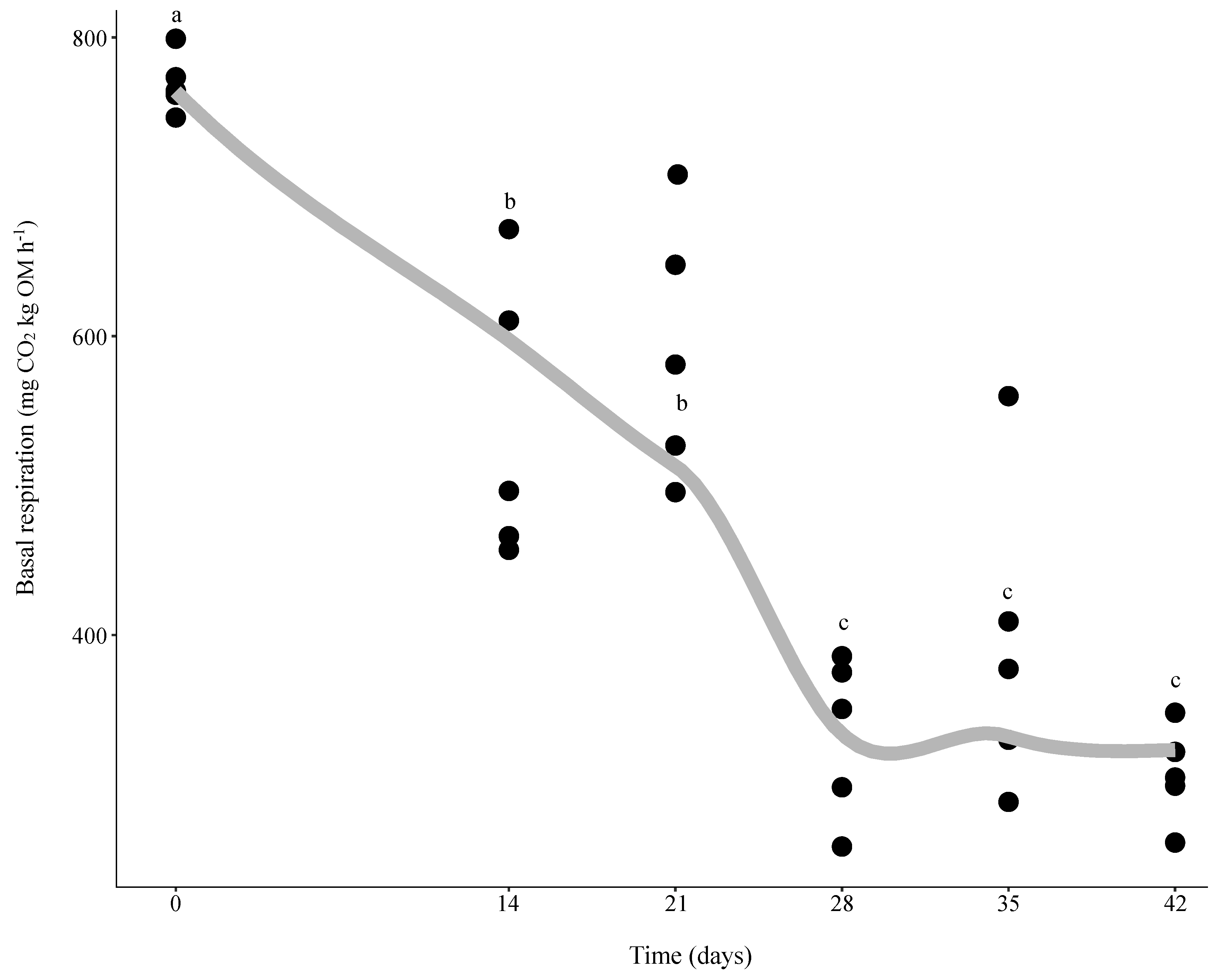
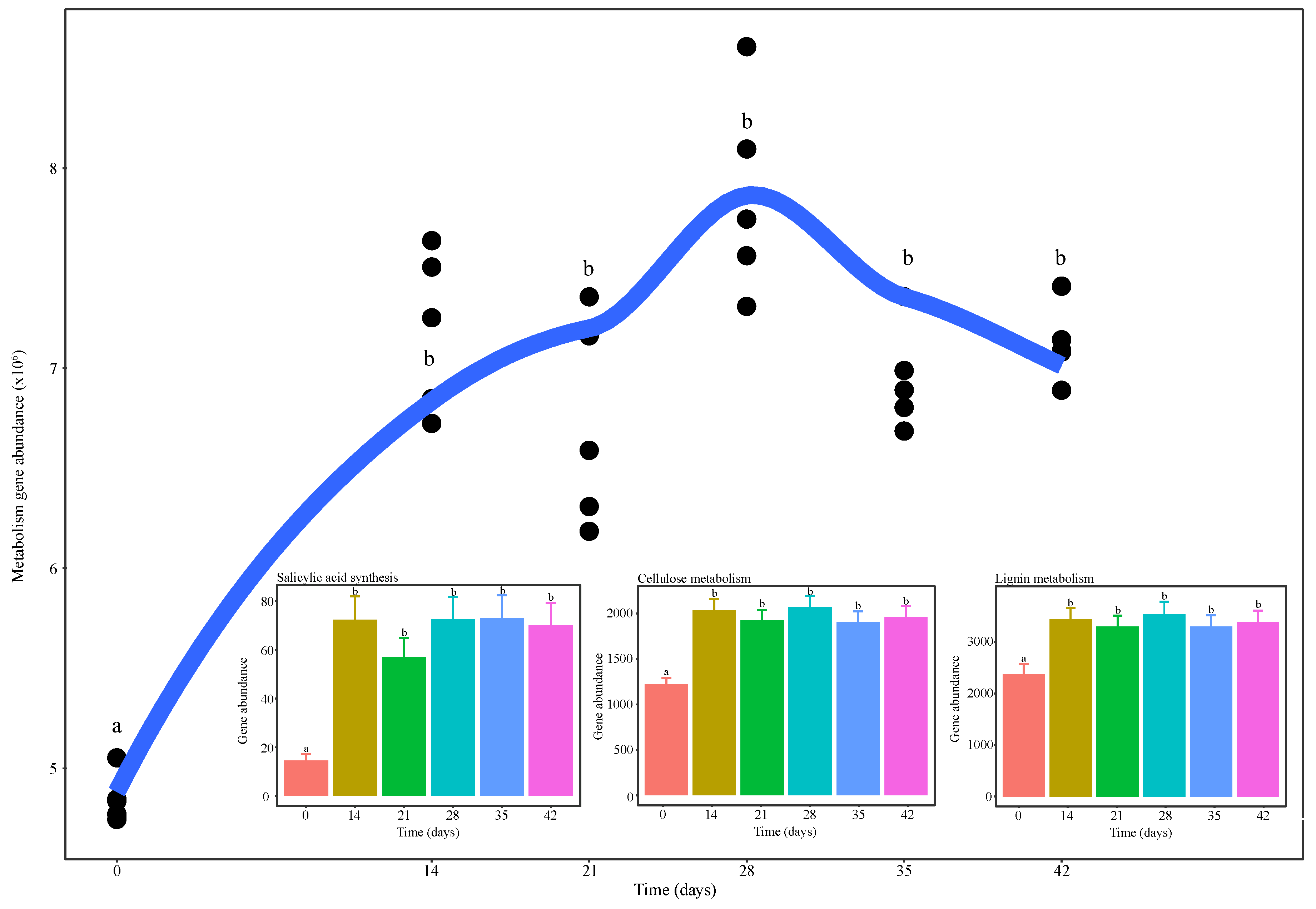
3.1. Changes in Microbial Activity during Vermicomposting of Distilled Grape Marc
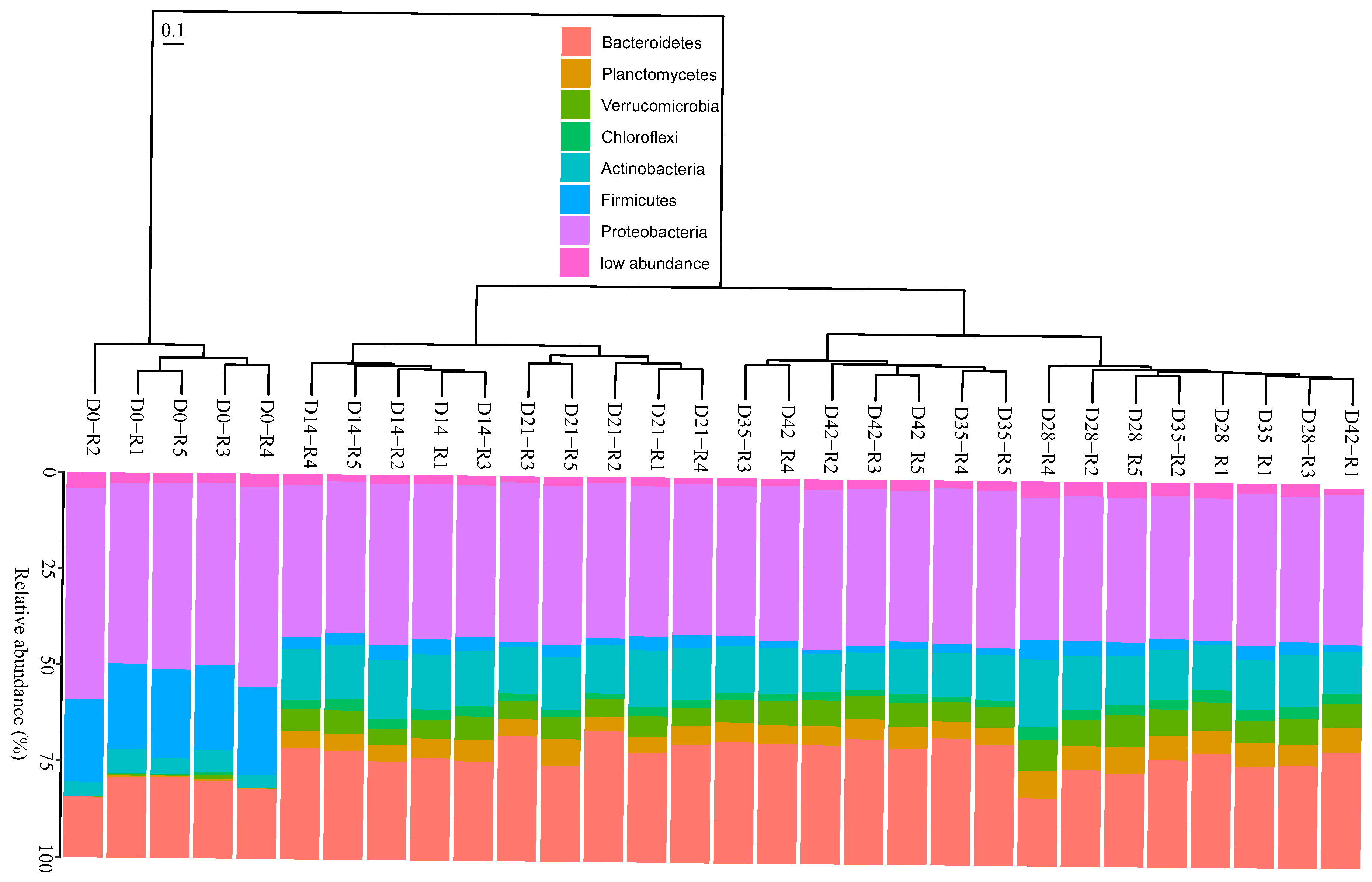
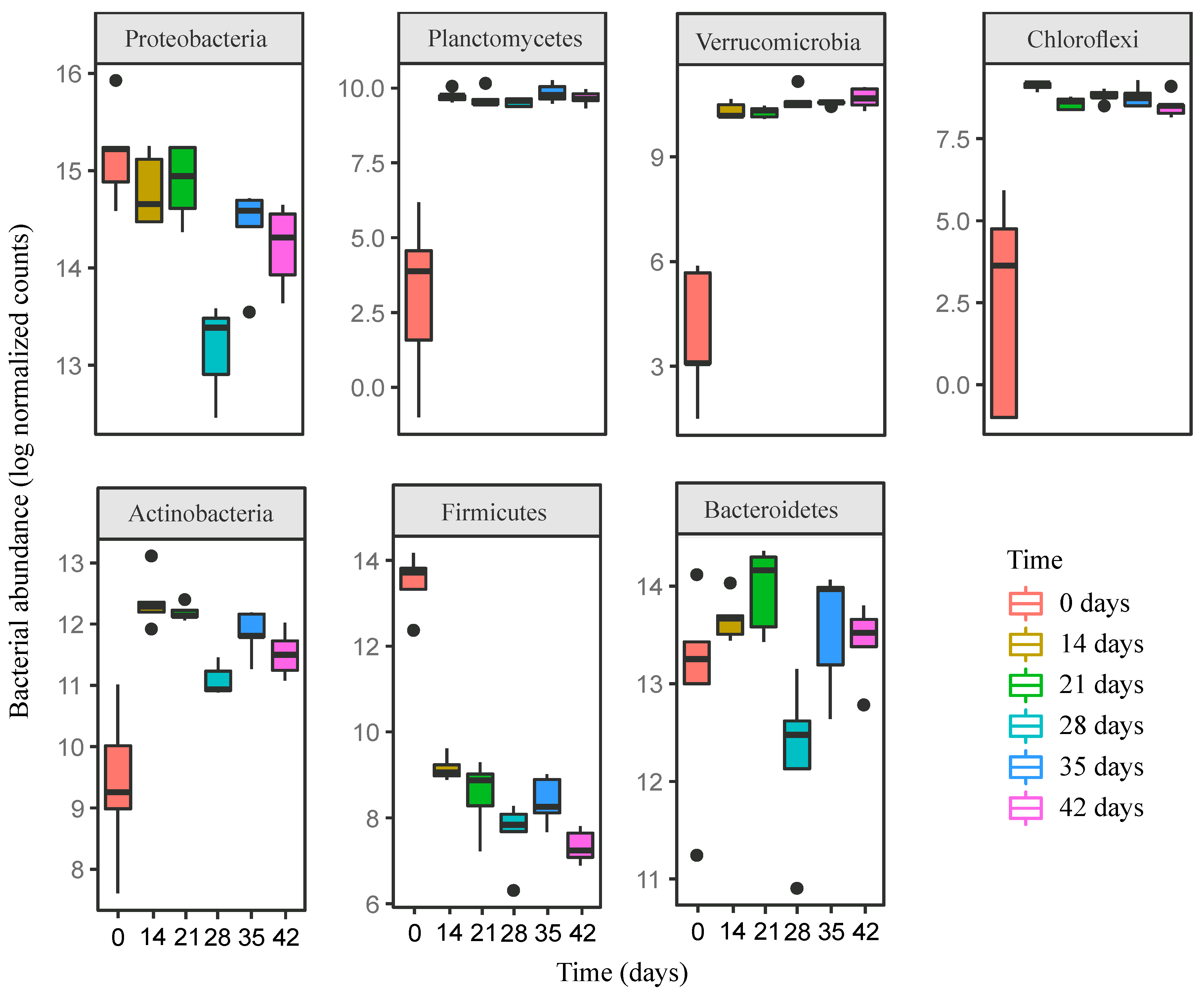
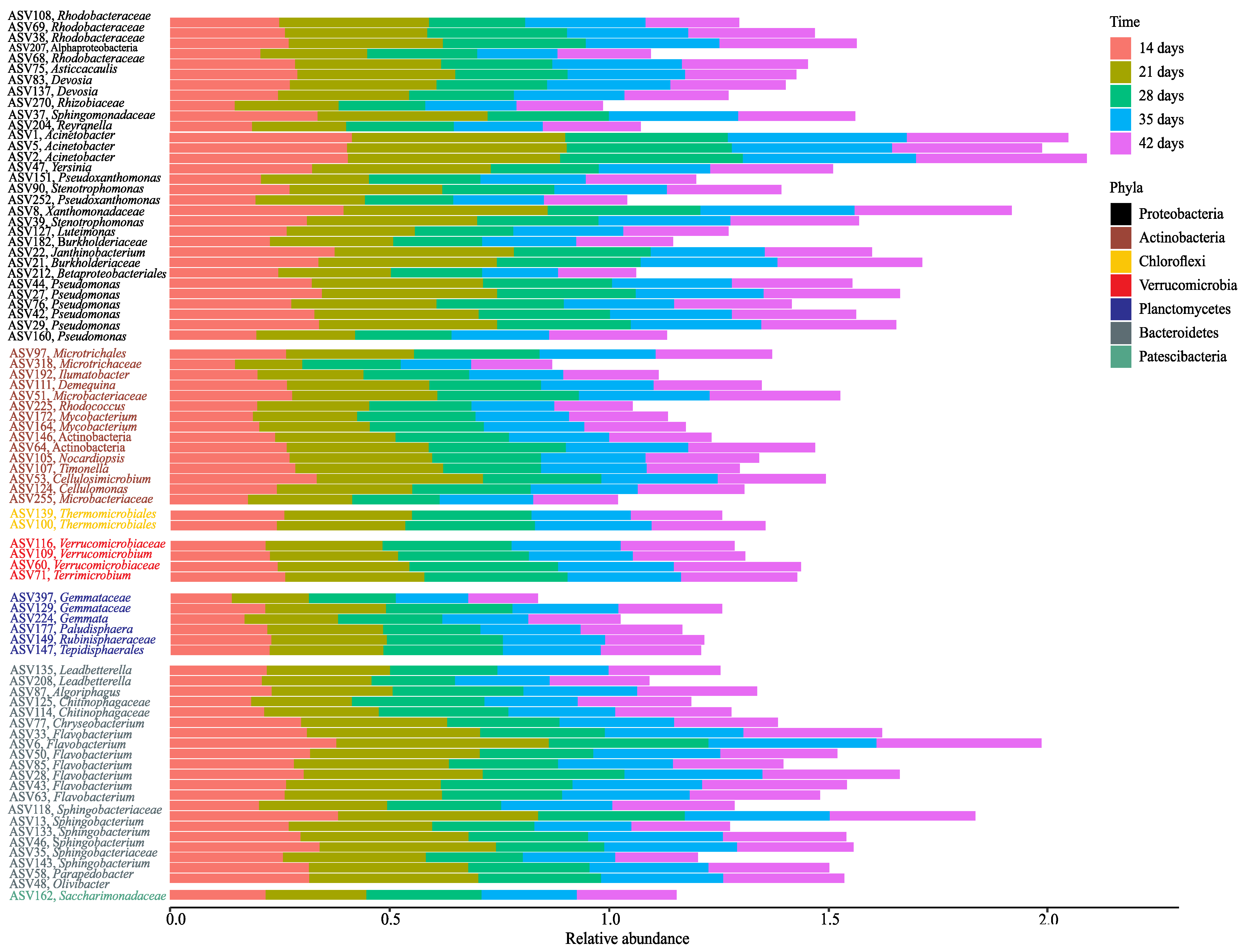
3.2. Changes in Bacterial Community Composition during Vermicomposting of Distilled Grape Marc
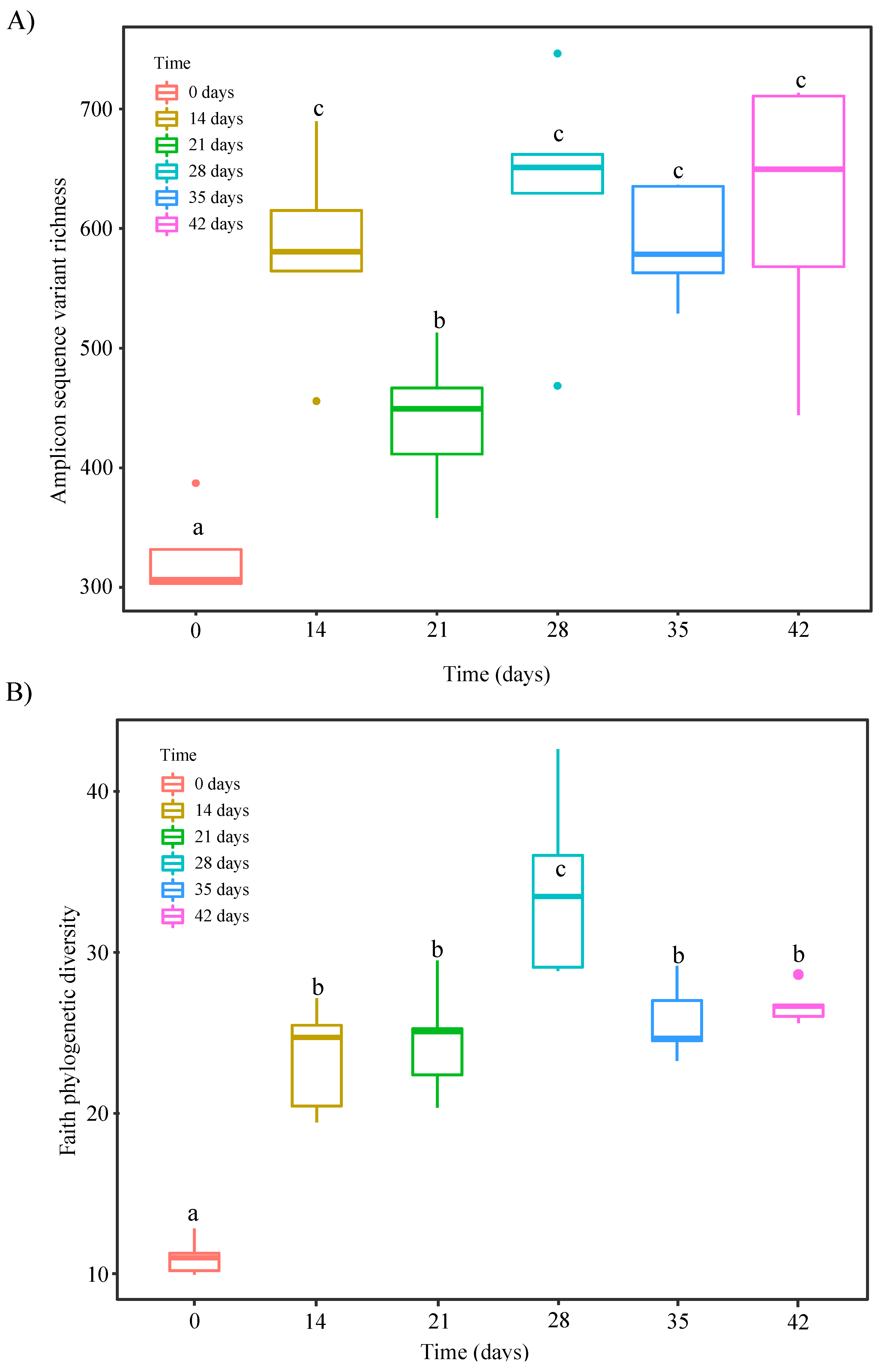
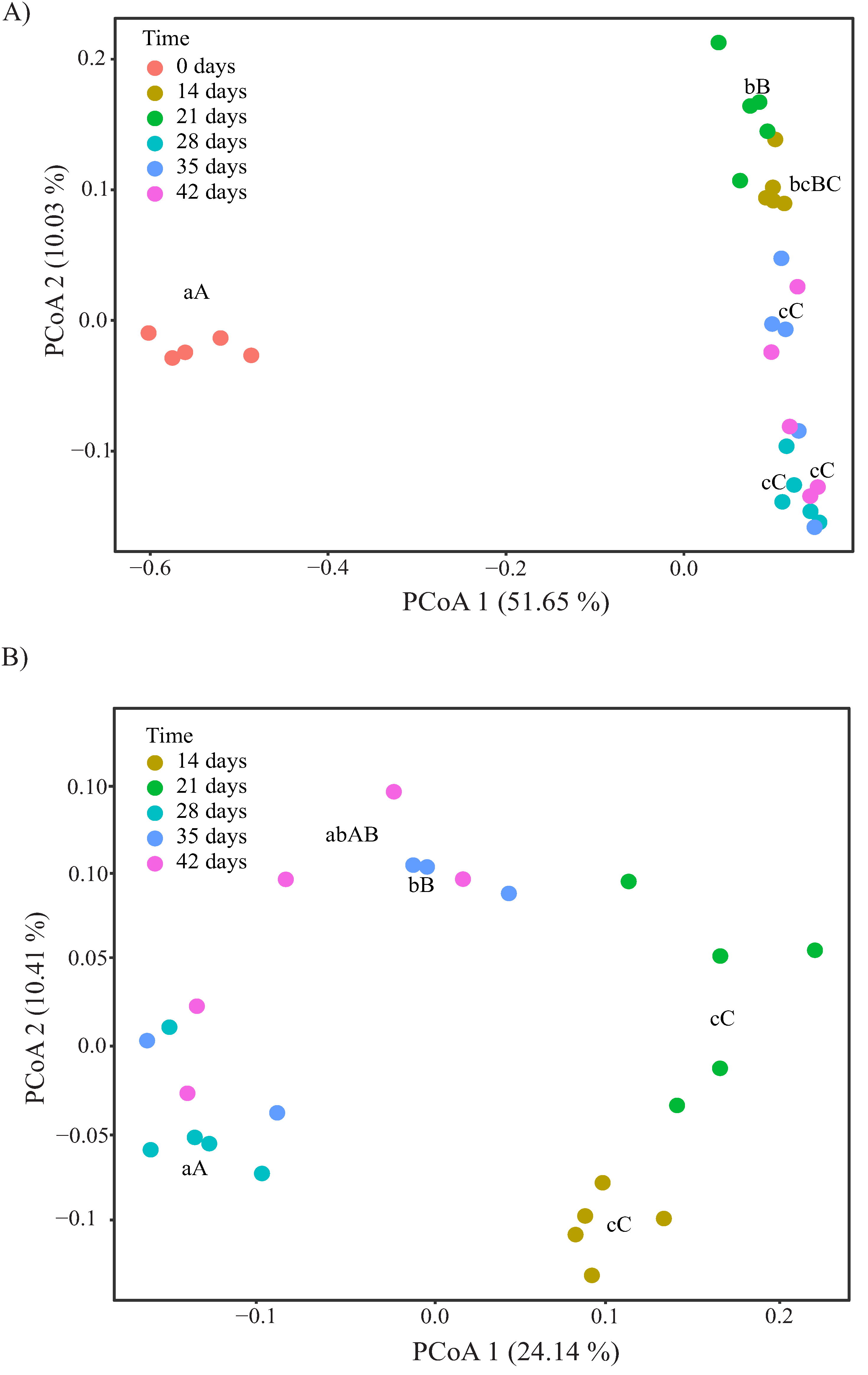
3.3. Changes in α- and β-Diversity during Vermicomposting of Distilled Grape Marc
3.4. Core bacterial Microbiome during Vermicomposting of Distilled Grape Marc
3.5. Functional Diversity of Bacterial Communities during Vermicomposting of Distilled Grape Marc
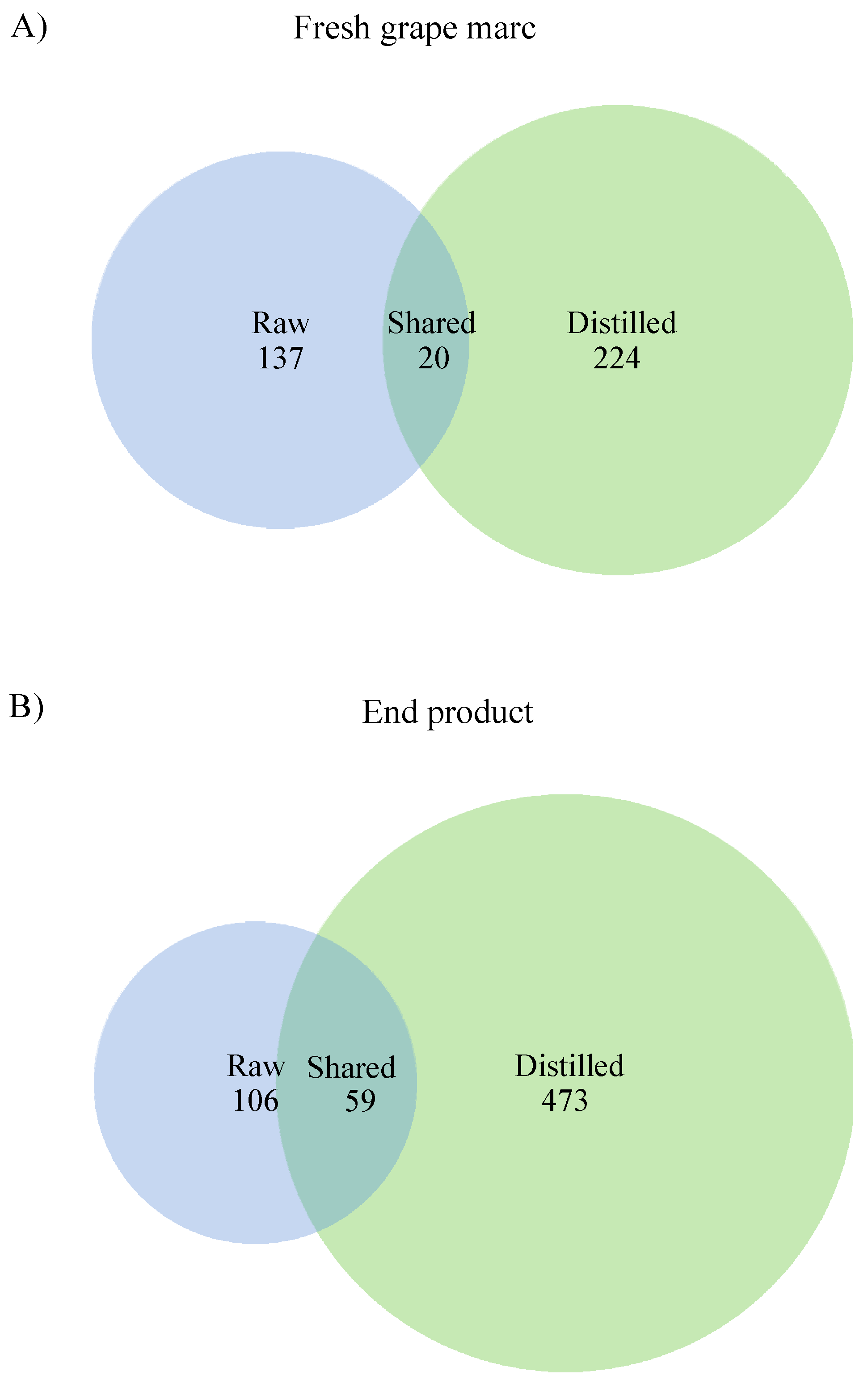
3.6. Comparison between Raw and Distilled Grape Marc Bacterial Community Composition and Diversity
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Corbin, K.R.; Hsieh, Y.S.Y.; Betts, N.S.; Byrt, C.S.; Henderson, M.; Stork, J.; DeBolt, S.; Fincher, G.B.; Burton, R.A. Grape marc as a source of carbohydrates for bioethanol: Chemical composition, pre-treatment and saccharification. Bioresour. Technol. 2015, 193, 76–83. [Google Scholar] [CrossRef]
- Domínguez, J.; Martínez-Cordeiro, H.; Alvarez-Casas, M.; Lores, M. Vermicomposting grape marc yields high quality organic biofertiliser and bioactive polyphenols. Waste Manag. Res. 2014, 32, 1235–1240. [Google Scholar] [CrossRef]
- Domínguez, J.; Martínez-Cordeiro, H.; Lores, M. Earthworms and Grape Marc: Simultaneous Production of a High-Quality Biofertilizer and Bioactive-Rich Seeds. In Grape and Wine Biotechnology; Morata, A., Loira, I., Eds.; Intech Open Science: Rijeka, Croatia, 2016; pp. 167–183. [Google Scholar]
- Domínguez, J.; Sánchez-Hernández, J.C.; Lores, M. Vermicomposting of Winemaking Byproducts. In Handbook of Grape Processing Byproducts; Galanakis, C.M., Ed.; Academic Press; Elsevier: London, UK, 2017; pp. 55–78. [Google Scholar]
