Anti-Contamination Strategies for Yeast Fermentations
Abstract
1. Introduction
1.1. Wine, Beer, and Bread
1.2. Biofuels
1.3. Protein Production
2. Microbial Contamination during Yeast Fermentation
2.1. Wine Fermentation
2.2. The Brewing Industry
2.3. Fuel Ethanol Fermentation
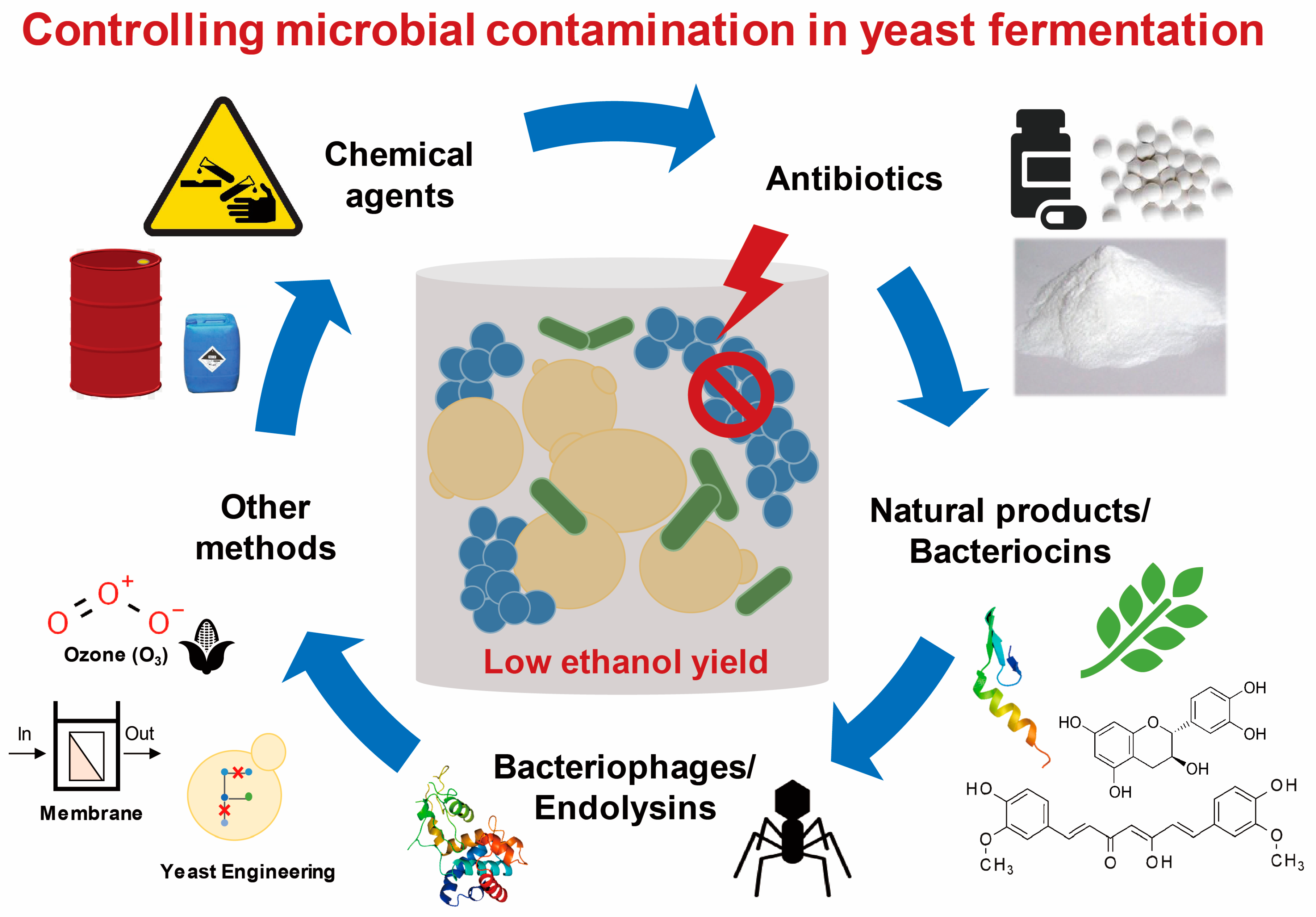
3. Anti-Bacterial Contamination Strategies in the Laboratory and Industry
3.1. Chemical Agents
3.2. Antibiotics
3.3. Natural Antimicrobial Compounds and Bacteriocins
3.4. Bacteriophages and Endolysins
3.5. Other Methods
4. Conclusions
Acknowledgments
Conflicts of Interest
References
- Feher, J.; Lengyel, G.; Lugasi, A. Cultural history of wine, the theoretical background of wine therapy. Orv. Hetil. 2005, 146, 2635–2639. [Google Scholar] [CrossRef]
- Barnett, J.A. A history of research on yeasts 2: Louis pasteur and his contemporaries, 1850-1880. Yeast 2000, 16, 755–771. [Google Scholar] [CrossRef]
- Dashko, S.; Zhou, N.; Compagno, C.; Piskur, J. Why, when, and how did yeast evolve alcoholic fermentation? FEMS Yeast Res. 2014, 14, 826–832. [Google Scholar] [CrossRef] [PubMed]
- Gray, W.D. Studies on the alcohol tolerance of yeasts. J. Bacteriol. 1941, 42, 561–574. [Google Scholar] [CrossRef] [PubMed]
- Vieira Gomes, A.M.; Souza Carmo, T.; Silva Carvalho, L.; Mendonca Bahia, F.; Parachin, N.S. Comparison of yeasts as hosts for recombinant protein production. Microorganisms 2018, 6. [Google Scholar] [CrossRef]
- Esteve-Zarzoso, B.; Manzanares, P.; Ramon, D.; Querol, A. The role of non-saccharomyces yeasts in industrial winemaking. Int. Microbiol. 1998, 1, 143–148. [Google Scholar]
- Beltran, G.; Torija, M.J.; Novo, M.; Ferrer, N.; Poblet, M.; Guillamon, J.M.; Rozes, N.; Mas, A. Analysis of yeast populations during alcoholic fermentation: A six year follow-up study. Syst. Appl. Microbiol. 2002, 25, 287–293. [Google Scholar] [CrossRef]
- Olesen, K.; Felding, T.; Gjermansen, C.; Hansen, J. The dynamics of the saccharomyces carlsbergensis brewing yeast transcriptome during a production-scale lager beer fermentation. FEMS Yeast Res. 2002, 2, 563–573. [Google Scholar]
- Holt, S.; Mukherjee, V.; Lievens, B.; Verstrepen, K.J.; Thevelein, J.M. Bioflavoring by non-conventional yeasts in sequential beer fermentations. Food Microbiol. 2018, 72, 55–66. [Google Scholar] [CrossRef]
- Randez-Gil, F.; Corcoles-Saez, I.; Prieto, J.A. Genetic and phenotypic characteristics of baker’s yeast: Relevance to baking. Annu. Rev. Food Sci. Technol. 2013, 4, 191–214. [Google Scholar] [CrossRef]
- Buijs, N.A.; Siewers, V.; Nielsen, J. Advanced biofuel production by the yeast saccharomyces cerevisiae. Curr. Opin. Chem. Biol. 2013, 17, 480–488. [Google Scholar] [CrossRef] [PubMed]
- Mohd Azhar, S.H.; Abdulla, R.; Jambo, S.A.; Marbawi, H.; Gansau, J.A.; Mohd Faik, A.A.; Rodrigues, K.F. Yeasts in sustainable bioethanol production: A review. Biochem. Biophys. Rep. 2017, 10, 52–61. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.G.; Jin, Y.S.; Cha, Y.L.; Seo, J.H. Bioethanol production from cellulosic hydrolysates by engineered industrial saccharomyces cerevisiae. Bioresour. Technol. 2017, 228, 355–361. [Google Scholar] [CrossRef] [PubMed]
- Stanley, D.; Bandara, A.; Fraser, S.; Chambers, P.J.; Stanley, G.A. The ethanol stress response and ethanol tolerance of saccharomyces cerevisiae. J. Appl. Microbiol. 2010, 109, 13–24. [Google Scholar] [CrossRef] [PubMed]
