Subcellular Location of Piscirickettsia salmonis Heat Shock Protein 60 (Hsp60) Chaperone by Using Immunogold Labeling and Proteomic Analysis
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Strains and Cell Line
2.2. Ethics Statement
2.3. In Silico P. salmonis Hsp60 Amino Acid Sequence Analyses
2.4. Immunogold Labelling of P. salmonis Hsp60
2.5. Subcellular Fractionation and Protein Extraction for Proteomic Analyses
2.6. Lysis of Protein Samples for Mass Spectrometry
2.7. Protein Identification and Analysis
2.8. Data Analysis
2.9. Generation of IgY Against P. salmonis Hsp60
2.10. Antigenicity Analyses of P. salmonis Hsp60 Synthetic Peptide by qRT-PCR
2.11. Inhibitory Activity of IgY Anti-Hsp60 in SHK-1 Cells
2.12. Statistical Analyses
3. Results
3.1. Labelling Patterns of In Vitro-Grown P. salmonis
3.2. Subcellular Fractionation and Proteomic Analyses
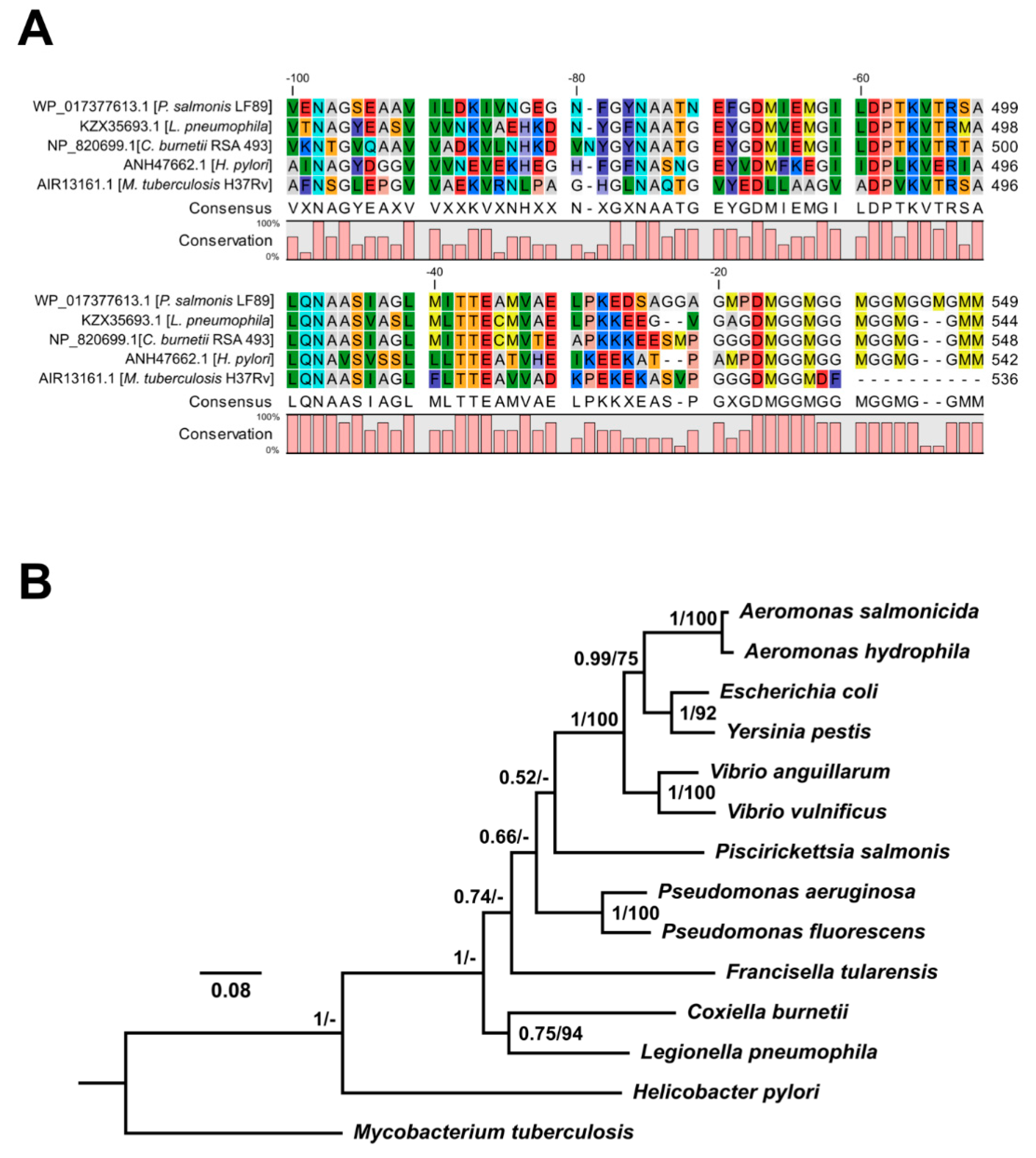
3.3. In Silico Analyses of P. salmonis Hsp60 Amino Acidic Sequence
3.4. Inhibitory Activity of IgY Anti-Hsp60 From P. salmonis
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Fryer, J.; Lannan, C.; Giovannoni, S.; Wood, N. Piscirickettsia salmonis gen. nov., sp. nov., the causative agent of an epizootic disease in salmonid fishes. Int. J. Syst. Evol. Microbiol. 1992, 42, 120–126. [Google Scholar] [CrossRef]