- Gómez-Brandón, M.; Lores, M.; Martinez-Cordeiro, H.; Dominguez, J. Effectiveness of vermicomposting for bioconversion of grape marc derived from red winemaking into a value-added product. Environ. Sci. Pollut. Res. 2019. [Google Scholar] [CrossRef]
- Gómez-Brandón, M.; Lores, M.; Insam, H.; Domínguez, J. Strategies for recycling and valorization of grape marc. Crit. Rev. Biotechnol. 2019, 39, 437–450. [Google Scholar] [CrossRef] [PubMed]
- Gómez-Brandón, M.; Aira, M.; Kolbe, A.R.; de Andrade, N.; Pérez-Losada, M.; Domínguez, J. Rapid bacterial community changes during vermicomposting of grape marc derived from red winemaking. Microorganisms 2019, 7, 473. [Google Scholar] [CrossRef] [PubMed]
- Bovo, B.; Nardi, T.; Fontana, F.; Carlot, M.; Giacomini, A.; Corich, V. Acidification of grape marc for alcoholic beverage production: Effects on indigenous microflora and aroma profile after distillation. Int. J. Food Microbiol. 2012, 152, 100–106. [Google Scholar] [CrossRef] [PubMed]
- Sousa, R.M.O.F.; Amaral, C.; Fernandes, J.M.C.; Fraga, I.; Semitela, S.; Braga, F.; Coimbra, A.M.; Dias, A.A.; Bezerra, R.M.; Sampaio, A. Hazardous impact of vinasse from distilled winemaking byproducts in terrestrial plants and aquatic organisms. Ecotoxicol. Environ. Saf. 2019, 183, 109493. [Google Scholar] [CrossRef]
- Dávila, I.; Robles, E.; Egüés, I.; Labidi, J.; Gullón-Estévez, P. The Biorefinery Concept for the Industrial Valorization of Grape-Processing Byproducts. In Handbook of Grape Processing Byproducts: Sustainable Solutions; Galanakis, C.M., Ed.; Academic Press: London, UK, 2017; pp. 29–49. [Google Scholar]
- Zhang, N.; Hoadley, A.; Patel, J.; Lim, S.; Li, C. Sustainable options for the utilization of solid residues from wine production. Waste Manag. 2017, 60, 173–183. [Google Scholar] [CrossRef]
- Pinter, I.F.; Fernández, A.S.; Martínez, L.E.; Riera, N.; Fernández, M.; Aguado, G.D.; Uliarte, E.M. Exhausted grape marc and organic residues composting with polyethylene cover: Process and quality evaluation as plant substrate. J. Environ. Manag. 2019, 246, 695–705. [Google Scholar] [CrossRef]