- Swidah, R.; Wang, H.; Reid, P.J.; Ahmed, H.Z.; Pisanelli, A.M.; Persaud, K.C.; Grant, C.M.; Ashe, M.P. Butanol production in s. Cerevisiae via a synthetic abe pathway is enhanced by specific metabolic engineering and butanol resistance. Biotechnol. Biofuels 2015, 8, 97. [Google Scholar] [CrossRef] [PubMed]
- Matsuda, F.; Ishii, J.; Kondo, T.; Ida, K.; Tezuka, H.; Kondo, A. Increased isobutanol production in saccharomyces cerevisiae by eliminating competing pathways and resolving cofactor imbalance. Microb. Cell Fact. 2013, 12, 119. [Google Scholar] [CrossRef]
- Lee, W.H.; Seo, S.O.; Bae, Y.H.; Nan, H.; Jin, Y.S.; Seo, J.H. Isobutanol production in engineered saccharomyces cerevisiae by overexpression of 2-ketoisovalerate decarboxylase and valine biosynthetic enzymes. Bioprocess. Biosyst. Eng. 2012, 35, 1467–1475. [Google Scholar] [CrossRef]
- Kim, S.J.; Seo, S.O.; Jin, Y.S.; Seo, J.H. Production of 2,3-butanediol by engineered saccharomyces cerevisiae. Bioresour. Technol. 2013, 146, 274–281. [Google Scholar] [CrossRef]
- Tamakawa, H.; Mita, T.; Yokoyama, A.; Ikushima, S.; Yoshida, S. Metabolic engineering of candida utilis for isopropanol production. Appl. Microbiol. Biotechnol. 2013, 97, 6231–6239. [Google Scholar] [CrossRef]
- Siripong, W.; Wolf, P.; Kusumoputri, T.P.; Downes, J.J.; Kocharin, K.; Tanapongpipat, S.; Runguphan, W. Metabolic engineering of pichia pastoris for production of isobutanol and isobutyl acetate. Biotechnol. Biofuels 2018, 11, 1. [Google Scholar] [CrossRef]
- Mattanovich, D.; Branduardi, P.; Dato, L.; Gasser, B.; Sauer, M.; Porro, D. Recombinant protein production in yeasts. Methods Mol. Biol. 2012, 824, 329–358. [Google Scholar] [PubMed]
- Nandy, S.K.; Srivastava, R.K. A review on sustainable yeast biotechnological processes and applications. Microbiol. Res. 2018, 207, 83–90. [Google Scholar] [CrossRef]
- Owens, D.R.; Landgraf, W.; Schmidt, A.; Bretzel, R.G.; Kuhlmann, M.K. The emergence of biosimilar insulin preparations—A cause for concern? Diabetes Technol. Ther. 2012, 14, 989–996. [Google Scholar] [CrossRef] [PubMed]
- Kumar, R.; Kumar, P. Yeast-based vaccines: New perspective in vaccine development and application. FEMS Yeast Res. 2019, 19. [Google Scholar] [CrossRef] [PubMed]
- Syed, Y.Y. Dtap5-hb-ipv-hib vaccine (vaxelis((r))): A review of its use in primary and booster vaccination. Paediatr. Drugs 2017, 19, 69–80. [Google Scholar] [CrossRef]
- Liu, Z.; Tyo, K.E.; Martinez, J.L.; Petranovic, D.; Nielsen, J. Different expression systems for production of recombinant proteins in saccharomyces cerevisiae. Biotechnol. Bioeng. 2012, 109, 1259–1268. [Google Scholar] [CrossRef]
- Gerngross, T.U. Advances in the production of human therapeutic proteins in yeasts and filamentous fungi. Nat. Biotechnol. 2004, 22, 1409–1414. [Google Scholar] [CrossRef]
- Mokdad-Gargouri, R.; Abdelmoula-Soussi, S.; Hadiji-Abbes, N.; Amor, I.Y.; Borchani-Chabchoub, I.; Gargouri, A. Yeasts as a tool for heterologous gene expression. Methods Mol. Biol. 2012, 824, 359–370. [Google Scholar]
- Loureiro, V.; Malfeito-Ferreira, M. Spoilage yeasts in the wine industry. Int. J. Food Microbiol. 2003, 86, 23–50. [Google Scholar] [CrossRef]
- Varela, C.; Borneman, A.R. Yeasts found in vineyards and wineries. Yeast 2017, 34, 111–128. [Google Scholar] [CrossRef]