- Mauel, M.J.; Ware, C.; Smith, P.A. Culture of Piscirickettsia salmonis on enriched blood agar. J. Vet. Diagn. Investig. 2008, 20, 213–214. [Google Scholar] [CrossRef] [PubMed]
- Yáñez, A.; Valenzuela, K.; Silva, H.; Retamales, J.; Romero, A.; Enriquez, R.; Figueroa, J.; Claude, A.; González, J.; Avendaño-Herrera, R.; et al. Broth medium for the successful culture of the fish pathogen Piscirickettsia salmonis. Dis. Aquat. Org. 2012, 97, 197–205. [Google Scholar] [CrossRef]
- Sernapesca. Informe Sanitario Salmonicultura en Centros Marinos—Año 1 Semestre 2016. 2016. Available online: http://www.sernapesca.cl/index.php?option=com_remository&Itemid=246&func=fileinfo&id=21746 (accessed on 21 October 2019).
- Rozas, M.; Enríquez, R. Piscirickettsiosis and Piscirickettsia salmonis in fish: A review. J. Fish Dis. 2014, 37, 163–188. [Google Scholar] [CrossRef] [PubMed]
- Sernapesca. Informe Uso de Antimicrobianos 2015. 2016. Available online: http://www.sernapesca.cl/presentaciones/Comunicaciones/Informe_Sobre_Uso_de_Antimicrobianos_2015.pdf (accessed on 21 October 2019).