- Paradelo, R.; Moldes, A.B.; Barral, M.T. Evoloution of organic matter during the mesophilic composting of lignocellulosic winery wastes. J. Environ. Manag. 2019, 116, 18–26. [Google Scholar] [CrossRef]
- Zhang, L.; Sun, X. Improving green waste composting by addition of sugarcane bagasse and exhausted grape marc. Bioresour. Technol. 2016, 218, 335–343. [Google Scholar] [CrossRef] [PubMed]
- Muhlack, R.A.; Potumarthi, R.; Jeffery, D.W. Sustainable wineries through waste valorization: A review of grape marc utilization for value-added products. Waste Manag. 2018, 72, 99–118. [Google Scholar] [CrossRef] [PubMed]
- García-Sánchez, M.; Taušnerová, H.; Hanč, A.; Tlustoš, P. Stabilization of different starting materials through vermicomposting in a continuous-feeding system: Changes in chemical and biological parameters. Waste Manag. 2017, 62, 33–42. [Google Scholar] [CrossRef] [PubMed]
- Částková, T.; Hanč, A. Change of the parameters of layers in a large-scale grape marc vermicomposting system with continuous feeding. Waste Manag. Res. 2018, 37, 826–832. [Google Scholar] [CrossRef] [PubMed]
- Lazcano, C.; Domínguez, J. The use of vermicompost in sustainable agriculture: Impact on plant growth and soil fertility. In Soil Nutrients; Miransari, M., Ed.; Nova Science Publishers: New York, NY, USA, 2011; pp. 230–254. [Google Scholar]
- Blouin, M.; Barrere, J.; Meyer, J.; Lartigue, S.; Barot, S.; Mathieu, J. Vermicompost significantly affects plant growth. A meta-analysis. Agron. Sustain. Dev. 2019, 39, 34. [Google Scholar] [CrossRef]
- Hanc, A.; Hrebeckova, T.; Kuzel, S. Vermicomposting of distillery residues in a vertical-flow windrow system. Waste Biomass Valor 2019, 10, 3647–3657. [Google Scholar] [CrossRef]
- Campanaro, S.; Treu, L.; Vendramin, V.; Bovo, B.; Giacomini, A.; Corich, V. Metagenomic analysis of the microbial community in fermented grape marc reveals that Lactobacillus fabifermentans is one of the dominant species: Insights into its genome structure. Appl. Microbiol. Biotechnol. 2014, 98, 6015–6037. [Google Scholar] [CrossRef]