- Dias, L.; Dias, S.; Sancho, T.; Stender, H.; Querol, A.; Malfeito-Ferreira, M.; Loureiro, V. Identification of yeasts isolated from wine-related environments and capable of producing 4-ethylphenol. Food Microbiol. 2003, 20, 567–574. [Google Scholar] [CrossRef]
- Rodrigues, N.; Goncalves, G.; Pereira-da-Silva, S.; Malfeito-Ferreira, M.; Loureiro, V. Development and use of a new medium to detect yeasts of the genera dekkera/brettanomyces. J. Appl. Microbiol. 2001, 90, 588–599. [Google Scholar] [CrossRef] [PubMed]
- Berbegal, C.; Spano, G.; Fragasso, M.; Grieco, F.; Russo, P.; Capozzi, V. Starter cultures as biocontrol strategy to prevent brettanomyces bruxellensis proliferation in wine. Appl. Microbiol. Biotechnol. 2018, 102, 569–576. [Google Scholar] [CrossRef] [PubMed]
- Schopp, L.M.; Lee, J.; Osborne, J.P.; Chescheir, S.C.; Edwards, C.G. Metabolism of nonesterified and esterified hydroxycinnamic acids in red wines by brettanomyces bruxellensis. J. Agric. Food Chem. 2013, 61, 11610–11617. [Google Scholar] [CrossRef] [PubMed]
- Steensels, J.; Daenen, L.; Malcorps, P.; Derdelinckx, G.; Verachtert, H.; Verstrepen, K.J. Brettanomyces yeasts—From spoilage organisms to valuable contributors to industrial fermentations. Int. J. Food Microbiol. 2015, 206, 24–38. [Google Scholar] [CrossRef]
- Obi, C. Brewery contaminants, challenges and remedies—A review. J. Microbiol. 2018, 31, 3926–3940. [Google Scholar]
- Suzuki, K. 125th anniversary review: Microbiological instability of beer caused by spoilage bacteria. J. Inst. Brew. 2011, 117, 131–155. [Google Scholar] [CrossRef]
- Hollerová, I.; Kubizniaková, P. Monitoring gram positive bacterial contamination in czech breweries. J. Inst. Brew. 2001, 107, 355–358. [Google Scholar] [CrossRef]
- Simpson, W.J. Ionophoric action of trans-isohumulone on lactobacillus brevis. Microbiology 1993, 139, 1041–1045. [Google Scholar] [CrossRef][Green Version]
- Simpson, W.J. Cambridge prize lecture. Studies on the sensitivity of lactic acid bacteria to hop bitter acids. J. Inst. Brew. 1993, 99, 405–411. [Google Scholar] [CrossRef]
- Simpson, W.J.; Fernandez, J.L. Mechanism of resistance of lactic acid bacteria to trans-isohumulone. J. Am. Soc. Brew. Chem. 1994, 52, 9–11. [Google Scholar] [CrossRef]
- Iijima, K.; Suzuki, K.; Asano, S.; Ogata, T.; Kitagawa, Y. Horc, a hop-resistance related protein, presumably functions in homodimer form. Biosci. Biotechnol. Biochem. 2009, 73, 1880–1882. [Google Scholar] [CrossRef] [PubMed]
- Iijima, K.; Suzuki, K.; Ozaki, K.; Yamashita, H. Horc confers beer-spoilage ability on hop-sensitive lactobacillus brevis abbc45cc. J. Appl. Microbiol. 2006, 100, 1282–1288. [Google Scholar] [CrossRef] [PubMed]
- Vriesekoop, F.; Krahl, M.; Hucker, B.; Menz, G. 125th anniversary review: Bacteria in brewing: The good, the bad and the ugly. J. Inst. Brew. 2012, 118, 335–345. [Google Scholar] [CrossRef]
- Lin, J.; Cao, Y.; Sun, J.; Lu, J. Monitoring spoilage bacteria and wild yeasts in eastern chinese breweries. J. Am. Soc. Brew. Chem. 2008, 66, 43–47. [Google Scholar] [CrossRef]
- Thelen, K.; Beimfohr, C.; Snaidr, J. Evaluation study of the frequency of different beer-spoiling bacteria using the vit analysis. Tech. Q. Master Brew. Assoc. Am 2006, 43, 31–35. [Google Scholar]