- Bohle, H.; Henríquez, P.; Grothusen, H.; Navas, E.; Bustamante, F.; Bustos, P.; Mancilla, M. The genome sequence of an oxytetracycline-resistant Isolate of the fish pathogen Piscirickettsia salmonis harbors a multidrug resistance plasmid. Genome Announc. 2017, 5, e01571-16. [Google Scholar] [CrossRef]
- Cartes, C.; Isla, A.; Lagos, F.; Castro, D.; Muñoz, M.; Yañez, A.; Haussmann, D.; Figueroa, J. Search and analysis of genes involved in antibiotic resistance in Chilean strains of Piscirickettsia salmonis. J. Fish Dis. 2016, 40, 1025–1039. [Google Scholar] [CrossRef]
- Sandoval, R.; Oliver, C.; Valdivia, S.; Valenzuela, K.; Haro, R.E.; Sánchez, P.; Olavarría, V.H.; Valenzuela, K.; Avendaño-Herrera, R.; Romero, A.; et al. Resistance-nodulation-division efflux pump acrAB is modulated by florfenicol and contributes to drug resistance in the fish pathogen Piscirickettsia salmonis. FEMS Microbiol. Lett. 2016, 363, fnw102. [Google Scholar] [CrossRef]
- Pulgar, R.; Travisany, D.; Zuñiga, A.; Maass, A.; Cambiazo, V. Complete genome sequence of Piscirickettsia salmonis LF-89 (ATCC VR-1361) a major pathogen of farmed salmonid fish. J. Biotechnol. 2015, 212, 30–31. [Google Scholar] [CrossRef]
- Yáñez, A.J.; Molina, C.; Haro, R.E.; Sanchez, P.; Isla, A.; Mendoza, J.; Rojas-Herrera, M.; Trombert, A.; Cárcamo, J.G.; Figueroa, J.; et al. Draft genome sequence of virulent strain AUSTRAL-005 of Piscirickettsia salmonis, the etiological agent of piscirickettsiosis. Genome Announc. 2014, 2, e00990-14. [Google Scholar] [CrossRef]
- Hartl, F.U.; Hayer-Hartl, M. Molecular chaperones in the cytosol: From nascent chain to folded protein. Science 2012, 295, 1852–1858. [Google Scholar] [CrossRef]
- Henderson, B.; Martin, A.C. Protein Moonlighting: A New Factor in Biology and Medicine; Portland Press Limited: London, UK, 2014. [Google Scholar]
- Garduño, R.A.; Garduño, E.; Hoffman, P.S. Surface-associated Hsp60 chaperonin of Legionella pneumophila mediates invasion in a HeLa cell model. Infect. Immun. 1998, 66, 4602–4610. [Google Scholar] [CrossRef] [PubMed]
- Hennequin, C.; Porcheray, F.; Waligora-Dupriet, A.-J.; Collignon, A.; Barc, M.-C.; Bourlioux, P.; Karjalainen, T. Hsp60 (Hsp60) of Clostridium difficile is involved in cell adherence. Microbiology 2001, 147, 87–96. [Google Scholar] [CrossRef] [PubMed]
- Ohue, R.; Hashimoto, K.; Nakamoto, M.; Furukawa, Y.; Masuda, T.; Kitabatake, N.; Tani, F. Bacterial heat shock protein 60, Hsp60, can induce the conversion of naïve T cells into a CD4+ CD25+ Foxp3-expressing phenotype. J. Innate Immun. 2011, 3, 605–613. [Google Scholar] [CrossRef] [PubMed]
- González-López, M.A.; Velázquez-Guadarrama, N.; Romero-Espejel, M.E.; Olivares-Trejo, J. Helicobacter pylori secretes the chaperonin Hsp60 (Hsp60), which binds iron. FEBS Lett. 2013, 587, 1823–1828. [Google Scholar] [CrossRef]
- Chong, A.; Lima, C.A.; Allan, D.S.; Nasrallah, G.K.; Garduño, R.A. The purified and recombinant Legionella pneumophila chaperonin alters mitochondrial trafficking and microfilament organization. Infect. Immun. 2009, 77, 4724–4739. [Google Scholar] [CrossRef]