- Yakushev, A.V.; Bubnov, I.A.; Semenov, A.M. Estimation of the effects of earthworms and initial substrates on the bacterial community in vermicomposts. Eurasian Soil Sci. 2011, 44, 1117–1124. [Google Scholar] [CrossRef]
- Aira, M.; Bybee, S.; Pérez-Losada, M.; Domínguez, J. Feeding on microbiomes: Effects of detritivory on the taxonomic and phylogenetic bacterial composition of animal manures. FEMS Microbiol. Ecol. 2015, 91, fiv117. [Google Scholar] [CrossRef]
- Aira, M.; Pérez-Losada, M.; Domínguez, J. Microbiome dynamics during cast ageing in the earthworm Aporrectodea caliginosa. Appl. Soil Ecol. 2019, 139, 56–63. [Google Scholar] [CrossRef]
- Kolbe, A.R.; Aira, M.; Gómez-Brandón, M.; Pérez-Losada, M.; Domínguez, J. Bacterial succession and functional diversity during vermicomposting of white grape marc (Vitis vinifera v. Albariño). Sci. Rep. 2019, 9, 7472. [Google Scholar] [CrossRef] [PubMed]
- Domínguez, J.; Aira, M.; Kolbe, A.R.; Gómez-Brandón, M.; Pérez-Losada, M. Changes in the composition and function of bacterial communities during vermicomposting may explain beneficial properties of vermicompost. Sci. Rep. 2019, 9, 9657. [Google Scholar] [CrossRef] [PubMed]
- Anderson, J.P.E. Soil Respiration. In Methods of Soil Analysis, Part 2, Chemical and Microbiological Properties; Page, A.L., Ed.; Agronomy monograph No. 9; ASA-SSSA: Madison, 1982; pp. 831–871. [Google Scholar]
- Kozich, J.J.; Westcott, S.L.; Baxter, N.T.; Highlander, S.K.; Schloss, P.D. Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl. Environ. Microbiol. 2013, 79, 5112–5120. [Google Scholar] [CrossRef] [PubMed]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Garrity, G.M.; Tiedje, J.M.; Cole, J.R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 2007, 73, 5261–5267. [Google Scholar] [CrossRef]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glockner, F.O. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 2013, 41, D590–D596. [Google Scholar] [CrossRef]