- Hussein, H.S.; Brasel, J.M. Toxicity, metabolism, and impact of mycotoxins on humans and animals. Toxicology 2001, 167, 101–134. [Google Scholar] [CrossRef]
- Rodriguez-Carrasco, Y.; Fattore, M.; Albrizio, S.; Berrada, H.; Manes, J. Occurrence of fusarium mycotoxins and their dietary intake through beer consumption by the european population. Food Chem. 2015, 178, 149–155. [Google Scholar] [CrossRef]
- Wolf-Hall, C.E. Mold and mycotoxin problems encountered during malting and brewing. Int. J. Food Microbiol. 2007, 119, 89–94. [Google Scholar] [CrossRef]
- Piacentini, K.C.; Savi, G.D.; Olivo, G.; Scussel, V.M. Quality and occurrence of deoxynivalenol and fumonisins in craft beer. Food Control. 2015, 50, 925–929. [Google Scholar] [CrossRef]
- Habler, K.; Geissinger, C.; Hofer, K.; Schuler, J.; Moghari, S.; Hess, M.; Gastl, M.; Rychlik, M. Fate of fusarium toxins during brewing. J. Agric. Food Chem. 2017, 65, 190–198. [Google Scholar] [CrossRef] [PubMed]
- Vegi, A.; Schwarz, P.; Wolf-Hall, C.E. Quantification of tri5 gene, expression, and deoxynivalenol production during the malting of barley. Int. J. Food Microbiol. 2011, 150, 150–156. [Google Scholar] [CrossRef]
- Reiss, J. Influence of the mycotoxins aflatoxin b1, rubratoxin b, patulin and diacetoxyscirpenol on the fermentation activity of baker’s yeast. Mycopathol. Mycol. Appl. 1973, 51, 337–345. [Google Scholar] [CrossRef] [PubMed]
- Klosowski, G.; Mikulski, D. The effect of raw material contamination with mycotoxins on the composition of alcoholic fermentation volatile by-products in raw spirits. Bioresour. Technol. 2010, 101, 9723–9727. [Google Scholar] [CrossRef] [PubMed]
- Lopes, M.L.; Paulillo, S.C.d.L.; Godoy, A.; Cherubin, R.A.; Lorenzi, M.S.; Giometti, F.H.C.; Bernardino, C.D.; Amorim Neto, H.B.d.; Amorim, H.V.d. Ethanol production in brazil: A bridge between science and industry. Braz. J. Microbiol. 2016, 47 (Suppl. S1), 64–76. [Google Scholar] [CrossRef]
- Bayrock, D.P.; Ingledew, W.M. Inhibition of yeast by lactic acid bacteria in continuous culture: Nutrient depletion and/or acid toxicity? J. Ind. Microbiol. Biotechnol. 2004, 31, 362–368. [Google Scholar] [CrossRef]
- Narendranath, N.V.; Hynes, S.H.; Thomas, K.C.; Ingledew, W.M. Effects of lactobacilli on yeast-catalyzed ethanol fermentations. Appl. Environ. Microbiol. 1997, 63, 4158–4163. [Google Scholar] [CrossRef]
- de Souza Liberal, A.T.; Basilio, A.C.; do Monte Resende, A.; Brasileiro, B.T.; da Silva-Filho, E.A.; de Morais, J.O.; Simoes, D.A.; de Morais, M.A., Jr. Identification of dekkera bruxellensis as a major contaminant yeast in continuous fuel ethanol fermentation. J. Appl. Microbiol. 2007, 102, 538–547. [Google Scholar] [CrossRef]
- Abbott, D.A.; Hynes, S.H.; Ingledew, W.M. Growth rates of dekkera/brettanomyces yeasts hinder their ability to compete with saccharomyces cerevisiae in batch corn mash fermentations. Appl. Microbiol. Biotechnol. 2005, 66, 641–647. [Google Scholar] [CrossRef]
- Ciani, M.; Maccarelli, F.; Fatichenti, F. Growth and fermentation behaviour of brettanomyces/dekkera yeasts under different conditions of aerobiosis. World J. Microbiol. Biotechnol. 2003, 19, 419–422. [Google Scholar] [CrossRef]
- Beckner, M.; Ivey, M.L.; Phister, T.G. Microbial contamination of fuel ethanol fermentations. Lett. Appl. Microbiol. 2011, 53, 387–394. [Google Scholar] [CrossRef] [PubMed]