- Nasrallah, G.K.; Riveroll, A.L.; Chong, A.; Murray, L.E.; Lewis, P.J.; Garduño, R.A. Legionella pneumophila requires polyamines for optimal intracellular growth. J. Bacteriol. 2012, 194, 3032. [Google Scholar] [CrossRef]
- Marshall, S.H.; Conejeros, P.; Zahr, M.; Olivares, J.; Gómez, F.; Cataldo, P.; Henríquez, V. Immunological characterization of a bacterial protein isolated from salmonid fish naturally infected with Piscirickettsia salmonis. Vaccine 2007, 25, 2095–2102. [Google Scholar] [CrossRef]
- Wilhelm, V.; Soza, C.; Martínez, R.; Rosemblatt, M.; Burzio, L.O.; Valenzuela, P.D. Production and immune response of recombinant Hsp60 and Hsp70 from the salmon pathogen Piscirickettsia salmonis. Biol. Res. 2005, 38, 69–82. [Google Scholar] [CrossRef]
- Wilhelm, V.; Miquel, A.; Burzio, L.O.; Rosemblatt, M.; Engel, E.; Valenzuela, S.; Parada, G.; Valenzuela, P.D. A vaccine against the salmonid pathogen Piscirickettsia salmonis based on recombinant proteins. Vaccine 2006, 24, 5083–5091. [Google Scholar] [CrossRef]
- Karatas, S.; Mikalsen, J.; Steinum, T.; Taksdal, T.; Bordevik, M.; Colquhoun, D. Real time PCR detection of Piscirickettsia salmonis from formalin-fixed paraffin-embedded tissues. J. Fish Dis. 2008, 31, 747–753. [Google Scholar] [CrossRef]
- Oliver, C.; Valenzuela, K.; Silva, H.; Haro, R.; Cortés, M.; Sandoval, R.; Pontigo, J.P.; Álvarez, C.; Figueroa, J.E.; Avendaño-Herrera, R.; et al. Effectiveness of egg yolk immunoglobulin against the intracellular salmonid pathogen Piscirickettsia salmonis. J. Appl. Microbiol. 2015, 119, 365–376. [Google Scholar] [CrossRef] [PubMed]
- Sievers, F.; Wilm, A.; Dineen, D.G.; Gibson, T.J.; Karplus, K.; Li, W.; Lopez, R.; McWilliam, H.; Remmert, M.; Soding, J.; et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 2011, 7, 539. [Google Scholar] [CrossRef] [PubMed]
- Campanella, J.J.; Bitincka, L.; Smalley, J. MatGAT: An application that generates similarity/identity matrices using protein or DNA sequences. BMC Bioinform. 2003, 4, 29. [Google Scholar] [CrossRef] [PubMed]
- Guindon, S.; Dufayard, J.F.; Lefort, V.; Anisimova, M.; Hordijk, W.; Gascuel, O. New algorithms and methods to estimate maximum-likelihood phylogenies: Assessing the performance of PhyML 3.0. Syst. Biol. 2010, 59, 307–321. [Google Scholar] [CrossRef] [PubMed]
- Ronquist, F.; Huelsenbeck, J.P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 2003, 19, 1572–1574. [Google Scholar] [CrossRef]
- Samudrala, R.; Heffron, F.; McDermott, J.E. Accurate prediction of secreted substrates and identification of a conserved putative secretion signal for type III secretion systems. PLoS Pathog. 2009, 5, e1000375. [Google Scholar] [CrossRef]
- Goldberg, T.; Rost, B.; Bromberg, Y. Computational prediction shines light on type III secretion origins. Sci. Rep. 2016, 6, 34516. [Google Scholar] [CrossRef]
- Eichinger, V.; Nussbaumer, T.; Platzer, A.; Jehl, M.A.; Arnold, R.; Rattei, T. EffectiveDB—Updates and novel features for a better annotation of bacterial secreted proteins and Type III, IV, VI secretion systems. Nucleic Acids Res. 2016, 44, D669–D674. [Google Scholar] [CrossRef]
- Löwer, M.; Schneider, G. Prediction of type III secretion signals in genomes of gram negative bacteria. PLoS ONE 2009, 4, e5917. [Google Scholar] [CrossRef]