- Langille, M.G.; Zaneveld, J.; Caporaso, J.G.; McDonald, D.; Knights, D.; Reyes, J.A.; Clemente, J.C.; Burkepile, D.E.; Vega Thurber, R.L.; Knight, R.; et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat. Biotechnol. 2013, 31, 814–821. [Google Scholar] [CrossRef]
- Kanehisa, M.; Sato, Y.; Furumichi, M.; Morishima, K.; Tanabe, M. New approach for understanding genome variations in KEGG. Nucleic Acids Res. 2019, 47, 590–595. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- McMurdie, P.J.; Holmes, S. Phyloseq: An R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 2013, 8, e61217. [Google Scholar] [CrossRef]
- Price, M.N.; Dehal, P.S.; Arkin, A.P. FastTree 2—Approximately maximum-likelihood trees for large alignments. PLoS ONE 2010, 5, e9490. [Google Scholar] [CrossRef] [PubMed]
- Faith, D.P. Conservation evaluation and phylogenetic diversity. Biol. Conserv. 1992, 61, 1–10. [Google Scholar] [CrossRef]
- Lozupone, C.; Knight, R. UniFrac: A new phylogenetic method for comparing microbial communities. Appl. Environ. MicroBiol. 2005, 71, 8228–8235. [Google Scholar] [CrossRef] [PubMed]
- Pinheiro, J.; Bates, D.; DebRoy, S.; Sarkar, D.; R Core Team. Nlme: Linear and Nonlinear Mixed Effects Models. In vol. R package version 3.1-120; The R Foundation: Vienna, Austria, 2015. [Google Scholar]
- Hothorn, T.; Bretz, F.; Westfall, P. Simultaneous inference in general parametric models. Biom. J. 2008, 50, 346–363. [Google Scholar] [CrossRef]
- Swinton, J. Vennerable: Venn and Euler Area-Proportional Diagrams. R package version 3.0/r82. 2013. Available online: https://R-Forge.R-project.org/projects/vennerable/ (accessed on 24 February 2020).
- R Core Team. A language and environment for statistical computing. In R Foundation for Statistical Computing; R Core Team: Vienna, Austria, 2014. [Google Scholar]
- Wickham, H. ggplot2: Elegant Graphics for Data Analysis; Springer-Verlag: New York, NY, USA, 2016. [Google Scholar]