- Bassi, A.P.G.; Meneguello, L.; Paraluppi, A.L.; Sanches, B.C.P.; Ceccato-Antonini, S.R. Interaction of saccharomyces cerevisiae-lactobacillus fermentum-dekkera bruxellensis and feedstock on fuel ethanol fermentation. Lett. Appl. Microbiol. 2018, 111, 1661–1672. [Google Scholar] [CrossRef]
- Pena-Moreno, I.C.; Castro Parente, D.; da Silva, J.M.; Andrade Mendonca, A.; Rojas, L.A.V.; de Morais Junior, M.A.; de Barros Pita, W. Nitrate boosts anaerobic ethanol production in an acetate-dependent manner in the yeast dekkera bruxellensis. J. Ind. Microbiol. Biotechnol. 2019, 46, 209–220. [Google Scholar] [CrossRef] [PubMed]
- Elshaghabee, F.M.; Bockelmann, W.; Meske, D.; de Vrese, M.; Walte, H.G.; Schrezenmeir, J.; Heller, K.J. Ethanol production by selected intestinal microorganisms and lactic acid bacteria growing under different nutritional conditions. Front. Microbiol. 2016, 7, 47. [Google Scholar] [CrossRef] [PubMed]
- Amorim, H.V.; Lopes, M.L.; de Castro Oliveira, J.V.; Buckeridge, M.S.; Goldman, G.H. Scientific challenges of bioethanol production in brazil. Appl. Microbiol. Biotechnol. 2011, 91, 1267–1275. [Google Scholar] [CrossRef]
- Tiukova, I.; Eberhard, T.; Passoth, V. Interaction of lactobacillus vini with the ethanol-producing yeasts dekkera bruxellensis and saccharomyces cerevisiae. Biotechnol. Appl. Biochem. 2014, 61, 40–44. [Google Scholar] [CrossRef] [PubMed]
- Santos, M.T.; Yokoya, F. Characteristics of yeast cell flocculation by lactobacillus fermentum. J. Ferment. Bioeng. 1993, 75, 151–154. [Google Scholar] [CrossRef]
- Carvalho-Netto, O.V.; Carazzolle, M.F.; Mofatto, L.S.; Teixeira, P.J.; Noronha, M.F.; Calderon, L.A.; Mieczkowski, P.A.; Argueso, J.L.; Pereira, G.A. Saccharomyces cerevisiae transcriptional reprograming due to bacterial contamination during industrial scale bioethanol production. Microb. Cell Fact. 2015, 14, 13. [Google Scholar] [CrossRef]
- Basso, L.C.; de Amorim, H.V.; de Oliveira, A.J.; Lopes, M.L. Yeast selection for fuel ethanol production in brazil. FEMS Yeast Res. 2008, 8, 1155–1163. [Google Scholar] [CrossRef]
- Costa, M.A.S.; Cerri, B.C.; Ceccato-Antonini, S.R. Ethanol addition enhances acid treatment to eliminate lactobacillus fermentum from the fermentation process for fuel ethanol production. Lett. Appl. Microbiol. 2018, 66, 77–85. [Google Scholar] [CrossRef]
- Barth, D.; de Souza Monteiro, A.R.; da Costa, M.M.; Virkajarvi, I.; Sacon, V.; Wilhelmsom, A. Desinfix tm 135 in fermentation process for bioethanol production. Braz. J. Microbiol. 2014, 45, 323–325. [Google Scholar] [CrossRef][Green Version]
- Broda, M.; Grajek, W. Ammonia disinfection of corn grains intended for ethanol fermentation. Acta Sci. Pol. Technol. Aliment. 2009, 8, 33–38. [Google Scholar]
- Salvi, D.A.; Aita, G.M.; Robert, D.; Bazan, V. Ethanol production from sorghum by a dilute ammonia pretreatment. J. Ind. Microbiol. Biotechnol. 2010, 37, 27–34. [Google Scholar] [CrossRef] [PubMed]