- Wang, Y.; Wei, X.; Bao, H.; Liu, S.L. Prediction of bacterial type IV secreted effectors by C-terminal features. BMC Genom. 2014, 15, 50. [Google Scholar] [CrossRef]
- Wang, J.; Yang, B.; Leier, A.; Marquez-Lago, T.T.; Hayashida, M.; Rocker, A.; Zhan, Y.; Akutsu, T.; Chou, K.C.; Strugnell, R.A.; et al. Bastion6: A bioinformatics approach for accurate prediction of type VI secreted effectors. Bioinformatics 2018, 34, 2546–2555. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Yao, Y.; Xu, H.H.; Hao, L.; Deng, Z.; Rajakumar, K.; Ou, H.Y. SecReT6: A web-based resource for type VI secretion systems found in bacteria. Environ. Microbiol. 2015, 17, 2196–2202. [Google Scholar] [CrossRef] [PubMed]
- Hanaichi, T.; Sato, T.; Iwamoto, T.; Malavasi-Yamashiro, J.; Hoshino, M.; Mizuno, N. A stable lead by modification of Sato’s method. J. Electron Spectrosc. 1986, 35, 304–306. [Google Scholar] [CrossRef]
- Thein, M.; Sauer, G.; Paramasivam, N.; Grin, I.; Linke, D. Efficient subfractionation of gram-negative bacteria for proteomics studies. J. Proteome Res. 2010, 9, 6135–6147. [Google Scholar] [CrossRef]
- Akita, E.; Nakai, S. Immunoglobulins from egg yolk: Isolation and purification. J. Food. Sci. 1992, 57, 629–634. [Google Scholar] [CrossRef]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT–PCR. Nucleic Acids Res. 2001, 29, e45. [Google Scholar] [CrossRef]
- Zügel, U.; Kaufmann, S.H. Role of heat shock proteins in protection from and pathogenesis of infectious diseases. Clin. Microbiol. Rev. 1999, 12, 19–39. [Google Scholar] [CrossRef]
- Scopio, A.; Johnson, P.; Laquerre, A.; Nelson, D.R. Subcellular localization and chaperone activities of Borrelia burgdorferi Hsp60 and Hsp70. J. Bacteriol. 1994, 176, 6449–6456. [Google Scholar] [CrossRef]
- Fernandez, R.C.; Logan, S.M.; Lee, S.; Hoffman, P.S. Elevated levels of Legionella pneumophila stress protein Hsp60 early in infection of human monocytes and L929 cells correlate with virulence. Infect. Immun. 1996, 64, 1968–1976. [Google Scholar] [CrossRef]
- Kovacs, E.; Török, Z.; Horváth, I.; Vigh, L. Heat stress induces association of the GroEL-analog chaperonin with thylakoid membranes in cyanobacterium, Synechocystis PCC 6803. Plant. Physiol. Biochem. 1994, 32, 285–293. [Google Scholar]
- Huesca, M.; Borgia, S.; Hoffman, P.; Lingwood, C.A. Acidic pH changes receptor binding specificity of Helicobacter pylori: A binary adhesion model in which surface heat shock (stress) proteins mediate sulfatide recognition in gastric colonization. Infect. Immun. 1996, 64, 2643–2648. [Google Scholar] [CrossRef] [PubMed]
- Ensgraber, M.; Loos, M. A 66-kilodalton heat shock protein of Salmonella typhimurium is responsible for binding of the bacterium to intestinal mucus. Infect. Immun. 1992, 60, 3072–3078. [Google Scholar] [CrossRef] [PubMed]
- Gomez, F.A.; Tobar, J.A.; Henríquez, V.; Sola, M.; Altamirano, C.; Marshall, S.H. Evidence of the presence of a functional Dot/Icm type IV-B secretion system in the fish bacterial pathogen Piscirickettsia salmonis. PLoS ONE 2013, 8, e54934. [Google Scholar] [CrossRef] [PubMed]