- Fierer, N.; Bradford, M.A.; Jackson, R.B. Toward an ecological classification of soil bacteria. Ecology 2007, 88, 1354–1364. [Google Scholar] [CrossRef]
- Domínguez, J.; Aira, M.; Gómez-Brandón, M. Vermicomposting: Earthworms Enhance the Work of Microbes. In Microbes at Work; Insam, H., Franke-Whittle, I., Goberna, M., Eds.; Springer: Berlin, Heidelberg, Germany, 2010; pp. 93–114. [Google Scholar]
- Gómez-Brandón, M.; Lores, M.; Domínguez, J. Species-specific effects of epigeic earthworms on microbial community structure during first stages of decomposition of organic matter. PLOS ONE 2012, 7, e31895. [Google Scholar] [CrossRef]
- Ho, A.; Di Lonardo, D.P.; Bodelier, P.L.E. Revising life strategy concepts in environmental microbial ecology. FEMS Microbiol Ecol 2017, 93, fix006. [Google Scholar] [CrossRef]
- Yang, W.; Yang, Z.; Guan, Y.; Zhai, C.; Shi, D.; Chen, J.; Wang, T.; Gu, S. Dose-dependent effect of compost amendment on soil bacterial community composition and co-occurrence network patterns in soybean agroecosystem. Arch. Agron. Soil Sci. 2019. [Google Scholar] [CrossRef]
- Gómez-Brandón, M.; Aira, M.; Lores, M.; Domínguez, J. Epigeic earthworms exert a bottleneck effect on microbial communities through gut associated processes. PLOS ONE 2011, 6, e24786. [Google Scholar] [CrossRef]
- Huang, K.; Li, F.; Wei, Y.; Chen, X.; Fu, X. Changes of bacterial and fungal community compositions during vermicomposting of vegetable wastes by Eisenia foetida. Bioresour. Technol. 2013, 150, 235–241. [Google Scholar] [CrossRef]
- Lv, B.; Xing, M.; Yang, J.; Zhang, L. Pyrosequencing reveals bacterial community differences in composting and vermicomposting on the stabilization of mixed sewage sludge and cattle dung. Appl. Microbiol. Biotechnol. 2015, 99, 10703–10712. [Google Scholar] [CrossRef] [PubMed]