- Narendranath, N.V.; Thomas, K.C.; Ingledew, W.M. Urea hydrogen peroxide reduces the numbers of lactobacilli, nourishes yeast, and leaves no residues in the ethanol fermentation. Appl. Environ. Microbiol. 2000, 66, 4187–4192. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Chang, I.S.; Kim, B.H.; Shin, P.K. Use of sulfite and hydrogen peroxide to control bacterial contamination in ethanol fermentation. Appl. Environ. Microbiol. 1997, 63, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Meneghin, S.P.; Reis, F.C.; de Almeida, P.G.; Ceccato-Antonini, S.R. Chlorine dioxide against bacteria and yeasts from the alcoholic fermentation. Braz. J. Microbiol. 2008, 39, 337–343. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Oliva-Neto, P.-d.; Yokoya, F. Effect of 3,4,4′-trichlorocarbanilide on growth of lactic acid bacteria contaminants in alcoholic fermentation. Bioresour. Technol. 1998, 63, 17–21. [Google Scholar] [CrossRef]
- Santos, M.C.; Nunes, C.; Saraiva, J.A.; Coimbra, M.A. Chemical and physical methodologies for the replacement/reduction of sulfur dioxide use during winemaking: Review of their potentialities and limitations. Eur. Food Res. Technol. 2012, 234, 1–12. [Google Scholar] [CrossRef]
- Morgan, S.C.; Haggerty, J.J.; Johnston, B.; Jiranek, V.; Durall, D.M. Response to sulfur dioxide addition by two commercial saccharomyces cerevisiae strains. Fermentation 2019, 5, 69. [Google Scholar] [CrossRef]
- Stevenson, D.D.; Simon, R.A. Sensitivity to ingested metabisulfites in asthmatic subjects. J. Allergy Clin. Immunol. 1981, 68, 26–32. [Google Scholar] [CrossRef]
- Threlfall, R.T.; Morris, J.R. Using dimethyldicarbonate to minimize sulfur dioxide for prevention of fermentation from excessive yeast contamination in juice and semi-sweet wine. J. Food Sci. 2002, 67, 2758–2762. [Google Scholar] [CrossRef]
- Stroppa, C.T.; Andrietta, M.G.S.; Andrietta, S.R.; Steckelberg, C.; Serra, G. Use of penicillin and monensin to control bacterial contamination of brazilian alcohol fermentations. Int. Sugar J. 2000, 102, 78–82. [Google Scholar]
- Aquarone, E. Penicillin and tetracycline as contamination control agents in alcoholic fermentation of sugar cane molasses. Appl. Microbiol. 1960, 8, 263–268. [Google Scholar] [CrossRef]
- Bayrock, D.P.; Thomas, K.C.; Ingledew, W.M. Control of lactobacillus contaminants in continuous fuel ethanol fermentations by constant or pulsed addition of penicillin g. Appl. Microbiol. Biotechnol. 2003, 62, 498–502. [Google Scholar] [CrossRef]
- Bischoff, K.M.; Liu, S.; Leathers, T.D.; Worthington, R.E.; Rich, J.O. Modeling bacterial contamination of fuel ethanol fermentation. Biotechnol. Bioeng. 2009, 103, 117–122. [Google Scholar] [CrossRef]
- Hynes, S.H.; Kjarsgaard, D.M.; Thomas, K.C.; Ingledew, W.M. Use of virginiamycin to control the growth of lactic acid bacteria during alcohol fermentation. J. Ind. Microbiol. Biotechnol. 1997, 18, 284–291. [Google Scholar] [CrossRef]
- Murphree, C.A.; Heist, E.P.; Moe, L.A. Antibiotic resistance among cultured bacterial isolates from bioethanol fermentation facilities across the united states. Curr. Microbiol. 2014, 69, 277–285. [Google Scholar] [CrossRef]
- Gyawali, R.; Ibrahim, S.A. Natural products as antimicrobial agents. Food Control. 2014, 46, 412–429. [Google Scholar] [CrossRef]
- Lucera, A.; Costa, C.; Conte, A.; Del Nobile, M.A. Food applications of natural antimicrobial compounds. Front. Microbiol. 2012, 3, 287. [Google Scholar] [CrossRef]
- Newman, D.J.; Cragg, G.M. Natural products as sources of new drugs from 1981 to 2014. J. Nat. Prod. 2016, 79, 629–661. [Google Scholar] [CrossRef]
- Salam, A.M.; Quave, C.L. Opportunities for plant natural products in infection control. Curr. Opin. Microbiol. 2018, 45, 189–194. [Google Scholar] [CrossRef] [PubMed]