- McDermott, J.E.; Corrigan, A.; Peterson, E.; Oehmen, C.; Niemann, G.; Cambronne, E.D.; Sharp, D.; Adkins, J.N.; Samudrala, R.; Heffron, F. Computational prediction of type III and IV secreted effectors in gram-negative bacteria. Infect. Immun. 2011, 79, 23–32. [Google Scholar] [CrossRef]
- Meyer, D.F.; Noroy, C.; Moumene, A.; Raffaele, S.; Albina, E.; Vachiery, N. Searching algorithm for type IV secretion system effectors 1.0: A tool for predicting type IV effectors and exploring their genomic context. Nucleic Acids Res. 2013, 41, 9218–9229. [Google Scholar] [CrossRef]
- Zou, L.; Nan, C.; Hu, F. Accurate prediction of bacterial type IV secreted effectors using amino acid composition and PSSM profiles. Bioinformatics 2013, 29, 3135–3142. [Google Scholar] [CrossRef]
- Valenzuela-Valderas, K.N.; Riveroll, A.L.; Robertson, P.; Murray, L.E.; Garduño, R.A. Legionella pneumophila chaperonin 60, an extra-and intra-cellular moonlighting virulence related factor. Moonlight. Proteins Novel Virulence Factors Bact. Infect. 2017, 111–134. [Google Scholar] [CrossRef]
- Guzman, F.; Fernando, G.; Henriquez, V.; Olivares, J.; Cardenas, C.; Marshall, S. Evaluation of immunogenic peptides derived from a highly protective antigenic protein of Piscirickettsia salmonis as a potential source for vaccine development. New Biotechnol. 2009, 25, S37. [Google Scholar] [CrossRef]
- Marshall, S.H.; Gómez, F.A.; Ramírez, R.; Nilo, L.; Henríquez, V. Biofilm generation by Piscirickettsia salmonis under growth stress conditions: A putative in vivo survival/persistence strategy in marine environments. Res. Microbiol. 2012, 163, 557–566. [Google Scholar] [CrossRef]
- Garrett, T.R.; Bhakoo, M.; Zhang, Z. Bacterial adhesion and biofilms on surfaces. Prog. Nat. Sci. 2008, 18, 1049–1056. [Google Scholar] [CrossRef]
- Hori, K.; Matsumoto, S. Bacterial adhesion: from mechanism to control. Biochem. Eng. J. 2010, 48, 424–434. [Google Scholar] [CrossRef]
- Van Hoek, M.L. Biofilms: An advancement in our understanding of Francisella species. Virulence 2013, 4, 833–846. [Google Scholar] [CrossRef] [PubMed]
- Murphy, K.; Park, A.J.; Hao, Y.; Brewer, D.; Lam, J.S.; Khursigara, C.M. Influence of O polysaccharides on biofilm development and outer membrane vesicle biogenesis in Pseudomonas aeruginosa PAO1. J. Bacteriol. 2014, 196, 1306–1317. [Google Scholar] [CrossRef] [PubMed]
- Grande, R.; Di Marcantonio, M.C.; Robuffo, I.; Pompilio, A.; Celia, C.; Di Marzio, L.; Paolino, D.; Codagnone, M.; Stoodley, P.; Hall-Stoodley, L.; et al. Helicobacter pylori ATCC 43629/NCTC 11639 outer membrane vesicles (OMVs) from biofilm and planktonic phase associated with extracellular DNA (eDNA). Front. Microbiol. 2015, 6, 1369. [Google Scholar] [CrossRef] [PubMed]
- Yonezawa, H.; Osaki, T.; Kurata, S.; Fukuda, M.; Kawakami, H.; Ochiai, K.; Hanawa, T.; Kamiya, S. Outer membrane vesicles of Helicobacter pylori TK1402 are involved in biofilm formation. BMC Microbiol. 2009, 9, 197. [Google Scholar] [CrossRef]
- Yonezawa, H.; Osaki, T.; Woo, T.; Kurata, S.; Zaman, C.; Hojo, F.; Hanawa, T.; Kato, S.; Kamiya, S. Analysis of outer membrane vesicle protein involved in biofilm formation of Helicobacter pylori. Anaerobe 2011, 17, 388–390. [Google Scholar] [CrossRef]