- Vivas, A.; Moreno, B.; Garcia-Rodriguez, S.; Benitez, E. Assessing the impact of composting and vermicomposting on bacterial community size and structure, and microbial functional diversity of an olive-mill waste. Bioresour. Technol. 2009, 100, 1319–1326. [Google Scholar] [CrossRef] [PubMed]
- Shade, A.; Handelsman, J. Beyond the Venn diagram: The hunt for a core microbiome. Environ. MicroBiol. 2012, 14, 4–12. [Google Scholar] [CrossRef] [PubMed]
- Aira, M.; Olcina, J.; Pérez-Losada, M.; Domínguez, J. Characterization of the bacterial communities of casts from Eisenia andrei fed with different substrates. Appl. Soil Ecol. 2016, 98, 103–111. [Google Scholar] [CrossRef]
- Cai, L.; Gong, X.; Sun, X.; Li, S.; Yu, X. Comparison of chemical and microbiological changes during the aerobic composting and vermicomposting of green waste. PLOS ONE 2018, 13, e0207494. [Google Scholar] [CrossRef]
- Galitskaya, P.; Biktasheva, L.; Saveliev, A.; Grigoryeva, T.; Boulygina, E.; Selivanovskaya, S. Fungal and bacterial successions in the process of co-composting of organic wastes as revealed by 454 pyrosequencing. PLOS ONE 2017, 12, e0186051. [Google Scholar] [CrossRef]
- Bedini, A.; Mercy, L.; Schneider, C.; Franken, P.; Lucic-Mercy, E. Unraveling the Initial Plant Hormone Signaling, Metabolic Mechanisms and Plant Defense Triggering the Endomycorrhizal Symbiosis Behavior. Front. Plant. Sci. 2018, 9, 1800. [Google Scholar] [CrossRef]
| Alpha Diversity | F5,24 | p( > F) | |
| Observed | 12.004 | <0.0001 | |
| Chao1 | 9.401 | <0.0001 | |
| Shannon | 54.031 | <0.0001 | |
| Faith PD | 26.615 | <0.0001 | |
| Beta Diversity | F5,24 | p-value | |
| Unifrac—unweighted | PCoA1 | 596.074 | <0.0001 |
| PCoA2 | 26.350 | <0.0001 | |
| Unifrac—weighted | PCoA1 | 456.28 | <0.0001 |
| PCoA2 | 6.101 | <0.0001 | |
| Bray–Curtis | PCoA1 | 3629.08 | <0.0001 |
| PCoA2 | 58.103 | <0.0001 | |
| Jaccard | PCoA1 | 2333.82 | <0.0001 |
| PCoA2 | 70.029 | <0.0001 | |
| Day 0 | End Process (91 days) | End Process (42 days) | ||
|---|---|---|---|---|
| Alpha Diversity | Raw Grape Marc | Distilled Grape Marc | Raw Grape Marc | Distilled Grape Marc |