- Sakamoto, K.; Konings, W.N. Beer spoilage bacteria and hop resistance. Int. J. Food Microbiol. 2003, 89, 105–124. [Google Scholar] [CrossRef]
- Muthaiyan, A.; Limayem, A.; Ricke, S.C. Antimicrobial strategies for limiting bacterial contaminants in fuel bioethanol fermentations. Prog. Energy Combust. Sci. 2011, 37, 351–370. [Google Scholar] [CrossRef]
- Liu, Q.; Meng, X.; Li, Y.; Zhao, C.N.; Tang, G.Y.; Li, H.B. Antibacterial and antifungal activities of spices. Int. J. Mol. Sci. 2017, 18. [Google Scholar] [CrossRef]
- Alves, M.J.; Ferreira, I.C.; Dias, J.; Teixeira, V.; Martins, A.; Pintado, M. A review on antimicrobial activity of mushroom (basidiomycetes) extracts and isolated compounds. Planta Med. 2012, 78, 1707–1718. [Google Scholar] [CrossRef]
- Bala, N.; Aitken, E.A.; Cusack, A.; Steadman, K.J. Antimicrobial potential of australian macrofungi extracts against foodborne and other pathogens. Phytother. Res. 2012, 26, 465–469. [Google Scholar] [CrossRef]
- Valverde, M.E.; Hernandez-Perez, T.; Paredes-Lopez, O. Edible mushrooms: Improving human health and promoting quality life. Int. J. Microbiol. 2015, 2015, 376387. [Google Scholar] [CrossRef]
- Taofiq, O.; Heleno, S.A.; Calhelha, R.C.; Alves, M.J.; Barros, L.; Barreiro, M.F.; Gonzalez-Paramas, A.M.; Ferreira, I.C. Development of mushroom-based cosmeceutical formulations with anti-inflammatory, anti-tyrosinase, antioxidant, and antibacterial properties. Molecules 2016, 21. [Google Scholar] [CrossRef]
- Gil, G.; del Monaco, S.; Cerrutti, P.; Galvagno, M. Selective antimicrobial activity of chitosan on beer spoilage bacteria and brewing yeasts. Biotechnol. Lett. 2004, 26, 569–574. [Google Scholar] [CrossRef]
- Haris, S.; Fang, C.; Bastidas-Oyanedel, J.R.; Prather, K.J.; Schmidt, J.E.; Thomsen, M.H. Natural antibacterial agents from arid-region pretreated lignocellulosic biomasses and extracts for the control of lactic acid bacteria in yeast fermentation. AMB Express 2018, 8, 127. [Google Scholar] [CrossRef]
- Brul, S.; Coote, P. Preservative agents in foods. Mode of action and microbial resistance mechanisms. Int. J. Food Microbiol. 1999, 50, 1–17. [Google Scholar] [CrossRef]
- Sang, Y.; Blecha, F. Antimicrobial peptides and bacteriocins: Alternatives to traditional antibiotics. Anim. Health Res. Rev. 2008, 9, 227–235. [Google Scholar] [CrossRef]
- Bischoff, K.M.; Skinner-Nemec, K.A.; Leathers, T.D. Antimicrobial susceptibility of lactobacillus species isolated from commercial ethanol plants. J. Ind. Microbiol. Biotechnol. 2007, 34, 739–744. [Google Scholar] [CrossRef]
- Müller-Auffermann, K.; Grijalva, F.; Jacob, F.; Hutzler, M. Nisin and its usage in breweries: A review and discussion. J. Inst. Brew. 2015, 121, 309–319. [Google Scholar] [CrossRef]
- Ogden, K. Nisin: A bacteriocin with a potential use in brewing. J. Inst. Brew. 1986, 92, 379–383. [Google Scholar] [CrossRef]
- Radler, F. Possible use of nisin in winemaking. I. Action of nisin against lactic acid bacteria and wine yeasts in solid and liquid media. Am. J. Enol. Vitic. 1990, 41, 1–6. [Google Scholar]
- Furfaro, L.L.; Payne, M.S.; Chang, B.J. Bacteriophage therapy: Clinical trials and regulatory hurdles. Front. Cell Infect. Microbiol. 2018, 8, 376. [Google Scholar] [CrossRef]