- Oliver, C.; Valenzuela, K.; Hernández, M.; Sandoval, R.; Haro, R.E.; Avendaño-Herrera, R.; Cárcamo, J.G.; Villar, M.T.; Artigues, A.; Garduño, R.; et al. Characterization and pathogenic role of outer membrane vesicles produced by the fish pathogen Piscirickettsia salmonis under in vitro conditions. Vet. Microbiol. 2016, 184, 94–101. [Google Scholar] [CrossRef]
- Oliver, C.; Hernández, M.A.; Tandberg, J.I.; Valenzuela, K.N.; Lagos, L.X.; Haro, R.E.; Sánchez, P.; Ruíz, P.A.; Sanhueza-Oyarzún, C.; Cortés, M.A.; et al. The proteome of biologically active membrane vesicles from Piscirickettsia salmonis LF-89 type strain identifies plasmid-encoded putative toxins. Front. Cell. Infect. Microbiol. 2017, 7, 420. [Google Scholar] [CrossRef]
- Zarankiewicz, T.; Madej, J.; Galli, J.; Bajzert, J.; Stefaniak, T. Inhibition of in vitro Histophilus somni biofilm production by recombinant Hsp60 antibodies. Pol. J. Vet. Sci. 2012, 15, 373–378. [Google Scholar] [CrossRef]


| Subcellular Location | Distribution of Epitopes (1) (%) |
|---|---|
| Cytoplasm | 1.49 ± 1.1 (61.2) |
| Cell envelope | 0.67 ± 0.73 (27.5) |
| Extracellular surface | 0.28 ± 0.48 (11.3) |
| Total | 2.43 ± 1.17 (100) |
| Cell Compartment | Protein Name | Xcor (1) | Sequence Coverage (%) (2) | Matched Peptides |
|---|---|---|---|---|
| Outer membrane | 60 kDa Chaperonin Hsp60 | 12,01 | 6,00 | 2 |
| Periplasm | 60 kDa Chaperonin Hsp60 | 19,30 | 9,00 | 3 |
| Inner membrane | 60 kDa Chaperonin Hsp60 | 6,23 | 7,00 | 2 |
| Cytoplasm | 60 kDa Chaperonin Hsp60 | 2329,85 | 51,00 | 41 |
| Outer membrane vesicles | 60 kDa Chaperonin Hsp60 | 286,72 | 48,53 | 14 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Oliver, C.; Sánchez, P.; Valenzuela, K.; Hernández, M.; Pontigo, J.P.; Rauch, M.C.; Garduño, R.A.; Avendaño-Herrera, R.; Yáñez, A.J. Subcellular Location of Piscirickettsia salmonis Heat Shock Protein 60 (Hsp60) Chaperone by Using Immunogold Labeling and Proteomic Analysis. Microorganisms 2020, 8, 117. https://doi.org/10.3390/microorganisms8010117
Oliver C, Sánchez P, Valenzuela K, Hernández M, Pontigo JP, Rauch MC, Garduño RA, Avendaño-Herrera R, Yáñez AJ. Subcellular Location of Piscirickettsia salmonis Heat Shock Protein 60 (Hsp60) Chaperone by Using Immunogold Labeling and Proteomic Analysis. Microorganisms. 2020; 8(1):117. https://doi.org/10.3390/microorganisms8010117
Chicago/Turabian StyleOliver, Cristian, Patricio Sánchez, Karla Valenzuela, Mauricio Hernández, Juan Pablo Pontigo, Maria C. Rauch, Rafael A. Garduño, Ruben Avendaño-Herrera, and Alejandro J. Yáñez. 2020. "Subcellular Location of Piscirickettsia salmonis Heat Shock Protein 60 (Hsp60) Chaperone by Using Immunogold Labeling and Proteomic Analysis" Microorganisms 8, no. 1: 117. https://doi.org/10.3390/microorganisms8010117
APA StyleOliver, C., Sánchez, P., Valenzuela, K., Hernández, M., Pontigo, J. P., Rauch, M. C., Garduño, R. A., Avendaño-Herrera, R., & Yáñez, A. J. (2020). Subcellular Location of Piscirickettsia salmonis Heat Shock Protein 60 (Hsp60) Chaperone by Using Immunogold Labeling and Proteomic Analysis. Microorganisms, 8(1), 117. https://doi.org/10.3390/microorganisms8010117