| Observed | 95.47 ± 7.66 | 326.26 ± 16.16 | 685.20 ± 38.87 | 617.14 ± 50.72 |
| Chao1 | 100.67 ± 9.54 | 353.41 ± 17.35 | 724.43 ± 46.91 | 637.15 ± 56.05 |
| Shannon | 2.40 ± 0.04 | 4.14 ± 0.07 | 5.39 ± 0.07 | 5.29 ± 0.08 |
| Faith PD | 10.76 ± 1.16 | 11.18 ± 0.51 | 40.62 ± 2.53 | 27.19 ± 0.52 |
| Relative Abundances—Phylum | ||||
| Proteobacteria | 58.89 ± 2.08 | 60.01 ± 3.67 | 52.00 ± 0.95 | 52.37 ± 1.366 |
| Firmicutes | 39.17 ± 2.05 | 18.87 ± 1.95 | 0.61 ± 0.05 | 0.45 ± 0.05 |
| Bacteroidetes | 1.09 ± 0.28 | 14.90 ± 2.82 | 30.10 ± 1.58 | 30.29 ± 1.66 |
| Actinobacteria | 0.78 ± 0.31 | 1.34 ± 0.27 | 9.73 ± 0.94 | 8.11 ± 0.55 |
| Verrucomicrobia | 0.01 ± 0.008 | 0.08 ± 0.03 | 3.13 ± 0.23 | 4.55 ± 0.14 |
| Planctomycetes | 0.05 ± 0.02 | not detected | 2.37 ± 0.20 | not detected |
| Chloroflexi | 0.023 ± 0.01 | not detected | 1.01 ± 0.13 | not detected |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gómez-Brandón, M.; Aira, M.; Santana, N.; Pérez-Losada, M.; Domínguez, J. Temporal Dynamics of Bacterial Communities in a Pilot-Scale Vermireactor Fed with Distilled Grape Marc. Microorganisms 2020, 8, 642. https://doi.org/10.3390/microorganisms8050642
Gómez-Brandón M, Aira M, Santana N, Pérez-Losada M, Domínguez J. Temporal Dynamics of Bacterial Communities in a Pilot-Scale Vermireactor Fed with Distilled Grape Marc. Microorganisms. 2020; 8(5):642. https://doi.org/10.3390/microorganisms8050642
Chicago/Turabian StyleGómez-Brandón, María, Manuel Aira, Natielo Santana, Marcos Pérez-Losada, and Jorge Domínguez. 2020. "Temporal Dynamics of Bacterial Communities in a Pilot-Scale Vermireactor Fed with Distilled Grape Marc" Microorganisms 8, no. 5: 642. https://doi.org/10.3390/microorganisms8050642
APA StyleGómez-Brandón, M., Aira, M., Santana, N., Pérez-Losada, M., & Domínguez, J. (2020). Temporal Dynamics of Bacterial Communities in a Pilot-Scale Vermireactor Fed with Distilled Grape Marc. Microorganisms, 8(5), 642. https://doi.org/10.3390/microorganisms8050642