- Roach, D.R.; Khatibi, P.A.; Bischoff, K.M.; Hughes, S.R.; Donovan, D.M. Bacteriophage-encoded lytic enzymes control growth of contaminating lactobacillus found in fuel ethanol fermentations. Biotechnol. Biofuels 2013, 6, 20. [Google Scholar] [CrossRef]
- Azam, A.H.; Tanji, Y. Bacteriophage-host arm race: An update on the mechanism of phage resistance in bacteria and revenge of the phage with the perspective for phage therapy. Appl. Microbiol. Biotechnol. 2019, 103, 2121–2131. [Google Scholar] [CrossRef]
- Ribelles, P.; Rodriguez, I.; Suarez, J.E. Lysa2, the lactobacillus casei bacteriophage a2 lysin is an endopeptidase active on a wide spectrum of lactic acid bacteria. Appl. Microbiol. Biotechnol. 2012, 94, 101–110. [Google Scholar] [CrossRef]
- Khatibi, P.A.; Roach, D.R.; Donovan, D.M.; Hughes, S.R.; Bischoff, K.M. Saccharomyces cerevisiae expressing bacteriophage endolysins reduce lactobacillus contamination during fermentation. Biotechnol. Biofuels 2014, 7, 104. [Google Scholar] [CrossRef]
- Kim, J.S.; Daum, M.A.; Jin, Y.S.; Miller, M.J. Yeast derived lysa2 can control bacterial contamination in ethanol fermentation. Viruses 2018, 10. [Google Scholar] [CrossRef]
- Rasmussen, M.L.; Koziel, J.A.; Jane, J.L.; Pometto, A.L., 3rd. Reducing bacterial contamination in fuel ethanol fermentations by ozone treatment of uncooked corn mash. J. Agric. Food Chem. 2015, 63, 5239–5248. [Google Scholar] [CrossRef]
- Mahboubi, A.; Cayli, B.; Bulkan, G.; Doyen, W.; De Wever, H.; Taherzadeh, M.J. Removal of bacterial contamination from bioethanol fermentation system using membrane bioreactor. Fermentation 2018, 4, 88. [Google Scholar] [CrossRef]
- Fernández de Ullivarri, M.; Mendoza, L.M.; Raya, R.R. Killer yeasts as biocontrol agents of spoilage yeasts and bacteria isolated from wine. In Proceedings of the BIO Web of Conferences, Mendoza, Argentina, 9–14 November 2014; Volume 3, p. 02001. [Google Scholar]
- Shaw, A.J.; Lam, F.H.; Hamilton, M.; Consiglio, A.; MacEwen, K.; Brevnova, E.E.; Greenhagen, E.; LaTouf, W.G.; South, C.R.; van Dijken, H.; et al. Metabolic engineering of microbial competitive advantage for industrial fermentation processes. Science 2016, 353, 583–586. [Google Scholar] [CrossRef]
| Fermentations | Major Contaminants | Contaminant’s Original Source | ||
|---|---|---|---|---|
| Yeast | Bacteria | Fungi | ||
| Wine fermentation | D. bruxellensis | grape | ||
| Brewing fermentation | L. brevis | Fusarium species | barley, malt, hops, or adjuncts | |
| Fuel-ethanol fermentation | D. bruxellensis | L. fermentum | cane juice and molasses | |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Seo, S.-O.; Park, S.-K.; Jung, S.-C.; Ryu, C.-M.; Kim, J.-S. Anti-Contamination Strategies for Yeast Fermentations. Microorganisms 2020, 8, 274. https://doi.org/10.3390/microorganisms8020274
Seo S-O, Park S-K, Jung S-C, Ryu C-M, Kim J-S. Anti-Contamination Strategies for Yeast Fermentations. Microorganisms. 2020; 8(2):274. https://doi.org/10.3390/microorganisms8020274
Chicago/Turabian StyleSeo, Seung-Oh, Sung-Kyun Park, Suk-Chae Jung, Choong-Min Ryu, and Jun-Seob Kim. 2020. "Anti-Contamination Strategies for Yeast Fermentations" Microorganisms 8, no. 2: 274. https://doi.org/10.3390/microorganisms8020274
APA StyleSeo, S.-O., Park, S.-K., Jung, S.-C., Ryu, C.-M., & Kim, J.-S. (2020). Anti-Contamination Strategies for Yeast Fermentations. Microorganisms, 8(2), 274. https://doi.org/10.3390/microorganisms8020274







